Abstract
The control of agricultural pest insects currently relies on broad-spectrum insecticides, which select for resistance in pest populations while also harming non-target species. In contrast, RNA interference (RNAi) has a species-dependent mode of action based on the delivery of double-stranded RNA (dsRNA) that precisely matches essential genes in pests, minimizing off-target effects. The successful application of RNAi requires the development of sprayable formulations that temporarily protect the dsRNA from environmental degradation (allowing uptake by pest insects) but also ensure the efficient release of the dsRNA within insect cells. Lipid nanoparticles (LNPs) based on pharmaceutical-grade lipids are currently too expensive for agricultural use, making the development of affordable and scalable dsRNA-LNP formulations essential for spray-induced gene silencing. Here we used technical-grade lipid components (available at the ton scale) and demonstrated the cost-effective production of structurally controlled dsRNA-LNP formulations by optimizing formulation recipes and scaling up the microfluidic mixing process. The dispersions contained spherical nanoparticles less than 100 nm in diameter, with a zeta potential exceeding + 20 mV, and an entropy-driven Gibbs free energy change for dsRNA-LNP decomplexation in the moderate range of approximately – 20 kJ/mol. The formulations protected dsRNA from RNase III degradation and hydrolysis at pH 4–11 for at least 24 h while allowing SDS-mediated dsRNA release. Our work provides insight into the structure–property correlations of inexpensive dsRNA-LNP formulations for sustainable RNAi-based pest management systems.
Similar content being viewed by others
Introduction
Agricultural pest insects cause major crop losses by direct damage and the transmission of plant diseases, thus threatening global food security1,2. Furthermore, one of the anticipated effects of climate change is an insect pest population boom that will drive expansion into new geographical areas3,4,5. The use of broad-spectrum insecticides to control native and invasive insect pests has a negative effect on ecologically beneficial non-target organisms and reduces overall biodiversity. In addition, the management of insecticide resistance in insect pests will become increasingly important for a growing number of pest species6,7. For example, the green peach aphid Myzus persicae, which vectors several plant pathogens, is one of the economically most relevant pests in numerous fruit and vegetable crops8. As of 2025, the Arthropod Pesticide Resistance Database lists more than 110 cases of field-evolved resistance in this species against the otherwise highly active neonicotinoid imidacloprid9. Detrimental off-target effects associated with such neonicotinoids have been observed among pollinators and other non-target organisms10,11,12,13. The persistence and bioaccumulation of some insecticides can lead to further negative impacts along the food chain14,15,16,17. New control methods that avoid harm to non-target species are therefore urgently needed.
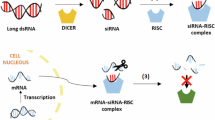
RNA interference (RNAi) is a gene-silencing mechanism triggered by double-stranded RNA (dsRNA) that precisely matches the target gene, offering a promising species-dependent plant protection alternative to chemical insecticides with a biochemical mode of action18,19,20. RNAi in agriculture initially focused on transgenic crops that express dsRNA, resulting in host-induced gene silencing (HIGS)21,22,23. However, regulatory restrictions reflecting the low public acceptance of genetically modified crops have driven research toward alternative RNAi delivery methods. Exogenously applied sprays containing dsRNA can also induce RNAi in insect pests, and the potential of spray-induced gene silencing (SIGS) was recently demonstrated by Greenlights Calantha, the first commercial RNAi-based plant protection product, which is used to control Colorado potato beetle (Leptinotarsa decemlineata)24,25,26. The RNAi mechanism can inhibit the production of proteins essential for development or survival by triggering the degradation of the corresponding mRNA, ultimately causing mortality27,28,29. Essential genes are involved in diverse physiological systems, including digestion, immunity, vesicular trafficking, and detoxification30,31,32,33,34,35,36,37,38. The design of dsRNAs to match unique sequences only found in the target pest minimizes or completely avoids effects in non-target species, including beneficial organisms39. Furthermore, any dsRNA remaining in the environment is broken down within days to weeks by numerous RNA-degrading enzymes, reducing risks of bioaccumulation in the environment or food chain40. The evolution of resistance to RNAi-based agents has been observed in laboratory settings but no field-evolved resistance has been reported thus far due to the novelty of this gene-silencing mechanism41.
Although the susceptibility of dsRNA to degradation is an advantage in the context of environmental safety, it is also an impediment to the successful deployment of SIGS in agriculture because premature degradation on the plant surface, or indeed in the insect gut, can reduce the efficacy of RNAi42. Lipid nanoparticles (LNPs) are widely used for the protection of nucleic acid medicines and are formed by the self-assembly of DNA or RNA with complexing ionic or ionizable lipids, dispersion stabilizers, and helper lipids acting as bilayer modifiers43,44,45,46. However, pharmaceutical-grade LNP materials are too expensive for agricultural applications. The development of cost-effective and scalable dsRNA-LNP formulations is therefore necessary for the agricultural use of SIGS.
Here we demonstrate the potential of cost-effective, technical-grade lipid components and the structure–process–property correlations of the resulting dsRNA-LNP formulations. By optimizing formulation recipes and microfluidic mixing parameters, we show it is possible to produce dsRNA-LNPs with controlled, spherical morphology, which can protect dsRNA from enzymatic degradation and hydrolysis, and also allow for dsRNA release in vitro. The development of dsRNA-LNP formulations based on technical lipids is a cost-effective solution with the potential to expand the use of RNAi in crop protection.
Results
Process–structure correlations of dsRNA-LNPs
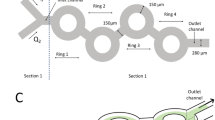
To improve the formation of complexes between dsRNA and LNPs, we developed a formulation toolbox comprising three inexpensive lipid components already available in large quantities, making them suitable for agricultural applications. We selected the quaternary fatty amine didodecyldimethylammonium bromide (DDAB) as a cationic reagent to from complexes with dsRNA, polyethylene glycol 15-hydroxystearate (Kolliphor HS-15), a PEGylated lipid acting as a dispersion stabilizer, and lecithin (derived from egg yolk), a natural amphoteric phospholipid mixture that modifies the molecular packing of LNPs. Encapsulation is thought to be initiated by a complexation reaction driven by ionic interactions between the cationic ammonium group of DDAB and the anionic phosphate groups of the dsRNA backbone. The rapidly decreasing concentration of organic solvent during mixing triggers self-assembly of the nucleated particles, incorporating the co-lipids and remaining DDAB via hydrophobic effects. First, we tested a lipid-dsRNA complex with a molar lipid ratio of 2:1:1 (DDAB:PEG-lipid:lecithin) in the mid-range of the ternary mixture diagram. We used an excess of this blend to form complexes with dsRNA, comparing lipid-to-phosphate (L:P) ratios of 4:1, 6:1 and 8:1 (described hereafter as L:P = 4, 6 and 8; Fig. 1A). Preliminary experiments revealed that the lipid blend mentioned above required at least L:P = 4 to achieve efficient particle formation (L:P = 3 resulted in precipitation). The complexes were produced by rapid microfluidic mixing of the lipid components (dissolved in ethanol) and an aqueous dsRNA solution using a serpentine channel specifically designed for efficient mixing (Fig. 1B). The relative flow rate of the dsRNA and lipid solutions during processing was 9:1, resulting in a dsRNA concentration of 0.2 mg/mL and a lipid concentration of 2.5–5.0 mM for L:P = 4–8 in 10% (v/v) ethanol. The total lipid concentration was comparable to pharmaceutical lipid blends typically used for the formulation of nucleic acid vaccines, although we omitted the cholesterol often found in pharmaceutical formulations 43. The microfluidic process was optimized for total flow rates (TFRs) of 4–16 mL/min. Visual observation of the formulations revealed a correlation between the L:P ratio and TFR during processing, resulting in different degrees of opalescence (Fig. 1C). Specifically, lower L:P ratios and slower mixing (lower TFRs) produced white, opalescent dispersions, whereas higher L:P ratios and faster mixing (higher TFRs) resulted in clear, blueish opalescent dispersions. These variations suggest differences in particle size within the dispersion, with higher L:P ratios and higher TFRs generating smaller particles.
Process-property correlation of dsRNA-LNP formulations. A Ternary phase diagram of lipid mixing at molar ratios of 2:1:1 (DDAB:PEG-lipid:lecithin) and total lipid-to-dsRNA phosphate group ratios of L:P = 4, 6 and 8. B Microfluidic 3D serpentine mixer chip with hydrodynamic flow focusing inlet for dsRNA-LNP formulation. C Images of dsRNA-LNP formulations at three L:P ratios and TFRs of 4–16 mL/min, showing lower opalescence at higher L:P and TFR values. D Agarose gel electrophoresis of dsRNA-LNP formulations. Absence of bands indicates complete complexation. E Gel after SDS decomplexation. Recovery of dsRNA bands confirms cargo integrity. F–K DLS and ELS measurements of size (Z-average), zeta potential and PDI (data are means ± standard deviation, n = 3) as functions of TFR (F–H) and L:P ratio (I–K). General trends are: higher flow rates lead to smaller particles with lower zeta potential and lower PDI whereas higher L:P ratios lead to smaller particles with a greater zeta potential and PDI. Corresponding particle size distributions and correlograms of DLS measurements are shown in Figure S1A, B.
The formulations were assessed for dsRNA complexation by agarose gel electrophoresis. Unformulated dsRNA appeared as a distinct band at ~ 550 bp (Fig. 1D). In contrast, no band was visible for the dsRNA-LNP formulations, indicating complete complexation and thus immobilization of the dsRNA (the complex does not migrate to the anode due to particle size and charge). Addition of the anionic surfactant sodium dodecylsulfate (SDS), which interacts with the cationic charge of DDAB and releases the dsRNA, led to the reappearance of a distinct band migrating slightly above the original ~ 550 bp dsRNA band observed in control samples without cationic or anionic constituents (Fig. 1E). The addition of SDS to the unformulated dsRNA reduced the intensity of the band relative to the untreated dsRNA. Formulations with L:P ratios of 4 and 6 showed near-identical band intensities after decomplexation, whereas a lower intensity was observed at L:P = 8, comparable to that observed for SDS-treated unformulated dsRNA. This reduction in intensity at higher L:P ratios may reflect incomplete decomplexation because the same amount of SDS was added to each sample and may not have been sufficient to completely neutralize the greater number of cationic lipids present in the L:P = 8 particles. The particle size, zeta potential and polydispersity index (PDI) were determined by dynamic light scattering (DLS) and electrophoretic light scattering (ELS) (Fig. 1F–K). The particle sizes were expressed as Z-average values from measurements in backscattering mode, and ranged from ~ 50 nm (L:P = 8, TFR = 16 mL/min) to ~ 100 nm (L:P = 4, TFR = 4 mL/min).
Higher flow rates therefore produced smaller particles, and higher L:P ratios produced smaller particles with a higher zeta potential and PDI. The particle sizes were consistent with the visual opalescence (Fig. 1C). Generally, particle sizes decreased at all L:P ratios with increasing flow rates, suggesting that the mixing channel architecture of our microfluidic chip requires a minimum flow rate to achieve sufficient turbulence and mixing performance for optimal nanoparticle formation kinetics. Interestingly, the zeta potential at a given L:P ratio appeared to be relatively independent of the TFR (Fig. 1G). However, higher L:P ratios at all flow rates resulted in a higher zeta potential. Across all formulations, the zeta potential ranged from approximately + 20 mV for high flow rates and low L:P ratios (L:P = 4, TFR = 16 mL/min) to + 35 mV for low flow rates and high L:P ratios (L:P = 8, TFR = 4 mL/min) (Fig. 1J). The PDI of all formulations was 0.1–0.2 but could be reduced by increasing the TFR and decreasing the L:P ratio (Fig. 1H, K). The recipe data and corresponding values are summarized in the supporting information (Figure S1A, B; Table S1).
These findings suggest that the morphology of dsRNA-LNP particles is strongly influenced by both the lipid content and mixing conditions. Variations in these parameters resulted in significant changes to particle structure and heterogeneity, as demonstrated by cryogenic transmission electron microscopy (cryo-TEM) in Fig. 2 (showing two representative images for each condition). For example, formulations with an L:P ratio of 4 mixed at a TFR of 4 mL/min contained heterogeneous particles, including agglomerates of spherical and non-spherical particles (100–400 nm) and larger structures (Fig. 2A). Increasing the TFR to 16 mL/min promoted the formation of more spherical particles, but these still formed agglomerates, including some with lamellar features (Fig. 2B). The lamellar structures were identified in TEM freeze-fracture replica experiments as dsRNA molecules (Figure S2). Notably, structural features that can be associated to free, uncomplexed dsRNA were not observed, indicating high dsRNA encapsulation efficiency. Nevertheless, some agglomerates that formed at an L:P ratio of 4 remained non-spherical and heterogeneous and larger than 200 nm, probably because insufficient lipids were available to support the formation of well-defined, spherical morphologies. The ratio between those larger agglomerates and small, well-dispersed particles was investigated by multi-angle dynamic light scattering (MADLS), comparing the intensity to the number and volume particle size distribution plots, which revealed that at least 95% of the particle population produced at high flow rates (TFR = 12 or 16 mL/min) were < 100 nm in diameter, with only occasional larger structures (Figure S3; Table S1).
Cryo-TEM images of dsRNA-LNP formulations prepared using different total lipid-to-dsRNA phosphate group ratios (L:P) and total flow rates (TFRs). Two micrographs of the same formulations are shown to give a representative impression of the particle populations. A L:P = 4, TFR = 4 mL/min. B L:P = 4, TFR = 16 mL/min. C L:P = 8, TFR = 4 mL/min. D L:P = 8, TFR = 16 mL/min. Increasing the lipid content and/or mixing speed enhances the formation of well-defined spherical particles. Small spherical particles (blue arrows), large merged agglomerates (orange arrows), and empty vesicles (white arrows) are highlighted.
Increasing the L:P ratio to 8 generated a higher proportion of small, well-defined spherical particles with lamellar features (Fig. 2C). At TFR = 4 mL/min, particles of 20–80 nm diameter were formed, similar in size to those at L:P = 4, but with a more uniform spherical morphology and fewer agglomerates. A notable fraction of empty spherical particles (15–70 nm) was observed, indicating excess lipids remained in the formulation after complete dsRNA encapsulation. At TFR = 16 mL/min, we observed even smaller loaded particles (15–50 nm) and fewer, smaller agglomerates than at TFR = 4 mL/min (Fig. 2D). We observed some empty particles, mostly 15–70 nm in diameter but also some larger structures. Increasing the TFR to 16 mL/min at L:P = 8 therefore reduced the particle size and ability to form larger agglomerates without compromising encapsulation efficiency. These results demonstrate that both lipid content and mixing conditions are key determinants of particle morphology, size distribution, and encapsulation behavior.
Stability of dsRNA-LNPs
Given that the lipid content and processing conditions influence the morphology and size of dsRNA-LNP systems, we conducted in vitro stability assays to evaluate the ability of the LNPs to provide temporary protection against dsRNA degradation. The stability of both unformulated and LNP-formulated dsRNA was assessed by agarose gel electrophoresis in the presence of RNase III and under a range of pH conditions (Fig. 3).
Stability of dsRNA and dsRNA-LNP formulations determined by agarose gel electrophoresis. A LNP formulations prepared using any of our L:P ratios and TFR parameters protect dsRNA from digestion by RNase III for at least 24 h, whereas unformulated dsRNA is fully degraded. B, C Non-formulated dsRNA exposed to pH 3–12 for 1 and 24 h. Reactions were quenched with ammonium acetate at each time point to stop further degradation. Unformulated dsRNA degrades rapidly at pH 2–3 and pH 12. D, E Encapsulation in LNPs (L:P = 4 or 8) protects dsRNA from degradation at pH 3–4 for up to 24 h, but offers no protection at pH 2 or 12. L4T4 = L:P 4 + TFR 4, L8T8 = L:P 8 + TFR 8. Control samples were not pH adjusted.
In the presence of RNase III, unformulated dsRNA was degraded within 24 h whereas all the dsRNA-LNP formulations we prepared, regardless of L:P ratio and TFR, provided good protection over the same duration (Fig. 3A). The stability of unformulated dsRNA varied across the pH range 2–12, with rapid degradation at pH 2–3 and pH 12, and partial degradation at pH 4, but stability at pH 10–11 (Fig. 3B, C). Encapsulation in LNPs with L:P ratios of 4 and 8, prepared at TFRs of 4 and 16 mL/min, significantly protected the dsRNA from acidic hydrolysis at pH 3–4, with intact dsRNA bands recoverable by decomplexation after 24 h (Fig. 3D, E). No additional protection was observed for any formulation at pH 12 compared to unformulated dsRNA.
The observation that L:P = 4 particles precipitated at pH 10 and L:P = 8 particles at pH 11, with subsequent dissolution of the formed solid at higher pH, may explain the lack of protection against such highly alkaline conditions. The isoelectric point of the particles is reached at a certain pH, correlating the respective amount of DDAB, resulting in precipitation due to the lack of inter-particle electrostatic repulsion. Dissolution following a further pH increase may proceed by ion replacement of dsRNA-bound DDAB, yielding sodium-dsRNA and DDAB hydroxide, thus exposing the cargo to the alkaline environment and facilitating degradation. However, these results demonstrate that LNP formulations provide strong protection against enzymatic digestion and significantly enhance stability under acidic conditions. Acidic to neutral pH conditions have been documented in coleopteran pests such as the Colorado potato beetle47,48, supporting the use of LNPs for efficient dsRNA delivery in pest management applications.
Scale-up of the formulation process
Having confirmed particle stability under relevant conditions and the protection of dsRNA, we next addressed the scalability of the formulation process to produce sufficient amounts for field trials. We evaluated the microfluidic mixing step using a hydrodynamic flow-focusing microfluidic chip design and continuous flow process pumps that achieve higher flow rates (Fig. 4A). The experimental design used a constant L:P ratio of 4 to conserve materials, while systematically varying the lipid mixture between 40–60% DDAB, 20–40% PEG-lipid and 20–40% lecithin (Fig. 4B). The compositions and DLS data are summarized in Table 1. The formulation process was run at a TFR of 50 mL/min, using a dsRNA:LNP flow ratio of 9:1, resulting in a batch size of 250 mL with a dsRNA concentration of 0.2 mg/mL.
Scale-up of dsRNA-LNP formulations and correlation of recipe with particle characteristics and thermodynamic stability. A Scaled-up microfluidic mixing using a TFR of 50 mL/min and a flow ratio of 45:5 mL/min (9:1 aqueous dsRNA:ethanolic lipid blend). B Ternary phase diagram of the three lipids used for the production of compositions 1–7. Composition 3 has a 2:1:1 molar ratio of the three lipid components and is therefore identical to the composition used in the initial small-scale formulations. C Cryo-TEM images of composition 3, revealing spherical particles of 20–90 nm (blue arrow) and particle agglomerates (orange arrow). The images show two micrographs from the same formulation. D Z-average, E zeta potential, and F polydispersity index (PDI) for compositions 1–7. G Isothermal titration calorimetry (ITC) profiles for SDS-induced dsRNA-LNP decomplexation. H Enthalpy–entropy compensation plot from ITC data, indicating entropy-dominated decomplexation at ΔG = –19.2 kJ/mol. Color code for DDAB content in panels D–H: black (60%), blue (50%), purple (40%). Detailed ITC data are provided in supplementary information Table S4.
Cryo-TEM analysis of LNP formulations produced using the scaled-up process revealed that the original 2:1:1 lipid blend (composition 3, 50% DDAB, 25% PEG-lipid and 25% lecithin) produced particles resembling those prepared at an L:P ratio of 8 and a TFR of 4 mL/min in the small-scale process (Fig. 4C). These particles were comparable in size and morphology, with a similar proportion of agglomerations, and featured a similar lamellar internal structure to the original small-scale formulation. However, one key difference was the absence of empty vesicles in the larger-scale process. DLS revealed that the LNPs from the scaled-up process increased gradually in size as the proportion of DDAB was reduced, from ~ 85 nm in composition 1 (60% DDAB) to ~ 110 nm in composition 7 (40% DDAB) (Fig. 4D). The quaternary amine lipid content also correlated with the observed zeta potential, which ranged from ~ 35 mV (composition 1) to ~ 28 mV (composition 6) (Fig. 4E). The mean particle size of composition 3 (~ 85 nm) was comparable to the formulation generated using the smaller-scale process at L:P = 4 and TFR = 8 mL/min, but the zeta potential of the particles from the scaled-up process was higher (~ 35 mV vs ~ 25 mV) for a comparable particle size. The PDIs of all compositions were in the range 0.21–0.27 and were therefore generally higher than those from the smaller-scale process (Fig. 4F).
Isothermal titration calorimetry (ITC) was applied to the larger-scale formulations to evaluate their decomplexation efficiency in the presence of SDS. The formulations were diluted to 1 mM total lipid concentration and titrated with 60 mM SDS in 10% ethanol (Fig. S4, Table S4). ITC profiles revealed that all reactions were exergonic with ΔG ranging from –18.0 kJ/mol (composition 5) to –20.3 kJ/mol (composition 4) and a mean value of –19.2 kJ/mol (Fig. 4G). The free Gibbs energy of decomplexation for formulations with lower amounts of DDAB and lecithin was generally less exergonic. Although this apparently makes the decomplexation reaction more exothermic, the effect on ΔG remains relatively constant due to a less negative –TΔS term. The enthalpy-entropy-compensation plot clearly shows that the decomplexation reactions are indeed entropy dominated (Fig. 4H)49.
Discussion
Our analysis of dsRNA-LNP systems was consistent with previous reports demonstrating that key formulation parameters, including lipid composition, L:P ratio, and processing conditions, influence the structural and functional properties of LNP systems for nucleic acid delivery in the context of both agricultural and pharmaceutical applications. Supporting our observations, studies of therapeutic LNPs loaded with siRNA and mRNA have shown that changes in lipid content and processing variables significantly affect particle size, encapsulation efficiency, and uniformity50,51. Similarly, a higher lipid content was found to yield smaller particles but also increased the formation of empty vesicles, whereas insufficient mixing led to partial aggregation52, similar to the agglomeration we observed under suboptimal conditions. LNP encapsulation significantly enhances the stability of dsRNA under acidic and alkaline conditions and provides strong protection against enzymatic digestion, both of which are relevant in the insect gut. Indeed, without protective formulation strategies, naked dsRNA has proven vulnerable to rapid degradation in numerous insect species53. Notably, our LNP system demonstrated molecular stability for more than 24 h in the presence of RNase III and maintained structural integrity for up to 24 h across the pH range 3–11. These stability profiles align with insect gut transit times during feeding and digestion, and are compatible with practical tank-mix preparation for field spray applications using water sources varying in pH. The improved dsRNA stability we observed is also consistent with reports describing nanocarriers based on cationic polymers or lipids, which can enhance the efficacy of RNAi in target pests54,55,56,57,58,59.
Building on fundamental understanding gained at the laboratory production scale, we scaled up the process by implementing a continuous high-throughput manufacturing approach based on an alternative microfluidic chip design suitable for higher flow rates. Operating at a high TFR of 50 mL/min while maintaining a constant L:P ratio of 4, the process remained robust at the pilot scale, providing well-defined spherical nanoparticle morphologies with only a few larger agglomerates, closely resembling those obtained in the smaller-scale process. Further process development and scale-up must focus on increasing the output to economically relevant levels. For example, the use of high-throughput heat exchangers to access larger solution reservoirs while eliminating pre-process heating could leverage material throughput as well as the parallelization of robust mixing units (microfluidic chips) designed for higher flow rates. Process chemistry could also be improved by producing concentrates with higher yields, facilitated by spray drying. Spraying applications typically require surfactants and other additives, such as wetting agents to ensure sufficient foliar coverage. Concentrated surfactants could affect the stability and morphology of dsRNA-LNP formulations, which must be assessed to prepare for diverse spraying scenarios. However, the particle morphologies we observed, using technical and agricultural grades of long dsRNA and lipids, were less defined and less homogeneous than those produced using pharmaceutical-grade mRNA-LNP systems60.
ITC is a powerful technique that can be used to determine the affinity, enthalpy and entropy of interactions between biologically active molecules and their partners, offering valuable insights into the formation and stability of dsRNA-LNP complexes. To our knowledge, this is the first use of ITC to characterize the decomplexation of dsRNA-LNP complexes in the presence of SDS, revealing the key thermodynamic parameters ΔH, –TΔS and ΔG relevant to stability, formulation and processing. The SDS-induced decomplexation of our dsRNA-LNP complexes is entropy-driven due to the released solvation energy of the lipid mixtures, whereby the exergonic state increases in complexes containing higher proportions of the cationic complexing reagent DDAB and with increasing PEG-lipid content at a constant amount of DDAB. Consistently, the ΔG of approximately − 20 kJ/mol falls within the lower/moderate range of interaction energies and is comparable to the non-charge interactions reported for DNA–protein and RNA–protein complexes, but weaker than the additional cationic charge interactions in those complexes61. In this context, additional systematic studies are needed to better understand the thermodynamic processes involved in the decomplexation of dsRNA from LNP formulations and to draw correlations between thermodynamic release characteristics and in vivo release processes.
In summary, these promising results support the use of microfluidic mixing technologies for the scalable production of dsRNA-LNP systems with well-controlled particle size and morphology, and the ability to protect and release dsRNA (in the presence of SDS), making them suitable for both greenhouse applications and field trials. However, the observed particle aggregation suggests that further optimization of mixing parameters and lipid composition may be required to completely prevent agglomeration and improve colloidal stability. Future research should address these formulation variables, including the key thermodynamic parameters that determine the protection and release of dsRNA during decomplexation, as well as the environmental stability and performance of these advanced formulations.
Methods
Microfluidic mixing
For lipid nanoparticle preparation, regardless of production scale, aqueous phases were prepared by dilution of a 523-bp model dsRNA (0.22 mg/mL, Greenlight Biosciences; see Supplementary Information) with ultra-pure water (Arium Pro, Sartorius) and subsequent sterile filtration (CHROMAFIL PF/PET45-25, Carl Roth). Stock solutions (0.4 M) of DDAB (CAS 3282-73-3; Thermo Fisher Scientific), Kolliphor HS-15 (CAS 70142-34-6; Sigma-Aldrich), and egg yolk lecithin (CAS 93685–90-6; Carl Roth) were prepared in 99.5% ethanol (VWR). For egg yolk lecithin, an average molecular weight of 744.38 g/mol was calculated according to data provided elsewhere62. Ethanolic phases with the desired lipid mixing ratios were prepared by mixing these stock solutions and subsequent dilution. Before microfluidic mixing, each solution was filter-sterilized using syringe filters (CHROMAFIL Xtra PTFE-20/25, Carl Roth).
Small-scale production
Small-scale microfluidic mixing was carried out using a 3D serpentine mixer chip (Fluidic 1079; Microfluidic ChipShop). A detailed description of the setup and process protocol is provided in the supplementary information (Figure S5, Table S2). Briefly, aqueous and ethanolic phases were equilibrated at 65 °C and aspirated rapidly, without bubbles, into 20-mL Luer-lock syringes. Filled syringes were then connected to the microfluidic setup and mounted to individual syringe pumps. The ethanolic flow (0.4–1.6 mL/min) was started first and the aqueous flow (3.6–14.4 mL/min) was added after a few seconds. The resulting dsRNA-LNP formulations were collected in fractions every 30 s. The system was then purged with ultra-pure water and ethanol and finally with air before running the next experiment. All formulations were produced in triplicate from individual lipid stock solutions.
Larger-scale production
For larger-scale microfluidic mixing, we used an Ibidi µSlide 3in1 chip (17,296,163; Thermo Fisher Scientific). Briefly, the aqueous and ethanolic phases, as well as ultra-pure water and ethanol, were kept in 1-L Schott flasks immersed in water baths at 65 °C. Solutions were equilibrated for 1 h and kept at that temperature during processing. Before preparing the dsRNA-LNP complexes, the system was purged with ultra-pure water and ethanol. The aqueous and ethanolic flows were started simultaneously, immediately after purging. The dsRNA-LNP formulations were collected in 50-mL fractions. The system was then purged with ultra-pure water and ethanol before running the next experiment. More detailed information is provided in the supplementary information (Figure S6, Table S3).
Agarose gel electrophoresis
For the analysis of complex formation and dsRNA release, as well as RNase III and pH stability, 10-µL samples were separated by 2% agarose gel electrophoresis in TAE buffer (40 mM Tris, 20 mM glacial acetic acid, 1 mM EDTA disodium salt) for 45 min at 80 V. Gels were prepared from NEEO ultra-quality agarose (Carl Roth). Unformulated dsRNA and formulated LNPs (4 µL) were diluted to 0.2 µg/µL dsRNA with 4 µL ultra-pure water and mixed with 1 µL 10 × BlueJuice loading dye (Thermo Fisher Scientific) and 1 µL 3 × SYBR Gold nucleic acid stain in DMSO (Thermo Fisher Scientific). Decomplexation was achieved by adding 1 µL 0.2% SDS (Carl Roth) and incubating for 20 min at room temperature. Results were visualized using the GelDoc XR + system and ImageLab software. Full-size gel pictures are shown in Figure S7.
RNase III assays
Samples of dsRNA or dsRNA-LNPs (20 µL, 0.2 µg/µL) were diluted with ultra-pure water (24 µL) and mixed with 1 µL ShortCut RNase III (2 U/µL; New England Biolabs) and 5 µL 10 × manganese (II) chloride (supplied with enzyme). The reactions were incubated at 37 °C, shaking at 500 rpm, for up to 24 h, before quenching with 5 µL proteinase K (0.8 U/µL; New England Biolabs). After incubation for 1 h, undiluted 8-µL samples were mixed with loading dye and nucleic acid stain and separated by agarose gel electrophoresis as described above.
Stability at different pH values
Samples of dsRNA or dsRNA-LNPs (2 mL) were adjusted to different pH values by adding 10 mM HCl or 10 mM NaOH and confirming the new value using a FiveEasy Plus pH meter with LE422 electrode (Mettler Toledo). Samples adjusted to pH 2–4 or 10–12 (tolerance ± 0.05 pH units) were incubated for up to 24 h at room temperature before 50-µL aliquots were quenched by adding 50 µL 5 mM ammonium acetate. Undiluted 8-µL samples were mixed with loading dye and nucleic acid stain and separated by agarose gel electrophoresis as described above.
DLS measurements
We used a Zetasizer Ultra Red (Malvern Panalytical) to determine the particle size using the Z-average, zeta potential and PDI from readings taken in backscattering mode using folded capillary zeta cells (DTS1070; Malvern Panalytical). For particle size measurements (MADLS) we used 4-mL polystyrene cuvettes (Carl Roth). All samples were equilibrated at room temperature for 1 h and mixed by inversion before measurement. All measurements were taken in triplicate and results were pooled. Data were processed using ZS Explorer v3.3 (Malvern Panalytical) and visualized using Origin2023b.
ITC measurements
For ITC, we used a MicroCal PEAQ-ITC (Malvern Panalytical) and dsRNA-LNP samples with a total lipid concentration of 1 mM. Titration with 60 mM SDS in 10% ethanol was carried out at 25 °C using a reference power of 10 µcal/s, 60 s initial delay, and 750 rpm stirring speed. The initial injection was 0.4 µL, followed by 37 injections of 1 µL each with 150-s intervals. The heat of dilution of the SDS control in 10% ethanol was subtracted. Data were analyzed using MicroCal PEAQ-ITC analysis software assuming one set of binding sites and forcing binding.
Cryo-TEM
We applied 7-µL samples to both sides of gold grids covered with perforated gold film (UltrAuFoil 1.2/1.3; Quantifoil Micro Tools). Excess sample was automatically blotted from the back of the grid with strips of filter paper for 1 s. The samples were rapidly plunge-frozen in liquid ethane (cooled to –180 °C) in a Cryobox (Carl Zeiss). Excess ethane was removed with filter paper. Samples were transferred immediately to a Gatan 626 cryo-transfer holder and into the pre-cooled cryo-electron microscope (Philips CM 120), operated at 120 kV and viewed under low-dose conditions. Images were recorded with an F216 2 K CMOC camera (TVIPS). Four images were averaged to one image to reduce noise.
Data availability
All data generated during this study are included in the article and the supplementary information.
References
Ma, C.-S. et al. Crop pest responses to global changes in climate and land management. Nat. Rev. Earth Environ. 6, 264–283. https://doi.org/10.1038/s43017-025-00652-3 (2025).
Savary, S. et al. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 3, 430–439. https://doi.org/10.1038/s41559-018-0793-y (2019).
Skendžić, S., Zovko, M., Živković, I. P., Lešić, V. & Lemić, D. The impact of climate change on agricultural insect pests. Insects 12, 440. https://doi.org/10.3390/insects12050440 (2021).
Subedi, B., Poudel, A. & Aryal, S. The impact of climate change on insect pest biology and ecology: implications for pest management strategies, crop production, and food security. J. Agric. Food Res. 14, 100733. https://doi.org/10.1016/j.jafr.2023.100733 (2023).
Deutsch, C. A. et al. Increase in crop losses to insect pests in a warming climate. Science 361, 916–919. https://doi.org/10.1126/science.aat3466 (2018).
Bass, C., Denholm, I., Williamson, M. S. & Nauen, R. The global status of insect resistance to neonicotinoid insecticides. Pestic. Biochem. Physiol. 121, 78–87. https://doi.org/10.1016/j.pestbp.2015.04.004 (2015).
Liang, J. et al. Insect resistance to insecticides: Causes, mechanisms, and exploring potential solutions. Arch. Insect Biochem. Physiol. 118, e70045. https://doi.org/10.1002/arch.70045 (2025).
Ali, J. et al. Peach–potato aphid Myzus persicae: Current management strategies, challenges, and proposed solutions. Sustainability 15, 11150. https://doi.org/10.3390/su151411150 (2023).
Mota-Sanchez, D. & Wise, J.C. The Arthropod Pesticide Resistance Database. Search tags: Genus [myzus], Active ingredient [imidacloprid], Type resistance [Field-evolved resistance]. (accessed on 19 Aug 2025). Available at http://www.pesticideresistance.org (2025).
van Lexmond, M. B., Bonmatin, J.-M., Goulson, D. & Noome, D. A. Worldwide integrated assessment on systemic pesticides: global collapse of the entomofauna: exploring the role of systemic insecticides. Environ. Sci. Pollut. Res. Int. 22, 1–4. https://doi.org/10.1007/s11356-014-3220-1 (2015).
Chagnon, M. et al. Risks of large-scale use of systemic insecticides to ecosystem functioning and services. Environ. Sci. Pollut. Res. Int. 22, 119–134. https://doi.org/10.1007/s11356-014-3277-x (2015).
Boily, M., Sarrasin, B., Deblois, C., Aras, P. & Chagnon, M. Acetylcholinesterase in honey bees (Apis mellifera) exposed to neonicotinoids, atrazine and glyphosate: Laboratory and field experiments. Environ. Sci. Pollut. Res. Int. 20, 5603–5614. https://doi.org/10.1007/s11356-013-1568-2 (2013).
Geiger, F. et al. Persistent negative effects of pesticides on biodiversity and biological control potential on European farmland. Basic Appl. Ecol. 11, 97–105. https://doi.org/10.1016/j.baae.2009.12.001 (2010).
Wintermantel, D. et al. Neonicotinoid-induced mortality risk for bees foraging on oilseed rape nectar persists despite EU moratorium. Sci. Total Environ. 704, 135400. https://doi.org/10.1016/j.scitotenv.2019.135400 (2020).
Limay-Rios, V. et al. Neonicotinoid insecticide residues in soil dust and associated parent soil in fields with a history of seed treatment use on crops in southwestern Ontario. Environ. Toxicol. Chem. 35, 303–310. https://doi.org/10.1002/etc.3257 (2016).
Samson-Robert, O., Labrie, G., Chagnon, M. & Fournier, V. Neonicotinoid-contaminated puddles of water represent a risk of intoxication for honey bees. PLoS ONE 9, e108443. https://doi.org/10.1371/journal.pone.0108443 (2014).
Ray, S. & Shaju, S. T. Bioaccumulation of pesticides in fish resulting toxicities in humans through food chain and forensic aspects. Environ. Anal. Health. Toxicol. 38, e2023017-0. https://doi.org/10.5620/eaht.2023017 (2023).
Cagliari, D. et al. Management of pest insects and plant diseases by non-transformative RNAi. Front. Plant Sci. 10, 1319. https://doi.org/10.3389/fpls.2019.01319 (2019).
Zotti, M. et al. RNA interference technology in crop protection against arthropod pests, pathogens and nematodes. Pest Manag. Sci. 74, 1239–1250. https://doi.org/10.1002/ps.4813 (2018).
Das, P. R. & Sherif, S. M. Application of exogenous dsRNAs-induced RNAi in agriculture: Challenges and triumphs. Front. Plant Sci. 11, 946. https://doi.org/10.3389/fpls.2020.00946 (2020).
Mao, Y.-B. et al. Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nat. Biotechnol. 25, 1307–1313. https://doi.org/10.1038/nbt1352 (2007).
Baum, J. A. et al. Control of coleopteran insect pests through RNA interference. Nat. Biotechnol. 25, 1322–1326. https://doi.org/10.1038/nbt1359 (2007).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811. https://doi.org/10.1038/35888 (1998).
Rodrigues, T., Sridharan, K., Manley, B., Cunningham, D. & Narva, K. Development of dsRNA as a sustainable bioinsecticide: From laboratory to field. In Crop Protection Products for Sustainable Agriculture 65–82 (American Chemical Society, 2021). https://doi.org/10.1021/bk-2021-1390.ch005.
Rodrigues, T. B. et al. First sprayable double-stranded RNA-based biopesticide product targets proteasome subunit beta type-5 in Colorado potato beetle (Leptinotarsa decemlineata). Front. Plant Sci. 12, 728652. https://doi.org/10.3389/fpls.2021.728652 (2021).
Graser, L. et al. Targeting the proteasome subunit PSMB5 by RNA interference induces proteasome dysfunction and mortality in the Colorado potato beetle (Leptinotarsa decemlineata). Sci. Rep. 15, 41183. https://doi.org/10.1038/s41598-025-28793-x (2025).
Zhu, K. Y. & Palli, S. R. Mechanisms, applications, and challenges of insect RNA interference. Annu. Rev. Entomol. 65, 293–311. https://doi.org/10.1146/annurev-ento-011019-025224 (2020).
Christiaens, O., Niu, J. & Nji Tizi Taning, C. RNAi in insects: A revolution in fundamental research and pest control applications. Insects 11, 415. https://doi.org/10.3390/insects11070415 (2020).
Cooper, A. M., Silver, K., Zhang, J., Park, Y. & Zhu, K. Y. Molecular mechanisms influencing efficiency of RNA interference in insects. Pest Manag. Sci. 75, 18–28. https://doi.org/10.1002/ps.5126 (2019).
Schmitt-Engel, C. et al. The iBeetle large-scale RNAi screen reveals gene functions for insect development and physiology. Nat. Commun. 6, 7822. https://doi.org/10.1038/ncomms8822 (2015).
Ulrich, J. et al. Large scale RNAi screen in Tribolium reveals novel target genes for pest control and the proteasome as prime target. BMC Genomics 16, 674. https://doi.org/10.1186/s12864-015-1880-y (2015).
Will, T. & Vilcinskas, A. The structural sheath protein of aphids is required for phloem feeding. Insect Biochem. Mol. Biol. 57, 34–40. https://doi.org/10.1016/j.ibmb.2014.12.005 (2015).
Kirfel, P., Vilcinskas, A. & Skaljac, M. Lysine acetyltransferase p300/CBP plays an important role in reproduction, embryogenesis and longevity of the pea aphid Acyrthosiphon pisum. Insects 11, 265. https://doi.org/10.3390/insects11050265 (2020).
Mehlhorn, S., Hunnekuhl, V. S., Geibel, S., Nauen, R. & Bucher, G. Establishing RNAi for basic research and pest control and identification of the most efficient target genes for pest control: A brief guide. Front. Zool. 18, 60. https://doi.org/10.1186/s12983-021-00444-7 (2021).
Cedden, D. & Bucher, G. The quest for the best target genes for RNAi-mediated pest control. Insect Mol. Biol. 34, 505–517. https://doi.org/10.1111/imb.12966 (2025).
Fishilevich, E. et al. RNAi targeting of rootworm troponin I transcripts confers root protection in maize. Insect Biochem. Mol. Biol. 104, 20–29. https://doi.org/10.1016/j.ibmb.2018.09.006 (2019).
Knorr, E. et al. Gene silencing in tribolium castaneum as a tool for the targeted identification of candidate RNAi targets in crop pests. Sci. Rep. 8, 2061. https://doi.org/10.1038/s41598-018-20416-y (2018).
Knorr, E. et al. Knockdown of genes involved in transcription and splicing reveals novel RNAi targets for pest control. Front. Agron. 3, 715823. https://doi.org/10.3389/fagro.2021.715823 (2021).
Silver, K., Cooper, A. M. & Zhu, K. Y. Strategies for enhancing the efficiency of RNA interference in insects. Pest Manag. Sci. 77, 2645–2658. https://doi.org/10.1002/ps.6277 (2021).
Bachman, P., Fischer, J., Song, Z., Urbanczyk-Wochniak, E. & Watson, G. Environmental fate and dissipation of applied dsRNA in soil, aquatic systems, and plants. Front. Plant Sci. 11, 21. https://doi.org/10.3389/fpls.2020.00021 (2020).
Mishra, S. & Jurat-Fuentes, J. L. Resistance to RNAi: mechanism, current status, and prospects. In RNA interference in Agriculture: basic science to applications (eds Smagghe, G. et al.) 367–382 (Springer, 2025). https://doi.org/10.1007/978-3-031-81549-2_16.
Velez, A. M. et al. Chapter one: insecticidal proteins and RNAi in the control of insects. In Advances in Insect Physiology Vol. 65 (ed. Jurat-Fuentes, J. L.) 1–54 (Elsevier, 2023). https://doi.org/10.1016/bs.aiip.2023.09.0.
Xie, J. et al. Microfluidic-based dsRNA delivery nanoplatform for efficient Spodoptera exigua control. J. Agric. Food Chem. 72, 12508–12515. https://doi.org/10.1021/acs.jafc.4c03307 (2024).
Su, C. et al. Delivery of methoprene-tolerant dsRNA to improve RNAi efficiency by modified liposomes for pest control. ACS Appl. Mater. Interfaces 15, 13576–13588. https://doi.org/10.1021/acsami.2c20151 (2023).
Kim, I. G. et al. Efficient delivery of globotriaosylceramide synthase siRNA using polyhistidine-incorporated lipid nanoparticles. Macromol. Biosci. 23, e2200423. https://doi.org/10.1002/mabi.202200423 (2023).
Lin, Y.-H., Huang, J.-H., Liu, Y., Belles, X. & Lee, H.-J. Oral delivery of dsRNA lipoplexes to German cockroach protects dsRNA from degradation and induces RNAi response. Pest Manag. Sci. 73, 960–966. https://doi.org/10.1002/ps.4407 (2017).
Felton, G. W., Workman, J. & Duffey, S. S. Avoidance of antinutritive plant defense: Role of midgut pH in Colorado potato beetle. J. Chem. Ecol. 18, 571–583. https://doi.org/10.1007/BF00987820 (1992).
Holtof, M., Lenaerts, C., Cullen, D. & Vanden Broeck, J. Extracellular nutrient digestion and absorption in the insect gut. Cell Tissue Res. 377, 397–414. https://doi.org/10.1007/s00441-019-03031-9 (2019).
Sugihara, G. & Hisatomi, M. Enthalpy-entropy compensation phenomenon observed for different surfactants in aqueous solution. J. Colloid Sci. 219, 31–36. https://doi.org/10.1006/jcis.1999.6378 (1999).
Kulkarni, J. A. et al. On the formation and morphology of lipid nanoparticles containing ionizable cationic lipids and siRNA. ACS Nano 12, 4787–4795. https://doi.org/10.1021/acsnano.8b01516 (2018).
Haque, M. A., Shrestha, A., Mikelis, C. M. & Mattheolabakis, G. Comprehensive analysis of lipid nanoparticle formulation and preparation for RNA delivery. Int. J. Pharm. X 8, 100283. https://doi.org/10.1016/j.ijpx.2024.100283 (2024).
Maeki, M., Uno, S., Niwa, A., Okada, Y. & Tokeshi, M. Microfluidic technologies and devices for lipid nanoparticle-based RNA delivery. J. Control. Release. 344, 80–96. https://doi.org/10.1016/j.jconrel.2022.02.017 (2022).
Singh, I. K., Singh, S., Mogilicherla, K., Shukla, J. N. & Palli, S. R. Comparative analysis of double-stranded RNA degradation and processing in insects. Sci. Rep. 7, 17059. https://doi.org/10.1038/s41598-017-17134-2 (2017).
Quilez-Molina, A. I., Niño Sanchez, J. & Merino, D. The role of polymers in enabling RNAi-based technology for sustainable pest management. Nat. Commun. 15, 9158. https://doi.org/10.1038/s41467-024-53468-y (2024).
Pugsley, C. E., Isaac, R. E., Warren, N. J. & Cayre, O. J. Recent advances in engineered nanoparticles for RNAi-mediated crop protection against insect pests. Front. Agron. 3, 652981. https://doi.org/10.3389/fagro.2021.652981 (2021).
Christiaens, O., Whyard, S., Vélez, A. M. & Smagghe, G. Double-stranded RNA technology to control insect pests: Current status and challenges. Front. Plant Sci. 11, 451. https://doi.org/10.3389/fpls.2020.00451 (2020).
Zhang, Y. et al. Liposome mediated double-stranded RNA delivery to silence ribosomal protein P0 in the tick Rhipicephalus haemaphysaloides. Ticks Tick Borne Dis. 9, 638–644. https://doi.org/10.1016/j.ttbdis.2018.01.015 (2018).
Castellanos, N. L., Smagghe, G., Sharma, R., Oliveira, E. E. & Christiaens, O. Liposome encapsulation and EDTA formulation of dsRNA targeting essential genes increase oral RNAi-caused mortality in the neotropical stink bug Euschistus heros. Pest Manag. Sci. 75, 537–548. https://doi.org/10.1002/ps.5167 (2019).
Huang, J.-H., Liu, Y., Lin, Y.-H., Belles, X. & Lee, H.-J. Practical use of RNA interference: Oral delivery of double-stranded RNA in liposome carriers for cockroaches. J. Vis. Exp. 135, 57385. https://doi.org/10.3791/57385 (2018).
Brader, M. L. et al. Encapsulation state of messenger RNA inside lipid nanoparticles. Biophys. J. 120, 2766–2770. https://doi.org/10.1016/j.bpj.2021.03.012 (2021).
Wilson, K. A., Kung, R. W., D´souza, S. & Wetmore, S. D. Anatomy of noncovalent interactions between the nucleobases or ribose and π-containing amino acids in RNA–protein complexes. Nucleic Acids Res. 49, 2213–2225. https://doi.org/10.1093/nar/gkab008 (2021).
Zhao, F., Li, R., Liu, Y. & Chen, H. Perspectives on lecithin from egg yolk: Extraction, physicochemical properties, modification, and applications. Front. Nutr. 9, 1082671. https://doi.org/10.3389/fnut.2022.1082671 (2022).
Acknowledgements
We acknowledge financial support provided by the German Federal Ministry of Agriculture, Food and Regional Identity (BMLEH) and the Federal Agency of Agriculture and Food (BLE) within the collaborative Project ViVe_Beet. We thank our collaboration partners from the Julius Kühn Institute (JKI) and Institute of Sugar Beet Research (IFZ). We would like to especially thank Malvern Panalytical for supporting the ITC measurements. The authors thank Ricard M. Twyman for professional editing of the manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL. German Federal Ministry of Agriculture, Food and Regional Identity (BMLEH) and the Federal Agency of Agriculture and Food (BLE)
Author information
Authors and Affiliations
Contributions
P.G. and C.He. designed and evaluated the experiments, prepared all figures, and drafted the manuscript. E.K. supported the gel electrophoresis experiments and helped to design the figures and draft the manuscript. F.S. carried out the cryo-TEM experiments and helped to draft the manuscript. A. L. supported the particle size and ITC measurements and helped to design the figures and draft the manuscript. C.Ha. helped to optimize the scaled-up continuous flow microfluidic process. T.S., A.V. and C.He. coordinated this project and helped to draft the manuscript. All authors gave final approval for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare that some of the data presented in this work has been used in a patent application (EP24220250.5, priority date: 16/12/2025; PCT/EP2025/087404).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Geisler, P., Knorr, E., Steiniger, F. et al. Microfluidic process-property correlations of dsRNA lipid nanoparticle formulations. Sci Rep 16, 9653 (2026). https://doi.org/10.1038/s41598-026-44095-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-44095-2