Abstract
Dolichylphosphate mannose synthase (DPMS) performs an essential function by synthesizing the activated lipid-linked mannose intermediate used in protein glycosylation pathways. In eukaryotes and archaea, DPMS catalyzes the transfer of mannose from GDP-mannose to dolichylphosphate to generate dolichylphosphate mannose (Dol-P-Man). Type-III DPMS from Pyrococcus furiosus (PfDPMS) has a catalytic domain attached to a GtrA-like transmembrane (TM) domain with an unusual topology. Here, we present crystallographic data from a crystal complex determined from an enzymatic reaction mixture that provides detailed information about donor- and acceptor binding in the active site prior to mannosyl transfer. We also present a new, unexpected structural state for the TM domain in which a Dol-P-Man molecule is bound “upside-down” with its mannosylphosphate headgroup positioned in a polar pocket between the TM helices. By generating a panel of TM-domain mutants, we confirm that the TM domain does not participate directly in the catalysis of mannosyl transfer and discuss the possibility of this domain providing moonlighting function to PfDPMS by translocating the Dol-P-Man product to the cell exterior.
Similar content being viewed by others
Introduction
Glycolipids play vital roles in a wide spectrum of fundamental biological processes across all domains of life. Biogenic membranes, i.e., the self-synthesizing membranes of the endoplasmic reticulum in eukaryotes and the plasma membrane in prokaryotes, are the sites of biogenesis for nascent membranes and proteins. In eukaryotes and archaea, the multifunctional glycolipid dolichylphosphate mannose (Dol-P-Man) serves as the only source of mannose for several critical glycosylation processes during protein maturation1,2. In eukaryotes, enzymatic processes in the ER lumen that depend on access to Dol-P-Man include3: (i) production of the lipid-linked oligosaccharide4 required for the protein N-glycosylation pathway5; (ii) synthesis of the mannosyl core of GPI anchors; (iii) O-mannosylation of α-dystroglycan; and (iv) protein C-mannosylation6.
Dol-P-Man is produced by the integral membrane enzyme dolichylphosphate mannose synthase (DPMS; EC 2.4.1.83), which catalyzes the transfer of guanosine diphosphate α-D-mannose (GDP-Man) to dolichylphosphate (Dol-P) on the cytosolic face of the ER membrane in eukaryotes1, or at the inner face of the plasma membrane in archaea. After synthesis, eukaryotic Dol-P-Man is translocated across the ER membrane, from the cytosolic face to the lumen where it serves as activated mannosyl donor for additional glycosylation step. Due to the energetically unfavorable movement of the polar headgroups of lipids and glycolipids across membrane bilayers, membrane translocation requires assistance from proteins. For instance, the activated Dol-P-linked glycosyl donors, Dol-P-Man7,8 and Dol-P-Glc9,10, are assembled in the cytosol and translocated to the ER lumen by an unknown mechanism.
In archaea, Dol-P-Man is synthesized on the cytoplasmic face of the plasma membrane where the glycolipid may act as mannose donor for specific cytoplasmic steps of the N-glycan-biosynthetic pathway11,12,13. While synthesis of archaeal N-glycan intermediates appears to be mainly confined to the cytoplasmic face of the plasma membrane, evidence exists for occasional glycosyltransfer reactions taking place on the external face. In the case of the euryarchaeon Haloferax volcanii, the enzyme AglS has been found to transfer mannose from Dol-P-Man to an N-glycan intermediate located on the external cell surface to complete the pentasaccharide glycan14, which presupposes that Dol-P-Man had been translocated across the plasma membrane by a yet unidentified protein.
We previously reported the first crystal structure of a DPMS, wild-type DPMS from the euryarchaeon Pyrococcus furiosus15. We showed that PfDPMS represents a new class of DPMS (type-III DPMS) that consists of a catalytic domain attached to a transmembrane (TM) domain with four transmembrane helices (TMHs). Furthermore, wild-type PfDPMS was determined as three complexes: with bound GDP and Mg2+ (5MLZ), with bound GDP-Man and Mn2+ (5MM0), and with the products GDP and Dol55-P-Man representing a post-catalysis state (5MM1). The three complexes allowed a preliminary description of the structural states relevant to catalysis of mannosyl transfer.
In the present work, we present a new PfDPMS complex produced from a crystallized enzyme-reaction mixture incubated at 25 °C. The resulting electron density is convoluted but reveals two major structural states that we have not observed before. The first state that could be identified is the active site with bound donor (GDP-Man) and traces of the Dol-P acceptor. This state shows the mannosyl group productively oriented for mannosyl transfer (pre-transfer state) and thereby provides a missing piece to our previous, preliminary delineation of structural reaction states.
The second state shows a flipped Dol-P-Man molecule inside the TM domain and appears to be unrelated to any other structural state that we have observed so far. By producing a set of TM-domain mutants, we show that removing the TM domain, or blocking the Dol-P-Man-binding site inside the TM domain, does not affect the catalytic competence of the enzyme, which confirms that the TM domain in type-III DPMS has a role other than supporting mannosyl transfer by binding of Dol-P-Man. Based on these observations, we also discuss a possible Dol-P-Man-translocation role for the TM domain.
Results
A crystallized reaction mixture shows traces of two major structural states, including a flipped Dol-P-Man
In the present study we crystallized a reaction mixture incubated at 25 °C where the enzyme’s activity remains reasonably high (Supplementary Fig. 1). The most similar acceptor substrate available for the reaction was 55-carbon Dol-P, which is two isoprenoid units shorter than the 65-carbon Dol-P used by P. furiosus16. The resulting crystal structure was deeply convoluted with traces of several reaction states, but with two states dominating the electron density (Supplementary Fig. 2). The first state that could be unambiguously identified was the binding of donor substrate, GDP-Man, which was bound in a productive orientation with the mannosyl group rotated to expose the mannosyl C1 atom to the point of nucleophilic attack by a Dol-P phosphate group, i.e., the pre-mannosyl transfer state (Fig. 1a, b). For this state, we observed residual electron density for what is likely to be the phosphate group of a disordered Dol55-P molecule (Fig. 1a, b). The acceptor phosphate group is coordinated by the conserved side chains of Ser135 in IFH1 and Arg117 in β-strand 5.
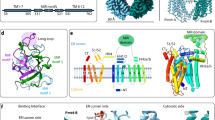
The second state of the convoluted electron density was unexpected. As noted earlier15, the four TMHs in PfDPMS adopt a unique and unusual arrangement with two pairs (or dimers) of TMHs, referred to as TMD1 (TMH1 and TMH2) and TMD2 (TMH3 and TMH4), oriented at a fixed angle of approximately 60° relative to each other (Fig. 1). The electron density most closely corresponded to a Dol-P-Man molecule oriented “upside-down” (flipped orientation) inside the TM domain, wedged between TMD1 and TMD2 with the mannosyl group bound in a polar pocket between TMD1 and TMD2.
An observation that was made in our original study but not ascribed any importance was the conformation of Phe338 in TMH4. In the previous structures, this side chain was found to have alternative conformations corresponding to a “flipped-up” and a “flipped-down” conformation (as in 5MLZ and 5MM0), or being only “flipped-down” (as in 5MM1). As realized in the present study, Phe338 assumes a strategic position at the TMD1/TMD2 interface (Fig. 1c, d) and is in the “flipped-up” position that allows access to the polar pocket and offers a binding platform for the isoprenoid chain of the flipped Dol-P-Man. A “flipped-down” Phe338 side chain would effectively block the path to the polar pocket.
Electron density for the ligands. At the left, overall ribbon drawing of PfDPMS with bound GDP-Man (donor) and Dol-P-Man (glycolipid product) as stick objects. The catalytic domain (GT2) is shown in gray. The two interface helices IFH1 and IFH2 are colored green. The TM dimer TMD1, including TMH1 and TMH2, is colored pink, and TMD2 in blue, including TMH3 and TMH4. The magnesium co-factor (dark purple) and the key side chains in the polar pocket are also shown. The upper left inset shows the GDP-Man (yellow) bound to the active site with the σA-weighted 2Fo-Fc map overlaid (a), and the same region overlaid by an annealing composite omit map, which shows more definition for Lys178 (b). The upper right inset shows the polar pocket with the mannosyl moiety of Dol-P-Man (orange) bound with either the 2Fo-Fc (c), or annealing composite omit map (d). In the view shown in (d), Asn277 that interacts with the mannosyl group is hidden behind the ligand. All maps were contoured at 1.0σ.
At the limited resolution of 2.9 Å, we only refined the B-factors of the ligands, not the occupancies. To evaluate the relative contribution of the two major ligand states we compared the B-factors, which are tightly correlated with occupancy, for the mannosyl moieties in GDP-Man (Fig. 1a) and Dol-P-Man (Fig. 1c). The isoprene chains are typically disordered even in structures determined at reasonably high resolution, making evaluation of the B-factors for this part of Dol-P-Man less useful for assessing the relative contribution to the electron density. The Wilson B-factor of the diffraction intensity data is an estimate of the global crystal disorder, which sets an overall baseline for the local atomic displacements in the model (atomic B-factors). The Wilson B-factor of the current crystal was 89.4 Å2, and the average atomic B-factors of the protein and ligands were 90.4 and 95.7 Å2, respectively (Table 1). The atomic B-factors for the mannosyl group in GDP-Man were ∼88–100 Å2, which is close to the average value for the protein (∼90 Å2). The atomic B-factors for the mannosyl of Dol-P-Man are slightly lower, 74–84 Å2, indicating a somewhat higher occupancy of this state. Based on the atomic B-factors, the relative occupancies of the two states are probably close to 45% for the GDP-Man state and 55% for the Dol-P-Man state.
Analysis of mechanistically relevant states of PfDPMS
Our previous PfDPMS structures included three complexes: GDP•Mg2+ (5MLZ); GDP-Man•Mn2+ (5MM0); and GDP•Dol-P-Man (5MM1). The latter structure, also produced from a crystallized reaction mixture, represented a post-transfer state with the two products, GDP and Dol-P-Man, bound at the active site. As discussed above, the new complex (GDP-Man•Dol-P/Dol-P-Man; 9EP0) contained nearly equal contribution from two reaction states: one that is relevant for mannosyl transfer, and an unrelated state with a flipped Dol-P-Man molecule inside the TM domain.
Combining the structural details observed in the active site of the three complexes GDP•Mg2+ (5MLZ), GDP-Man•Mn2+ (5MM1) and GDP-Man•Dol-P (9EP0) provides a detailed picture of the binding events that take place immediately before and after mannosyl transfer, including discrete substrate- and product-binding states (Fig. 2). In the GDP-Man•Mn2+ complex (5MM0; Fig. 2a), the diphosphate group is directly coordinated by the metal co-factor, and indirectly by the metal-coordinating residues Asp91 and Gln93. Lys178 and Asp89 interacts with the mannosyl group of the donor substrate. The loop between β6 and α5 (residues 163–178) containing Phe177 (“back door”) and the acceptor loop (A-loop, “front door”, comprising residues 202–210) are ordered in a “closed” conformation that closes the active site.
In the new PfDPMS complex (GDP-Man•Dol-P/Dol-P-Man; 9EP0; Fig. 2b), the β6/α5 loop and the acceptor loop have rearranged leading to an opening close to Phe177. The metal co-factor is bound as observed in the GDP-Man•Mn2+ complex. GDP-Man remains bound by Lys178 and Asp89 but has rotated to assume an orientation suitable for mannosyl transfer to the Dol-P phosphate group, and residual signal is also observed for a Dol-P molecule in the electron density. The data from these experimentally determined structures suggest that the state shown in Fig. 2b corresponds to the pre-transfer state.
Comparison of the active site in relevant reaction states. (a) GDP-Man•Mn2+ complex (pink; 5MM0); (b) GDP-Man•Dol-P complex (orange; 9EP0); and (c) GDP•Dol-P-Man complex (blue; 5MM1).
Following mannosyl transfer (GDP•Dol-P-Man complex; 5MM1; Fig. 2c), a post-transfer state is observed where the β6/α5 loop and the acceptor loop remain open to allow for GDP departure. The metal co-factor has been released, Lys178 does not make any interaction with either GDP or Dol-P-Man, and the diphosphate group in GDP has rotated towards the “back door” exit. Dol-P-Man is still bound in the active site, secured only by its phosphate group that is bound by Arg117, Arg131 and Ser135.
Analysis of the TM domain with a bound flipped Dol-P-Man molecule
We were first reluctant to assign the density as a flipped Dol-P-Man molecule, but upon closer inspection the probability of this observation being correct increased. As mentioned above, this state of the convoluted structure is quite well occupied. Furthermore, the packing of protein monomers in the crystal lattice is spacious around the TMHs where the Dol-P-Man is bound, allowing us to exclude that crystal contacts in this region are responsible for the flipped Dol-P-Man orientation.
The angle of approximately 60° between TMD1 and TMD2 is interesting (Fig. 1). We confirmed that the conformation of TMD1 and TMD2 and the angle between them are conserved in these enzymes by determining the structure of the related DPMS from Pyrococcus abyssi, which showed the identical TMD1/TMD2 orientation (unpublished data). All four TMHs would contribute to the Dol-P-Man binding, but three side chains can offer direct polar interactions with the mannosyl and phosphate groups in Dol-P-Man (Fig. 1c, d): Asn248 (TMH1) and Asn277 (TMH2) are both positioned to be able to interact with the mannosyl group while Arg341 (TMH4) can bind the phosphate group. While the earlier PfDPMS structures featured a few water molecules at the TMD1/TMD2 interface, this new structure shows that the same interface can bind large polar groups like mannose.
Site-directed mutagenesis of the TM domain
In our earlier study, we reported that a variant of PfDPMS lacking the TM domain, PfDPMS Δ230–352, retained full catalytic activity, here referred to as PF_Δ115. While this would seem to support a conclusion that the TM domain is dispensable for DPMS activity, removing the TM domain may also introduce compensatory effects that may rescue or increase turnover, e.g., increased acceptor access and/or elimination of rate-limiting acceptor-substrate binding or product release events.
To further evaluate the role of the TM domain for DPMS activity, we designed a new set of variant enzymes targeting the TM domain (Supplementary Table 1). Specifically, we produced: (1) a deletion variant retaining half of TMH1 referred to as PF_Δ2 (Δ242–352); and (2) PF_Δ3, which corresponds to PF_Δ2 with the additional single point mutations K236L and S238I; (3) a variant representing a TM-domain chimera of PfDPMS and Saccharomyces cerevisiae DPM1 (ScDPM1) referred to as PF_CHIM (replacing the TM domain residues 226–352 in PfDPMS by the single TMH present in ScDPM1); (4) a single point mutation replacing the conserved Asn277 in the polar pocket by methionine (PF_N277M); and (5) replacing Phe338 that gates the opening to the polar pocket by a glutamine side chain (PF_F338Q). To monitor stable membrane insertion by GFP fluorescence, all mutants lacking all or most of the TM domain were cloned and produced using the 2GFP-T vector while the single point mutants (N277M and F338Q) were cloned in the vector pNIC28-Bsa4 adding only a His6 tag.
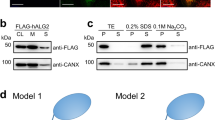
TM-domain variants target to the host membrane and show similar activity as the wild-type enzyme (Fig. 3), possibly except for PF_Δ2 that is slightly destabilized due to unfavorable membrane interactions by Lys236 and Ser238. This problem was solved by introducing the point mutations K236L and S238I to generate PF_Δ3. The result shows that DPMS activity is surprisingly resistant to manipulation of the TM domain, which further supports our proposal that the TM domain in PfDPMS is dispensable for Dol-P-Man synthesis and has a function beyond membrane tethering, acceptor binding and transition-state stabilization.
Activity of TM-domain variants. Comparison of enzymatic activity at 30 °C for wild-type and TM-domain variants of PfDPMS produced using two expression systems 2GFPT (prefix 2GFPT) and pNIC28-Bsa4 (prefix PN28). Descriptions of the individual variants are available in Supplementary Table 1. Wild-type PfDPMS was expressed in both vectors to allow direct comparison of activities across constructs. Data represent the mean ± SD from three independent experiments. Light gray bars correspond to the enzymatic reaction, dark gray bars are controls without acceptor substrate, and the white bars are controls without acceptor and enzyme.
Distribution of type-III DPMS
Unlike eukaryotic DPMS of type I or II, type-III DPMS, represented by the founding member PfDPMS, is a monomeric protein (Fig. 4). To define sequence patterns unique to the type-III DPMS, several functional sequence motifs were considered, including motifs of the active site, substrate-binding sites, and motifs related to the TM domain (see Methods). According to this rigorous definition, type-III DPMS only occurs in archaea, and only in a limited number of taxonomic orders.
Different known classes of DPMSs. Schematic representation of different types of DPMSs. Type-I is represented by Saccharomyces cerevisiae DPMS and is predicted to be DPM1-DPM2 heterodimer17,18. Type-II is a DPM1-DPM2-DPM3 heterotrimer represented by human DPMS19,20. Type-III is a monomer as observed for P. furiosus DPMS.
Type-III DPMS is most represented in the archaeal kingdom Methanobacteriati (former Euryarchaeota) where most examples belong to the order Thermococcales (Fig. 5, Supplementary Fig. 3 and Supplementary Table 2). Of the 49 sequences analyzed, only members of the genera Thermococcus, Pyrococcus and Palaeococcus show sequence identities > 70% to PfDPMS. Other orders of Methanobacteriati that contain relevant members, albeit with lower sequence identity to PfDPMS (40–50%), include Thermoplasmatales, Methanococcales, Methanosarcinales, and Candidatus Hadarchaeales.
Type-III DPMS sequences with 40–50% sequence identity to the Thermococcales group are also found in the kingdom Thermoproteati (phylum Thermoproteota, formerly known as Crenarchaeota). While most of the Thermoproteati members belong to the orders Desulfurococcales and Thermoproteales, isolated examples are also found in the orders Acidilobales and Sulfolobales, and in the provisional phyla Candidatus Bathyarchaeota and Candidatus Nezhaarchaeales. In addition to Methanobacteriati and Thermoproteati, possible DPMS type-III sequences appear in the kingdom Nanobdellati (syn. DPANN), specifically in the phylum Nanobdellota and the provisional phylum Candidatus Micrarchaeota.
Thermococcales type-III DPMS have an arginine (PfDPMS Arg341 in TMH4) positioned in the polar pocket that coordinates the phosphate group in the flipped Dol-P-Man molecule. This arginine is replaced by an asparagine side chain in all other sequences analyzed. Replacing arginine by asparagine in this position would increase the size of the polar pocket while retaining its polar character.
One of the asparagine side chains in the polar pocket that interact with the mannosyl headgroup of the flipped Dol-P-Man in PfDPMS (Asn248, TMH1) is conserved in 46 of 49 sequences (Supplementary Fig. 3). In the Thermoproteales members Pyrobaculum islandicum, Pyrobaculum neutrophilum and Thermoproteus thermophilus this asparagine is replaced by alanine, but this is compensated for by the presence of another polar residue (asparagine or threonine) in TMH3 on the opposite side of the polar pocket, which would restore the function. The phenylalanine side chain that gates the access to the polar pocket in PfDPMS (Phe338) is present in all sequences except for the species Candidatus Nanoclepta minutus (order Nanobdellales) and Pyrobaculum islandicum (order Thermoproteales) that instead have tyrosine in this position, which is expected to be able to perform the same function as Phe338.
Unrooted cladogram. Archaeal members with protein sequences that fulfill the definition of a type-III DPMS. The sequences are named starting with the taxonomic order followed by genus or species. For detailed sequence information, see Supplementary Fig. 3 and Supplementary Table 2.
We conclude that while relatively few archaeal members have type-III DPMS, those that fit the definition are all hyperthermophiles that grow at temperatures of 80–100 °C, or higher. We also note that there are hyperthermophilic archaea that do not have a DPMS that is sufficiently similar in sequence to PfDPMS to be unequivocally assigned to the type-III group but nonetheless may have been variations on the type-III theme, which further emphasizes the vast diversity in adaptation mechanisms for similar cellular processes in the archaeal domain of life.
Comparison with GtrA flippases
The TM domain of PfDPMS has distinct similarities with the family of small membrane transporters belonging to the GtrA family (InterPro IPR007267). The GtrA family includes small integral membrane proteins with 2 to 4 TMHs that participate in the glycosylation of cell surface glycoconjugates. Examples of GtrA members include (i) the founding member GtrA from the Shigella flexneri bacteriophage X (SfxGtrA; UniProt Q9T1D7), which translocates lipid-linked glucose across the cytoplasmic membrane and causes bacterial cell lysis21; (ii) the Escherichia coli ArnE/ArnF (previously PmrL/PmrM) proposed to act as a translocator of undecaprenylphosphate α-L-Ara4N across the inner membrane as part of the polymyxin resistance pathway22; (iii) the Mycobacterium tuberculosis protein Rv3789 (UniProt P9WMS9) proposed to translocate decaprenylphosphate arabinose to the periplasm23, although this hypothesis has been challenged24; and (iv) GtcA from Listeria monocytogenes serovar 1/a (UniProt P58412) and Bacillus subtilis strain 168 (UniProt P39602) proposed to translocate undecaprenylphosphate galactose for galactosylation of the cell-wall polymers wall teichoic acid (WTA) and lipoteichoic acid (LTA)25.
The sequence identity between GtrA and GtrA-like proteins is low, but they all display similar membrane topology (Fig. 6). The sequence motifs that define the polar pocket (Asn248, Asn277, Arg341) and the gating Phe338 in type-III DPMSs are absent in GtrA/GtrA-like TM domains, which suggests that a possible Dol-P-Man translocation mechanism in type-III DPMS must be different from that of GtrA/GtrA-like TM domains.
Alignment of selected GtrA proteins. Sequence alignment of GtrA from Shigella flexneri bacteriophage X (GTRA_BPSFX; Q9T1D7), Rv3789 from Mycobacterium tuberculosis (AGBR_MYCTU; P9WMS9), and the TM domain of PfDPMS (Q8U4M3_PYRFU). Global identities are in red filled boxes, local identities with light-red shaded background, residues with global similarity are enclosed by red frames, and residues that show group similarity are shown as red characters. Red bars below the sequences correspond to TM helices.
Discussion
The new PfDPMS crystal complex provides important information regarding proper positioning of donor and acceptor loop prior to mannosyl transfer and clarifies the involvement of the β6/α5 loop (back door) and acceptor (front door) in shaping the overall architecture of the active site to allow substrate binding and product departure. Based on the available structural states of the active site in PfDPMS, we can outline the key structural determinants for mannosyl transfer.
Step 1: Binding of donor and metal cofactor (Figs. 2a and 7: step 1). Prior to encounter with a Dol-P acceptor molecule, GDP-Man and metal ion are bound in the active site. Release of the donor and metal ion is prevented by the acceptor loop (pink bow), which is folded over the “front door” of the active site. Step 2: Binding of acceptor (Figs. 2b and 7: step 2). IFH2 is displaced in response to entry of acceptor, which promotes the β6/α5 loop and acceptor loop to undergo conformational changes to open the active site and the “back door”. Arg131 and Ser135 coordinate the Dol-P phosphate group. Computational calculations combined with NMR measurements have suggested that the isoprenoid chain of Dol-P is oriented at an angle of 75° with the membrane plane26, which would provide a pre-organized conformation of the dolichyl chain for docking with TMD2. Step 3: Nucleophilic attack and mannosyl transfer to produce GDP and Dol-P-Man (Figs. 2c and 7: step 3). GDP-Man orients to expose the mannosyl C1 atom to the Dol-P phosphate group. The Dol-P phosphate group is pre-activated for nucleophilic attack and attacks with one of its oxygen atoms at the α-mannosyl C1 and the new bond is formed with inverted configuration at C127. Following formation of products, GDP is released via the “back door” and the metal cofactor released through the “front door”.
Key steps of mannosyl transfer. Coloring scheme: Catalytic domain, blue; TMD1, red; TMD2, pink; IF helices, green; GDP, orange oval; Man, yellow oval; Dol-P, gray chain with green circle; metal ion, purple circle; Phe177, gray oval; polar pocket, light orange circle.
As previously observed and currently confirmed by mutants missing the TM domain, mannose transfer does not depend on TMD1 and TMD2. Whereas in the absence of the TM domain the mutant enzyme most likely releases the Dol-P-Man product into the same leaflet of the membrane. Our results indicate that the wild-type enzyme could facilitate the flipping of Dol-P-Man prior to its release. Whereas crystal packing is not responsible for the “flipped” Dol-P-Man, we cannot completely exclude that it might be due to the enzyme being in detergent and not in a more membrane-like environment. Nonetheless, the 9EP0 crystal structure clearly shows that the TM domain of type-III DPMS can accommodate a flipped Dol-P-Man and provide suitable interactions and spatial requirements for the mannosyl and phosphate moieties as well as part of the isoprenoid chain. Furthermore, the retained wild-type activity of the TM-domain mutants shows that the function of the TM domain is de-coupled from mannosyl transfer, suggesting a role of the TM domain unrelated to DPMS activity.
This raises the question of whether the function of the type-III TM domain is to translocate Dol-P-Man. We find it difficult to rationalize the experimental data without at least considering the possibility of the flipped Dol-P-Man being an authentic state relevant to a translocation mechanism. The unique TMD1/TMD2 arrangement provides a support for the isoprenoid chain of the Dol-P acceptor and of the newly synthesized unflipped Dol-P-Man (5MM1) (Fig. 8). The close spatial proximity between the active site and the path leading to the polar pocket would suggest a high probability that upon release from the active site, the Dol-P-Man mannosylphosphate moiety could flip down and interact inside the TM domain.
If this is the case, it would imply that type-III DPMS can moonlight as a Dol-P-Man synthase and translocator. Protein moonlighting refers to the ability of a single polypeptide chain to perform multiple, distinct biological functions28,29,30–31, which would make the monomeric type-III DPMS fit the description of a bona fide moonlighting protein. Importantly, moonlighting activity has been acknowledged as a problematic trait that can effectively impede the possibility to identify new biogenic flippases3. Examples of other moonlighting membrane proteins include the G-protein-coupled receptor opsin, which displays moonlighting characteristics in its ability to function as a constitutive, ATP-independent scramblase that translocates phospholipids bidirectionally across retinal disc membranes32–33. Another example is the lipid-biosynthetic enzyme MprF that synthesizes lysyl-phosphatidylglycerol using its C-terminal domain and translocates the product using the N-terminal domain34.
Energy-independent translocation of glycolipids across biogenic membranes is considered to be protein-assisted processes driven by facilitated diffusion. The driving force is the chemical energy gradient created by unequal concentrations of the glycolipid on the two sides of the membrane that drives translocation downhill the concentration gradient, from the cytoplasmic face where the glycolipid is synthesized to the other side (exterior side in the case of archaea) where it is consumed. The assistance provided by the protein lowers the energy barrier enough for the polar headgroup to traverse the hydrophobic interior of the membrane. In the case of type-III DPMS, the polar pocket would be suitable to provide such assistance.
Comparison of Dol-P-Man bound in unflipped and flipped orientation. Superimposition of the PfDPMS complex with unflipped Dol-P-Man (blue; GDP•Dol-P-Man complex; 5MM1) and the new complex with flipped Dol-P-Man (orange; GDP-Man•Dol-P/Dol-P-Man complex; 9EP0). For clarity, the structures are shown as ribbon representations with only the ligands (GDP, GDP-Man and Dol-P-Man) and the gating Phe338 drawn as stick objects.
The membrane proteins responsible for translocating lipid-linked sugar intermediates into the ER lumen without the use of ATP have consistently eluded identification and/or functional confirmation. The reasons are manifold, but a general problem is designing in-vitro translocation assays with topological labeling that is sufficiently sensitive. As an example, RFT1 was proposed to be responsible for translocation of Dol-PP-Man5GlcNAc2 (M5-DLO) already 25 years ago35, but later studies failed to conclusively prove or disprove the hypothesis. In 2024, Chen and co-workers designed an RFT1-specific M5-DLO-translocation in-vitro assay in proteoliposomes (PRLs) that offered support for the M5-DLO-translocating capacity of Rft136. This elegant assay employed specific α-mannosidases to trim the Man5 headgroup of M5-DLO to determine whether the headgroup was exposed (outward-facing in PRLs) or buried (inward-facing).
Dol-P-Man translocation is considered a general requirement in all organisms that use this activated Dol-P intermediate. As for RFT1, the search for the eukaryotic protein responsible for Dol-P-Man translocation has been ongoing for decades, and candidates for this function have included DPMS itself as well as other proteins. Dol-P-Man translocation activity was first proposed for yeast DPMS37–38, but later studies failed to confirm or disprove this claim39, leaving the matter unresolved. In 2009, new evidence of Dol-P-Man translocation activity was reported40. Even though the responsible protein was not identified, it was confirmed that the translocation is energy-independent, protein-assisted, and distinct from the protein translocating M5-DLO into the ER lumen (RFT1).
For yeast, it has also been established that the catalytic domain of yeast DPMS (DPM1) is not responsible of Dol-P-Man translocation3941,. In light of this, the possibility of Dol-P-Man translocation by type-III DPMS would seem to present an exciting opportunity to solve this longstanding question, however, based on the limited sequence similarity between the eukaryotic type-I/II (DPM2 and DPM3 subunits) and type-III TM domains, we expect the eukaryotic TM domains to be structurally (and probably functionally) distinct, leaving the question unresolved.
Earlier experiments have shown that soluble analogs of Dol-P-Man can be scrambled across mammalian microsomal membranes42, and that Dol-P-Man can be scrambled in proteoliposomes reconstituted with mammalian ER membrane by an energy-independent and protein-assisted process40. These findings show that in the model systems used, the protein responsible for Dol-P-Man translocation in eukaryotes can translocate Dol-P-Man in both directions, and similar scrambling activity has been proposed for RFT14344.
A mechanism of facilitated diffusion of a metabolic intermediate like Dol-P-Man would be expected to be evolutionarily optimized towards translocation from the cytoplasmic side to the exterior (ER lumen for eukaryotes) where mannosyltransferases continuously consume the intermediate. This would leave little unconsumed Dol-P-Man at the exterior/lumenal face to be back-translocated, making the process appear unidirectional under normal conditions. Accumulation of Dol-P-Man on the outer/lumenal face may however happen because of malfunctioning regulatory control of DPMS activity. Even if scrambling is observed in different membrane systems in vitro, it is uncertain if back-translocation of the Dol-P-Man intermediate is a significant process during normal operation in vivo. Regardless, the observation of Dol-P-Man scrambling in eukaryotic systems raises the question of whether PfDPMS would be able to translocate Dol-P-Man in both directions using the same structural determinants?
The eukaryotic TMEM16 scramblases feature a large hydrophilic membrane-exposed groove that spans the membrane and opens up at the membrane surface, and this groove has been proposed to (1) provide a path for the lipid headgroup during translocation45, and (2) to induce membrane thinning at the groove’s opening of up to 30% to further facilitate translocation46–47. Opening and closure of the groove is regulated differently in different TMEM16 scramblases, and different translocation mechanisms have been proposed to be possible depending on the lipid, including translocation through the open groove, as well as on the outside (“out-of-the-groove” transport), facilitated by membrane thinning47. TMEM16 scramblases translocate phospholipids in both directions but are, unlike the protein responsible for Dol-P-Man translocation, relatively unspecific with respect to lipid48, ranging from phospholipids with small (e.g.., phosphatidylserine etc49.) to very large headgroups (e.g., PEGylated head groups50). Based on recent theoretical modeling, RFT1 is predicted to sequester M5-DLO headgroup within a hydrophilic central cavity while the lipid tail interacts with a hydrophobic groove, making the groove distinct from that of TMEM16-type scramblases44.
Although structurally unrelated to TMEM16 scramblases and RFT1, the unique splayed-out structure of TMD1 and TMD2 in PfDPMS also creates a groove that opens at one face of the membrane, however, the groove has a mixed hydrophobicity/hydrophilicity profile where the mannosyl group can be guided by hydrophilic interactions along the interior face of TMD1 and the isoprenoid chain by hydrophobic interactions provided by mainly by TMH4 in TMD2. The fixed relative position of TMD1 and TMD2 would preclude a mechanism involving large conformational changes such as the alternating-access transport mechanism suggested for RFT1 based on theoretical modeling44. The presence of a groove may however indicate that protein-mediated membrane thinning could be required in Dol-P-Man translocation by type-III DPMS. The optimal growth temperature of P. furiosus is 100 °C and its membrane contains a mixture of diether and tetraether lipids51 that provides a highly rigid and stable cytoplasmic membrane. Thus, any discussion on the possibility of induced membrane thinning as part of a Dol-P-Man mechanism would need to take into account the rigid nature of hyperthermophilic archaeal membranes, however not much is known about membrane remodeling in these archaea.
Conclusions
We have provided detailed information about the structural determinants of the active site that enable mannosyl transfer in archaeal type-III DPMS. Our phylogenetic analysis shows that this type of DPMS is a trait of hyperthermophilic archaea, and especially common in the order Thermococcales of the archaeal kingdom Methanobacteriati. Based on the experimentally determined structure of the PfDPMS complex with a flipped Dol-P-Man molecule bound, we hypothesize that the TM domain in type-III DPMS moonlights as a Dol-P-Man translocator. This unique TM structure features a hydrophilic groove that opens at the exterior face of the membrane and a polar pocket that is suitably designed to guide and shield the mannosylphosphate headgroup during translocation. The high temperatures at which these organisms live require highly specialized adaptations of both the membrane composition and the membrane proteins. A chimeric structure including both Dol-P-Man synthase and translocator would serve to increase the probability of successful translocation by bringing the nascent Dol-P-Man near the translocation machinery and reduce the risk of thermal degradation of the metabolic intermediate before being delivered to the mannosyltransferases operating at the cell surface. Furthermore, consumption of the activated mannosyl intermediate by external mannosyltransferases must be equally efficient to safeguard S-layer-protein glycosylation and cell survival.
Methods
Protein production
The production of full-length PfDPMS for crystallization was performed as described previously15. Briefly, the gene coding for PfDPMS in the expression vector pNIC28-Bsa4 was expressed in Escherichia coli strain C41 (DE3) and extracted from the membrane using 25 mM K2HPO4 (pH 7.2), 150 mM NaCl, and 5% (v/v) glycerol supplemented with 1% n-dodecyl-β-D-maltoside (DDM, Anatrace), followed by centrifugation at 30,000 r.p.m. (104,600 g) using a 45 Ti rotor mounted in a Beckman Coulter Optima ultracentrifuge. The resulting supernatant was loaded onto a Ni-NTA agarose column (Invitrogen), and the detergent exchanged to 0.05% N,N-dimethyldodecylamine N-oxide (LDAO, Anatrace) in 25 mM K2HPO4 (pH 7.2), 150 mM NaCl, 30–50 mM imidazole, 5–10% (v/v) glycerol during the wash step. The same buffer but with 500 mM imidazole was used to elute the bound protein fraction, followed by size-exclusion chromatography (SEC) on a HiLoad™ 16/60 Superdex™ 200 prep grade column (GE Healthcare Life Sciences). The SEC buffer contained 50 mM 4-(2-hydroxyethyl)−1-piperazineethanesulfonic acid (Hepes) pH 7.5, 150 mM NaCl, 5–10% (v/v) glycerol, and 0.05% LDAO. Recovered PfDPMS was further concentrated using a Vivaspin20® concentrator (MWCO 50 or 100 kDa) to a final concentration of 15–20 mg/ml. For activity assays, PfDPMS was expressed in Escherichia coli C43 (DE3) using a pNIC28-Bsa4 vector carrying the full-length PfDPMS gene. Proteins were extracted and centrifuged as above, but using 1% 3-dodecylamido-N, N’-dimethylpropyl amine oxide (LAPAO, Anatrace) instead of 1% DDM. The supernatant was loaded onto Ni-NTA agarose beads and washed first with a buffer containing 25 mM potassium phosphate (pH 7.2), 150 mM NaCl, 30 mM imidazole, 5% glycerol, 0.14% LAPAO and then with a buffer containing 25 mM potassium phosphate (pH 7.2), 300 mM NaCl, 50 mM imidazole, 5% glycerol, 0.1% LAPAO. Proteins were eluted in 25 mM potassium phosphate (pH 7.2), 150 mM NaCl, 500 mM imidazole, 5% glycerol, 0.07% LAPAO. Ethylenediaminetetraacetic acid (EDTA) and ethylene glycol-bis(β-aminoethyl ether)-N,N,N’,N’-tetraacetic acid (EGTA) were added to the eluate to a final concentration of 10 mM each and the mixture was incubated for 1 h at room temperature. Proteins were then fractionated on a HiLoad™ 16/60 Superdex™ 200 prep grade column in 50 mM Hepes pH 7.5, 150 mM NaCl, 5% glycerol, and 0.07% LAPAO.
Glycosyltransferase batch reaction for crystallization
A reaction mixture was prepared by mixing 15.6 mg/ml PfDPMS in 50 mM Hepes (pH 7.5), 150 mM NaCl, 10% (v/v) glycerol, 0.05% LDAO with 5 mM GDP-Man, 5 mM Mg2+ and 0.8 mM Dol55-P and incubated for 40 min at 25 °C. Following incubation, the reaction mixture was set up for crystallization using the sitting drop vapor diffusion method and mixed with a reservoir solution containing 0.2 M calcium chloride dehydrate, 0.1 M Hepes pH 7.5 and 53% (v/v) PEG 400.
Site-directed mutagenesis
All mutagenesis experiments were performed using the codon-optimized pfdpms gene previously described by Gandini et al. (2017). Two cloning strategies were employed depending on the type of modification. For transmembrane (TM) domain manipulation, the 2GFP-T vector was used, whereas point mutations (N277M and F338Q) were introduced into the pNIC28-Bsa4 vector. Gene-specific primers were designed to incorporate the desired mutations (listed in Supplementary Tables 2 and 3). The wild-type pfdpms sequence in pNIC28 served as the template for generating the single mutants N277M and F338Q.
To generate the PF_CHIM construct, an E. coli codon-optimized gene encoding Saccharomyces cerevisiae dolichol phosphate mannose synthase (ScDpm1, UniProt P14020) was synthesized by Integrated DNA Technologies (https://www.idtdna.com). The GeneArt™ Seamless Cloning Kit (Thermo Fisher Scientific) was used to replace the region corresponding to amino acids 226–240 in PF_Δ2 with the corresponding 226–257 segment from ScDpm1. For construction of the 2GFPT-PF_CHIM variant, the plasmid 2GFPT-PF_Δ2 served as the template, and cloning was carried out using a two-step seamless cloning strategy. In the first step, the region corresponding to amino acids 226–240 in PF_Δ2 was deleted by PCR amplification using the following primers:
Pf-Sc_fwd: 5′-TGGTATTCTAATAACATTGGAAGTGGATAA-3′.
Pf-Sc_rev: 5′-TTGAATACCATCAGACGGTAAATGTG-3′.
In the second step, the segment corresponding to amino acids 226–257 from ScDpm1 was amplified from a synthesized ScDpm1 gene using the following primers:
Pf-Sc_TM_fwd: 5′-GTCTGATGGTATTCAAGTTTGGAGCAA-3′.
Pf-Sc_TM_rev: 5′-TGTTATTAGAATACCAGGTGATAAAGC-3′.
The resulting PCR fragments were assembled using the GeneArt™ Seamless Cloning Kit, to generate the chimeric 2GFPT-PF_CHIM construct. All final plasmid constructs were confirmed by Sanger sequencing and transformed into E. coli C41 (DE3) cells for recombinant protein expression and purification, following the procedure described above, except that the EDTA and EGTA treatment steps were omitted during purification.
Activity assay using GDP-Glo™ glycosyltransferase assay
Dol-P used for assays was solubilized in a 2:1 (v/v) chloroform: methanol (CHCl₃:MeOH) mixture in a small glass vial containing a magnetic stir bar was dried under a gentle N2 stream. Once the organic solvent had evaporated, a 1:1 ratio of SEC buffer and 2% LAPAO dissolved in H2O were added in amounts to yield the desired concentration. The glass vial was wrapped in aluminum foil, and the solution was stirred on a magnetic stirred and sonicated in a bath sonicator, taking turns, until the solution was clear. All steps were performed at ambient temperature, and the solubilized Dol-P solution was stored at − 20 °C until use.
Enzyme activity of wild type and mutant PfDPMS constructs was measured using GDP-Glo™ Glycosyltransferase Assay (Promega). Reactions were set up in a 40 µl final volume of 50 mM Hepes, 150 mM NaCl, 5% glycerol, 0.07% LAPAO with 0.01 nmol PfDPMS and 0.1 nmol acceptor lipid (either Dol55-P, Dol95-P, or undecaprenyl phosphate (UndP), all from Larodan) added. For reactions in bicelles, PfDPMS in 5% bicelles (MemMagic® Bicelle Screen Kit, MemX Biosciences LLC) were prepared according to manufacturer’s instructions before setting up the reaction. After 30 min at room temperature, 1 nmol GDP-Man and 5 nmol MnCl2 were added. For control reactions, either the acceptor or both the acceptor and PfDPMS were replaced with buffer. Reactions were incubated at either 60 °C, 30 °C–25 °C for 3 h. Reaction products were then assayed as per manufacturer’s instructions, with minor modifications. A GDP standard curve was prepared by serially diluting a 10 mM GDP stock to generate concentrations of 10, 5, 2.5, 1.25, 0.625, 0.3125, and 0 µM in assay buffer. 25 µl of each dilution was transferred to a solid white 96-well plate for luminescence detection. The GDP-Glo working solution was prepared by adding 1 µl GDP-Glo enzyme to 74 µl enzyme dilution buffer. The GDP detection reagent was made by mixing 1/100 volume of the working solution with nucleotide detection reagent. Following completion of the glycosyltransferase reaction, 25 µl of the reaction mixture was added to each well, followed by 25 µl of GDP detection reagent. Plates were mixed briefly, incubated at room temperature in dark for 1 h, and luminescence was recorded using a microplate reader. All reactions were performed in triplicate, and enzyme activity was calculated based on the GDP standard curve.
Crystallographic analysis
Synchrotron data were collected on a rod-shaped crystal at beamline I03 (Diamond Light Source, UK) and processed and scaled using the XDS software package52 at 2.9 Å resolution. The crystal belonged to space group C2221 with unit cell dimensions of a = 88.18 Å, b = 145.73 Å, c = 99.43 Å, α = β = γ = 90°, and one molecule in the asymmetric unit (Table 1). Phasing was performed using molecular replacement in Phenix53 using the wild-type PfDPMS model 5MM115 as template. To remove model bias in the electron density, composite omit maps applying simulated annealing, including Cartesian molecular dynamics with a starting temperature of 5000 K, were calculated using Phenix53. The resulting map is shown in Fig. 2b, d and Supplementary Fig. 2. The composite omit map is of better quality than the normal 2Fo-Fc map, which is due to the iterative process and the high-temperature annealing step that effectively removes model bias and increases phase accuracy. Model adjustment was done with Coot54, and refinement continued with Phenix53 (Table 1). The refinement protocol included reciprocal-space refinement of x, y,z coordinates and initial group B-factor refinement. Individual B-factors were refined in the last two refinement rounds.
Sequence analyses
To analyze the distribution of type-III DPMS, we first defined the functional sequence motifs unique for this type of DPMS. Specifically, we define a type-III DPMS as containing (1) 89DADxQHPP96 (where x is Leu, Phe, or Tyr) defining the structural determinants required for binding of the metal co-factor and the donor’s diphosphate group15; (2) Arg117 (motif 116SRY118), Arg131, Ser135 and Lys178 (178KIL180) that coordinate the phosphate group in Dol-P and Dol-P-Man (not flipped); (3) Asn248 and Asn277 that interact with the mannosyl group in the polar pocket and Phe338 that gates the path to the polar pocket and serves as a binding platform for the isoprenoid chain in flipped but not in unflipped Dol-P-Man. Initial alignment was performed using Clustal Omega v.1.2.45556, followed by manual adjustment guided by known structural information. The adjusted alignment was then used to generate a phylogenetic tree using Clustal Omega. The sequences of GtrA from Shigella flexneri bacteriophage X (GTRA_BPSFX; Q9T1D7), Rv3789 from Mycobacterium tuberculosis (AGBR_MYCTU; P9WMS9), and the TM domain of PfDPMS (Q8U4M3_PYRFU) were aligned using TM-Aligner (https://lms.snu.edu.in/TM-Aligner/index.php57).
Visualization
Figures 2, 3 and 4 and S1 were prepared using PyMOL (The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC). The sequence alignments in Figs. 8 and S2 were visualized using ESPript 3.x/ENDscript 2.x (https://espript.ibcp.fr/ESPript/cgi-bin/ESPript.cgi58). The phylogenetic tree generated by Clustal Omega was visualized as an unrooted circular cladogram using iTOL v. 759.
Data availability
The coordinates and structure factors have been deposited in the Protein Data Bank (www.rcsb.org) with the accession number 9EP0.
References
Hirschberg, C. B. & Snider, M. D. Topography of glycosylation in the rough endoplasmic reticulum and Golgi apparatus. Annu. Rev. Biochem. 56, 63–87. https://doi.org/10.1146/annurev.bi.56.070187.000431 (1987).
Sanyal, S. & Menon, A. K. Flipping lipids: Why an’ What’s the reason for?. ACS Chem. Biol. 4, 895–909. https://doi.org/10.1021/cb900163d (2009).
Rush, J. S. Role of flippases in protein glycosylation in the endoplasmic reticulum. Lipid Insights 8, 45–53. https://doi.org/10.4137/LPI.S31784 (2016).
Burda, P. & Aebi, M. The dolichol pathway of N-linked glycosylation. Biochim. Biophys. Acta 1426, 239–257. https://doi.org/10.1016/S0304-4165(98)00127-5 (1999).
Lommel, M. & Strahl, S. Protein O-mannosylation: Conserved from bacteria to humans. Glycobiology 19, 816–828. https://doi.org/10.1093/glycob/cwp066 (2009).
Doucey, M.-A., Hess, D., Cacan, R. & Hofsteenge, J. Protein C-mannosylation is enzyme-catalysed and uses dolichyl-phosphate-mannose as a precursor. Mol. Cell. Biol. 9, 291–300. https://doi.org/10.1091/mbc.9.2.291 (1998).
Rush, J. S. & Waechter, C. J. Functional reconstitution into proteoliposomes and partial purification of a rat liver ER transport system for a water-soluble analogue of mannosylphosphoryldolichol. Biochemistry 43, 7643–7652. https://doi.org/10.1021/bi036083o (2004).
Anand, M. et al. Requirement of the Lec35 gene for all known classes of monosaccharide-P-dolichol-dependent glycosyltransferase reactions in mammals. Mol. Biol. Cell 12, 487–501. https://doi.org/10.1091/mbc.12.2.487 (2001).
Rush, J. S., van Leyen, K., Ouerfelli, O., Wolucka, B. & Waechter, C. J. Transbilayer movement of Glc-P-dolichol and its function as a glucosyl donor: Protein-mediated transport of a water-soluble analog into sealed ER vesicles from pig brain. Glycobiology 8, 1195–1205 (1998).
Skrabanek, L. A. & Menon, A. K. Phylogenetic profiling identifies glucosyl-phosphoryl dolichol scramblase candidates. bioRxiv (2017). https://doi.org/10.1101/209106
Zhu, B. C. & Laine, R. A. Dolichyl-phosphomannose synthase from the archae Thermoplasma acidophilum. Glycobiology 6, 811–816. https://doi.org/10.1093/glycob/6.8.811 (1996).
Urushibata, Y., Ebisu, S. & Matsui, I. A thermostable dolichol phosphoryl mannose synthase responsible for glycoconjugate synthesis of the hyperthermophilic archaeon Pyrococcus horikoshii. Extremophiles 12, 665–676. https://doi.org/10.1007/s00792-008-0173-7 (2008).
Eichler, J. & Imperiali, B. Stereochemical divergence of polyprenol phosphate glycosyltransferases. Trends Biochem. Sci. 43, 10–17. https://doi.org/10.1016/j.tibs.2017.10.008 (2019).
Cohen-Rosenzweig, C., Yurist-Doutsch, S. & Eichler, J. AglS, a novel component of the Haloferax volcanii N-glycosylation pathway, is a dolichol phosphate-mannose mannosyltransferase. J. Bacteriol. 194, 6909–6916. https://doi.org/10.1128/JB.01716-12 (2012).
Gandini, R., Reichenbach, T., Tan, T. C. & Divne, C. Structural basis for dolichylphosphate mannose biosynthesis. Nat. Commun. 8, 120. https://doi.org/10.1038/s41467-017-00187-2 (2017).
Chang, M. -, Imperiali, B., Eichler, J. & Guan, Z. N-linked glycans are assembled on highly reduced dolichol phosphate carriers in the hyperthermophilic archaea Pyrococcus furiosus. PLoS One 10, e0130482. https://doi.org/10.1371/journal.pone.0130482 (2015).
Orlean, P., Albright, C. & Robbins, P. W. Cloning and sequencing of the DPM1 gene encoding dolichol-phosphate mannose synthase from Saccharomyces cerevisiae. J. Biol. Chem. 263, 17499–17507. https://doi.org/10.1016/s0021-9258(19)77863-6 (1988).
Piłsyk, S. et al. Yil102c-A is a functional homologue of the DPMII subunit of dolichyl phosphate mannose synthase in Saccharomyces cerevisiae. Int. J. Mol. Sci. 21, 8938. https://doi.org/10.3390/ijms21238938 (2020).
Maeda, Y., Tanaka, S., Hino, J., Kangawa, K. & Kinoshita, T. Human dolichol-phosphate-mannose synthase consists of three subunits, DPM1, DPM2 and DPM3. EMBO J. 19, 2475–2482. https://doi.org/10.1093/emboj/19.11.2475 (2000).
Ashida, H., Maeda, Y. & Kinoshita, T. DPM1, the catalytic subunit of dolichol-phosphate mannose synthase, is tethered to and stabilized on the endoplasmic reticulum membrane by DPM3. J. Biol. Chem. 281, 896–904. https://doi.org/10.1074/jbc.M511311200 (2006).
Guan, S., Bastin, D. A. & Verma, N. K. Functional analysis of the O antigen glucosylation gene cluster of Shigella flexneri bacteriophage SfX. Microbiology 145, 1263–1273. https://doi.org/10.1099/13500872-145-5-1263 (1999).
Yan, A., Guan, Z. & Raetz, C. R. H. An undecaprenyl phosphate-aminoarabinose flippase required for polymyxin resistance in Escherichia coli. J. Biol. Chem. 282, 36077–36089. https://doi.org/10.1074/jbc.M706172200 (2007).
Larrouy-Maumus, G. et al. A small multidrug resistance-like transporter involved in the arabinosylation of arabinogalactan and lipoarabinomannan in mycobacteria. J. Biol. Chem. 287, 39933–39941. https://doi.org/10.1074/jbc.M112.400986 (2012).
Kolly, G. S. et al. GtrA protein Rv3789 is required for arabinosylation of arabinogalactan in Mycobacterium tuberculosis. J. Bacteriol. 17, 3686–3697. https://doi.org/10.1128/JB.00628-15 (2015).
Rismondo, J., Haddad, T. F. M., Shen, Y., Loessner, M. J. & Gründling, A. GtcA is required for LTA glycosylation in Listeria monocytogenes serovar 1/2a and Bacillus subtilis. Cell Surf. 6, 100038. https://doi.org/10.1016/j.tcsw.2020.100038 (2020).
Zhou, G. P. & Troy, I. I. NMR study of the preferred membrane orientation of polyisoprenols (dolichol) and the impact of their complex with polyisoprenyl recognition sequence peptides on membrane structure. Glycobiology 15, 347–359. https://doi.org/10.1093/glycob/cwi016 (2005).
Herscovics, A., Warren, C. D. & Jeanloz, R. W. Anomeric configuration of the dolichyl-D-mannosyl phosphate formed in calf pancreas microsomes. J. Biol. Chem. 250, 8079–8084. https://doi.org/10.1016/S0021-9258(19)40818-1 (1975).
Campbell, R. M. & Scanes, C. G. Endocrine peptides “moonlighting” as immune modulators: Roles for somatostatin and GH-releasing factor. J. Endocrinol. 147, 383–396. https://doi.org/10.1677/joe.0.1470383 (1995).
Jeffery, C. J. Moonlighting proteins: Old proteins learning new tricks. Trends Genet. 19, 415–417. https://doi.org/10.1016/S0168-9525(03)00167-7 (2003).
Jeffery, C. J. Multifunctional proteins: Examples of gene sharing. Ann. Med. 35, 28–35. https://doi.org/10.1080/07853890310004101 (2003).
Singh, N. & Bhalla, N. Moonlighting proteins. Annu. Rev. Genet. 54, 265–285. https://doi.org/10.1146/annurev-genet-030620-102906 (2020).
Menon, I. et al. Opsin is a phospholipid flippase. Curr. Biol. 21, 149–153. https://doi.org/10.1016/j.cub.2010.12.031 (2011).
Goren, M. A. et al. Constitutive phospholipid scramblase activity of a G protein-coupled receptor. Nat. Commun. 5, 5115. https://doi.org/10.1038/ncomms6115 (2015).
Slavetinsky, C. J. et al. Sensitizing Staphylococcus aureus to antibacterial agents by decoding and blocking the lipid flippase MprF. eLife 11, e66376. https://doi.org/10.7554/eLife.66376 (2022).
Helenius, J. et al. Translocation of lipid-linked oligosaccharides across the ER membrane requires Rft1 protein. Nature (447), 415–450. https://doi.org/10.1038/415447a (2002).
Chen, S. et al. Rft1 catalyzes lipid-linked oligosaccharide translocation across the ER membrane. Nat. Commun. 15, 5157. https://doi.org/10.1038/s41467-024-48999-3 (2024).
Haselbeck, A. & Tanner, W. Dolichyl phosphate-mediated mannosyl transfer through liposomal membranes. Proc. Natl. Acad. Sci. USA. 79, 1520–1524 (1982). https://doi.org/10.1073/pnas.79.5.1520
Haselbeck, A. & Tanner, W. Further evidence for dolichyl phosphate-mediated glycosyl translocation through membranes. FEMS Microbiol. Lett. 21, 305–308. https://doi.org/10.1111/j.1574-6968.1984.tb00327.x (1984).
Schutzbach, J. S. & Zimmerman, J. W. Yeast dolichyl-phosphomannose synthase: Reconstitution of enzyme activity with phospholipids. Biochem. Cell Biol. 70, 460–465. https://doi.org/10.1139/o92-070 (1992).
Sanyal, S. & Menon, A. K. Stereoselective transbilayer translocation of mannosyl phosphoryl dolichol by an endoplasmic reticulum flippase. Proc. Natl. Acad. Sci. USA 107, 11289–11294 (2010). https://doi.org/10.1073/pnas.1002408107
Zimmerman, J. W. & Robbins, P. W. The hydrophobic domain of dolichyl-phosphate-mannose synthase is not essential for enzyme activity or growth in Saccharomyces cerevisiae. J. Biol. Chem. 268, 16764–16753 (1993).
Rush, J. S. & Waechter, C. J. Transmembrane movement of a water-soluble analogue of mannosylphosphoryldolichol is mediated by an endoplasmic reticulum protein. J. Cell Biol. 130, 529–536. https://doi.org/10.1083/jcb.130.3.529 (1995).
Hirata, E. et al. J. Biol. Chem. 300, 107584 https://doi.org/10.1016/j.jbc.2024.107584 (2024).
Chiduza, G. N., Sakata, K., Noor, F. & Menon, A. K. Proposed mechanism for Rft1-mediated scrambling of a dolichol-linked oligosaccharide. bioRxiv preprint https://doi.org/10.64898/2025.12.07.692794
Kalienkova, V., Clerico Mosina, V. & Paulino, C. The groovy TMEM16 family: Molecular mechanisms of lipid scrambling and ion conduction. J. Mol. Biol. 433, 166941. https://doi.org/10.1016/j.jmb.2021.166941 (2021).
Feng, Z. et al. Cell. Calcium 121, 102896 https://doi.org/10.1016/j.ceca.2024.102896 (2024).
Feng, Z., Alvarenga, O. E. & Accardi, A. Structural basis of closed groove scrambling by a TMEM16 protein. Nat. Struct. Mol. Biol. 31, 1468–1481. https://doi.org/10.1038/s41594-024-01284-9 (2024).
Ballesteros, A. & Swartz, K. J. Lipids surf the groove in scramblases. Proc. Natl. Acad. Sci. USA 115, 7648–7650 (2018). https://doi.org/10.1073/pnas.1809472115
Suzuki, J. et al. Calcium-dependent phospholipid scramblase activity of TMEM16 protein family members. J. Biol. Chem. 288, 13305–13316. https://doi.org/10.1074/jbc.M113.457937 (2013).
Malvezzi, M., Andra, K. K., Pandey, K. & Accardi, A. Out-of-the-groove transport of lipids by TMEM16 and GPCR scramblases. Proc. Natl. Acad. Sci. USA 115, E7033–E7042 (2018).
Lobasso, S. et al. Coupled TLC and MALDI-TOF/MS analyses of the lipid extract of the hyperthermophilic archaeon Pyrococcus furiosus. Archaea 2012 (1-10), 2012. https://doi.org/10.1155/2012/957852 (2012).
Kabsch, W. X. D. S. Acta Crystallogr. D Biol. Crystallogr. 66,125–132 https://doi.org/10.1107/S0907444909047337 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221. https://doi.org/10.1107/S0907444909052925 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132. https://doi.org/10.1107/S0907444904019158 (2004).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539. https://doi.org/10.1038/msb.2011.75 (2011).
Madeira, F. et al. The EMBL-EBI job dispatcher sequence analysis tools framework in 2024. Nucl. Acids Res. 52, W521–W525. https://doi.org/10.1093/nar/gkae241 (2024).
Bhat, B., Ganai, N. A., Andrabi, S. M., Shah, R. A. & Singh, A. TM-Aligner: Multiple sequence alignment tool for transmembrane proteins with reduced time and improved accuracy. Sci. Rep. 7, 12543. https://doi.org/10.1038/s41598-017-13083-y (2017).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucl. Acids Res. 42, W320–W324. https://doi.org/10.1093/nar/gku316 (2014).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v6: Recent updates to the phylogenetic tree display and annotation tool. Nucl. Acids Res. 52, W78–W82. https://doi.org/10.1093/nar/gkae268 (2024).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033. https://doi.org/10.1126/science.1218231 (2012).
Strong, M. et al. Toward the structural genomics of complexes: Crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 8060–8065 (2006). https://doi.org/10.1073/pnas.0602606103
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21. https://doi.org/10.1107/S0907444909042073 (2010).
Acknowledgements
The research leading to these results has been enabled by support from the European Community’s Seventh Framework Programme (FP7/2007–2013) under BioStruct-X (grant agreement no 283570). We thank the beamline staff scientists for support during data collection at beamline I03 at Diamond Light Source (UK). Part of this work was facilitated by the Protein Science Facility at Karolinska Institutet/SciLifeLab (http://psf.ki.se). The 2GFP-T plasmid was a gift from Scott Gradia (Addgene plasmid #29716; http://n2t.net/addgene:29716; RRID: Addgene_29716).
Funding
Open access funding provided by Royal Institute of Technology.
Author information
Authors and Affiliations
Contributions
R.G., M.M.K., D.K., T.R. and C.D. conducted the experiments; R.G., M.M.K., D.K., T.R. and C.D. designed the study; R.G., M.M.K., D.K., T.R. and C.D. wrote the manuscript. All authors reviewed and completed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gandini, R., Keskitalo, M.M., Reichenbach, T. et al. Crystallographic data for Pyrococcus furiosus dolichylphosphate mannose synthase suggest that the enzyme could flip its glycolipid product. Sci Rep 16, 9076 (2026). https://doi.org/10.1038/s41598-026-44343-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-44343-5