Abstract
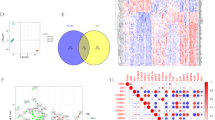
Diabetic kidney disease (DKD) is a leading cause of renal failure. Inflammation of the renal tubules and interstitium is a critical factor in the progression of DKD; however, the key regulatory genes and characteristics of the immune microenvironment remain poorly understood. This study aims to identify key inflammatory biomarkers in the renal tubule tissues of DKD patients and to elucidate their potential immunoregulatory mechanisms. By integrating multiple GEO transcriptome datasets and employing differential expression analysis, weighted gene co-expression network analysis (WGCNA), and machine learning algorithms (LASSO, Random Forest), we identified arachidonate 5-lipoxygenase (ALOX5) as a crucial feature gene of renal tubular inflammation in DKD. Clinical correlation analysis revealed that ALOX5 is significantly upregulated in DKD tissues, with high expression closely associated with decreased glomerular filtration rate and infiltration of M1 macrophages. Additionally, combining single-cell sequencing pseudotime analysis and multiplex immunohistochemistry (mIHC), we demonstrated that ALOX5 and its partner protein ALOX5AP are primarily expressed in CD68\(^+\) macrophages infiltrating the renal interstitium. They exhibit a high degree of co-localization with NF-\(\kappa\)B/p65, iNOS, and CYSLTR1, suggesting that they may mediate the pro-inflammatory polarization of macrophages through the leukotriene-NF-\(\kappa\)B axis. Finally, based on molecular docking and ADMET analysis, we screened the natural small molecule honokiol as a potential inhibitor of ALOX5, which possesses favorable pharmacokinetic properties. This study suggests that ALOX5 is a potential biomarker of immune microenvironment imbalance in DKD and provides a rationale for further investigation of targeted anti-inflammatory strategies, with honokiol as a candidate compound.
Similar content being viewed by others
Data availability
The datasets generated and analyzed during the current study are available in the GEO repository (Accession numbers: GSE104954, GSE30529, GSE47184, GSE99325, GSE209781). Requests for additional data or materials should be directed to J.Y. (email: yuanjinger@126.com)
References
Tziomalos, K. & Athyros, V. G. Diabetic nephropathy: New risk factors and improvements in diagnosis. Rev. Diabetic Stud. RDS 12, 110 (2015).
Alicic, R. Z., Rooney, M. T. & Tuttle, K. R. Diabetic kidney disease: Challenges, progress, and possibilities. Clin. J. Am. Soc. Nephrol. 12, 2032–2045 (2017).
Pan, X. et al. The burden of diabetes-related chronic kidney disease in china from 1990 to 2019. Front. Endocrinol. 13, 892860 (2022).
de Boer, I. H. et al. Diabetes management in chronic kidney disease: a consensus report by the american diabetes association (ada) and kidney disease: Improving global outcomes (kdigo). Diabetes Care 45, 3075–3090 (2022).
Tuttle, K. R. et al. Molecular mechanisms and therapeutic targets for diabetic kidney disease. Kidney Int. 102, 248–260 (2022).
Mori, Y. et al. Kim-1 mediates fatty acid uptake by renal tubular cells to promote progressive diabetic kidney disease. Cell Metab. 33, 1042–1061 (2021).
Wang, X. et al. The role of tubulointerstitial markers in differential diagnosis and prognosis in patients with type 2 diabetes and biopsy proven diabetic kidney disease. Clin. Chim. Acta 547, 117448 (2023).
Williams, B., Cliff, C. L., Lee, K., Squires, P. & Hills, C. The role of the nlrp3 inflammasome in mediating glomerular and tubular injury in diabetic nephropathy. Front. Physiol. 13, 907504 (2022).
Aranda-Rivera, A. K. et al. Involvement of inflammasome components in kidney disease. Antioxidants 11, 246 (2022).
Xia, W. et al. Molecular subtypes and prognostic signature rooted in disulfidptosis highlight tumor microenvironment in lung adenocarcinoma. Chin. J. Cancer Res. 37, 796 (2025).
Zhang, P. et al. Metabolic reprogramming signature predicts immunotherapy efficacy in lung adenocarcinoma: Targeting slc25a1 to overcome immune resistance. Chin. J. Cancer Res. 37, 1000 (2025).
Yao, L. et al. Mitochondrial dysfunction in diabetic tubulopathy. Metabolism 131, 155195 (2022).
Gilbert, R. E. Proximal tubulopathy: Prime mover and key therapeutic target in diabetic kidney disease. Diabetes 66, 791–800 (2017).
Wada, J. & Makino, H. Inflammation and the pathogenesis of diabetic nephropathy. Clin. Sci. 124, 139–152 (2013).
Youssef, N., Noureldein, M. H., Riachi, M. E., Haddad, A. & Eid, A. A. Macrophage polarization and signaling in diabetic kidney disease: a catalyst for disease progression. Am. J. Physiol. Renal Physiol. 326, F301–F312 (2024).
Klessens, C. Q. et al. Macrophages in diabetic nephropathy in patients with type 2 diabetes. Nephrol. Dial. Transplant. 32, 1322–1329 (2017).
Li, H.-D. et al. Roles and crosstalks of macrophages in diabetic nephropathy. Front. Immunol. 13, 1015142 (2022).
Cantero-Navarro, E. et al. Role of macrophages and related cytokines in kidney disease. Front. Med. 8, 688060 (2021).
Koc, M. et al. Toll-like receptor expression in monocytes in patients with chronic kidney disease and haemodialysis: relation with inflammation. Nephrol. Dial. Transplant. 26, 955–963 (2011).
Ito, S. et al. Effects of a ccr2 antagonist on macrophages and toll-like receptor 9 expression in a mouse model of diabetic nephropathy. Am. J. Physiol. Renal Physiol. 321, F757–F770 (2021).
Luo, M., Jones, S. M., Peters-Golden, M. & Brock, T. G. Nuclear localization of 5-lipoxygenase as a determinant of leukotriene b4 synthetic capacity. Proc. Natl. Acad. Sci. 100, 12165–12170 (2003).
Funk, C. D. Prostaglandins and leukotrienes: advances in eicosanoid biology. Science 294, 1871–1875 (2001).
Crosslin, D. R. et al. Genetic effects in the leukotriene biosynthesis pathway and association with atherosclerosis. Hum. Genet. 125, 217–229 (2009).
Dixon, R. et al. Requirement of a 5-lipoxygenase-activating protein for leukotriene synthesis. Nature 343, 282–284 (1990).
Ichiyama, T. et al. Cysteinyl leukotrienes induce monocyte chemoattractant protein 1 in human monocytes/macrophages. Clin. Exp. Allergy 35, 1214–1219 (2005).
Haneda, Y. et al. Leukotriene d4 enhances tumor necrosis factor-\(\alpha\)-induced vascular endothelial growth factor production in human monocytes/macrophages. Cytokine 55, 24–28 (2011).
Otunctemur, A. et al. Beneficial effects montelukast, cysteinyl-leukotriene receptor antagonist, on renal damage after unilateral ureteral obstruction in rats. Int. Braz. J. Urol. 41, 279–287 (2015).
Chen, X., Xie, H., Liu, Y., Ou, Q. & Deng, S. Interference of alox5 alleviates inflammation and fibrosis in high glucose-induced renal mesangial cells. Exp. Ther. Med. 25, 34 (2022).
Lim, C. S., Veltri, B., Kashon, M., Porter, D. W. & Ma, Q. Multi-walled carbon nanotubes induce arachidonate 5-lipoxygenase expression and enhance the polarization and function of m1 macrophages in vitro. Nanotoxicology 17, 249–269 (2023).
Hoque, A. et al. Increased 5-lipoxygenase expression and induction of apoptosis by its inhibitors in esophageal cancer: a potential target for prevention. Carcinogenesis 26, 785–791 (2005).
Tennant, C. M., Seale, J. P. & Temple, D. M. Effects of a 5-lipoxygenase inhibitor, rev-5901, on leukotriene and histamine release from human lung tissue in-vitro. J. Pharm. Pharmacol. 39, 309–311 (1987).
Tse, A.K.-W., Wan, C.-K., Shen, X.-L., Yang, M. & Fong, W.-F. Honokiol inhibits tnf-\(\alpha\)-stimulated nf-\(\kappa\)b activation and nf-\(\kappa\)b-regulated gene expression through suppression of ikk activation. Biochem. Pharmacol. 70, 1443–1457 (2005).
Hamasaki, Y. et al. Inhibition of leukotriene synthesis by honokiol in rat basophilic leukemia cells. Int. Arch. Allergy Immunol. 110, 278–281 (1996).
Woroniecka, K. I. et al. Transcriptome analysis of human diabetic kidney disease. Diabetes 60, 2354–2369 (2011).
Grayson, P. C. et al. Metabolic pathways and immunometabolism in rare kidney diseases. Ann. Rheum. Dis. 77, 1226–1233 (2018).
Ju, W. et al. Defining cell-type specificity at the transcriptional level in human disease. Genome Res. 23, 1862–1873 (2013).
Shved, N. et al. Transcriptome-based network analysis reveals renal cell type-specific dysregulation of hypoxia-associated transcripts. Sci. Rep. 7, 8576 (2017).
Lv, Z. et al. Trail induces podocyte panoptosis via death receptor 5 in diabetic kidney disease. Kidney Int. 107, 317–331 (2025).
Langfelder, P. & Horvath, S. Wgcna: an r package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Chen, H. & Boutros, P. C. Venndiagram: a package for the generation of highly-customizable venn and euler diagrams in r. BMC Bioinformatics 12, 35 (2011).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterprofiler: an r package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 16, 284–287 (2012).
Kanehisa, M. & Goto, S. Kegg: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
The gene ontology resource. enriching a gold mine. Nucleic Acids Res. 49, D325–D334 (2021).
Aran, D., Hu, Z. & Butte, A. J. xcell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 220 (2017).
Friedman, J. H., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Sarica, A., Cerasa, A. & Quattrone, A. Random forest algorithm for the classification of neuroimaging data in alzheimer’s disease: a systematic review. Front. Aging Neurosci. 9, 329 (2017).
Robin, X. et al. proc: an open-source package for r and s+ to analyze and compare roc curves. BMC Bioinf. 12, 77 (2011).
Harrell, F. E. Jr., Lee, K. L. & Mark, D. B. Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat. Med. 15, 361–387 (1996).
Tang, R. et al. Single-cell transcriptomics uncover hub genes and cell-cell crosstalk in patients with hypertensive nephropathy. Int. Immunopharmacol. 125, 111104 (2023).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Lake, B. B. et al. An atlas of healthy and injured cell states and niches in the human kidney. Nature 619, 585–594 (2023).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Acknowledgements
We are grateful to Guizhou University and Guizhou Provincial People’s Hospital for providing the valuable research platform and support during the course of this study.
Funding
This work was supported by the National Natural Science Foundation of China (Grant Nos. 82360148); Guizhou Science & Technology Department (Grant No. QKHCG2023-ZD010); Talent Fund of Guizhou Provincial People’s Hospital (Grant No. [2022]-1); Guizhou Provincial People’s Hospital Research Fund, Youth Fund (Grant No. GZSYQN[2021] 18); Guizhou Provincial Medical Research Joint Fund for High-quality Development of Health and Health Care in 2024 (Grant No. 2024GZYXKYJJXM0007); and the General Program of Guizhou Provincial Department of Science and Technology (Grant No. Qiankehe Basic MS [2025] 494).
Author information
Authors and Affiliations
Contributions
W.L. and Y.D. conceived and designed the study. Y.Z. and J.Y. supervised the project and acquired funding. W.L., Y.D., and Q.X. developed the methodology. W.L., L.Z., Y.Z., and J.Y. designed the bioinformatics analyses. W.L., L.Z., Y.Z., and X.X. performed data processing, statistical analyses, and visualization. Y.D., L.L., and F.Y. curated the data and assisted with validation. Y.Z., J.Y., D.A., and L.L. coordinated patient recruitment and sample collection. W.L., Y.D., D.Y., K.Y., and D.A. performed the multiplex immunohistochemistry experiments. X.L. and L.J.L. provided technical support for imaging and interpretation. W.L. drafted the manuscript. Y.D., Y.Z., and J.Y. revised the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All kidney biopsy samples were collected after obtaining informed consent from the patients. This study was approved by the Ethics Committee of Guizhou Provincial People’s Hospital (Approval No: 2022-65) and was conducted in strict accordance with the Declaration of Helsinki and relevant ethical guidelines for human research.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lu, W., Deng, Y., Zhai, L. et al. Integrated machine learning and multi-omics analysis identifies ALOX5 as a potential therapeutic target for tubulointerstitial inflammation in diabetic kidney disease. Sci Rep (2026). https://doi.org/10.1038/s41598-026-44445-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-44445-0