Abstract
Co-infection with Mycoplasma pneumoniae (MP) represents a clinically significant complication that often leads to prolonged hospital stay, increased mortality risk, and a higher demand for mechanical ventilation. This study constructed a risk prediction model for early identification of SARS-CoV-2 and MP co-infections. We retrospectively analyzed SARS-CoV-2 patients admitted between December 2022 and February 2023. Patients were stratified into co-infection and mono-infection groups based on MP antibody results. LASSO regression screened 55 variables, followed by multicollinearity checks. Restricted cubic splines (RCS) analyzed nonlinear relationships between continuous variables and infection risk. Conventional logistic and integrated RCS-logistic models were constructed and compared. LASSO identified seven predictors: age, globulin, anion gap, blood urea nitrogen (BUN), uric acid, prothrombin time (PT), and thrombin time (TT). Multivariate analysis showed globulin, anion gap, uric acid, and TT were independent risk factors, whereas BUN was protective. RCS revealed significant nonlinear associations between globulin, PT, and TT levels. The RCS-logistic model outperformed the conventional linear model, with a higher AUC of 0.827, better calibration (Brier score = 0.169), and greater net clinical benefit on decision curve analysis. This model enables early risk assessment and optimizes treatment, offering a methodological reference for predicting co-infections with emerging respiratory pathogens.
Similar content being viewed by others

Introduction
Coronavirus disease 2019 (COVID-19) is the third global coronavirus pandemic in the 21 st century and has become a major public health event1. The pathogenicity of the virus has decreased2, and global prevention and control measures have been relaxed. However, the number of infected individuals has continued to increase. As of August 17, 2025, the WHO reported 778,510,178 confirmed cases and 7,100,227 deaths3. Epidemiological data show that SARS-CoV-2 infection is frequently accompanied by other pathogens, which may affect the diagnosis, treatment, prognosis, and even increase mortality4,5. Common pathogens that cause co-infections include bacteria, such as Streptococcus pneumoniae, and atypical pathogens, with Mycoplasma pneumoniae (MP) being particularly prevalent6. A meta-analysis showed that approximately 7% of patients hospitalized with COVID-19 had bacterial co-infections, with MP among the most common pathogens in patients with SARS-CoV-2 co-infections7.
Chest CT imaging manifestations of COVID-19 are polymorphic, including ground-glass opacity, consolidation, reticular opacity, and nodules8. The clinical and imaging features of MP pneumonia often overlap with those of COVID-19, leading to potential under-recognition of co-infection1. However, the two conditions require different treatment strategies. A large multi-center retrospective study showed that, compared with mono-infection, patients with COVID-19 and MP co-infection had a significantly higher mortality rate, increased mechanical ventilation requirements, and longer hospital stay9. Therefore, inadequate or delayed identification and treatment of co-infections are key risk factors, often leading to improper antibacterial use, prolonged disease course, poor prognosis, and an increased risk of fatal complications such as severe pneumonia.
Traditional logistic regression models often assume a linear relationship between predictors and outcomes, limiting their ability to depict complex associations. Although pure machine-learning models can capture nonlinearity, they lack interpretability and are prone to overfitting in small clinical datasets. However, till date, no existing model has adequately combined predictive accuracy, flexibility for nonlinear relationships, and clinical interpretability, and this represents a critical gap in the management of SARS-CoV-2 and MP co-infections. To address this, this study aimed to construct a restricted cubic spline (RCS)-based logistic regression model to flexibly capture the nonlinear relationship between predictors and co-infection risk, taking into account both prediction accuracy and clinical interpretability. This approach supports early identification, appropriate intervention, and disease surveillance of patients co-infected with SARS-CoV-2 and MP, while providing a stable, interpretable, and clinically applicable framework for managing co-infections caused by emerging or recurrent respiratory pathogens with overlapping symptoms but different treatment strategies. Thus, this study provides a methodological reference for optimizing precise identification and intervention strategies for co-infections.
Methods
Data source
A total of 312 patients with COVID-19 were retrospectively enrolled from the Department of Respiratory and Critical Care Medicine at Mindong Hospital Affiliated to Fujian Medical University in Ningde between December 26, 2022, and February 14, 2023. The diagnosis of SARS-CoV-2 infection was based on the criteria outlined in the “Diagnosis and Treatment Protocol for COVID-19” issued by the National Health Commission of the People’s Republic of China. MP infection was diagnosed by the detection of serum total antibodies using the particle agglutination method. After applying these criteria, 242 eligible patients with COVID-19 were included. Given that all enrolled patients presented with acute respiratory symptoms and imaging findings of pneumonia, patients with an MP antibody titer of > 1:160 (indicating acute infection) were assigned to the co-infection group, whereas those with a negative titer (< 1:40) were assigned to the non-co-infection group. Ethical approval for the study was obtained from the Institutional Review Board of Mindong Hospital Affiliated to Fujian Medical University in Ningde (Approval No. K2026011501), which waived the need for informed consent because of the retrospective nature of the study. The study was conducted in accordance with the principles of the Declaration of Helsinki.
Data and variables
General clinical data were collected for all enrolled patients, including sex, age, body mass index, smoking and alcohol use history, length of hospital stay, COVID-19 vaccination status, presence of lung imaging lesions, and underlying diseases (hypertension, diabetes, coronary heart disease, chronic obstructive pulmonary disease, pneumoconiosis). Laboratory parameters included complete blood count, biochemistry, coagulation profile, and brain natriuretic peptide. Blood tests were performed on the first day of admission, and when multiple test results were available for a specific variable, the first measurement was used for the analysis.
Statistical analysis
Continuous variables with a normal distribution were expressed as mean ± standard deviation, and group differences were analyzed using the t-test. Continuous variables that were not normally distributed were presented as medians and interquartile ranges (M [P25, P75]), and group comparisons were performed using the Mann–Whitney U test. Categorical data were expressed as counts and percentages (n, %), and differences between groups were analyzed using the chi-square test. All statistical tests were two-sided, and a P-value of < 0.05 was considered statistically significant. Data organization and statistical analyses were performed using IBM SPSS (version 26.0) and R (version 4.3.0).
Model construction and evaluation
To identify key variables and mitigate the risk of overfitting, LASSO regression was used to select features from 55 predictive variables. The optimal regularization parameter (λ) was determined through 10-fold cross-validation, and the variable combination corresponding to λ.1SE was chosen based on the “one standard error” rule. Subsequently, pairwise correlation analysis was conducted on the selected variables to assess multicollinearity. For continuous variables included in the model, RCS and likelihood ratio tests were employed to evaluate nonlinear relationships with the outcome, with statistical significance set at P < 0.05. If a nonlinear association was detected, knots were placed at the 25th, 50th, and 75th percentiles for RCS fitting; otherwise, the variable was included as a linear term. Based on this, an RCS-logistic regression model was constructed and compared with a conventional linear logistic model. A nomogram was then developed based on the final model. Model performance was evaluated using the following metrics: discrimination (area under the curve [AUC], sensitivity, and specificity), calibration (calibration curve and Brier score), and clinical utility (decision curve analysis, DCA).
Results
Baseline characteristics
The demographic and clinical characteristics of patients in the co-infection and non-co-infection groups are shown in Table 1. Significant differences were observed between the two groups for age, globulin (GLB), anion gap (Ag), blood urea nitrogen (BUN), uric acid (UA), prothrombin time (PT), and thrombin time (TT) (P < 0.05), while no significant differences were found for the remaining variables, including sex, underlying diseases, laboratory parameters, and imaging indicators (P > 0.05).
Predictor variable selection: LASSO regression analysis
LASSO regression was employed to select variables from 55 candidate predictors, mitigating the risk of model overfitting and identifying the most predictive variables. The optimal regularization parameters were determined via 10-fold cross-validation (λ.min = 0.047 and λ.1SE = 0.052; Fig. 1a,b). The model corresponding to λ.1SE yielded a cross-validated binomial deviance of 1.335. This value falls within one standard error (0.017) of the minimum deviance (1.330), providing a more concise and strongly regularized model with seven nonzero coefficient variables. Therefore, the model corresponding to λ.1SE was selected as the final variable set. The seven features incorporated into the predictive model and their respective LASSO coefficients are: TT (β = 0.096), PT (β = 0.049), Ag (β = 0.033), BUN (β = −0.020), age (β = −0.006), UA (β = 0.002), GLB (β = 0.001). The ranking of variable importance is shown in Fig. 1c.
Feature selection and variable importance analysis using the LASSO regression model. (a) Selection of the optimal tuning parameter (\(\lambda\)) in the LASSO model using 10-fold cross-validation based on the minimum criteria and 1-standard error (1-SE) criteria. The dotted vertical lines represent the optimal \(\lambda\)values. (b) LASSO coefficient profiles of the candidate variables against the log (\(\lambda\)) sequence. The legend indicates the specific \(\lambda\) values (min and 1-SE) selected for the final model. (c) Relative importance ranking of the significant predictors. Red bars indicate positive coefficients (risk factors), whereas blue bars indicate negative coefficients (protective factors).
Multivariate logistic regression and collinearity analysis
Seven variables were screened using LASSO and included in multivariate logistic regression. The results are summarized in Table 2. GLB, Ag, UA, and TT were independent risk factors for MP infection in patients with COVID-19, while BUN was an independent protective factor. The multicollinearity analysis results are shown in Supplementary Table 1, with all variables having a variance inflation factor below 2, indicating no significant multicollinearity among the selected variables.
Test of correlation and nonlinear relationship among variables
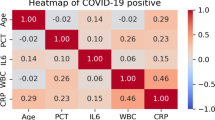
The results of the pairwise correlation analysis of the variables are shown in Fig. 2a. BUN was significantly positively correlated with UA (r = 0.504, P < 0.001), whereas TT was negatively correlated with GLB (r = −0.311, P < 0.001). RCS analysis indicated that GLB, PT, and TT had significant nonlinear associations with the outcome (P = 0.0003, 0.0008, and 0.001, respectively), whereas age, BUN, Ag, and UA conformed to the linear hypothesis (P < 0.05) (Fig. 2b–h). We used standardized odds ratios to compare the relative impacts of various variables on the risk of MP co-infection (Table 3). Specifically, GLB > 28.36 g/L, PT < 11.84 s, and TT ≥ 13.29 s were significantly associated with an increased risk of MP infection.
Correlation analysis of predictors and restricted cubic spline (RCS) analysis for continuous variables. (a) Spearman correlation heatmap of the identified predictor variables. The color intensity corresponds to the magnitude of the correlation coefficient, indicating no severe multicollinearity among the features. (b–h) RCS plots visualizing the nonlinear associations between continuous predictors and the risk of Mycoplasma pneumoniae co-infection: (b) Age, (c) thrombin time, (d) prothrombin time, (e) anion gap, (f) uric acid, (g) globulin, and (h) blood urea nitrogen. The solid lines represent the estimated odds ratios, and the shaded areas represent the 95% confidence intervals.
Model evaluation
All seven predictors selected by LASSO regression were included as linear terms in the logistic regression model. The model achieved an AUC value of 0.789, with a sensitivity and specificity of 63.6% and 86.0%, respectively. Three variables, GLB, PT, and TT, were included as nonlinear terms using three-node RCS, whereas the remaining predictors, age, BUN, Ag, and UA, were included as linear terms. An RCS-logistic regression model was constructed and demonstrated superior performance, with an AUC of 0.827, sensitivity of 75.2%, and specificity of 77.7%. The receiver operating characteristic curves for the two prediction models are shown in Fig. 3a. Calibration evaluation showed that the Brier score of the RCS-logistic model was lower (0.169 vs. 0.189), and the calibration curve was closer to the ideal diagonal (Fig. 3b). DCA (Fig. 3c) further indicated that, within a threshold probability range of 0.1–0.6, the RCS-logistic model provided greater clinical net benefits.
Performance evaluation and comparison between the standard logistic model and the restricted cubic spline (RCS)-logistic model. (a) Receiver operating characteristic curves illustrating discrimination performance. The RCS-logistic model (blue line) achieved a higher area under the curve (AUC = 0.827) than the standard logistic model (yellow line, AUC = 0.789). (b) Calibration curves assessing the agreement between predicted and observed probabilities. The diagonal dashed line represents the ideal calibration. The RCS-logistic model demonstrated superior calibration, with a lower Brier score (0.1687) than the standard model (0.1891). (c) Decision curve analysis (DCA) estimating the net clinical benefit across different threshold probabilities. The RCS-logistic model consistently provided a higher standardized net benefit than the standard model across a wide range of thresholds.
Construction of the predictive model nomogram
The aforementioned novel clinical risk prediction model for SARS-CoV-2 and Mycoplasma co-infection is presented as a nomograms (Fig. 4a,b). The higher the total score calculated based on the nomogram, the higher the risk of Mycoplasma infection in patients with COVID-19.
Nomograms for predicting SARS-CoV-2 and Mycoplasma co-infection using logistic and spline-based regression approaches. (a) Nomogram based on the standard logistic regression model, assuming linear relationships for all predictors. (b) Nomogram based on the RCS-logistic model. In this model, globulin, prothrombin time, and thrombin time were modeled using RCS with three knots to account for nonlinear relationships, whereas age, blood urea nitrogen, anion gap, and uric acid were included as linear terms.
Discussion
Since the onset of the SARS-CoV-2 pandemic, global infections have persisted, complicating clinical management with challenges such as bacterial co-infection10. Historical data from H1N1 and H7N9 epidemics confirm that concurrent bacterial infections significantly exacerbate disease severity11,12,13,14. Specifically, co-infection with MP in patients with COVID-19 is common and associated with adverse outcomes, including increased mortality, intensive care requirements, and prolonged hospital stays compared to mono-infection1,7. Consequently, constructing a risk prediction model for SARS-CoV-2 and MP co-infection is clinically imperative. This model enables the early identification of patients who are at high risk, minimizes missed diagnoses, and guides personalized interventions to prevent complications.
Our study identified coagulation dysregulation as a hallmark of co-infection, characterized by prolonged TT but notably shortened PT (< 11.84 s). Although the wide confidence interval for the PT subgroup reflected the limited sample size (lower clinical prevalence) and multivariable analysis yielded a borderline P-value (P = 0.052), PT was retained in the final model via LASSO selection because it clinically suggests an intensely hypercoagulable state. This finding aligns with the evidence that co-infection amplifies the thrombosis risk beyond that of mono-infection15. Mechanistically, SARS-CoV-2 drives endothelial dysfunction and thrombin bursts via cytokine storms16,17,18,19, whereas MP activates coagulation pathways20. We postulate a synergistic effect, in which initial hypercoagulability (shortened PT) may coexist with or progress to consumptive coagulopathy (prolonged TT). Thus, these opposing coagulation abnormalities may reflect a severely disrupted coagulation-inflammatory network in co-infected patients.
Age was another important feature of this study. Although co-infection risk was associated with age, the pattern contradicted the common expectation that older people are more susceptible, aligning with the epidemiological characteristics of MP infection21. Possible explanations include: first, differences in communication and behavioral patterns, as Mycoplasma is primarily transmitted through droplets. Young people have frequent social interactions and greater mobility, and their exposure opportunities considerably increase in collective environments such as schools22, while older adults have a relatively limited range of activities and lower risk of exposure. Second, differences in immune memory may play a role, as older adults may have partial cross-immunity from previous infections, whereas younger individuals generally lack such immune experience. Third, diagnosis in older patients is often insufficient, since clinical manifestations can overlap with underlying diseases, leading to atypical infections being easily missed and infection rates underestimated23. Therefore, incorporating age can enhance the model’s ability to identify populations at high risk of co-infections and guide early clinical intervention.
Ag is a low-cost, routine detection index. Elevated Ag often indicates metabolic acidosis and can reflect pathological conditions such as insufficient tissue perfusion or renal insufficiency24. In patients with COVID-19, elevated Ag levels have been associated with an increased risk of severe illness and mortality25,26. This study identified a considerable nonlinear relationship between Ag and the risk of secondary infection: as Ag levels rise, the risk of infection initially increases but levels off after a certain critical value, exhibiting a “saturation effect.” This result confirms Ag as a marker of disease severity and clarifies its dose-response with infection risk. It suggests that clinical intervention should target precise thresholds, with timely correction of tissue perfusion and renal function before Ag reaches critical levels to help prevent secondary infections. When the Ag increases significantly, a more comprehensive critical care management approach should be adopted. This study expands the understanding of the role of Ag in secondary COVID-19 infections and provides a more refined basis for clinical risk assessment.
UA, an end product of purine metabolism, is an important endogenous antioxidant27. SARS-CoV-2 infection can promote the generation of reactive oxygen species and induce oxidative stress, thereby causing cellular and tissue damage28. In response, the body activates its antioxidant defense system. Elevated serum UA can inhibit virus-induced oxidative stress, thereby reducing cell damage. Recent studies have further demonstrated that in cases of severe Mycoplasma infection, the body can compensate by increasing serum UA concentrations through multi-system regulatory mechanisms29. Co-infection with two pathogens may intensify oxidative stress, leading to a compensatory increase in blood UA levels. Therefore, hyperuricemia can serve as a potential biomarker for early warning of the risk of co-infection.
γ-globulin, a member of the GLB family, is essentially an immunoglobulin (antibody) and plays a core role in humoral immunity30. SARS-CoV-2 infection activates B cells to produce antibodies, and co-infection further increases antibody synthesis, resulting in elevated serum GLB levels. The increase in GLB levels observed in this study reflects the body’s enhanced humoral immunity to pathogens. Although the effect size is relatively small, GLB remains an important marker of immune status and may enhance the accuracy of the prediction model in identifying co-infection risks when considered alongside other indicators.
BUN, a metabolic byproduct reflecting renal and hepatic function, was negatively correlated with MP co-infection risk in our study, contrasting with reports of elevated BUN in severe COVID-19 cases31. This discrepancy can be attributed to several factors. First, given the reduced virulence of the current SARS-CoV-2 strains and lower incidence of acute kidney injury, most patients maintain normal baseline BUN levels. However, infection-induced anorexia likely reduces protein intake and subsequent BUN production. Additionally, since both pathogens can impair hepatic function32,33, co-infection may suppress urea synthesis. Consistent with this, previous studies on MP-related acute respiratory distress syndrome reported no significant BUN elevation34, and some severe cases had lower BUN levels than did mild ones35. Despite these differences, BUN has demonstrated significant predictive power. Adhering to a data-driven approach, we retained this variable to optimize the model performance and prevent bias by excluding statistically significant predictors.
The prediction model constructed in this study can accurately identify patients at high risk of COVID-19 with MP infection, supporting the targeted detection, avoiding unnecessary screening, optimizing resource allocation, and reducing the medical burden while maintaining diagnostic and treatment quality. All indicators included for this model are from routine clinical tests, providing a solid basis for broader application. Its importance extends beyond current use in the post-pandemic era; whether seasonal surges of endemic pathogens (such as influenza or respiratory syncytial virus) occur or a new type of respiratory virus outbreak occurs again in the future (such as the Nipah virus disease discovered recently in India), we need to pay attention to the possibility of concurrent Mycoplasma or other bacterial infections and investigate whether such co-infections will lead to a worse prognosis. A discussion about these issues can provide methodological references and help predict potential co-infection scenarios to guide clinical decision-making and antimicrobial stewardship.
This study has a few limitations. First, it was based on single-center retrospective data for a small sample collected from a specific region (Fujian Province, China) during a unique timeframe (the initial Omicron wave, Dec 2022–Feb 2023). 2qsfThis spatiotemporal limitation may have introduced selection bias, restricting the model’s generalizability to regions with different MP prevalences or to future infection waves driven by new viral variants. Second, driven by urgent triage needs during the COVID-19 surge, we lacked a control group with MP mono-infection, thus preventing the definitive separation of the synergistic effects of co-infection from MP-specific features. Third, the model lacked external validation, although our interpretable RCS-logistic framework helped mitigate overfitting risks. Fourth, single-point serology may result in misclassification of recently resolved MP infections as active co-infections, alongside potential false negatives during the window period. Moving forward, our team plans to initiate multicenter prospective studies that include nucleic acid testing and more comprehensive control groups (including MP mono-infection) to externally validate the model and explore underlying pathophysiological mechanisms in depth.
Conclusion
The RCS-logistic model showed strong predictive ability for SARS-CoV-2 and MP co-infection, offering clinicians a valuable tool for early risk assessments and optimized treatment strategies. This modeling framework provides a methodological approach for predicting new or recurrent co-infections with respiratory pathogens.
Data availability
The datasets analyzed during the current study are available from the corresponding author on reasonable request.
References
Huang, A. C. C., Huang, C. G., Yang, C. T. & Hu, H. C. Concomitant infection with COVID-19 and Mycoplasma pneumoniae. Biomed. J. 43, 458–461. https://doi.org/10.1016/j.bj.2020.07.002 (2020).
Suzuki, R. et al. Attenuated fusogenicity and pathogenicity of SARS-CoV-2 Omicron variant. Nature 603, 700–705. https://doi.org/10.1038/s41586-022-04462-1 (2022).
WHO COVID-19 Dashboard. https://data.who.int/dashboards/covid19/deaths?n=c.
Ruiz-Bastián, M., Falces-Romero, I., Ramos-Ramos, J. C., de Pablos, M. & García-Rodríguez, J. Bacterial co-infections in COVID-19 pneumonia in a tertiary care hospital: Surfing the first wave. Diagn. Microbiol. Infect. Dis. 101, 115477. https://doi.org/10.1016/j.diagmicrobio.2021.115477 (2021).
Hoque, M. N. et al. Diversity and genomic determinants of the microbiomes associated with COVID-19 and non-COVID respiratory diseases. Gene Rep. 23, 101200. https://doi.org/10.1016/j.genrep.2021.101200 (2021).
Hoque, M. N. et al. Microbial co-infections in COVID-19: Associated microbiota and underlying mechanisms of pathogenesis. Microb. Pathog. 156, 104941. https://doi.org/10.1016/j.micpath.2021.104941 (2021).
Lansbury, L., Lim, B., Baskaran, V. & Lim, W. S. Co-infections in people with COVID-19: A systematic review and meta-analysis. J. Infect. 81, 266–275. https://doi.org/10.1016/j.jinf.2020.05.046 (2020).
Ai, T. et al. Correlation of chest CT and RT-PCR testing for coronavirus disease 2019 (COVID-19) in China: A report of 1014 cases. Radiology 296, E32–E40. https://doi.org/10.1148/radiol.2020200642 (2020).
Rangroo, R. et al. The severity of the co-infection of mycoplasma pneumoniae in COVID-19 patients. Cureus 14, e24563. https://doi.org/10.7759/cureus.24563 (2022).
Paget, C. & Trottein, F. Mechanisms of bacterial superinfection post-influenza: A role for unconventional T cells. Front. Immunol. 10, 336. https://doi.org/10.3389/fimmu.2019.00336 (2019).
Jia, L. et al. Mechanisms of severe mortality-associated bacterial co-infections following influenza virus infection. Front. Cell. Infect. Microbiol. 7, 338. https://doi.org/10.3389/fcimb.2017.00338 (2017).
Dhanoa, A., Fang, N. C., Hassan, S. S., Kaniappan, P. & Rajasekaram, G. Epidemiology and clinical characteristics of hospitalized patients with pandemic influenza A (H1N1) 2009 infections: the effects of bacterial coinfection. Virol. J. 8, 501. https://doi.org/10.1186/1743-422X-8-501 (2011).
Klein, E. Y. et al. The frequency of influenza and bacterial coinfection: A systematic review and meta-analysis. Influenza and Other Respiratory Viruses 10, 394–403. https://doi.org/10.1111/irv.12398 (2016).
Yang, M. et al. Bacterial coinfection is associated with severity of avian influenza A (H7N9), and procalcitonin is a useful marker for early diagnosis. Diagn. Microbiol. Infect. Dis. 84, 165–169. https://doi.org/10.1016/j.diagmicrobio.2015.10.018 (2015).
Zha, L. et al. Clinical features and outcomes of adult COVID-19 patients co-infected with Mycoplasma pneumoniae. J. Infect. 81, e12–e15. https://doi.org/10.1016/j.jinf.2020.07.010 (2020).
Zhang, L. et al. D-dimer levels on admission to predict in-hospital mortality in patients with Covid-19. J. Thromb. Haemost. 18, 1324–1329. https://doi.org/10.1111/jth.14820 (2020).
Tang, N., Li, D., Wang, X. & Sun, Z. Abnormal coagulation parameters are associated with poor prognosis in patients with novel coronavirus pneumonia. J. Thromb. Haemost. 18, 844–847. https://doi.org/10.1111/jth.14768 (2020).
Bryce, C. et al. Pathophysiology of SARS-CoV-2: The Mount Sinai COVID-19 autopsy experience. Mod. Pathol. 34, 1456–1467. https://doi.org/10.1038/s41379-021-00793-y (2021).
Wiersinga, W. J. & van der Poll, T. Immunopathophysiology of human sepsis. EBioMedicine 86, 104363. https://doi.org/10.1016/j.ebiom.2022.10436 (2022).
Qiu, J., Ge, J., Cao, L. & D-dimer The risk factor of children’s severe mycoplasma pneumoniae pneumonia. Front. Pediatr. 10, 828437. https://doi.org/10.3389/fped.2022.828437 (2022).
Chen, Q., Hu, T., Wu, L. & Chen, L. Clinical features and biomarkers for early prediction of refractory mycoplasma pneumoniae pneumonia in children. Emerg. Med. Int. 2024, 9328177. https://doi.org/10.1155/2024/9328177 (2024).
Kant, R. et al. Critical insights from recent outbreaks of Mycoplasma pneumoniae: decoding the challenges and effective interventions strategies. Int. J. Infect. Dis. 147, 107200. https://doi.org/10.1016/j.ijid.2024.107200 (2024).
Nicolson, G. L. Pathogenic mycoplasma infections in chronic illnesses: General considerations in selecting conventional and integrative treatments. Int. J. Clin. Med. 10, 477–522. https://doi.org/10.4236/ijcm.2019.1010041 (2019).
Achanti, A. & Szerlip, H. M. Acid-base disorders in the critically ill patient. Clin. J. Am. Soc. Nephrol. 18, 102–112. https://doi.org/10.2215/CJN.04500422 (2022).
Zemlin, A. E. et al. The association between acid-base status and clinical outcome in critically ill COVID-19 patients admitted to intensive care unit with an emphasis on high anion gap metabolic acidosis. Ann. Clin. Biochem. 60, 86–91. https://doi.org/10.1177/00045632221134687 (2022).
Huang, Y., Qu, J. & Zhen, P. Non-linear association between anion gap and 28-day mortality in critically ill patients with COVID-19: A cohort study from MIMIC IV database. J. Thorac. Dis. 17, 4662–4671. https://doi.org/10.21037/jtd-2024-1964 (2025).
Allegrini, S., Garcia-Gil, M., Pesi, R., Camici, M. & Tozzi, M. G. The good, the bad and the new about uric acid in cancer. Cancers. 14, 4959. https://doi.org/10.3390/cancers14194959 (2022).
Aydin Bahat, K. The effect of uric acid level on the severity of COVID-19. Acta Clin. Croat. 63, 251–259. https://doi.org/10.20471/acc.2024.63.02.1 (2024).
Pan, C., Chen, Y., Wang, S., Li, M. & Qu, S. The study of routine laboratory factors in children with mycoplasma pneumoniae pneumonia: serum uric acid may have anti-inflammatory effect. J. Clin. Lab. Anal. 35, e24026. https://doi.org/10.1002/jcla.24026 (2021).
Primorac, D. et al. Adaptive immune responses and immunity to SARS-CoV-2. Front. Immunol. 13, 848582. https://doi.org/10.3389/fimmu.2022.848582 (2022).
Ghahramani, S. et al. Laboratory features of severe vs. non-severe COVID-19 patients in Asian populations: a systematic review and meta-analysis. Eur. J. Med. Res. 25, 30. https://doi.org/10.1186/s40001-020-00432-3 (2020).
Papagiouvanni, I. et al. COVID-19 and liver injury: An ongoing challenge. World J. Gastroenterol. 29, 257–271. https://doi.org/10.3748/wjg.v29.i2.257 (2023).
Yu, Y., Ji, R., Xia, Y. & Liu, F. Multicenter analysis of clinical characteristics and risk factors for liver injury in severe Mycoplasma pneumoniae pneumonia. Pediatr. Infect. Dis. J. 45, 236–243. https://doi.org/10.1097/INF.0000000000005037 (2026).
Chaabane, N., Coupez, E., Buscot, M. & Souweine, B. Acute respiratory distress syndrome related to Mycoplasma pneumoniae infection. Respir. Med. Case Rep. 20, 89–91. https://doi.org/10.1016/j.rmcr.2016.11.016 (2016).
Ding, W. et al. Clinical features and risk factors for severe macrolide-resistant Mycoplasma pneumoniae pneumonia induced by 23 S rRNA A2063G mutation: A retrospective observational study. Eur. J. Clin. Microbiol. Infect. Dis. 44, 2475–2486. https://doi.org/10.1007/s10096-025-05215-4 (2025).
Author information
Authors and Affiliations
Contributions
KY, YS, XH, YD and JX conceived and designed the study. XC, BS, QZ, HL and LZ performed the data analyses. KY, YS, XH wrote the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Ethical approval for the study was obtained from the Institutional Review Board of Mindong Hospital Affiliated to Fujian Medical University in Ningde (Approval No. K2026011501). The study was conducted in accordance with the principles of the Declaration of Helsinki.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ye, K., Su, Y., Hu, X. et al. A retrospective clinical risk prediction model for co‑infection with Mycoplasma pneumoniae in patients with COVID‑19 based on restricted cubic splines. Sci Rep 16, 14206 (2026). https://doi.org/10.1038/s41598-026-44539-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-44539-9