Abstract
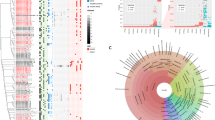
Pneumonia remains a leading cause of global mortality. Conventional diagnostic approaches frequently fail to distinguish microbial colonization from true infection in the lower respiratory tract, complicating clinical decision-making and contributing to antibiotic overuse. Improved diagnostic strategies are urgently needed. In this prospective, single-center study, deep sputum specimens were collected from patients with respiratory colonization (n = 17) and infectious pneumonia (n = 27) admitted to the neurosurgical ICU of Huashan Hospital. Metagenomic next-generation sequencing (mNGS) and metatranscriptomic profiling were performed to characterize both the pulmonary microbiota and the host immune response. These features were subsequently integrated to construct a diagnostic model. Microbial community profiling revealed reduced alpha diversity and enrichment of metabolically active pathogenic taxa in the infection group, consistent with a dysbiotic state permissive to invasion. In contrast, the colonization group demonstrated a more balanced microbial ecosystem. Transcriptomic analyses identified 2232 differentially expressed host genes between the two groups. The colonization group showed marked activation of the Wnt, MAPK, chemokine, and focal adhesion pathways, which are functionally implicated in epithelial barrier maintenance and early immune homeostasis. A multi-omics diagnostic model incorporating seven gene features (ANKRD52, ZC3HAV1L, SERPINE3, CDPF1, ZNF720, TAGLN3, and LRRC15) achieved a discrimination between colonization and infection (AUC = 0.951 in the training cohort; 0.875 in the validation set). By jointly analyzing the pulmonary microbiome and host transcriptome, this study provides insight into host–microbe interactions distinguishing colonization from infection and presents a predictive model with potential clinical relevance.
Similar content being viewed by others
Data availability
The raw sequencing data generated in this study have been deposited in the Genome Sequence Archive (GSA) under accession number CRA032964. The data can be accessed via the following link: https://ngdc.cncb.ac.cn/gsa/browse/CRA032964.All analysis scripts (R code) used for statistical analysis, machine learning modeling, and figure generation are publicly available on GitHub at https://github.com/fuzhangfan-lgtm/colonization-and-infectious-pneumonia.git.The study protocol is available from the corresponding author upon reasonable request.
References
Abdalla, S., Abd-Allah, F. & Abdel Aziz, M. I. Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet 385(117–171), 2015. https://doi.org/10.1016/s0140-6736(14)61682-2 (2015).
Jones, B. E. et al. Incidence and outcomes of non–ventilator-associated hospital-acquired pneumonia in 284 US hospitals using electronic surveillance criteria. JAMA Netw. Open 6, e2314185-e2314185. https://doi.org/10.1001/jamanetworkopen.2023.14185 (2023).
Li, W. et al. Incidence and risk factors of ventilator-associated pneumonia in the intensive care unit: A systematic review and meta-analysis. J. Thorac. Dis. 16, 5518–5528. https://doi.org/10.21037/jtd-24-150 (2024).
Sun, Y. et al. Incidence of community-acquired pneumonia in urban China: A national population-based study. Vaccine 38, 8362–8370. https://doi.org/10.1016/j.vaccine.2020.11.004 (2020).
Gray, J. & Coupland, L. J. The increasing application of multiplex nucleic acid detection tests to the diagnosis of syndromic infections. Epidemiol. Infect. 142, 1–11. https://doi.org/10.1017/s0950268813002367 (2014).
Fan, R. R. et al. Nasopharyngeal pneumococcal density and evolution of acute respiratory illnesses in young children, Peru, 2009-2011. Emerg. Infect. Dis. 22, 1996–1999. https://doi.org/10.3201/eid2211.160902 (2016).
Esposito, S. et al. DNA bacterial load in children with bacteremic pneumococcal community-acquired pneumonia. Eur. J. Clin. Microbiol. Infect. Dis. 32, 877–881. https://doi.org/10.1007/s10096-013-1821-0 (2013).
Platts-Mills, J. A., Liu, J. & Houpt, E. R. New concepts in diagnostics for infectious diarrhea. Mucosal Immunol. 6, 876–885. https://doi.org/10.1038/mi.2013.50 (2013).
Fischer, N. et al. Evaluation of unbiased next-generation sequencing of RNA (RNA-seq) as a diagnostic method in Influenza virus-positive respiratory samples. J. Clin. Microbiol. 53, 2238–2250. https://doi.org/10.1128/jcm.02495-14 (2015).
Fu, Z. et al. Pathogen quantitative efficacy of different spike-in internal controls and clinical application in central nervous system infection with metagenomic sequencing. Microbiol. Spectr. 11, e0113923. https://doi.org/10.1128/spectrum.01139-23 (2023).
Wilson, M. R. et al. Diagnosing Balamuthia mandrillaris encephalitis with metagenomic deep sequencing. Ann. Neurol. 78, 722–730. https://doi.org/10.1002/ana.24499 (2015).
Suarez, N. M. et al. Superiority of transcriptional profiling over procalcitonin for distinguishing bacterial from viral lower respiratory tract infections in hospitalized adults. J. Infect. Dis. 212, 213–222. https://doi.org/10.1093/infdis/jiv047 (2015).
Tsalik, E. L., McClain, M. & Zaas, A. K. Moving toward prime time: Host signatures for diagnosis of respiratory infections. J. Infect. Dis. 212, 173–175. https://doi.org/10.1093/infdis/jiv032 (2015).
Langelier, C. et al. Integrating host response and unbiased microbe detection for lower respiratory tract infection diagnosis in critically ill adults. Proc. Natl. Acad. Sci. U. S. A. 115, E12353-e12362. https://doi.org/10.1073/pnas.1809700115 (2018).
Diao, Z. et al. Validation of a metagenomic next-generation sequencing assay for lower respiratory pathogen detection. Microbiol. Spectr. 11, e0381222. https://doi.org/10.1128/spectrum.03812-22 (2023).
Sulaiman, I. et al. Microbial signatures in the lower airways of mechanically ventilated COVID-19 patients associated with poor clinical outcome. Nat. Microbiol. 6, 1245–1258. https://doi.org/10.1038/s41564-021-00961-5 (2021).
Zhou, Z. et al. Heightened innate immune responses in the respiratory tract of COVID-19 patients. Cell Host Microbe 27, 883-890.e882. https://doi.org/10.1016/j.chom.2020.04.017 (2020).
Han, D. et al. Metagenomic fingerprints in bronchoalveolar lavage differentiate pulmonary diseases. npj Digit. Med. 8, 599. https://doi.org/10.1038/s41746-025-01977-5 (2025).
Sun, Q. et al. Clinical implement of probe-capture metagenomics in sepsis patients: A multicentre and prospective study. Clin. Transl. Med. 15, e70297. https://doi.org/10.1002/ctm2.70297 (2025).
Tao, Y. et al. Diagnostic performance of metagenomic next-generation sequencing in pediatric patients: A retrospective study in a large children’s medical center. Clin. Chem. 68, 1031–1041. https://doi.org/10.1093/clinchem/hvac067 (2022).
Steen, C. B., Liu, C. L., Alizadeh, A. A. & Newman, A. M. Profiling cell type abundance and expression in bulk tissues with CIBERSORTx. Methods Mol. Biol. 2117, 135–157. https://doi.org/10.1007/978-1-0716-0301-7_7 (2020).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782. https://doi.org/10.1038/s41587-019-0114-2 (2019).
Whitney, A. W. A direct method of nonparametric measurement selection. IEEE Trans. Comput. C–20, 1100–1103. https://doi.org/10.1109/T-C.1971.223410 (1971).
Collins, G. S. et al. TRIPOD+AI statement: Updated guidance for reporting clinical prediction models that use regression or machine learning methods. BMJ 385, e078378. https://doi.org/10.1136/bmj-2023-078378 (2024).
Ageevets, V. A. et al. Emergence of carbapenemase-producing Gram-negative bacteria in Saint Petersburg, Russia. Int. J. Antimicrob. Agents. 44, 152–155. https://doi.org/10.1016/j.ijantimicag.2014.05.004 (2014).
Huang, L. et al. Risk factors for imipenem-nonsusceptible Acinetobacter nosocomialis bloodstream infection. J. Microbiol. Immunol. Infect. 47, 311–317. https://doi.org/10.1016/j.jmii.2013.02.002 (2014).
Pace, N. R. Structure and synthesis of the ribosomal ribonucleic acid of prokaryotes. Bacteriol. Rev. 37, 562–603. https://doi.org/10.1128/br.37.4.562-603.1973 (1973).
Bonamy, C., Hirschbein, L. & Szulmajster, J. Synthesis of ribosomal ribonucleic acid during sporulation of Bacillus subtilis. J. Bacteriol. 113, 1296–1306. https://doi.org/10.1128/jb.113.3.1296-1306.1973 (1973).
Zhu, B. et al. Uncoupling of macrophage inflammation from self-renewal modulates host recovery from respiratory viral infection. Immunity 54, 1200-1218.e1209. https://doi.org/10.1016/j.immuni.2021.04.001 (2021).
Wheaton, A. K., Agarwal, M., Jia, S. & Kim, K. K. Lung epithelial cell focal adhesion kinase signaling inhibits lung injury and fibrosis. Am. J. Physiol. Lung Cell Mol. Physiol. 312, L722-l730. https://doi.org/10.1152/ajplung.00478.2016 (2017).
Hu, X. et al. PI3K-Akt-mTOR/PFKFB3 pathway mediated lung fibroblast aerobic glycolysis and collagen synthesis in lipopolysaccharide-induced pulmonary fibrosis. Lab. Invest. 100, 801–811. https://doi.org/10.1038/s41374-020-0404-9 (2020).
Spragge, F. et al. Microbiome diversity protects against pathogens by nutrient blocking. Science 382, eadj3502. https://doi.org/10.1126/science.adj3502 (2023).
Heilbronner, S., Krismer, B., Brötz-Oesterhelt, H. & Peschel, A. The microbiome-shaping roles of bacteriocins. Nat. Rev. Microbiol. 19, 726–739. https://doi.org/10.1038/s41579-021-00569-w (2021).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457. https://doi.org/10.1038/nmeth.3337 (2015).
Aran, D., Hu, Z. & Butte, A. J. xCell: Digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 220. https://doi.org/10.1186/s13059-017-1349-1 (2017).
Sturm, G. et al. Comprehensive evaluation of transcriptome-based cell-type quantification methods for immuno-oncology. Bioinformatics 35, i436–i445. https://doi.org/10.1093/bioinformatics/btz363 (2019).
Strehl, C., Ehlers, L., Gaber, T. & Buttgereit, F. Glucocorticoids—all-rounders tackling the versatile players of the immune system. Front. Immunol. 10, 1744. https://doi.org/10.3389/fimmu.2019.01744 (2019).
Idris, T. et al. Akt-driven TGF-β and DKK1 secretion impairs F508del cystic fibrosis airway epithelium polarity. Am. J. Respir. Cell Mol. Biol. 71, 81–94. https://doi.org/10.1165/rcmb.2023-0408OC (2024).
Zhou, F., Onizawa, S., Nagai, A. & Aoshiba, K. Epithelial cell senescence impairs repair process and exacerbates inflammation after airway injury. Respir. Res. 12, 78. https://doi.org/10.1186/1465-9921-12-78 (2011).
Zhou, X. et al. Mast cell chymase impairs bronchial epithelium integrity by degrading cell junction molecules of epithelial cells. Allergy 74, 1266–1276. https://doi.org/10.1111/all.13666 (2019).
Le, Y., Zhou, Y., Iribarren, P. & Wang, J. Chemokines and chemokine receptors: Their manifold roles in homeostasis and disease. Cell. Mol. Immunol. 1, 95–104 (2004).
Svitkina, T. M. & Borisy, G. G. Arp2/3 complex and actin depolymerizing factor/cofilin in dendritic organization and treadmilling of actin filament array in lamellipodia. J. Cell Biol. 145, 1009–1026. https://doi.org/10.1083/jcb.145.5.1009 (1999).
MacDonald, B. T., Tamai, K. & He, X. Wnt/beta-catenin signaling: Components, mechanisms, and diseases. Dev. Cell 17, 9–26. https://doi.org/10.1016/j.devcel.2009.06.016 (2009).
Engelman, J. A., Luo, J. & Cantley, L. C. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat. Rev. Genet. 7, 606–619. https://doi.org/10.1038/nrg1879 (2006).
Yuan, T. L. & Cantley, L. C. PI3K pathway alterations in cancer: Variations on a theme. Oncogene 27, 5497–5510. https://doi.org/10.1038/onc.2008.245 (2008).
Song, C. et al. Evidence for the critical role of the PI3K signaling pathway in particulate matter-induced dysregulation of the inflammatory mediators COX-2/PGE(2) and the associated epithelial barrier protein filaggrin in the bronchial epithelium. Cell Biol. Toxicol. 36, 301–313. https://doi.org/10.1007/s10565-019-09508-1 (2020).
Pollard, T. D. & Cooper, J. A. Actin, a central player in cell shape and movement. Science 326, 1208–1212. https://doi.org/10.1126/science.1175862 (2009).
Sun, T. et al. TAZ is required for lung alveolar epithelial cell differentiation after injury. JCI Insight https://doi.org/10.1172/jci.insight.128674 (2019).
Loo, L. et al. Fibroblast-expressed LRRC15 is a receptor for SARS-CoV-2 spike and controls antiviral and antifibrotic transcriptional programs. PLoS Biol. 21, e3001967. https://doi.org/10.1371/journal.pbio.3001967 (2023).
Song, T. Y. et al. Tumor evolution selectively inactivates the core microRNA machinery for immune evasion. Nat. Commun. 12, 7003. https://doi.org/10.1038/s41467-021-27331-3 (2021).
Huang, W. et al. ZC3HAV1 promotes the proliferation and metastasis via regulating KRAS in pancreatic cancer. Aging (Albany NY) 13, 18482–18497. https://doi.org/10.18632/aging.203296 (2021).
Funding
This work was supported by Research grants from the Shanghai Science and Technology Committee (20Y11900400) and the National Key Research and Development Program of China (2022YFC2009802, 2022YFC2009800, 2022YFC2009801).
Author information
Authors and Affiliations
Contributions
Z.F.F. and Y.H.S. wrote the main manuscript text. H.J.Y. and J.H. prepared Figs. 1, 2 and 3. Q.H.L., Q.Z., Y.Z., N.J., J.W.A., J.L.J., and W.H.Z. contributed to data analysis and interpretation. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors have read the journal’s policy on disclosure of potential conflicts of interest and have none to declare.
Ethical approval and consent to participants
All participants in the parent trial agreed to participate in this study and provided written informed consent (Ky2020-1304).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fu, Z., Sun, Y., Yao, H. et al. A diagnostic model based on pulmonary microbiota and host gene expression to distinguish colonization from pneumonia. Sci Rep (2026). https://doi.org/10.1038/s41598-026-44972-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-44972-w