Abstract
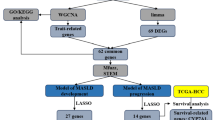
Type 2 diabetes mellitus (T2DM) is a major risk factor for metabolic dysfunction-associated steatotic liver disease (MASLD), and their convergence presents a significant therapeutic challenge. Palmatine, an isoquinoline alkaloid with lipid-regulating and anti-inflammatory properties, is a promising candidate; however, its multi-target mechanisms remain undefined. To address this, we employed a sequential, multi-omics bioinformatics workflow to generate a testable mechanistic hypothesis, followed by rigorous experimental validation. First, integrative analyses—including target prediction, differential gene expression, pathway enrichment, and machine learning—converged to identify five core targets: ADRB2, BCL3, EGR1, FOS, and MAP3K8. Molecular docking predicted strong binding, and single-cell sequencing contextualized their expression within specific hepatic cell types. To experimentally test this predictive framework, we evaluated palmatine in a rat model of T2DM-associated MASLD. Palmatine treatment significantly improved liver function(reduced ALT, AST), attenuated inflammation (lowered TNF-α, IL-6) and oxidative stress (increased SOD, GSH; decreased MDA), ameliorated glycolipid metabolism (reduced TC, TG, LDL-C, and GLU), and reduced hepatic steatosis and fibrosis. Mechanistically, confirming the bioinformatic prediction, palmatine downregulated the expression of the five key targets and concurrently suppressed the activation of critical apoptotic executers (Caspase-3, Caspase-8, GSDME). These findings demonstrate that palmatine alleviates MASLD by modulating a novel target network to inhibit hepatocyte apoptosis, providing a robust, hypothesis-driven preclinical foundation for its therapeutic development.
Similar content being viewed by others
Data availability
Data will be available upon request from the corresponding author.
Abbreviations
- ACSL1:
-
Acyl-CoA Synthetase Long-Chain Family Member 1
- ADRB2:
-
Adrenoceptor Beta 2
- AKT:
-
AKT Serine/Threonine Kinase
- ALT:
-
Alanine Aminotransferase
- AMPK:
-
Liver Adenosine Monophosphate-Activated Protein Kinase
- ASK1:
-
Apoptosis Signal-regulating Kinase 1
- AST:
-
Aspartate Aminotransferase
- ATF4:
-
Activating Transcription Factor 4
- BCA:
-
Bicinchoninic Acid Assay
- BCL3:
-
BCL3 Transcription Coactivator
- BSA:
-
Bovine Serum Albumin
- BP:
-
BiologicalProcess
- β-arrestin2:
-
Beta-Arrestin 2
- CC:
-
CellularComponent
- cAMP:
-
Cyclic Adenosine Monophosphate
- CDS:
-
CellDynamicSimulation
- C/EBP-α:
-
CCAAT/Enhancer-Binding Protein Alpha
- cGAS-STING:
-
cyclic GMP-AMP synthase-stimulator of interferon genes
- DAMPs:
-
Damage-Associated Molecular Patterns
- DAPI:
-
4’6-Diamidino-2-phenylindole
- DEGs:
-
Differentially Expressed Genes
- DNL:
-
De Novo Lipogenesis
- DT:
-
Decision Tree
- ECL:
-
Enhanced Chemiluminescence
- EGR1:
-
Early Growth Response 1
- eIF2α:
-
eukaryotic Initiation Factor 2 alpha
- ERK:
-
Extracellular Signal-Regulated Kinase
- FADD:
-
Fas Associated Via Death Domain
- FBG:
-
Fasting Blood Glucose
- FFA:
-
Free Fatty Acids
- FAS:
-
FAS Cell Surface Death Receptor
- FOS:
-
AP-1 Transcription Factor Subunit
- FXR:
-
Farnesoid X Receptor Agonist
- GF-β1:
-
Transforming Growth Factor-beta 1
- GBM:
-
GradientBoostingMachine
- GEO:
-
Gene Expression Omnibus Database
- Gi:
-
G inhibitory protein
- GLM:
-
GeneralizedLinearModel
- GLP-1:
-
Glucagon-Like Peptide-1 Receptor Agonist
- GLU:
-
Glucose
- GSDME:
-
Gasdermin E
- GSH:
-
glutathione
- GO:
-
Gene Ontology
- HMGB1:
-
High Mobility Group Box 1
- HE:
-
Hematoxylin and Eosin Staining
- HepG2:
-
Hepatoma Cell Line G2
- HSCs:
-
Hepatic Stellate Cells
- HRP:
-
Horseradish Peroxidase
- IGT:
-
Impaired Glucose Tolerance
- IL-1β:
-
Interleukin-1 Beta
- IL-6:
-
Interleukin-6
- IF:
-
Immunofluorescence Staining
- IR:
-
Insulin Resistance
- JNK:
-
c-Jun N-terminal Kinase
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- KNN:
-
K-NearestNeighbors
- LASSO:
-
LeastAbsoluteShrinkageandSelectionOperator
- LDL-C:
-
Low-Density Lipoprotein Cholesterol
- mTOR:
-
Mechanistic Target of Rapamycin
- mTORC1:
-
Mechanistic Target of Rapamycin Complex 1
- MAP3K8:
-
Mitogen-Activated Protein Kinase Kinase Kinase 8
- MDA:
-
Malondialdehyde
- MF:
-
MolecularFunction
- MyD88:
-
Myeloid Differentiation Primary Response 88
- NAFLD:
-
Non-Alcoholic Fatty Liver Disease
- MASLD:
-
Metabolic dysfunction-associated steatotic liver disease
- NASH:
-
Non-Alcoholic Steatohepatitis
- NF-κB:
-
Nuclear Factor kappa-light-chain-enhancer of activated B cells
- NLRP3:
-
NACHT LRR and PYD domains-containing protein 3
- NNET:
-
NeuralNetwork
- OTCC:
-
Optimal Temperature Cutting Compound
- Pal:
-
palmatine
- PERK:
-
PKR-like Endoplasmic Reticulum Kinase
- PI3K:
-
Phosphatidylinositol 3-Kinase
- PBS:
-
Phosphate Buffered Saline
- PKA:
-
cAMP-dependent Protein Kinase
- PDE4:
-
Phosphodiesterase 4
- PPAR-γ:
-
Peroxisome Proliferator-Activated Receptor Gamma
- PUMA:
-
p53 Upregulated Modulator of Apoptosis
- PVDF:
-
Polyvinylidene Fluoride
- RIPA:
-
Radioimmunoprecipitation Assay Buffer
- ROC:
-
Receiver Operating Characteristic Curve
- RF:
-
RandomForest
- SOD:
-
Superoxide Dismutase
- Sem:
-
Semaglutide
- Smad3:
-
Mothers Against Decapentaplegic Homolog 3
- SREBP-1c:
-
Sterol Regulatory Element-Binding Protein 1c
- STZ:
-
Streptozotocin
- ssGSEA:
-
Single-Sample Gene Set Enrichment Analysis
- SVM:
-
SupportVectorMachine
- TBA:
-
Total Bile Acids
- T-C:
-
Total Cholesterol
- TAK1:
-
TGF-β-Activated Kinase 1
- T-G:
-
Triglycerides
- TLR4:
-
Toll-Like Receptor 4
- TNF-α:
-
Tumor Necrosis Factor Alpha
- T2DM:
-
Type 2 Diabetes Mellitus
- WB:
-
Western Blot
- XGB:
-
eXtremeGradientBoosting
References
Younossi, Z. M. et al. Global epidemiology of nonalcoholic fatty liver disease-Meta-analytic assessment of prevalence, incidence, and outcomes. Hepatology 64 (1), 73–84 (2016).
Konings, L. A. M. et al. Pharmacological treatment options for metabolic dysfunction-associated steatotic liver disease in patients with type 2 diabetes mellitus: A systematic review. Eur. J. Clin. Invest. 55(4), e70003 (2025).
EASL-EASD-EASO Clinical Practice Guidelines on the. Management of Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD). Obes. Facts. 17 (4), 374–444 (2024).
Eslam, M., Sanyal, A. J. & George, J. A Consensus-Driven Proposed Nomenclature for Metabolic Associated Fatty Liver Disease. Gastroenterology 158 (7), 1999–2014e1 (2020).
Ji, Y., Wang, Q., Jiang, Y. & Liu, B. Global epidemiology of T2DM in patients with NAFLD or MAFLD: the real situation may be even more serious. BMC Med. 22 (1), 476 (2024).
Cao, L. et al. Global epidemiology of type 2 diabetes in patients with NAFLD or MAFLD: a systematic review and meta-analysis. BMC Med. 22 (1), 101 (2024).
Stefan, N. & Cusi, K. A global view of the interplay between non-alcoholic fatty liver disease and diabetes. Lancet Diabetes Endocrinol. 10 (4), 284–296 (2022).
Wang, Y. et al. Analysis of Immune and Inflammatory Microenvironment Characteristics of Noncancer End-Stage Liver Disease. J. Interferon Cytokine Res. 43 (2), 86–97 (2023).
Duell, P. B. et al. Nonalcoholic Fatty Liver Disease and Cardiovascular Risk: A Scientific Statement From the American Heart Association. Arterioscler. Thromb. Vasc Biol. 42 (6), e168–e185 (2022).
Sarker, J., Okpara, E., De Los Santos, B., Kim, M. & Kim, K. Comparative effectiveness of GLP-1 receptor agonists, SGLT2 inhibitors and DPP-4 inhibitors on liver outcomes in metabolic dysfunction-associated steatotic liver disease: A retrospective cohort study. Diabetes Obes. Metab. 28(4), 3155–3164 (2026).
Tong, G. et al. The efficacy of sulodexide combined with Jinshuibao for treating early diabetic nephropathy patients. Int. J. Clin. Exp. Med. 13 (11), 8308–8317 (2020).
Cegla, J. Liraglutide safety and efficacy in patients with non-alcoholic steatohepatitis (LEAN): a multicentre, double-blind, randomized, placebo-controlled phase 2 study. Ann. Clin. Biochem. 53 (4), 518 (2016).
Li, W. et al. The signaling pathways of selected traditional Chinese medicine prescriptions and their metabolites in the treatment of diabetic cardiomyopathy: a review. Front. Pharmacol. 15, 1416403 (2024).
Choi, S. et al. Convergent metabolic pathways in MASH therapeutics: An AMPK-centric analysis. J. Cell. Mol. Med. 30(2), e71023 (2026).
Tian, X. et al. Palmatine ameliorates high fat diet induced impaired glucose tolerance. Biol. Res. 53 (1), 39 (2020).
Nwabueze, O. P. et al. M. P., Comparative Studies of Palmatine with Metformin and Glimepiride on the Modulation of Insulin Dependent Signaling Pathway In Vitro, In Vivo & Ex Vivo. Pharmaceuticals (Basel) 15, (11). (2022).
Long, J. et al. Palmatine: A review of its pharmacology, toxicity and pharmacokinetics. Biochimie 162, 176–184 (2019).
Yin, X., Liu, Z. & Wang, J. Tetrahydropalmatine ameliorates hepatic steatosis in nonalcoholic fatty liver disease by switching lipid metabolism via AMPK-SREBP-1c-Sirt1 signaling axis. Phytomedicine 119, 155005 (2023).
Cheng, J. J. et al. Palmatine Protects Against MSU-Induced Gouty Arthritis via Regulating the NF-κB/NLRP3 and Nrf2 Pathways. Drug Des. Devel Ther. 16, 2119–2132 (2022).
Zhang, N. et al. Discovery and development of palmatine analogues as anti-NASH agents by activating farnesoid X receptor (FXR). Eur. J. Med. Chem. 245 (Pt 1), 114886 (2023).
Choi, J. S. et al. Coptis chinensis alkaloids exert anti-adipogenic activity on 3T3-L1 adipocytes by downregulating C/EBP-α and PPAR-γ. Fitoterapia 98, 199–208 (2014).
Zhang, S., Liu, K., Liu, Y., Hu, X. & Gu, X. The role and application of bioinformatics techniques and tools in drug discovery. Front. Pharmacol. 16, 1547131 (2025).
Altman, R. B. & Klein, T. E. Challenges for biomedical informatics and pharmacogenomics. Annu. Rev. Pharmacol. Toxicol. 42, 113–133 (2002).
Qian, Y., Wang, Q., Yin, L. & Lu, A. Y. CF-DTI: coarse-to-fine feature extraction for enhanced drug-target interaction prediction. Health Inf. Sci. Syst. 13 (1), 55 (2025).
Ru, J. et al. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J. Cheminform. 6, 13 (2014).
Wang, X. et al. PharmMapper 2017 update: a web server for potential drug target identification with a comprehensive target pharmacophore database. Nucleic Acids Res. 45 (W1), W356–w360 (2017).
Fang, S. et al. HERB: a high-throughput experiment- and reference-guided database of traditional Chinese medicine. Nucleic Acids Res. 49 (D1), D1197–d1206 (2021).
Wu, Y. et al. SymMap: an integrative database of traditional Chinese medicine enhanced by symptom mapping. Nucleic Acids Res. 47 (D1), D1110–d1117 (2019).
UniProt. the Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 53 (D1), D609–d617 (2025).
Cheng, M. et al. Exploring the mechanism of PPCPs on human metabolic diseases based on network toxicology and molecular docking. Environ. Int. 196, 109324 (2025).
He, Q. et al. Exploring the mechanism of curcumin in the treatment of colon cancer based on network pharmacology and molecular docking. Front. Pharmacol. 14, 1102581 (2023).
Clough, E. & Barrett, T. The Gene Expression Omnibus Database. Methods Mol. Biol. 1418, 93–110 (2016).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43(7), e47 (2015).
Rosario, S. R. et al. Pan-cancer analysis of transcriptional metabolic dysregulation using The Cancer Genome Atlas. Nat. Commun. 9 (1), 5330 (2018).
Qin, H. et al. Integrated machine learning survival framework develops a prognostic model based on inter-crosstalk definition of mitochondrial function and cell death patterns in a large multicenter cohort for lower-grade glioma. J. Transl Med. 21 (1), 588 (2023).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods. 12 (5), 453–457 (2015).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177(7), 1888–1902e21 (2019).
Zappia, L. & Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. Gigascience 7, (7). (2018).
Xiong, X. et al. Landscape of Intercellular Crosstalk in Healthy and NASH Liver Revealed by Single-Cell Secretome Gene Analysis. Mol. Cell. 75 (3), 644–660e5 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32 (4), 381–386 (2014).
La Manno, G. et al. V., RNA velocity of single cells. Nature 560 (7719), 494–498 (2018).
Ning, Y., Xu, F., Xin, R. & Yao, F. Palmatine regulates bile acid cycle metabolism and maintains intestinal flora balance to maintain stable intestinal barrier. Life Sci. 262, 118405 (2020).
Ning, N. et al. Hypolipidemic Effect and Mechanism of Palmatine from Coptis chinensis in Hamsters Fed High-Fat diet. Phytother Res. 29 (5), 668–673 (2015).
Tong, J., Ding, Y. & Zhang, D. Mechanisms of Actinidia chinensis Planch roots in the treatment of breast cancer based on network pharmacology and molecular docking. Med. (Baltim). 104(31), e43560 (2025).
de Oliveira, A. S., Muniz Seif, E. J. & da Silva Junior, P. I. In silico prospection of receptors associated with the biological activity of U1-SCTRX-lg1a: an antimicrobial peptide isolated from the venom of Loxosceles gaucho. Silico Pharmacol. 12 (1), 15 (2024).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53 (D1), D672–d677 (2025).
Cheng, X. et al. Quercetin: A promising therapy for diabetic encephalopathy through inhibition of hippocampal ferroptosis. Phytomedicine 126, 154887 (2024).
Singh, A. B., Dong, B., Xu, Y., Zhang, Y. & Liu, J. Identification of a novel function of hepatic long-chain acyl-CoA synthetase-1 (ACSL1) in bile acid synthesis and its regulation by bile acid-activated farnesoid X receptor. Biochim. Biophys. Acta Mol. Cell. Biol. Lipids. 1864 (3), 358–371 (2019).
Schuster-Gaul, S. et al. ASK1 inhibition reduces cell death and hepatic fibrosis in an Nlrp3 mutant liver injury model. JCI Insight 5, (2). (2020).
Hotamisligil, G. S. Endoplasmic reticulum stress and the inflammatory basis of metabolic disease. Cell 140 (6), 900–917 (2010).
Pan, J. et al. Fatty acid activates NLRP3 inflammasomes in mouse Kupffer cells through mitochondrial DNA release. Cell. Immunol. 332, 111–120 (2018).
Friedman, S. L. Mechanisms of hepatic fibrogenesis. Gastroenterology 134 (6), 1655–1669 (2008).
Wang, F. et al. Canonical Wnt signaling promotes HSC glycolysis and liver fibrosis through an LDH-A/HIF-1α transcriptional complex. Hepatology 79 (3), 606–623 (2024).
Duan, Y. et al. Crosstalk in extrahepatic and hepatic system in NAFLD/NASH. Liver Int. 44 (8), 1856–1871 (2024).
Ren, W. et al. Glutamine Metabolism in Macrophages: A Novel Target for Obesity/Type 2 Diabetes. Adv. Nutr. 10 (2), 321–330 (2019).
Liu, J., Wang, G., Jia, Y. & Xu, Y. GLP-1 receptor agonists: effects on the progression of non-alcoholic fatty liver disease. Diabetes Metab. Res. Rev. 31 (4), 329–335 (2015).
He, J. et al. Graveoline attenuates D-GalN/LPS-induced acute liver injury via inhibition of JAK1/STAT3 signaling pathway. Biomed. Pharmacother. 177, 117163 (2024).
Hindson, J. Obeticholic acid for the treatment of NASH. Nat. Rev. Gastroenterol. Hepatol. 17 (2), 66 (2020).
Guicciardi, M. E. & Gores, G. J. Bile acid-mediated hepatocyte apoptosis and cholestatic liver disease. Dig. Liver Dis. 34 (6), 387–392 (2002).
Angelim, M. & Moraes-Vieira, P. M. Glycolysis modulates efferocytosis in a noncanonical manner. Nat. Metab. 5 (3), 360–361 (2023).
Hambright, H. G., Meng, P., Kumar, A. P. & Ghosh, R. Inhibition of PI3K/AKT/mTOR axis disrupts oxidative stress-mediated survival of melanoma cells. Oncotarget 6 (9), 7195–7208 (2015).
Yu, J. et al. Extracellular vesicles derived from menstrual blood-derived mesenchymal stem cells suppress inflammatory atherosclerosis by inhibiting NF-κB signaling. BMC Med. 23 (1), 565 (2025).
Mohammadpour, H., MacDonald, C. R., McCarthy, P. L., Abrams, S. I. & Repasky, E. A. β2-adrenergic receptor signaling regulates metabolic pathways critical to myeloid-derived suppressor cell function within the TME. Cell. Rep. 37 (4), 109883 (2021).
Zhang, Q., Didonato, J. A., Karin, M. & McKeithan, T. W. BCL3 encodes a nuclear protein which can alter the subcellular location of NF-kappa B proteins. Mol. Cell. Biol. 14 (6), 3915–3926 (1994).
Wu, F., Zhang, P. & Zhou, G. The involvement of EGR1 in neuron apoptosis in the in vitro model of spinal cord injury via BTG2 up-regulation. Neurol. Res. 45 (7), 646–654 (2023).
Li, X., Meng, Y., Wu, P., Zhang, Z. & Yang, X. Angiotensin II and Aldosterone stimulating NF-kappaB and AP-1 activation in hepatic fibrosis of rat. Regul. Pept. 138 (1), 15–25 (2007).
Wu, K. C. et al. Tpl2 kinase regulates inflammation but not tumorigenesis in mice. Toxicol. Appl. Pharmacol. 418, 115494 (2021).
Fleischmann, A., Jochum, W., Eferl, R., Witowsky, J. & Wagner, E. F. Rhabdomyosarcoma development in mice lacking Trp53 and Fos: tumor suppression by the Fos protooncogene. Cancer Cell. 4 (6), 477–482 (2003).
Schonthaler, H. B., Guinea-Viniegra, J. & Wagner, E. F. Targeting inflammation by modulating the Jun/AP-1 pathway. Ann. Rheum. Dis. 70 (Suppl 1), i109–i112 (2011).
Liu, S. et al. Melatonin mitigates aflatoxin B1-induced liver injury via modulation of gut microbiota/intestinal FXR/liver TLR4 signaling axis in mice. J. Pineal Res. 73(2), e12812 (2022).
Bhat, A. A. et al. The pyroptotic role of Caspase-3/GSDME signalling pathway among various cancer: A Review. Int. J. Biol. Macromol. 242 (Pt 2), 124832 (2023).
Li, L. et al. Photodynamic therapy induces human esophageal carcinoma cell pyroptosis by targeting the PKM2/caspase-8/caspase-3/GSDME axis. Cancer Lett. 520, 143–159 (2021).
Li, R. Y. et al. Cisplatin-induced pyroptosis is mediated via the CAPN1/CAPN2-BAK/BAX-caspase-9-caspase-3-GSDME axis in esophageal cancer. Chem. Biol. Interact. 361, 109967 (2022).
Li, Y. et al. PD-L1 Regulates Platelet Activation and Thrombosis via Caspase-3/GSDME Pathway. Front. Pharmacol. 13, 921414 (2022).
Mai, F. Y. et al. Caspase-3-mediated GSDME activation contributes to cisplatin- and doxorubicin-induced secondary necrosis in mouse macrophages. Cell. Prolif. 52(5), e12663 (2019).
Mouasni, S. & Tourneur, L. FADD at the Crossroads between Cancer and Inflammation. Trends Immunol. 39 (12), 1036–1053 (2018).
Werner, M. H., Wu, C. & Walsh, C. M. Emerging roles for the death adaptor FADD in death receptor avidity and cell cycle regulation. Cell. Cycle. 5 (20), 2332–2338 (2006).
Scott, F. L. et al. The Fas-FADD death domain complex structure unravels signalling by receptor clustering. Nature 457 (7232), 1019–1022 (2009).
Lavrik, I. et al. The active caspase-8 heterotetramer is formed at the CD95 DISC. Cell. Death Differ. 10 (1), 144–145 (2003).
Li, J. et al. Integrating transcriptomics and network pharmacology to reveal the mechanisms of total Rhizoma Coptidis alkaloids against nonalcoholic steatohepatitis. J. Ethnopharmacol. 322, 117600 (2024).
Lin, G. S. et al. Palmatine attenuates hepatocyte injury by promoting autophagy via the AMPK/mTOR pathway after alcoholic liver disease. Drug Dev. Res. 83 (7), 1613–1622 (2022).
Ai, X. et al. Berberis dictyophylla F. inhibits angiogenesis and apoptosis of diabetic retinopathy via suppressing HIF-1α/VEGF/DLL-4/Notch-1 pathway. J. Ethnopharmacol. 296, 115453 (2022).
Liu, Z. et al. Palmatine ameliorates cisplatin-induced acute kidney injury through regulating Akt and NF-κB/MAPK pathways. Arab. J. Chem. 17 (5), 105731 (2024).
Zhang, W. et al. Antiviral effect of palmatine against infectious bronchitis virus through regulation of NF-κB/IRF7/JAK-STAT signalling pathway and apoptosis. Br. Poult. Sci. 65 (2), 119–128 (2024).
Selase, A., Cynthia, D. A. & Newman, O. et al. Palmatine modulates triple negative mammary carcinoma by regulating the endogenous function of P53, P21 and Mdm2. Biomed. Pharmacol. J. 14(2), 943–954 (2021).
Acknowledgements
Not applicable.
Funding
The author(s) declare that financial support was received for the research, authorship, and/or publication of this article. This research was funded by the National Natural Science Foundation of China (grant numbers 81873231 and 82474381).
Author information
Authors and Affiliations
Contributions
Huasen Yang: Writing–original draft, Formal analysis, Visualization. Zhoujing Shi: Software, Validation. Yazhi Qi: Investigation, Validation. Shuchang Bao: Investigation, Validation. Chaochong Li: Software, Conceptualization. Junhui Mei: Software, Conceptualization. Mingshuang Sun: Software, Conceptualization. Yusheng Han: Supervision, Writing–review and editing. Boyan Ma: Funding acquisition, Project administration. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
This study is reported in accordance with the ARRIVE guidelines and was approved by the Ethics Committee of Heilongjiang University of Traditional Chinese Medicine (Institutional Animal Use License: SYXK-2020-004; ethical approval number: 2023113008).
Euthanasia method
Intravenous injection of Zoletil™50, followed by cervical dislocation to induce death after anesthesia.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, H., Shi, Z., Qi, Y. et al. Palmatine ameliorates MASLD in type 2 diabetes by modulating hepatic apoptosis and inflammation. Sci Rep (2026). https://doi.org/10.1038/s41598-026-45476-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-45476-3