Abstract
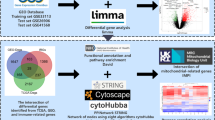
Colorectal cancer (CRC) is one of the most common malignant tumors worldwide. Patients with different immunophenotypes of CRC could achieve different effect of immunotherapy and yield different prognosis. With the advancement of bioinformatics, multi-omics analysis of the variations at both genomics and epigenomics levels helps a lot to provide a molecular basis for immunophenotype. Gene expression and clinical data of CRC patients were obtained from The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO). We calculated Spearman correlation of CD274 (programmed cell death ligand-1, PD-L1) and IFNG (interferon gamma, IFN-γ) expressions with immune cell fraction, and screened different immune cell types with CD274 and IFNG by Lasso regression analysis. Multi-omics analysis was exploited to screen out candidate genes with differential in genetic and epigenetic landscapes between two CRC subtypes with the greatest difference in immune infiltration. Finally, a risk scoring model was established and the role of candidate genes in prognosis and oncoimmunology was evaluated at the pan-cancer level. Two CRC types (cluster A and cluster B) including five subtypes (subclusters A1, A2, B1, B2A, and B2B) were identified by unsupervised clustering analysis. Somatic mutations, CNVs, and DNA methylation differed between subcluster A2 and B2, and analysis of DEGs correlated with CRC immune phenotypes identified FUT9 and MS4A3 as key genes related to CRC immune-phenotypes and prognosis. Furthermore, FUT9 was validated to act as a key gene related to CRC immune escape in vitro. The present study established a risk model for CRC immunophenotyping and prognosis, and highlighted the significance of FUT9 and MS4A3 in oncoimmunology of CRC.
Similar content being viewed by others
Data availability
All data supporting the conclusions of this article are included within the article and its supplementary files. Further inquiries can be directed to the corresponding authors.
References
Sanchez-Gundin, J., Fernandez-Carballido, A. M., Martinez-Valdivieso, L., Barreda-Hernandez, D. & Torres-Suarez, A. I. New trends in the therapeutic approach to metastatic colorectal cancer. Int. J. Med. Sci. 15, 659–665. https://doi.org/10.7150/ijms.24453 (2018).
Ferlay, J. et al. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, E359-386. https://doi.org/10.1002/ijc.29210 (2015).
Van Cutsem, E., Cervantes, A., Nordlinger, B., Arnold, D., Group, E. G. W. Metastatic colorectal cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 25(Suppl 3), iii1-9. https://doi.org/10.1093/annonc/mdu260 (2014).
Prendergast, G. C. & Jaffee, E. M. Cancer immunologists and cancer biologists: Why we didn’t talk then but need to now. Cancer Res. 67, 3500–3504. https://doi.org/10.1158/0008-5472.CAN-06-4626 (2007).
Tauriello, D. V., Calon, A., Lonardo, E. & Batlle, E. Determinants of metastatic competency in colorectal cancer. Mol. Oncol. 11, 97–119. https://doi.org/10.1002/1878-0261.12018 (2017).
Xu, X. et al. Multi-omics analysis to identify driving factors in colorectal cancer. Epigenomics 12, 1633–1650. https://doi.org/10.2217/epi-2020-0073 (2020).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782. https://doi.org/10.1038/s41587-019-0114-2 (2019).
Koboldt, D. C. et al. VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576. https://doi.org/10.1101/gr.129684.111 (2012).
Mayakonda, A., Lin, D. C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756. https://doi.org/10.1101/gr.239244.118 (2018).
Sun, W. et al. TSVdb: A web-tool for TCGA splicing variants analysis. BMC Genomics 19, 405. https://doi.org/10.1186/s12864-018-4775-x (2018).
Larsen, S. J., do Canto, L. M., Rogatto, S. R. & Baumbach, J. CoNVaQ: A web tool for copy number variation-based association studies. BMC Genomics 19, 369. https://doi.org/10.1186/s12864-018-4732-8 (2018).
Quinlan, A. R. & Hall, I. M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842. https://doi.org/10.1093/bioinformatics/btq033 (2010).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47. https://doi.org/10.1093/nar/gkv007 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7. https://doi.org/10.1186/1471-2105-14-7 (2013).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612. https://doi.org/10.1038/ncomms3612 (2013).
Qi, X. et al. ceRNA in cancer: Possible functions and clinical implications. J. Med. Genet. 52, 710–718. https://doi.org/10.1136/jmedgenet-2015-103334 (2015).
Tay, Y., Rinn, J. & Pandolfi, P. P. The multilayered complexity of ceRNA crosstalk and competition. Nature 505, 344–352. https://doi.org/10.1038/nature12986 (2014).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558. https://doi.org/10.1038/s41591-018-0136-1 (2018).
Cen, B. et al. Mutant APC promotes tumor immune evasion via PD-L1 in colorectal cancer. Oncogene 40, 5984–5992. https://doi.org/10.1038/s41388-021-01972-6 (2021).
Liu, Y. T. & Sun, Z. J. Turning cold tumors into hot tumors by improving T-cell infiltration. Theranostics 11, 5365–5386. https://doi.org/10.7150/thno.58390 (2021).
He, Q. et al. Genome-wide prediction of cancer driver genes based on SNP and cancer SNV data. Am. J. Cancer Res. 4, 394–410 (2014).
Fernandez-Rozadilla, C. et al. A genome-wide association study on copy-number variation identifies a 11q11 loss as a candidate susceptibility variant for colorectal cancer. Hum. Genet. 133, 525–534. https://doi.org/10.1007/s00439-013-1390-4 (2014).
Yang, Z. et al. Multiomics analysis on DNA methylation and the expression of both messenger RNA and microRNA in lung adenocarcinoma. J. Cell. Physiol. 234, 7579–7586. https://doi.org/10.1002/jcp.27520 (2019).
Liang, Y., Buckley, T. R., Tu, L., Langdon, S. D. & Tedder, T. F. Structural organization of the human MS4A gene cluster on Chromosome 11q12. Immunogenetics 53, 357–368. https://doi.org/10.1007/s002510100339 (2001).
Kutok, J. L., Yang, X., Folkerth, R. & Adra, C. N. Characterization of the expression of HTm4 (MS4A3), a cell cycle regulator, in human peripheral blood cells and normal and malignant tissues. J. Cell. Mol. Med. 15, 86–93. https://doi.org/10.1111/j.1582-4934.2009.00925.x (2011).
Nakajima, T. et al. Identification of granulocyte subtype-selective receptors and ion channels by using a high-density oligonucleotide probe array. J. Allergy Clin. Immunol. 113, 528–535. https://doi.org/10.1016/j.jaci.2003.12.036 (2004).
Donato, J. L. et al. Human HTm4 is a hematopoietic cell cycle regulator. J. Clin. Invest. 109, 51–58. https://doi.org/10.1172/JCI14025 (2002).
Heller, G. et al. EVI1 promotes tumor growth via transcriptional repression of MS4A3. J. Hematol. Oncol. 8, 28. https://doi.org/10.1186/s13045-015-0124-6 (2015).
Zhao, H. et al. MS4A3 promotes differentiation in chronic myeloid leukemia by enhancing common β-chain cytokine receptor endocytosis. Blood 139, 761–778. https://doi.org/10.1182/blood.2021011802 (2022).
Tu, Z., Lin, Y. N. & Lin, C. H. Development of fucosyltransferase and fucosidase inhibitors. Chem. Soc. Rev. 42, 4459–4475. https://doi.org/10.1039/c3cs60056d (2013).
Kashiwazaki, H. et al. Mice lacking alpha1,3-fucosyltransferase 9 exhibit modulation of in vivo immune responses against pathogens. Pathol. Int. 64, 199–208. https://doi.org/10.1111/pin.12159 (2014).
Seagle, B. L. et al. Discovery of candidate tumor biomarkers for treatment with intraperitoneal chemotherapy for ovarian cancer. Sci. Rep. 6, 21591. https://doi.org/10.1038/srep21591 (2016).
Liu, S. et al. Differentially expressed mRNAs and their long noncoding RNA regulatory network with Helicobacter pylori-associated diseases including atrophic gastritis and gastric cancer. BioMed. Res. Int. 2020, 3012193. https://doi.org/10.1155/2020/3012193 (2020).
Auslander, N. et al. An integrated computational and experimental study uncovers FUT9 as a metabolic driver of colorectal cancer. Mol. Syst. Biol. 13, 956. https://doi.org/10.15252/msb.20177739 (2017).
Blanas, A. et al. FUT9-driven programming of colon cancer cells towards a stem cell-like state. Cancers (Basel) https://doi.org/10.3390/cancers12092580 (2020).
Wirta, E. V. et al. Immunoscore in mismatch repair-proficient and -deficient colon cancer. J. Pathol. Clin. Res. 3, 203–213. https://doi.org/10.1002/cjp2.71 (2017).
Pages, F. et al. In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J. Clin. Oncol. 27, 5944–5951. https://doi.org/10.1200/JCO.2008.19.6147 (2009).
Anitei, M. G. et al. Prognostic and predictive values of the immunoscore in patients with rectal cancer. Clin. Cancer Res. 20, 1891–1899. https://doi.org/10.1158/1078-0432.CCR-13-2830 (2014).
Francini, E. et al. The prognostic value of CD3 + tumor-infiltrating lymphocytes for stage II colon cancer according to use of adjuvant chemotherapy: A large single-institution cohort study. Transl. Oncol. 14, 100973. https://doi.org/10.1016/j.tranon.2020.100973 (2021).
Kroemer, M. et al. Investigation of the prognostic value of CD4 T cell subsets expanded from tumor-infiltrating lymphocytes of colorectal cancer liver metastases. J. Immunother. Cancer https://doi.org/10.1136/jitc-2020-001478 (2020).
Wang, W. et al. Correlation between schistosomiasis and CD8 + T cell and stromal PD-L1 as well as the different prognostic role of CD8 + T cell and PD-L1 in schistosomal-associated colorectal cancer and non-schistosomal-associated colorectal cancer. World J. Surg. Oncol. 19, 321. https://doi.org/10.1186/s12957-021-02433-w (2021).
Acknowledgements
We acknowledge public databases including TCGA and GEO for providing their platforms and contributors for uploading their meaningful datasets. We thank Charlesworth Author Services for the English language editing of the article.
Funding
This work was supported by the National Natural Science Foundation of China (grant number: 82303594), Zhejiang Medical and Health Science and Technology Plan (grant number: 2024XY091 and 2019RC175) and Zhejiang Provincial County Level Advantageous Disciplines of Traditional Chinese Medicine Construction Plan (2023-XK-D040).
Author information
Authors and Affiliations
Contributions
ZF and XX conceived the idea and handled bioinformatic analyses, designed and monitored the research. MZ wrote the main manuscript text and prepared the figures and tables. MZ, HD and YH measured *FUT9* expression, MZ and YH assessed and confirmed the staining results in clinical tissue samples. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Tissue microarray chip comprising 94 human CRC and 86 paired adjacent normal tissue samples was obtained from Sanmen People’s Hospital. The study protocol was approved by Sanmen People’s Hospital Ethics Committee (2025-064). All experiments were performed in compliance with the relevant regulations, and all patients provided written informed consent.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, M., Dong, H., Hu, Y. et al. Integrative multi-omics analysis identified FUT9 and MS4A3 as novel immune-phenotype and prognosis biomarkers for colorectal cancer and analyze the role of FUT9 in oncoimmunology. Sci Rep (2026). https://doi.org/10.1038/s41598-026-45508-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-45508-y