Abstract
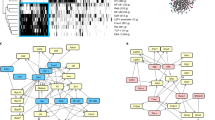
The basic leucine zipper ATF-like transcription factor 3 (BATF3) has been implicated in the pathogenesis of type 1 diabetes mellitus (T1DM), where it may influence immune regulation and pancreatic β-cell homeostasis. Nevertheless, the upstream molecular mechanisms governing BATF3 expression remain largely undefined. Bioinformatic analyses of GEO and UCSC databases were conducted to identify transcription factors potentially regulating BATF3 (GEO: GSE9006 PBMC microarray; newly diagnosed T1D, n = 43; healthy controls, n = 24). Clinical samples (PBMC, n = 30) from T1DM patients and healthy controls were analyzed by qPCR to assess BATF3 and candidate transcription factor expression. Lentiviral transduction and siRNA-mediated knockdown were applied to examine BATF3 regulation and its impact on CD8⁺ T-cell function. Transcription factor–promoter interactions were validated using dual-luciferase reporter assays and ChIP-qPCR. EGR1, EGR2, EGR3, and c-MYC were identified as differentially expressed transcription factors in GSE9006, with c-MYC emerging as the central regulator. Clinical analysis demonstrated significantly elevated expression of c-MYC and BATF3 in T1DM patients compared with healthy controls (n = 30, p < 0.05). In vitro assays confirmed that c-MYC binds to the BATF3 promoter region approximately 1–2 kb upstream of the transcription start site, thereby promoting BATF3 transcription, enhancing CD8⁺ T-cell proliferation, and inhibiting apoptosis (CD8⁺ T cells isolated from PBMCs of healthy children). ChIP-qPCR further localized the primary c-MYC binding site to the − 1,214 to − 1,203 bp region relative to the BATF3 transcription start site. c-MYC, a critical regulator of BATF3, is markedly elevated in T1DM patients. By driving BATF3 transcription, it promotes CD8⁺ T-cell expansion and limits apoptosis, jointly contributing to pediatric T1DM pathogenesis. These observations highlight the c-MYC–BATF3 axis as a mechanistic pathway relevant to pediatric T1DM and a potential biomarker framework.
Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Gillespie, K. M. Type 1 diabetes: pathogenesis and prevention. CMAJ 175 (2), 165–170 (2006).
Barnett, R. Type 1 diabetes. Lancet 391 (10117), 195 (2018).
Mathieu, C., Gillard, P. & Benhalima, K. Insulin analogues in type 1 diabetes mellitus: getting better all the time. Nat. Rev. Endocrinol. 13 (7), 385–399 (2017).
Sims, E. K. et al. 100 years of insulin: celebrating the past, present and future of diabetes therapy. Nat. Med. 27 (7), 1154–1164 (2021).
Powers, A. C. Type 1 diabetes mellitus: much progress, many opportunities. J. Clin. Invest., 131(8). (2021).
Ejaz, S. & Wilson, T. Managing type 1 diabetes – a journey from starvation to insulin pump. Minerva Endocrinol. 38 (2), 123–131 (2013).
von Scholten, B. J. et al. Current and future therapies for type 1 diabetes. Diabetologia 64 (5), 1037–1048 (2021).
Guilliams, M., Tamura, T. & Decrypting, D. C. development. Nat Immunol, 20(9): pp. 1090–1092. (2019).
Lukowski, S. W. et al. Absence of Batf3 reveals a new dimension of cell state heterogeneity within conventional dendritic cells. iScience 24 (5), 102402 (2021).
Ferris, S. T., Carrero, J. A. & Unanue, E. R. Antigen presentation events during the initiation of autoimmune diabetes in the NOD mouse. J. Autoimmun. 71, 19–25 (2016).
Ataide, M. A. et al. BATF3 programs CD8(+) T cell memory. Nat. Immunol. 21 (11), 1397–1407 (2020).
Qiu, Z. et al. Batf3 Expression by CD8 T Cells Critically Regulates the Development of Memory Populations. J. Immunol. 205 (4), 901–906 (2020).
Murphy, T. L., Tussiwand, R. & Murphy, K. M. Specificity through cooperation: BATF-IRF interactions control immune-regulatory networks. Nat. Rev. Immunol. 13 (7), 499–509 (2013).
Zhong, T. et al. TGF-beta-mediated crosstalk between TIGIT(+) Tregs and CD226(+)CD8(+) T cells in the progression and remission of type 1 diabetes. Nat. Commun. 15 (1), 8894 (2024).
Unanue, E. R., Ferris, S. T. & Carrero, J. A. The role of islet antigen presenting cells and the presentation of insulin in the initiation of autoimmune diabetes in the NOD mouse. Immunol. Rev. 272 (1), 183–201 (2016).
Ferris, S. T. et al. A minor subset of Batf3-dependent antigen-presenting cells in islets of Langerhans is essential for the development of autoimmune diabetes. Immunity 41 (4), 657–669 (2014).
Klebanoff, C. A. et al. Central memory self/tumor-reactive CD8 + T cells confer superior antitumor immunity compared with effector memory T cells. Proc. Natl. Acad. Sci. U S A. 102 (27), 9571–9576 (2005).
Carrero, J. A., Ferris, S. T. & Unanue, E. R. Macrophages and dendritic cells in islets of Langerhans in diabetic autoimmunity: a lesson on cell interactions in a mini-organ. Curr. Opin. Immunol. 43, 54–59 (2016).
Chiou, J. et al. Interpreting type 1 diabetes risk with genetics and single-cell epigenomics. Nature 594 (7863), 398–402 (2021).
Huang, D. & Ovcharenko, I. Enhancer-silencer transitions in the human genome. Genome Res. 32 (3), 437–448 (2022).
Bhatt, B., Garcia-Diaz, P. & Foight, G. W. Synthetic transcription factor engineering for cell and gene therapy. Trends Biotechnol. 42 (4), 449–463 (2024).
Zhuang, J. J. et al. Current strategies and progress for targeting the undruggable transcription factors. Acta Pharmacol. Sin. 43 (10), 2474–2481 (2022).
Zhong, S., Ge, J. & Yu, J. Y. Icariin prevents cytokine-induced beta-cell death by inhibiting NF-kappaB signaling. Exp. Ther. Med. 16 (3), 2756–2762 (2018).
Zandi, M. et al. The Impact of STAT3 rs1053005 Variation on Type 1 Diabetes Mellitus Susceptibility: Association Study and in Silico Analysis. Immunol. Invest. 51 (6), 1908–1919 (2022).
Clark, M. et al. The Role of T Cell Receptor Signaling in the Development of Type 1 Diabetes. Front. Immunol. 11, 615371 (2020).
Tontonoz, P. & Spiegelman, B. M. Fat and beyond: the diverse biology of PPARgamma. Annu. Rev. Biochem. 77, 289–312 (2008).
Puigserver, P. et al. Insulin-regulated hepatic gluconeogenesis through FOXO1-PGC-1alpha interaction. Nature 423 (6939), 550–555 (2003).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28 (1), 27–30 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28 (11), 1947–1951 (2019).
Kanehisa, M. et al. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51 (D1), D587–D592 (2023).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51 (D1), D638–D646 (2023).
Shen, W. et al. AnimalTFDB 4.0: a comprehensive animal transcription factor database updated with variation and expression annotations. Nucleic Acids Res. 51 (D1), D39–D45 (2023).
Rauluseviciute, I. et al. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 52(D1), D174–D182 (2024).
Lechleitner, M. et al. [Diagnosis and insulin therapy of type 1 diabetes mellitus (Update 2023)]. Wien Klin. Wochenschr. 135 (Suppl 1), 98–105 (2023).
Atkinson, M. A., Eisenbarth, G. S. & Michels, A. W. Type 1 diabetes. Lancet 383 (9911), 69–82 (2014).
Moreno, A. M. et al. CXCR3 expression in regulatory T cells drives interactions with type I dendritic cells in tumors to restrict CD8(+) T cell antitumor immunity. Immunity 56 (7), 1613–1630e5 (2023).
Han, S. et al. Genes and transcription factors related to the adverse effects of maternal type I diabetes mellitus on fetal development. Mol. Cell. Probes. 43, 64–71 (2019).
Donati, G. & Amati, B. MYC and therapy resistance in cancer: risks and opportunities. Mol. Oncol. 16 (21), 3828–3854 (2022).
Zhu, S. et al. RNA pull-down confocal nanoscanning (RP-CONA) detects quercetin as pri-miR-7/HuR interaction inhibitor that decreases alpha-synuclein levels. Nucleic Acids Res. 49 (11), 6456–6473 (2021).
Madden, S. K. et al. Taking the Myc out of cancer: toward therapeutic strategies to directly inhibit c-Myc. Mol. Cancer. 20 (1), 3 (2021).
Linsley, P. S., Greenbaum, C. J. & Nepom, G. T. Uncovering Pathways to Personalized Therapies in Type 1 Diabetes. Diabetes 70 (4), 831–841 (2021).
Dang, C. V. MYC on the path to cancer. Cell 149 (1), 22–35 (2012).
Hann, S. R. MYC cofactors: molecular switches controlling diverse biological outcomes. Cold Spring Harb Perspect. Med. 4 (9), a014399 (2014).
Marchingo, J. M. et al. Quantitative analysis of how Myc controls T cell proteomes and metabolic pathways during T cell activation. Elife, 9. (2020).
Zhu, C. et al. Tim-3 adaptor protein Bat3 is a molecular checkpoint of T cell terminal differentiation and exhaustion. Sci. Adv., 7(18). (2021).
Chou, C. et al. c-Myc-induced transcription factor AP4 is required for host protection mediated by CD8 + T cells. Nat. Immunol. 15 (9), 884–893 (2014).
Di Lorenzo, T. P., Peakman, M. & Roep, B. O. Translational mini-review series on type 1 diabetes: Systematic analysis of T cell epitopes in autoimmune diabetes. Clin. Exp. Immunol. 148 (1), 1–16 (2007).
Ahmed, S. et al. Standardizing T-Cell Biomarkers in Type 1 Diabetes: Challenges and Recent Advances. Diabetes 68 (7), 1366–1379 (2019).
D’Avola, A. et al. Spotlight on New Therapeutic Opportunities for MYC-Driven Cancers. Onco Targets Ther. 16, 371–383 (2023).
Chabanon, R. M. & Postel-Vinay, S. A Novel Synthetic Lethal Approach to Target MYC-Driven Cancers. Cancer Res. 82 (6), 969–971 (2022).
Pelengaris, S., Khan, M. & Evan, G. c-MYC: more than just a matter of life and death. Nat. Rev. Cancer. 2 (10), 764–776 (2002).
Han, H. et al. Small-Molecule MYC Inhibitors Suppress Tumor Growth and Enhance Immunotherapy. Cancer Cell. 36 (5), 483–497e15 (2019).
Shi, X. et al. JQ1: a novel potential therapeutic target. Pharmazie 73 (9), 491–493 (2018).
Megiorni, F. et al. OTX015 Epi-Drug Exerts Antitumor Effects in Ovarian Cancer Cells by Blocking GNL3-Mediated Radioresistance Mechanisms: Cellular, Molecular and Computational Evidence. Cancers (Basel), 13(7). (2021).
Cidado, J. et al. AZD4573 Is a Highly Selective CDK9 Inhibitor That Suppresses MCL-1 and Induces Apoptosis in Hematologic Cancer Cells. Clin. Cancer Res. 26 (4), 922–934 (2020).
Joshi, H. et al. The Pharmacological Implications of Flavopiridol: An Updated Overview. Molecules, 28(22). (2023).
Gnanaprakasam, J., Sherman, J. W. & Wang, R. MYC and HIF in shaping immune response and immune metabolism. Cytokine Growth Factor. Rev. 35, 63–70 (2017).
Riquelme, M. I. & Lubovac-Pilav, Z. Gene Co-Expression Network Analysis for Identifying Modules and Functionally Enriched Pathways in Type 1 Diabetes. PLoS One. 11 (6), e0156006 (2016).
Funding
This study was supported by Nantong Bureau of Science and Technology [grant numbers MS22022016] and, in part, by Health Commission of Nantong [grant numbers MA2021002]. Additional funding were provided by the Priority Academic Program Development of Nantong Talent Center [grant numbers 2022-Ⅲ-609] and grants from Jiangsu Provincial Research Hospital [grant numbers YJXYY202204].
Author information
Authors and Affiliations
Contributions
Ying Zhao and Zhicheng Tang contributed equally to this manuscript as co-first authors. Professor Weixia Yang participated in the design and revision of the overall research approach, the modification of the manuscript, and the provision of funding.Ying Zhao performed bioinformatics analysis, cell experiments, and wrote the manuscripts. Zhicheng Tang performed the plasmid design, lentivirus constructing and contributing to the manuscript writing. Ying Tao participated in clinical sample collection and cell experiments. Sihui Zhao participated in clinical sample collection, clinical sample processing, and data analysis. Qijie Ding participated in the collection of general information of study subjects and sample collection.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical statement
This study was approved by the Ethics Committee of the Affiliated Hospital of Nantong University(Approval Number: 2021-K085-01) on August 10, 2021. And it was conducted in accordance with the principles of the Declaration of Helsinki. Written informed consent was obtained from all participants or their legal guardians prior to participation in the study. The confidentiality of patient data was ensured throughout the study, and all personal identifiers were removed to protect the privacy of the individuals involved.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, Y., Tang, Z., Tao, Y. et al. c-MYC enhances transcription of the type 1 diabetes mellitus associated gene BATF3 via promoter binding. Sci Rep (2026). https://doi.org/10.1038/s41598-026-45579-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-45579-x