Abstract
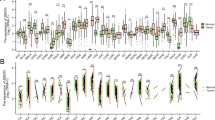
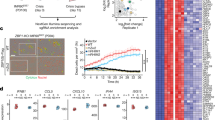
Z-DNA binding protein 1 (ZBP1) is a critical factor that induces various forms of cell death and inflammatory responses. However, it has not been comprehensively analyzed in malignancies. The present study aimed to elucidate the oncogenic role of ZBP1 in human malignancies and examine its prospective function in renal cell carcinoma. The pathogenic significance of ZBP1 in malignancies was clarified using a multi-omics analysis and single-cell analysis utilizing bioinformatics methods. The function of ZBP1 in renal cell carcinoma was validated using in vivo and in vitro studies. In this study, we proved that ZBP1 high expression was significantly correlated with worse prognosis in kidney renal clear cell carcinoma and acute myeloid leukemia. Elevated ZBP1 levels correlated with cancer-associated fibroblast invasion. Furthermore, the single-cell analysis revealed that ZBP1 was significantly expressed in CD4+ T cells, CD8+ T cells, and regulatory T cells. ZBP1 may participate in cancer by regulating the inflammatory pathways. Cellular tests demonstrated that downregulating ZBP1 expression significantly inhibited renal cell carcinoma cell proliferation, migration, and invasion. The transplanted tumor model indicated that suppressing ZBP1 expression significantly reduced tumor progression. ZBP1 is a promising biomarker related to the prognosis of pan-cancer, especially renal cell carcinoma.
Similar content being viewed by others
Data availability
Availability of the data and materials, through public databases [TCGA (https://www.cancer.gov/ccg/research/genome-sequencing/tcga), GEO (https://www.ncbi.nlm.nih.gov/geo/), TISCH2 (https://tisch.compbio.cn/home/)] and analysis tools are outlined in the Methods section.
Reference
Kuriakose, T. & Kanneganti, T. D. ZBP1: Innate sensor regulating cell death and inflammation. Trend. Immunol. 39, 123–134. https://doi.org/10.1016/j.it.2017.11.002 (2018).
Hao, Y. et al. ZBP1: A powerful innate immune sensor and double-edged sword in host immunity. Int. J. Mol. Sci. https://doi.org/10.3390/ijms231810224 (2022).
Oh, S. & Lee, S. Recent advances in ZBP1-derived PANoptosis against viral infections. Front. Immunol. 14, 1148727. https://doi.org/10.3389/fimmu.2023.1148727 (2023).
Zhang, T. et al. Influenza virus Z-RNAs induce ZBP1-mediated necroptosis. Cell 180, 1115-1129 e1113. https://doi.org/10.1016/j.cell.2020.02.050 (2020).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci. Immunol. https://doi.org/10.1126/sciimmunol.aag2045 (2016).
Kesavardhana, S. et al. ZBP1/DAI ubiquitination and sensing of influenza vRNPs activate programmed cell death. J. Exp. Med. 214, 2217–2229. https://doi.org/10.1084/jem.20170550 (2017).
Jiao, H. et al. ADAR1 averts fatal type i interferon induction by ZBP1. Nature 607, 776–783. https://doi.org/10.1038/s41586-022-04878-9 (2022).
Li, S. et al. SARS-CoV-2 Z-RNA activates the ZBP1-RIPK3 pathway to promote virus-induced inflammatory responses. Cell. Res. 33, 201–214. https://doi.org/10.1038/s41422-022-00775-y (2023).
Devos, M. et al. Sensing of endogenous nucleic acids by ZBP1 induces keratinocyte necroptosis and skin inflammation. J. Exp. Med. https://doi.org/10.1084/jem.20191913 (2020).
Lei, Y. et al. Cooperative sensing of mitochondrial DNA by ZBP1 and cGAS promotes cardiotoxicity. Cell 186, 3013-3032 e3022. https://doi.org/10.1016/j.cell.2023.05.039 (2023).
Enzan, N. et al. ZBP1 protects against mtDNA-induced myocardial inflammation in failing hearts. Circ. Res. 132, 1110–1126. https://doi.org/10.1161/CIRCRESAHA.122.322227 (2023).
Zhang, T. et al. ADAR1 masks the cancer immunotherapeutic promise of ZBP1-driven necroptosis. Nature 606, 594–602. https://doi.org/10.1038/s41586-022-04753-7 (2022).
Baik, J. Y. et al. ZBP1 not RIPK1 mediates tumor necroptosis in breast cancer. Nat. Commun. 12, 2666. https://doi.org/10.1038/s41467-021-23004-3 (2021).
Rui, R., Zhou, L. & He, S. Cancer immunotherapies: Advances and bottlenecks. Front. Immunol. 14, 1212476. https://doi.org/10.3389/fimmu.2023.1212476 (2023).
Bejarano, L., Jordão, M. J. C. & Joyce, J. A. Therapeutic targeting of the tumor microenvironment. Cancer. Discov. 11, 933–959. https://doi.org/10.1158/2159-8290.CD-20-1808 (2021).
Biffi, G. & Tuveson, D. A. Diversity and biology of cancer-associated fibroblasts. Physiol. Rev. 101, 147–176. https://doi.org/10.1152/physrev.00048.2019 (2021).
Kennel, K. B., Bozlar, M., De Valk, A. F. & Greten, F. R. Cancer-associated fibroblasts in inflammation and antitumor immunity. Clin. Cancer Res. 29, 1009–1016. https://doi.org/10.1158/1078-0432.CCR-22-1031 (2023).
Zheng, M. & Kanneganti, T. D. The regulation of the ZBP1-NLRP3 inflammasome and its implications in pyroptosis, apoptosis, and necroptosis (PANoptosis). Immunol. Rev. 297, 26–38. https://doi.org/10.1111/imr.12909 (2020).
Karki, R. & Kanneganti, T. D. ADAR1 and ZBP1 in innate immunity, cell death, and disease. Trends Immunol. 44, 201–216. https://doi.org/10.1016/j.it.2023.01.001 (2023).
Wang, Q. et al. ZBP1 pathway promotes tumor immunogenicity in the combination of anti-HER2 therapy and epigenetic therapy. Cell Rep. 44, 116314. https://doi.org/10.1016/j.celrep.2025.116314 (2025).
Nassour, J. et al. Telomere-to-mitochondria signalling by ZBP1 mediates replicative crisis. Nature 614, 767–773. https://doi.org/10.1038/s41586-023-05710-8 (2023).
Yan, J., Wan, P., Choksi, S. & Liu, Z. G. Necroptosis and tumor progression. Trends Cancer 8, 21–27. https://doi.org/10.1016/j.trecan.2021.09.003 (2022).
Műzes, G. & Sipos, F. PANoptosis as a two-edged sword in colorectal cancer: A pathogenic mechanism and therapeutic opportunity. Cells https://doi.org/10.3390/cells14100730 (2025).
Cho, M. G. et al. MRE11 liberates cGAS from nucleosome sequestration during tumorigenesis. Nature 625, 585–592. https://doi.org/10.1038/s41586-023-06889-6 (2024).
Gong, Z. et al. Machine learning identifies TIME subtypes linking EGFR mutations and immune states in lung adenocarcinoma. NPJ Digit. Med. 8, 796. https://doi.org/10.1038/s41746-025-02172-2 (2025).
Zhang, P. et al. Metabolic reprogramming signature predicts immunotherapy efficacy in lung adenocarcinoma: targeting SLC25A1 to overcome immune resistance. Chin. J. Cancer Res. 37, 1000–1019. https://doi.org/10.21147/j.issn.1000-9604.2025.06.11 (2025).
Zhang, H. et al. Define cancer-associated fibroblasts (CAFs) in the tumor microenvironment: new opportunities in cancer immunotherapy and advances in clinical trials. Mol. Cancer 22, 159. https://doi.org/10.1186/s12943-023-01860-5 (2023).
Liao, Z., Tan, Z. W., Zhu, P. & Tan, N. S. Cancer-associated fibroblasts in tumor microenvironment-accomplices in tumor malignancy. Cell Immunol. 343, 103729. https://doi.org/10.1016/j.cellimm.2017.12.003 (2019).
Ou, Q., Power, R. & Griffin, M. D. Revisiting regulatory T cells as modulators of innate immune response and inflammatory diseases. Front. Immunol. 14, 1287465. https://doi.org/10.3389/fimmu.2023.1287465 (2023).
Ohkura, N. & Sakaguchi, S. Transcriptional and epigenetic basis of Treg cell development and function: Its genetic anomalies or variations in autoimmune diseases. Cell Res. 30, 465–474. https://doi.org/10.1038/s41422-020-0324-7 (2020).
Guan, X. et al. Microglial CMPK2 promotes neuroinflammation and brain injury after ischemic stroke. Cell Rep. Med. 5, 101522. https://doi.org/10.1016/j.xcrm.2024.101522 (2024).
Qing, F. & Liu, Z. Interferon regulatory factor 7 in inflammation, cancer and infection. Front. Immunol. 14, 1190841. https://doi.org/10.3389/fimmu.2023.1190841 (2023).
Liang, M. et al. OAS1 induces endothelial dysfunction and promotes monocyte adhesion through the NFκB pathway in atherosclerosis. Arch. Biochem. Biophys. 763, 110222. https://doi.org/10.1016/j.abb.2024.110222 (2025).
Bi, Q. et al. Tumor-associated inflammation: The tumor-promoting immunity in the early stages of tumorigenesis. J. Immunol. Res. 2022, 3128933. https://doi.org/10.1155/2022/3128933 (2022).
Nomoto, D. et al. Fusobacterium nucleatum promotes esophageal squamous cell carcinoma progression via the NOD1/RIPK2/NF-κB pathway. Cancer Lett. 530, 59–67. https://doi.org/10.1016/j.canlet.2022.01.014 (2022).
Marandi, Y. et al. NLRP3-inflammasome activation is associated with epithelial-mesenchymal transition and progression of colorectal cancer. Iran. J. Basic Med. Sci. 24, 483–492. https://doi.org/10.22038/ijbms.2021.52355.11835 (2021).
Deng, Q. et al. NLRP3 inflammasomes in macrophages drive colorectal cancer metastasis to the liver. Cancer Lett. 442, 21–30. https://doi.org/10.1016/j.canlet.2018.10.030 (2019).
Chen, L. et al. NF-κB p65 and SETDB1 expedite lipopolysaccharide-induced intestinal inflammation in mice by inducing IRF7/NLR-dependent macrophage M1 polarization. Int. Immunopharmacol. 115, 109554. https://doi.org/10.1016/j.intimp.2022.109554 (2023).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: An enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47, W556–W560. https://doi.org/10.1093/nar/gkz430 (2019).
Chen, F., Chandrashekar, D. S., Varambally, S. & Creighton, C. J. Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers. Nat. Commun. 10, 5679. https://doi.org/10.1038/s41467-019-13528-0 (2019).
Unberath, P. et al. Searching of clinical trials made easier in cBioPortal using patients’ genetic and clinical profiles. Appl. Clin. Inform. 13, 363–369. https://doi.org/10.1055/s-0042-1743560 (2022).
Cerami, E. et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404. https://doi.org/10.1158/2159-8290.CD-12-0095 (2012).
Bardou, P., Mariette, J., Escudié, F., Djemiel, C. & Klopp, C. Jvenn: An interactive venn diagram viewer. BMC Bioinform. 15, 293. https://doi.org/10.1186/1471-2105-15-293 (2014).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucl. Acid. Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Han, Y. et al. TISCH2: Expanded datasets and new tools for single-cell transcriptome analyses of the tumor microenvironment. Nucleic Acids Res. 51, D1425-d1431. https://doi.org/10.1093/nar/gkac959 (2023).
Sun, X. et al. Comprehensive analysis of SLC35A2 in pan-cancer and validation of its role in breast cancer. J. Inflamm. Res. 16, 3381–3398. https://doi.org/10.2147/JIR.S419994 (2023).
Funding
This research was funded by the National Natural Science Foundation of China (No: 82560514) and the "Medical Excellence Award" funded by the Creative Research Development Grant from the First Affiliated Hospital of Guangxi Medical University.
Author information
Authors and Affiliations
Contributions
C.L. and C.S. designed the experiments; C.L., J.N. and Y.L. performed the experiments; C.L. and J.N. prepared all figures; C.L., J.N., Y.L., C.W. and W.L. analyzed the data; C.S. and B.S. supervised the work; C.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interests.
Ethics
The study received approval from the First Affiliated Hospital of Guangxi Medical University Ethics Committee (Approval No. 2024-S770-01). The written informed consent of the patient and/or their legal guardian was obtained before the tissue specimens were obtained. All animal experiments were conducted under the approval and supervision of the Ethics Committee of the First Affiliated Hospital of Guangxi Medical University (approval number: 2024-D0291).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, C., Nong, J., Li, Y. et al. Integration of multi-omics and single-cell analysis reveals ZBP1 as a prognostic biomarker of renal cell carcinoma. Sci Rep (2026). https://doi.org/10.1038/s41598-026-45826-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-45826-1