Abstract
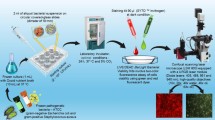
Healthcare-associated infections (HCAIs) contribute significantly to global mortality, driven by the increasing antimicrobial resistance. Rapid, high-throughput bacterial detection is crucial for infection control and patient care. We report a real-time, multiplex lamp-based Photoionization Detector (PID) assisted by AI-image-based analysis for bacterial identification. Using four lamps with varying ionization energies, the sensor selectively ionizes VOCs emitted by bacteria, producing four distinct current curves for each target species (Escherichia coli, Staphylococcus aureus, Pseudomonas aeruginosa, and Klebsiella pneumoniae). These curves were transformed into image representations, capturing their spectral patterns for bacterial differentiation. A pre-trained ResNet-18 Convolutional Neural Network (CNN) within a Few-Shot Learning (FSL) framework extracted key features, enabling accurate (> 88%) bacterial differentiation even with limited labeled data. This sensor detected bacterial concentrations as low as 10² CFU and distinguished contamination levels. The synergistic integration of PID sensing with AI-driven analysis offers a powerful approach to rapid bacterial diagnostics, demonstrating strong potential for clinical implementation and improved patient care. This study marks an early step toward AI-based VOC sensing, where FSL acts as a proof-of-concept under data scarcity.
Similar content being viewed by others
Data availability
All processed data are available in the main text or the supplementary materials.
References
Balasubramanian Id, R., Van Boeckel, T. P., Carmeli, Y., Cosgrove, S. & Laxminarayan, R. Global Incidence in Hospital-Associated Infections Resistant to Antibiotics: An Analysis of Point Prevalence Surveys from 99 Countries. PLoS Med. 20, e1004178. https://doi.org/10.1371/journal.pmed.1004178 (2023).
Abban, M. K., Ayerakwa, E. A., Mosi, L. & Isawumi, A. The Burden of Hospital Acquired Infections and Antimicrobial Resistance. Heliyon 9, e20561. https://doi.org/10.1016/j.heliyon.2023.e20561 (2023).
Horan, T. C., Andrus, M. & Dudeck, M. A. CDC/NHSN Surveillance Definition of Health Care-Associated Infection and Criteria for Specific Types of Infections in the Acute Care Setting. Am. J. Infect. Control. 36, 309–332. https://doi.org/10.1016/j.ajic.2008.03.002 (2008).
Klevens, R. M. et al. Estimating Health Care-Associated Infections and Deaths in U.S. Hospitals, 2002. Public. Health Rep. 122, 160–166. https://doi.org/10.1177/003335490712200205 (2007).
European Centre for Disease Prevention and Control (ECDC). Point Prevalence Survey of Healthcare-Associated Infections and Antimicrobial Use in European Long-Term Care Facilities; ; (2025).
Sandu, A. M. et al. Healthcare-Associated Infections: The Role of Microbial and Environmental Factors in Infection Control-A Narrative Review. 14, 933–971, (2025). https://doi.org/10.1007/s40121-025-01143-0
Pogue, J. M., Kaye, K. S., Cohen, D. A. & Marchaim, D. Appropriate Antimicrobial Therapy in the Era of Multidrug-Resistant Human Pathogens. Clin. Microbiol. Infect. 21, 302–312. https://doi.org/10.1016/J.CMI.2014.12.025 (2015).
Timbrook, T. T. et al. The Effect of Molecular Rapid Diagnostic Testing on Clinical Outcomes in Bloodstream Infections: A Systematic Review and Meta-Analysis. Clin. Infect. Dis. 64, 15–23. https://doi.org/10.1093/CID/CIW649 (2017).
Lambregts, M. M. C., Bernards, A. T., van der Beek, M. T., Visser, L. G. & de Boer, M. G. Time to Positivity of Blood Cultures Supports Early Re-Evaluation of Empiric Broad-Spectrum Antimicrobial Therapy. PLoS One. 14 https://doi.org/10.1371/JOURNAL.PONE.0208819 (2019).
Rajapaksha, P. et al. A Review of Methods for the Detection of Pathogenic Microorganisms. Analyst 144, 396–411. https://doi.org/10.1039/C8AN01488D (2019).
Váradi, L. et al. Methods for the Detection and Identification of Pathogenic Bacteria: Past, Present, and Future. Chem. Soc. Rev. 46, 4818–4832. https://doi.org/10.1039/C6CS00693K (2017).
Schulz, S., Dickschat, J. S. & Bacterial Volatiles The Smell of Small Organisms. Nat. Prod. Rep. 24, 814–842. https://doi.org/10.1039/B507392H (2007).
Epping, R. & Koch, M. On-Site Detection of Volatile Organic Compounds (VOCs). Molecules 28, 1598. https://doi.org/10.3390/MOLECULES28041598 (2023).
Bos, L. D. J., Sterk, P. J. & Schultz, M. J. Volatile Metabolites of Pathogens: A Systematic Review. PLoS Pathog. 9, e1003311. https://doi.org/10.1371/journal.ppat.1003311 (2013).
Ratiu, I. A. et al. An Optimistic Vision of Future: Diagnosis of Bacterial Infections by Sensing Their Associated Volatile Organic Compounds. Crit. Rev. Anal. Chem. 50, 501–512. https://doi.org/10.1080/10408347.2019.1663147 (2020).
Filipiak, W. et al. Molecular Analysis of Volatile Metabolites Released Specifically by Staphylococcus Aureus and Pseudomonas Aeruginosa. BMC Microbiol. 12 https://doi.org/10.1186/1471-2180-12-113 (2012).
Filipiak, W. et al. GC-MS Profiling of Volatile Metabolites Produced by Klebsiella Pneumoniae. Front. Mol. Biosci. 9, 1019290. https://doi.org/10.3389/fmolb.2022.1019290 (2022).
Hewett, K. et al. De Lacy Costello, B. Towards the Identification of Antibiotic-Resistant Bacteria Causing Urinary Tract Infections Using Volatile Organic Compounds Analysis-A Pilot Study. Antibiotics 9, 797. https://doi.org/10.3390/antibiotics9110797 (2020).
Luo, H. et al. Rapid Identification of Carbapenemase-Producing Klebsiella Pneumoniae Using Headspace Solid-Phase Microextraction Combined with Gas Chromatography-Mass Spectrometry. Infect. Drug Resist. 16, 2601–2609. https://doi.org/10.2147/IDR.S404742 (2023).
Kunze-Szikszay, N. et al. Headspace Analyses Using Multi-Capillary Column-Ion Mobility Spectrometry Allow Rapid Pathogen Differentiation in Hospital-Acquired Pneumonia Relevant Bacteria. BMC Microbiol. 21, 69. https://doi.org/10.1186/S12866-021-02102-8/FIGURES/3 (2021).
Dias, T. et al. A Lab-Made E-Nose-MOS Device for Assessing the Bacterial Growth in a Solid Culture Medium. Biosens. (Basel). 13 https://doi.org/10.3390/bios13010019 (2023).
Reidt, U. et al. Detection of Microorganisms with an Electronic Nose for Application under Microgravity Conditions. Gravitational Space Res. 8, 1–17. https://doi.org/10.2478/gsr-2020-0001 (2020).
Reidt, U. et al. Detection of Microorganisms Onboard the International Space Station Using an Electronic Nose. Gravitational Space Res. 5, 89–111. https://doi.org/10.2478/gsr-2017-0013 (2017).
Bous, M. et al. Detection of Volatile Organic Compounds in Headspace of Klebsiella Pneumoniae and Klebsiella Oxytoca Colonies. Front. Pediatr. 11, 1151000. https://doi.org/10.3389/fped.2023.1151000 (2023).
Kunze, N. et al. Detection and Validation of Volatile Metabolic Patterns over Different Strains of Two Human Pathogenic Bacteria during Their Growth in a Complex Medium Using Multi-Capillary Column-Ion Mobility Spectrometry (MCC-IMS). Appl. Microbiol. Biotechnol. 97, 3665–3676. https://doi.org/10.1007/s00253-013-4762-8 (2013).
Zhu, J., Bean, H. D., Kuo, Y. M. & Hill, J. E. Fast Detection of Volatile Organic Compounds from Bacterial Cultures by Secondary Electrospray Ionization-Mass Spectrometry. J. Clin. Microbiol. 48, 4426–4431. https://doi.org/10.1128/JCM.00392-10/FORMAT/EPUB (2010).
Li, H. & Zhu, J. Differentiating Antibiotic-Resistant Staphylococcus Aureus Using Secondary Electrospray Ionization Tandem Mass Spectrometry. Anal. Chem. 90, 12108–12115. https://doi.org/10.1021/ACS.ANALCHEM.8B03029/ASSET/IMAGES/LARGE/AC-2018-030292_0006.JPEG (2018).
Agbroko, S. O., Covington, J. A. & Novel Low-Cost, Portable PID Sensor for the Detection of Volatile Organic Compounds. Sens. Actuators B Chem. 275, 10–15. https://doi.org/10.1016/j.snb.2018.07.173 (2018).
Rees, C. A. et al. Detection of High-Risk Carbapenem-Resistant Klebsiella Pneumoniae and Enterobacter Cloacae Isolates Using Volatile Molecular Profiles. Sci. Rep. 8, 13297. https://doi.org/10.1038/s41598-018-31543-x (2018).
Rees, C. A., Franchina, F. A., Nordick, K. V., Kim, P. J. & Hill, J. E. Expanding the Klebsiella Pneumoniae Volatile Metabolome Using Advanced Analytical Instrumentation for the Detection of Novel Metabolites. J. Appl. Microbiol. 122, 785–795. https://doi.org/10.1111/jam.13372 (2017).
Ródenas García, M. et al. Review of Low-Cost Sensors for Indoor Air Quality: Features and Applications. Appl. Spectrosc. Rev. 57, 747–779. https://doi.org/10.1080/05704928.2022.2085734 (2022).
Pang, X. et al. Low-Cost Photoionization Sensors as Detectors in GC × GC Systems Designed for Ambient VOC Measurements. Sci. Total Environ. 664, 771–779. https://doi.org/10.1016/j.scitotenv.2019.01.348 (2019).
Jian, R. S., Sung, L. Y. & Lu, C. J. Measuring Real-Time Concentration Trends of Individual VOC in an Elementary School Using a Sub-Ppb Detection ΜGC and a Single GC–MS Analysis. Chemosphere 99, 261–266. https://doi.org/10.1016/J.CHEMOSPHERE.2013.10.094 (2014).
Tait, E., Perry, J. D., Stanforth, S. P. & Dean, J. R. Identification of Volatile Organic Compounds Produced by Bacteria Using HS-SPME-GC-MS. J. Chromatogr. Sci. 52, 363–373. https://doi.org/10.1093/chromsci/bmt042 (2014).
Almasoud, N. et al. Discrimination of Bacteria Using Whole Organism Fingerprinting: The Utility of Modern Physicochemical Techniques for Bacterial Typing. Analyst 146, 770–788. https://doi.org/10.1039/D0AN01482F (2021).
Locke, A., Fitzgerald, S. & Mahadevan-Jansen, A. Advances in Optical Detection of Human-Associated Pathogenic Bacteria. Molecules 25, 5256. https://doi.org/10.3390/MOLECULES25225256 (2020).
Wang, L. et al. Applications of Raman Spectroscopy in Bacterial Infections: Principles, Advantages, and Shortcomings. Front. Microbiol. 12, 683580. https://doi.org/10.3389/FMICB.2021.683580/TEXT (2021).
Chícharo, A. et al. Precision-Engineered Plasmonic Nanostar Arrays for High-Performance SERS Sensing. Adv. Opt. Mater. 13, e01275. https://doi.org/10.1002/ADOM.202501275 (2025).
Zhang, C. et al. Volatilomics Analysis of Jasmine Tea during Multiple Rounds of Scenting Processes. Foods 12, 812. https://doi.org/10.3390/FOODS12040812/S1 (2023).
Chen, G. F., Lai, C. H. & Chen, W. H. Principal Component Analysis and Mapping to Characterize the Emission of Volatile Organic Compounds in a Typical Petrochemical Industrial Park. Aerosol Air Qual. Res. 20, 465–476. https://doi.org/10.4209/AAQR.2019.07.0365 (2020).
Arora, M. et al. Machine Learning Approaches to Identify Discriminative Signatures of Volatile Organic Compounds (VOCs) from Bacteria and Fungi Using SPME-DART-MS. Metabolites 12, 232. https://doi.org/10.3390/METABO12030232 (2022).
Zhou, B. et al. A Lightweight Convolutional Neural Network for Bacterial Identification Based on Raman Spectra. RSC Adv. 12, 26463. https://doi.org/10.1039/d2ra03722j (2022).
Liu, Y. et al. Rapid and Accurate Identification of Marine Microbes with Single-Cell Raman Spectroscopy. Analyst 145, 3297. https://doi.org/10.1039/c9an02069a (2020).
Zeng, W., Xiao, Z., Zeng, W. & Xiao, Z. Few-Shot Learning Based on Deep Learning: A Survey. Mathematical Biosciences and Engineering 21, 679–711, (2024). https://doi.org/10.3934/MBE.2024029
Gharoun, H., Momenifar, F., Chen, F. & Gandomi, A. H. Meta-Learning Approaches for Few-Shot Learning: A Survey of Recent Advances. ACM Comput. Surv. 56, 1–41. https://doi.org/10.1145/3659943/ASSET/A4843494-3BFB-4FA5-B394-E26841DD7583/ASSETS/IMAGES/LARGE/CSUR-2023-0062-F27.JPG (2024).
Mi, F. et al. Recent Advancements in Microfluidic Chip Biosensor Detection of Foodborne Pathogenic Bacteria: A Review. Anal. Bioanal Chem. 414, 2883. https://doi.org/10.1007/S00216-021-03872-W (2022).
Gangwar, R. et al. Plasma Functionalized Carbon Interfaces for Biosensor Application: Toward the Real-Time Detection of Escherichia Coli O157: H7. ACS Omega. 7, 21025–21034. https://doi.org/10.1021/ACSOMEGA.2C01802/ASSET/IMAGES/MEDIUM/AO2C01802_M001.GIF (2022).
Izadi, M. & Arvand, M. An Aptamer-Functionalized AuNPs/RGO Nanocomposite Biosensor for Ultrasensitive Detection of Foodborne Pathogen E. Coli O157:H7. Sci. Rep. 16, 2701, (2026). https://doi.org/10.1038/s41598-025-32516-7
Costa, S. P. et al. A Microfluidic Platform Combined with Bacteriophage Receptor Binding Proteins for Multiplex Detection of Escherichia Coli and Pseudomonas Aeruginosa in Blood. Sens. Actuators B Chem. 376, 132917. https://doi.org/10.1016/J.SNB.2022.132917 (2023).
Muller, V. et al. Identification of Pathogenic Bacteria in Complex Samples Using a Smartphone Based Fluorescence Microscope. RSC Adv. 8, 36493. https://doi.org/10.1039/c8ra06473c (2018).
Storer, M. K., Hibbard-Melles, K., Davis, B. & Scotter, J. Detection of Volatile Compounds Produced by Microbial Growth in Urine by Selected Ion Flow Tube Mass Spectrometry (SIFT-MS). J. Microbiol. Methods. 87, 111–113. https://doi.org/10.1016/j.mimet.2011.06.012 (2011).
Allardyce, R. A., Hill, A. L. & Murdoch, D. R. The Rapid Evaluation of Bacterial Growth and Antibiotic Susceptibility in Blood Cultures by Selected Ion Flow Tube Mass Spectrometry. Diagn. Microbiol. Infect. Dis. 55, 255–261. https://doi.org/10.1016/j.diagmicrobio.2006.01.031 (2006).
He, K., Zhang, X., Ren, S. & Sun, J. Deep Residual Learning for Image Recognition. Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition 770–778, (2016). https://doi.org/10.1109/CVPR.2016.90
Miranda, A. & De Beule, P. A. A. Atmospheric Photoionization Detector with Improved Photon Efficiency: A Proof of Concept for Application of a Nanolayer Thin-Film Electrode. Sensors 21, 7738. https://doi.org/10.3390/s21227738 (2021).
Boots, A. W., Bos, L. D., van der Schee, M. P., van Schooten, F. J. & Sterk, P. J. Exhaled Molecular Fingerprinting in Diagnosis and Monitoring: Validating Volatile Promises. Trends Mol. Med. 21, 633–644. https://doi.org/10.1016/J.MOLMED.2015.08.001 (2015).
Zscheppank, C., Wiegand, H. L., Lenzen, C., Wingender, J. & Telgheder, U. Investigation of Volatile Metabolites during Growth of Escherichia Coli and Pseudomonas Aeruginosa by Needle Trap-GC-MS. Anal. Bioanal Chem. 406, 6617–6628. https://doi.org/10.1007/s00216-014-8111-2 (2014).
Goeminne, P. C. et al. Detection of Pseudomonas Aeruginosa in Sputum Headspace through Volatile Organic Compound Analysis. Respir Res. 13, 87. https://doi.org/10.1186/1465-9921-13-87 (2012).
Davis, T. J. et al. Pseudomonas Aeruginosa Volatilome Characteristics and Adaptations in Chronic Cystic Fibrosis Lung Infections. mSphere 5, e00843–20, (2020). https://doi.org/10.1128/mSphere
Neyshabur, B., Sedghi, H. & Zhang, C. What Is Being Transferred in Transfer Learning? In Proceedings of the 34th Conference on Neural Information Processing Systems (NeurIPS 2020), Vancouver, Canada.; Neural information processing systems foundation, ; Vol. 2020-December. (2020).
Snell, J., Swersky, K. & Zemel, T. R. Prototypical Networks for Few-Shot Learning. In Proceedings of the Advances in Neural Information Processing Systems; 30. (2017).
Cheng, H., Garrick, D. J. & Fernando, R. L. Efficient Strategies for Leave-One-out Cross Validation for Genomic Best Linear Unbiased Prediction. J. Anim. Sci. Biotechnol. 8, 38. https://doi.org/10.1186/S40104-017-0164-6/TABLES/5 (2017).
Beleites, C. & Salzer, R. Assessing and Improving the Stability of Chemometric Models in Small Sample Size Situations. Anal. Bioanal Chem. 390, 1261–1271. https://doi.org/10.1007/S00216-007-1818-6/FIGURES/11 (2008).
Kornblith, S., Shlens, J. & Le, Q. V. Do Better ImageNet Models Transfer Better? In Proceedings of the Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition; IEEE Computer Society, Vol. 2019-June, pp. 2656–2666.
Kim, H. E. et al. Transfer Learning for Medical Image Classification: A Literature Review. BMC Med. Imaging. 22, 69. https://doi.org/10.1186/S12880-022-00793-7 (2022).
Dong, G. & Liu, H. Feature Engineering for Machine Learning and Data Analytics; CRC Press, Taylor and Francis, ISBN 1351721275. (2018).
Fitzgerald, S., Duffy, E., Holland, L. & Morrin, A. Multi-Strain Volatile Profiling of Pathogenic and Commensal Cutaneous Bacteria. Sci. Rep. 10, 17971. https://doi.org/10.1038/s41598-020-74909-w (2020).
Skvarc, M., Stubljar, D., Rogina, P. & Kaasch, A. J. Non-Culture-Based Methods to Diagnose Bloodstream Infection: Does It Work? Eur. J. Microbiol. Immunol. (Bp). 3, 97. https://doi.org/10.1556/EUJMI.3.2013.2.2 (2013).
Yagupsky, P. & Nolte, F. S. Quantitative Aspects of Septicemia. Clin. Microbiol. Rev. 3, 269–279. https://doi.org/10.1128/CMR.3.3.269 (1990).
Kranz, J. et al. European Association of Urology Guidelines on Urological Infections: Summary of the 2024 Guidelines. Eur. Urol. 86, 27–41. https://doi.org/10.1016/J.EURURO.2024.03.035 (2024).
Hay, A. D. et al. Microbiological Diagnosis of Urinary Tract Infection by NHS and Research Laboratories. Pediatrics 125, 335–341. https://doi.org/10.1542/PEDS.2008-1455 (2016).
Ferone, M., Gowen, A., Fanning, S. & Scannell, A. G. M. Microbial Detection and Identification Methods: Bench Top Assays to Omics Approaches. Compr. Rev. Food Sci. Food Saf. 19, 3106–3129. https://doi.org/10.1111/1541-4337.12618 (2020).
Guo, Y., Gao, W., Yang, H., Ma, C. & Sui, S. De-Escalation of Empiric Antibiotics in Patients with Severe Sepsis or Septic Shock: A Meta-Analysis. Heart Lung. 45, 454–459. https://doi.org/10.1016/J.HRTLNG.2016.06.001 (2016).
Smart, A. et al. Sniffing out Resistance - Rapid Identification of Urinary Tract Infection-Causing Bacteria and Their Antibiotic Susceptibility Using Volatile Metabolite Profiles. J. Pharm. Biomed. Anal. 167, 59–65. https://doi.org/10.1016/J.JPBA.2019.01.044 (2019).
Roda, B. et al. New Analytical Platform Based on Field-Flow Fractionation and Olfactory Sensor to Improve the Detection of Viable and Non-Viable Bacteria in Food. Anal. Bioanal Chem. 408, 7367–7377. https://doi.org/10.1007/S00216-016-9836-X/FIGURES/8 (2016).
Palma, S. I. C. J. et al. A. Machine Learning for the Meta-Analyses of Microbial Pathogens’ Volatile Signatures. Sci. Rep. 8, 3360. https://doi.org/10.1038/s41598-018-21544-1 (2018).
Shauloff, N. et al. Sniffing Bacteria with a Carbon-Dot Artificial Nose. Nanomicro Lett. 13, 112. https://doi.org/10.1007/s40820-021-00610-w (2021).
Henry, J., Endres, J. L., Sadykov, M. R., Bayles, K. W. & Svechkarev, D. Fast and Accurate Identification of Pathogenic Bacteria Using Excitation-Emission Spectroscopy and Machine Learning. Sens. Diagnostics. 3, 1253–1262. https://doi.org/10.1039/d4sd00070f (2024).
Song, C., Ristenpart, T. & Shmatikov, V. Machine Learning Models That Remember Too Much. In Proceedings of the Proceedings of the ACM Conference on Computer and Communications Security; Association for Computing Machinery, pp. 587–601. (2017).
Sibila, O. et al. Airway Bacterial Load and Inhaled Antibiotic Response in Bronchiectasis. Am. J. Respir Crit. Care Med. 200, 33–41. https://doi.org/10.1164/RCCM.201809-1651OC (2019).
Burd, E. M. & Kehl, K. S. A Critical Appraisal of the Role of the Clinical Microbiology Laboratory in the Diagnosis of Urinary Tract Infections. J. Clin. Microbiol. 49, 34–38. https://doi.org/10.1128/JCM.00788-11 (2011).
Yagupsky, P. & Nolte, F. S. Quantitative Aspects of Septicemia. Clin. Microbiol. Rev. 3, 269–279. https://doi.org/10.1128/CMR.3.3.269 (1990).
Funding
The authors acknowledge the financial support of the project SMARTgNOSTICS, with the reference n.º C644915155-00000024, co-funded by Component C5 – Capitalisation and Business Innovation under the Portuguese Resilience and Recovery Plan, through the NextGenerationEU Fund.
Author information
Authors and Affiliations
Contributions
S.P.C., H.M., and F.G. designed and performed the experimental work. A.C. and F.Y. performed algorithm development; S.P.C., A.C., and F.Y. analyzed the data. S.P.C., H.M., and A.C. wrote the manuscript. All authors read, reviewed, and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Costa, S.P., Cardoso, A., Mahmoodnia, H. et al. Bacterial species differentiation via real-time detection of microbial volatile organic compounds using a wavelength multiplexed photoionization detector and AI image-based analysis. Sci Rep (2026). https://doi.org/10.1038/s41598-026-46818-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-46818-x