Abstract
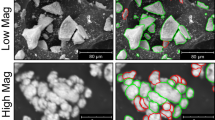
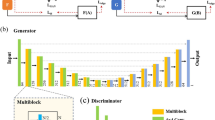
Scanning Electron Microscope (SEM) image analysis plays a vital role in semiconductor thin-film characterisation. In particular, defect detection and classification are performed using SEM images. However, conventional methods rely on labelled datasets or handcrafted features for classification, which limits their generalisation in real-world industrial inspection settings. The present work proposes a self-supervised multi-resolution learning framework for label-agnostic morphology representation learning and clustering of semiconductor thin-film defects. It uses a SEM image Dataset (4591 images) obtained from industrial wafer inspection. The framework starts with image pre-processing to remove acquisition artifacts. It employs a multi-resolution image pyramid for capturing surface morphologies at fine, intermediate, and coarse spatial scales. A shared-weight convolutional encoder that ensures alignment between embeddings across the three resolutions is trained (on an unlabelled dataset) and utilised for unsupervised defect morphology grouping. The framework learns morphological representations without defect labels. However, defect labels are used only for post-hoc evaluation by normalized mutual information (NMI) and visualization. Intrinsic clustering metrics and low-dimensional visualization are used to assess the algorithm’s efficacy. Experimental results reveal that the proposed method, gray level co-occurrence matrix (GLCM), local binary patterns (LBP), wavelet-based features, and principal component analysis (PCA) on raw pixels obtained a silhouette score of 0.50, 0.43, 0.31, 0.45, and 0.22, respectively. While normalized mutual information (NMI) values remained moderate across the models. These results reflect the label-agnostic nature of the proposed SSL framework. Further, UMAP and t-SNE visualizations confirm the coherent manifold structure and the effectiveness of morphology-driven grouping. These results demonstrate the robust, scale-invariant quality of the proposed self-supervised multi-resolution learning framework for defect clustering.
Similar content being viewed by others
Data availability
The data that support the findings of this study are openly available in [zenodo] at [https://zenodo.org/records/10715190](https:/zenodo.org/records/10715190) reference number24.
References
Ho, S. & Ejikeme, E. I. Investigation of nanostructured thin films using scanning electron microscopy technique. Int. J. Eng. Trends Technol. https://doi.org/10.14445/22315381/IJETT-V73I2P103 (2025) (Article P103).
Sivaraj, S. et al. A comprehensive review on current performance, challenges and progress in thin-film solar cells. Energies 15(22), 8688. https://doi.org/10.3390/en15228688 (2022).
Ren, Z. et al. State of the art in defect detection based on machine vision. Int. J. of Precis. Eng. and Manuf.-Green Tech 9, 661–691. https://doi.org/10.1007/s40684-021-00343-6 (2022).
Haralick, R. M., Shanmugam, K. & Dinstein, I. Textural Features for Image Classification, in IEEE Transactions on Systems, Man, and Cybernetics, vol. SMC-3, no. 6, pp. 610–621, https://doi.org/10.1109/TSMC.1973.4309314(1973).
Ojala, T., Pietikainen, M. & Maenpaa, T. Multiresolution gray-scale and rotation invariant texture classification with local binary patterns, in IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 24, no. 7, pp. 971–987, https://doi.org/10.1109/TPAMI.2002.1017623(2002).
Unser, M. Texture classification and segmentation using wavelet frames, in IEEE Transactions on Image Processing, vol. 4, no. 11, pp. 1549–1560, https://doi.org/10.1109/83.469936(1995).
Brian, L., DeCost, E. A. & Holm,. A computer vision approach for automated analysis and classification of microstructural image data. Comput. Mater. Sci. https://doi.org/10.1016/j.commatsci.2015.08.011 (2015).
Maxim Ziatdinov, O. et al. Deep Learning of Atomically Resolved Scanning Transmission Electron Microscopy Images: Chemical Identification and Tracking Local Transformations. ACS Nano, https://doi.org/10.1021/acsnano.7b07504.
Taekyeong Park, Y. et al. Long-tailed detection and classification of wafer defects from scanning electron microscope images robust to diverse image backgrounds and defect scales. Engineering Applications of Artificial Intelligence 162, 112342. https://doi.org/10.1016/j.engappai.2025.112342 (2025).
Chen, T., Kornblith, S., Norouzi, M. & Hinton, G. A simple framework for contrastive learning of visual representations. In Proceedings of the 37th International Conference on Machine Learning (ICML’20), Vol. 119. JMLR.org, Article 149, 1597–1607. https://arxiv.org/abs/2002.05709(2020).
Grill, J. B., Strub, F., Altché, F., Tallec, C. & Richemond, P. H. Elena Buchatskaya, Carl Doersch, Bernardo Avila Pires, Zhaohan Daniel Guo, Mohammad Gheshlaghi Azar, Bilal Piot, Koray Kavukcuoglu, Rémi Munos, and Michal Valko. Bootstrap your own latent a new approach to self-supervised learning. In Proceedings of the 34th International Conference on Neural Information Processing Systems (NIPS ‘20). Curran Associates Inc., Red Hook, NY, USA, Article 1786, 21271–21284. (2020).
Chen, X. & He, K. Exploring Simple Siamese Representation Learning, IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Nashville, TN, USA, 2021, pp. 15745–15753, Nashville, TN, USA, 2021, pp. 15745–15753, https://doi.org/10.1109/CVPR46437.2021.01549(2021).
Ma, J. et al. Review of wafer surface defect detection methods. Electronics 12(8), 1787. https://doi.org/10.3390/electronics12081787 (2023).
Kim, T. Y. et al. Deep learning-based detection of defects in wafer buffer zone during semiconductor packaging process. Multiscale Sci. Eng. 6, 25–32. https://doi.org/10.1007/s42493-024-00103-z (2024).
López de la Rosa, F., Gómez-Sirvent, J. L., Morales, R., Sánchez-Reolid, R. & Fernández-Caballero, A. Defect detection and classification on semiconductor wafers using two-stage geometric transformation-based data augmentation and SqueezeNet lightweight convolutional neural network. Computers & Industrial Engineering 183, 109549. https://doi.org/10.1016/j.cie.2023.109549 (2023).
Zheng, J., Dang, J. & Zhang, T. Wafer surface defect detection based on feature enhancement and predicted box aggregation. Electronics 12(1), 76. https://doi.org/10.3390/electronics12010076 (2023).
Zheng, J. & Zhang, T. Wafer surface defect detection based on background subtraction and faster R-CNN. Micromachines 14(5), 905. https://doi.org/10.3390/mi14050905 (2023).
Kim, M., Tak, J. & Shin, J. A deep learning model for wafer defect map classification: Perspective on classification performance and computational volume. Phys. Status Solidi B 261, 2300113. https://doi.org/10.1002/pssb.202300113 (2024).
Mayank Jariya, P., Kumar, R., Devi, B. & Singh Silicon wafer defect pattern detection using machine learning, Materials Today: Proceedings, ISSN 2214–7853, https://doi.org/10.1016/j.matpr.2023.04.233(2023).
Chien, J.-C., Wu, M.-T. & Lee, J.-D. Inspection and classification of semiconductor wafer surface defects using CNN deep learning networks. Appl. Sci. 10(15), 5340. https://doi.org/10.3390/app10155340 (2020).
Lee, J. et al. Enhancing Confidence and Interpretability of a CNN-Based Wafer Defect Classification Model Using Temperature Scaling and LIME. Micromachines (Basel) 16(9), 1057. https://doi.org/10.3390/mi16091057 (2025).
Yoon, S. & Kang, S. Semi-automatic wafer map pattern classification with convolutional neural networks. Comput. Ind. Eng. 166, 0360–8352. https://doi.org/10.1016/j.cie.2022.107977 (2022).
Manivannan, S. Semi-supervised imbalanced classification of wafer bin map defects using a Dual-Head CNN. Expert Sys. Appl. 238, 122301. https://doi.org/10.1016/j.eswa.2023.122301 (2024).
Dataset available at: https://zenodo.org/records/10715190
Funding
Open access funding provided by Symbiosis International (Deemed University).
Author information
Authors and Affiliations
Contributions
U.K. conceived the study and contributed to the experimental design. C.K.C. assisted in methodology development and technical guidance. C.S. contributed to data analysis and interpretation of results. U.K and A.V. supervised the research work and coordinated the overall project. S.S. contributed to software work and validation of results. D.V. assisted in data collection, visualization, and manuscript preparation. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Krishnamoorthy, U., Chan, C.K., Sonawane, C. et al. Self-supervised multi-resolution learning for label-agnostic morphology representation and clustering of semiconductor thin-film SEM defects. Sci Rep (2026). https://doi.org/10.1038/s41598-026-46947-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-46947-3