Abstract
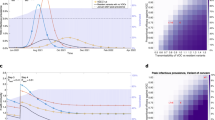
During the global COVID-19 pandemic, different SARS-CoV-2 variants of concern (VOCs) emerged, causing massive transmission waves that affected infection outcomes, economies and public health. From January to July 2021 in Uruguay, the transmission wave was primarily driven by the Gamma VOC amidst a largely immunologically naive population. Following the implementation of a nationwide vaccination campaign, the detection of the Delta VOC in July 2021 did not trigger a significant transmission wave, even in the face of increased mobility during the winter holidays (late June to mid-July). We present a comprehensive analysis of the dynamics of SARS-CoV-2 in Uruguay until September 2021. By analysing 1792 viral genomes, we integrate genomic data with vaccination records, variant surveillance and epidemiological information at both regional and global scales. Our study highlights the role of timely vaccination combined with real-time genomic surveillance programmes in effectively monitoring and mitigating critical phases of the COVID-19 epidemic.
Similar content being viewed by others
Data availability
SARS-CoV-2 genomic consensus sequences generated in this study can be found in the EpiCoV/GISAID database, available at: https://gisaid.org. The accession numbers and metadata available can be found in Table S3. Additionally, the South America build for the background dataset used for the phylodynamic analysis was retrieved from EpiCoV/GISAID on December 31, 2021. Phylogenetic data generated in this study were deposited in Mendeley Data (https://data.mendeley.com/preview/2y4xjcm84s?a=ba557727-f20b-4bac-a08e-a49b60ae3b0a). Epidemiological and statistical data from the COVID-19 pandemic and population mobility in Uruguay (Grupo Uruguayo Interdisciplinario de Análisis de Datos de COVID-19) are available at: https://guiad-covid.github.io/. Vaccination data (Catálogo de datos abiertos, Ministerio de Salud Pública) is available at: https://catalogodatos.gub.uy/dataset/vacunacion-por-covid-19. Global statistics are available at: https://ourworldindata.org/. All data analysis was performed using the following open source software: PoreCov pipeline is available through the Nextflow system at: https://github.com/replikation/poreCov. Pangolin is available through Bioconda at: https://github.com/cov-lineages/pangolin; Nextclade CLI is available through Docker hub at: https://hub.docker.com/r/nextstrain/nextclade. Custom R scripts for tables and data visualization are available in the project GitHub repository at: https://github.com/Ceci07/SARS-CoV-2_URU.
References
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020).
Lu, R. et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 395, 565–574 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
WHO. Tracking SARS-CoV-2 variants. https://www.who.int/activities/tracking-SARS-CoV-2-variants (2022).
Hanly, P., Ahern, M., Sharp, L., Ursul, D. & Loughnane, G. The cost of lost productivity due to premature mortality associated with COVID-19: A Pan-European study. Eur. J. Health Econ. 23, 249–259 (2022).
Yeyati, E. L. & Filippini, F. Social and economic impact of COVID-19.
Harvey, W. T. et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat. Rev. Microbiol. 19, 409–424 (2021).
Elizondo, V. et al. SARS-CoV-2 genomic characterization and clinical manifestation of the COVID-19 outbreak in Uruguay. Emerg. Microbes Infect. 10, 51–65 (2021).
Salazar, C. et al. Multiple introductions, regional spread and local differentiation during the first week of COVID-19 epidemic in Montevideo. Uruguay. Preprint at https://doi.org/10.1101/2020.05.09.086223 (2020).
Moreno, P. et al. An effective COVID-19 response in South America: The Uruguayan Conundrum. Preprint at https://doi.org/10.1101/2020.07.24.20161802 (2020).
Mir, D. et al. Recurrent dissemination of SARS-CoV-2 through the Uruguayan-Brazilian Border. Front. Microbiol. 12, 653986 (2021).
Faria, N. R. et al. Genomics and epidemiology of the P.1 SARS-CoV-2 lineage in Manaus, Brazil. Science 372, 815–821 (2021).
Naveca, F. G. et al. COVID-19 in Amazonas, Brazil, was driven by the persistence of endemic lineages and P.1 emergence. Nat. Med. 27, 1230–1238 (2021).
Chen, R. E. et al. Resistance of SARS-CoV-2 variants to neutralization by monoclonal and serum-derived polyclonal antibodies. Nat. Med. 27, 717–726 (2021).
Wang, Z. et al. mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants. Nature 592, 616–622 (2021).
Cherian, S. et al. Convergent evolution of SARS-CoV-2 spike mutations, L452R, E484Q and P681R, in the second wave of COVID-19 in Maharashtra. India. Preprint at https://doi.org/10.1101/2021.04.22.440932 (2021).
del Rio, C., Malani, P. N. & Omer, S. B. Confronting the Delta variant of SARS-CoV-2, summer 2021. JAMA 326, 1001–1002 (2021).
Atherstone, C. J. et al. COVID-19 Epidemiology during delta variant dominance period in 45 high-income countries, 2020–2021. Emerg. Infect. Dis. J. CDC 29(9). https://doi.org/10.3201/eid2909.230142 (2023).
Singh, J., Rahman, S. A., Ehtesham, N. Z., Hira, S. & Hasnain, S. E. SARS-CoV-2 variants of concern are emerging in India. Nat. Med. 27, 1131–1133 (2021).
Attwood, S. W., Hill, S. C., Aanensen, D. M., Connor, T. R. & Pybus, O. G. Phylogenetic and phylodynamic approaches to understanding and combating the early SARS-CoV-2 pandemic. Nat. Rev. Genet. 23, 547–562 (2022).
Rego, N. et al. Emergence and spread of a B.1.1.28-derived P.6 lineage with Q675H and Q677H spike mutations in Uruguay. Viruses 13, 1801 (2021).
Rego, N. et al. Real-time genomic surveillance for SARS-CoV-2 variants of concern, Uruguay. Emerg. Infect. Dis. 27, 2957–2960 (2021).
Rambaut, A. et al. COVID-19 Genomics Consortium UK (CoG-UK). Preliminary genomic characterisation of an emergent SARS-CoV-2 lineage in the UK defined by a novel set of spike mutations. virological.org https://virological.org/t/preliminary-genomic-characterisation-of-an-emergent-sars-cov-2-lineage-in-the-uk-defined-by-a-novel-set-of-spike-mutations/563 (2020).
Salazar, C. et al. Case report: Early transcontinental import of SARS-CoV-2 variant of concern 202012/01 (B.1.1.7) From Europe to Uruguay. Front. Virol. 1, (2021).
Bolze, A. et al. SARS-CoV-2 variant Delta rapidly displaced variant Alpha in the United States and led to higher viral loads. Cell Rep. Med. 3, 100564 (2022).
Layan, M. et al. Impact and mitigation of sampling bias to determine viral spread: Evaluating discrete phylogeography through CTMC modeling and structured coalescent model approximations. Virus Evol. 9, vead010 (2023).
Freed, N. E., Vlková, M., Faisal, M. B. & Silander, O. K. Rapid and inexpensive whole-genome sequencing of SARS-CoV-2 using 1200 bp tiled amplicons and Oxford Nanopore Rapid Barcoding. Biol. Methods Protocols 5, bpaa014 (2020).
Tyson, J. R. et al. Improvements to the ARTIC multiplex PCR method for SARS-CoV-2 genome sequencing using nanopore. Preprint at https://doi.org/10.1101/2020.09.04.283077 (2020).
Brandt, C. et al. poreCov-an easy to use, fast, and robust workflow for SARS-CoV-2 genome reconstruction via nanopore sequencing. Front. Genet. 12, 711437 (2021).
Rambaut, A. et al. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 5, 1403–1407 (2020).
O’Toole, Á. et al. Assignment of epidemiological lineages in an emerging pandemic using the Pangolin tool. Virus Evol. 7, veab064 (2021).
Aksamentov, I., Roemer, C., Hodcroft, E. B. & Neher, R. A. Nextclade: Clade assignment, mutation calling and quality control for viral genomes. J. Open Source Softw. 6, 3773 (2021).
Nguyen, L.-T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016).
Sagulenko, P., Puller, V. & Neher, R. A. TreeTime: Maximum-likelihood phylodynamic analysis. Virus Evol. 4 (2018).
Acknowledgements
Authors would like to acknowledge the invaluable efforts of all the members of the Interinstitutional Working Group (IiWG) and the institutions involved in the COVID-19 testing and sample/data collection in Uruguay during the pandemic. Authors would also like to acknowledge the Grupo Uruguayo Interdisciplinario de Análisis de Datos de COVID-19 (https://guiad-covid.github.io/) for collecting and openly sharing COVID-19 epidemiological data of Uruguay. We also acknowledge all the authors and institutions around the world that shared SARS-CoV-2 genomic sequences through the EpiCoV/GISAID database. A detailed description of the authors of sequences used in this study is available in EPI_SET_240229hw (https://doi.org/10.55876/gis8.240229hw).
Disclaimer
The opinions expressed in this article are those of the authors and do not reflect the view of the National Institutes of Health, the Department of Health and Human Services, or the United States government.
Funding
FOCEM COF 03/11, G4 program, Institut Pasteur de Montevideo funded by the BSE of Uruguay, Fondo de Solidaridad para Proyectos Innovadores, Sociedad Civil, Francofonía y Desarrollo Humano, Ambassade de France, Centro Latinoamericano de Biotecnología, Agencia Nacional de Investigación e Innovación, International Programme, British Embassy Montevideo, Uruguay and Fundación Manuel Perez, Facultad de Medicina, Universidad de la República.
Author information
Authors and Affiliations
Consortia
Contributions
CS: conceptualization, data generation, analysis, manuscript drafting, editing, and review. NST: conceptualization, analysis, manuscript editing, and review. IF: analysis, manuscript drafting and review. AC, MP, PP: data generation and manuscript review. AM, VB, RC, BR, MA, MM: data generation. NR, TFC, LS: funding, conceptualization, manuscript editing and review. GI: funding, conceptualization, analysis, manuscript drafting. GM: funding, conceptualization, work lab coordination and supervision, manuscript drafting, editing and review. PM: funding, conceptualization, work lab coordination and supervision, manuscript drafting, editing and review.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Salazar, C., Trovão, N.S., Ferrés, I. et al. Impact of timely vaccination and genomic surveillance on controlling consecutive waves of SARS-CoV-2 variants. Sci Rep (2026). https://doi.org/10.1038/s41598-026-48131-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-48131-z