Abstract
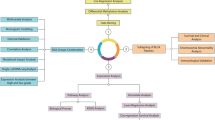
Considering that the circulating proteome represents a primary source of candidate biomarkers and therapeutic targets, we conducted a large-scale Mendelian randomization (MR) study to identify plasma proteins potentially involved in the pathogenesis and treatment of common urological cancers (UCs), including bladder cancer (BC), prostate cancer (PC), renal cell carcinoma (RCC), and testicular cancer (TC). Cis-protein quantitative trait loci (cis-pQTLs) were derived from two large-scale genome-wide association studies (GWASs) of plasma proteomes. GWAS for UCs were obtained from FinnGen and pan-UKBB meta-analysis, FinnGen and UK Biobank. Colocalization analysis and summary data-based MR (SMR) were performed to evaluate the robustness of the associations. Further evaluations involved Bulk RNA-seq differential expression and single cell-type expression analysis, protein–protein interaction, and druggability evaluation. For BC, we identified four protein markers: one associated with increased risk (PSCA) and three with decreased risk (GSTM1, GSTM3, GSTM4), which are mainly expressed in pericyte, urothelial, and NK cells in bladder tumors. Regarding PC, we found 15 protein markers: seven linked to increased risk (AGER, ALAD, CHMP2B, PEX14, ZG16B, PPP1R14A, SERPINA3) and eight to decreased risk (BTN2A1, CEACAM21, DNAJB9, MSMB, PYGL, HLA-E, SOD2, TOR1AIP1), enriched in epithelial cells, monocytes/macrophages in prostate tumors. The strongest evidence from MR and colocalization analyses supported GSTM4 (BC), SOD2 and CHMP2B (PC) as causal markers. Notably, seven identified proteins have been previously targeted by drugs for other cancers and immune disorders, indicating their potential therapeutic relevance in UCs. This study highlights several proteins with predictive value for BC and PC risk and supports their utility in biomarker discovery and drug development.
Similar content being viewed by others
Acknowledgements
The authors sincerely thank the authors who shared the original dataset in this study. This study was funded by 1.3.5 project for disciplines of excellence, West China Hospital, Sichuan University (ZYYC25003), the National Natural Science Fund of China (32471519, 82422015, 82270720 and 82400904).
Funding
National Natural Science Fund of China, 32471519, 82422015,82270720 and 82400904, 1.3.5 project for disciplines of excellence, West China Hospital, Sichuan University, ZYYC25003.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lyu, X., Peng, L., Fan, Y. et al. Identification of novel protein markers and therapeutic targets for common urological cancers by integrating large-scale human plasma proteome with the genome. Sci Rep (2026). https://doi.org/10.1038/s41598-026-49333-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-49333-1