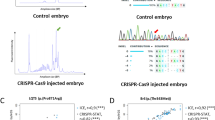
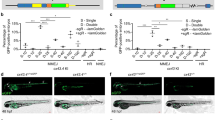
Abstract
Zebrafish serve as a valuable model organism for studying human genetic diseases. While generating knockout lines is relatively straightforward, introducing precise disease-specific genetic variants by knock-in (KI) remains challenging. KI lines, however, enable more accurate studies of molecular and physiological consequences of genetic diseases. Their generation is often hampered by low editing efficiency (EE) and potential off-target effects. Here, we optimized conventional CRISPR–Cas9-mediated homology-directed repair (HDR) strategies for precise KI of genetic variants in zebrafish and compared their efficacy with prime editing, a recently developed technique that is not yet commonly used. Using next-generation sequencing, we determined KI EE by HDR for six unique base-pair substitutions in three different zebrafish genes. We assessed the effect of (1) varying Cas9 amounts, (2) HDR templates with chemical modifications to improve integration efficiency, (3) different microinjection procedures and (4) introduction of additional synonymous guide-blocking variants in the HDR template. Increasing Cas9 amounts augmented KI EE, with optimal injected amounts of Cas9 between 200 pg and 800 pg. The use of Alt-R HDR templates further increased KI EE, while guide-blocking modifications did not. Injecting components directly into the cell was not superior to injections into the yolk. Prime editing, however, increased EE up to fourfold and expanded the F0 founder pool for four targets compared with conventional HDR editing, with fewer off-target effects. Therefore, prime editing is a very promising methodology for improving the creation of precise genomic edits in zebrafish, facilitating the modeling of human diseases.
This is a preview of subscription content, access via your institution
Access options






Similar content being viewed by others
Data availability
All output values of BATCH-GE for IF and EE are provided and summarized in the supplementary tables. The raw fastq files that support the findings of this study are available from the corresponding authors upon request.
References
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Adhish, M. & Manjubala, I. Effectiveness of zebrafish models in understanding human diseases—a review of models. Heliyon 9, e14557 (2023).
Zang, L., Torraca, V., Shimada, Y. & Nishimura, N. Editorial: zebrafish models for human disease studies. Front. Cell Dev. Biol. 10, 861941 (2022).
El-brolosy, M. A., Kontarakis, Z., Rossi, A. & Kuenne, C. Genetic compensation triggered by mutant mRNA degradation. Nature 568, 193–197 (2019).
Serobyan, V. et al. Transcriptional adaptation in Caenorhabditis elegans. eLife https://doi.org/10.7554/elife.50014 (2020).
Ma, Z. et al. PTC-bearing mRNA elicits a genetic compensation response via Upf3a and COMPASS components. Nature 568, 259–263 (2019).
Pottie, L. et al. Loss of zebrafish atp6v1e1b, encoding a subunit of vacuolar ATPase, recapitulates human ARCL type 2C syndrome and identifies multiple pathobiological signatures. PLoS Genet. 17, e1009603 (2021).
Yang, H. et al. Methods favoring homology-directed repair choice in response to CRISPR/Cas9 induced-double strand breaks. Int. J. Mol. Sci. 21, 6461 (2020).
Salsman, J. & Dellaire, G. Precision genome editing in the CRISPR era. Biochem. Cell Biol. 95, 187–201 (2017).
Shakirova, A., Karpov, T., Komarova, Y. & Lepik, K. In search of an ideal template for therapeutic genome editing: a review of current developments for structure optimization. Front. Genome Ed. 5, 1068637 (2023).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Chai, R. et al. Gene editing by SSB/CRISPR–Cas9 ribonucleoprotein in bacteria. Int. J. Biol. Macromol. 278, 135065 (2024).
Bai, H. et al. CRISPR/Cas9-mediated precise genome modification by a long ssDNA template in zebrafish. BMC Genomics https://doi.org/10.1186/s12864-020-6493-4 (2020).
Chen, F. et al. High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases. Nat. Methods 8, 753–755 (2011).
Boel, A. et al. CRISPR/Cas9-mediated homology-directed repair by ssODNs in zebrafish induces complex mutational patterns resulting from genomic integration of repair-template fragments. Dis. Models Mech. 11, dmm035352 (2018).
Prill, K. & Dawson, J. F. Homology-directed repair in zebrafish: witchcraft and wizardry? Front. Mol. Biosci. 7, 595474 (2020).
Prykhozhij, S. V. & Berman, J. N. Mutation knock-in methods using single-stranded dna and gene editing tools in zebrafish. Methods Mol. Biol. 2707, 279–303 (2024).
Prykhozhij, S. V. et al. Optimized knock-in of point mutations in zebrafish using CRISPR/Cas9. Nucleic Acids Res. 46, 17 (2018).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Zhao, Z., Shang, P., Mohanraju, P. & Geijsen, N. Prime editing: advances and therapeutic applications. Trends Biotechnol. 41, 1000–1012 (2023).
Gistelinck, C. et al. Zebrafish type I collagen mutants faithfully recapitulate human type I collagenopathies. Proc. Natl Acad. Sci. USA 115, E8037–E8046 (2018).
Vierstraete, J. et al. Accurate quantification of homologous recombination in zebrafish: brca2 deficiency as a paradigm. Sci. Rep. 7, 16518 (2017).
Vierstraete, J., Fieuws, C., Willaert, A., Vral, A. & Claes, K. B. M. Zebrafish as an in vivo screening tool to establish PARP inhibitor efficacy. DNA Repair 97, 103023 (2021).
Van Damme, T. et al. Mutations in ATP6V1E1 or ATP6V1A cause autosomal-recessive cutis laxa. Am. J. Hum. Genet. 100, 216–227 (2017).
Richardson, M. E. et al. Strong functional data for pathogenicity or neutrality classify BRCA2 DNA-binding-domain variants of uncertain significance. Am. J. Hum. Genet. 108, 458–468 (2021).
Chen, Q., Zhang, Y. & Yin, H. Recent advances in chemical modifications of guide RNA, mRNA and donor template for CRISPR-mediated genome editing. Adv. Drug Deliv. Rev. 168, 246–258 (2021).
Schubert, M. S. et al. Optimized design parameters for CRISPR Cas9 and Cas12a homology-directed repair. Sci. Rep. 11, 19482 (2021).
Okamoto, S., Amaishi, Y., Maki, I., Enoki, T. & Mineno, J. Highly efficient genome editing for single-base substitutions using optimized ssODNs with Cas9-RNPs. Sci. Rep. 9, 4811 (2019).
Concordet, J. P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Liang, X., Potter, J., Kumar, S., Ravinder, N. & Chesnut, J. D. Enhanced CRISPR/Cas9-mediated precise genome editing by improved design and delivery of gRNA, Cas9 nuclease, and donor DNA. J. Biotechnol. 241, 136–146 (2017).
O'Brien, A. R., Wilson, L. O. W., Burgio, G. & Bauer, D. C. Unlocking HDR-mediated nucleotide editing by identifying high-efficiency target sites using machine learning. Sci. Rep. 9, 2788 (2019).
Wang, K. et al. Efficient generation of orthologous point mutations in pigs via CRISPR-assisted ssODN-mediated homology-directed repair. Mol. Ther. Nucleic Acids 5, e396 (2016).
Mao, Z., Bozzella, M., Seluanov, A. & Gorbunova, V. Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair 7, 1765–1771 (2008).
Ferree, P. L., Deneke, V. E. & Di Talia, S. Measuring time during early embryonic development. Semin. Cell Dev. Biol. 55, 80–88 (2016).
Aksoy, Y. A. et al. Chemical reprogramming enhances homology-directed genome editing in zebrafish embryos. Commun. Biol. 2, 198 (2019).
Yuan, S. & Sun, Z. Microinjection of mRNA and morpholino antisense oligonucleotides in zebrafish embryos. J. Vis. Exp. https://doi.org/10.3791/1113 (2009).
Kroll, F. et al. A simple and effective F0 knockout method for rapid screening of behaviour and other complex phenotypes. eLife https://doi.org/10.7554/elife.59683 (2021).
Cartegni, L., Chew, S. L. & Krainer, A. R. Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nat. Rev. Genet. 3, 285–298 (2002).
Cartegni, L., Wang, J., Zhu, Z., Zhang, M. Q. & Krainer, A. R. ESEfinder: a web resource to identify exonic splicing enhancers. Nucleic Acids Res. 31, 3568–3571 (2003).
Findlay, G. M. et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature 562, 217–222 (2018).
Iwata, H. & Gotoh, O. Comparative analysis of information contents relevant to recognition of introns in many species. BMC Genomics 12, 45 (2011).
Jaganathan, K. et al. Predicting splicing from primary sequence with deep learning. Cell 176, 535–548 (2019).
Kim, K. et al. Highly efficient RNA-guided base editing in mouse embryos. Nat. Biotechnol. 35, 435–437 (2017).
Zong, Y. et al. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 35, 438–440 (2017).
Li, G. et al. Highly efficient and precise base editing in discarded human tripronuclear embryos. Protein Cell 8, 776–779 (2017).
Zhang, Y. et al. Programmable base editing of zebrafish genome using a modified CRISPR–Cas9 system. Nat. Commun. 8, 118 (2017).
Ponnienselvan, K. et al. Reducing the inherent auto-inhibitory interaction within the pegRNA enhances prime editing efficiency. Nucleic Acids Res. 51, 6966–6980 (2023).
Petri, K. et al. CRISPR prime editing with ribonucleoprotein complexes in zebrafish and primary human cells. Nat. Biotechnol. 40, 189–193 (2022).
Zhang, W. et al. Enhancing CRISPR prime editing by reducing misfolded pegRNA interactions. eLife 12, RP90948 (2024).
Mathis, N. et al. Predicting prime editing efficiency and product purity by deep learning. Nat. Biotechnol. 41, 1151–1159 (2023).
Tsai, H. H. et al. Whole genomic analysis reveals atypical non-homologous off-target large structural variants induced by CRISPR–Cas9-mediated genome editing. Nat. Commun. 14, 5183 (2023).
Aleström, P. et al. Zebrafish: housing and husbandry recommendations. Lab Anim. 54, 213–224 (2020).
Westerfield, M. The Zebrafish Book; A Guide for the Laboratory Use of Zebrafish (Danio rerio) 5th edn (Univ. of Oregon Press, 2007).
Chow, R. D., Chen, J. S., Shen, J. & Chen, S. A web tool for the design of prime-editing guide RNAs. Nat. Biomed. Eng. 5, 190–194 (2021).
De Leeneer, K. et al. Flexible, scalable, and efficient targeted resequencing on a benchtop sequencer for variant detection in clinical practice. Hum. Mutat. 36, 379–387 (2015).
Boel, A. et al. BATCH-GE: batch analysis of next-generation sequencing data for genome editing assessment. Sci. Rep. 6, 30330 (2016).
Acknowledgements
We thank the Zebrafish Facility Ghent Core at Ghent University, and particularly K. Vermeulen for the diligent care for the zebrafish. We also thank the VIB Protein Core for the generation of the prime editor and J. K. Joung for the donation of the pET-PE2-His plasmid (Addgene plasmid no. IK1822). This research was funded by a grant from the Research Foundation – Flanders (grant no. FWOOPR2020009501) to B.C. and A.W., a Concerted Research Action grant from the Ghent University Special Research Fund (grant no. BOF GOA019-21) to B.C. and a Ghent University Special Research Fund (grant no. BOF21/DOC/242) to E.D.N. B.C. is a senior clinical investigator of the Research Foundation – Flanders.
Author information
Authors and Affiliations
Contributions
Conceptualization: A.B., A.W., B.C. and K.B.M.C. Experimental procedures: M.V., E.D.N. and H.D.S. Data analysis: M.V., E.D.N. and H.D.S. Visualization and writing: M.V. and E.D.N. Original draft preparation: M.V. and E.D.N. Review and editing: M.V., E.D.N., A.W., B.C. and K.B.M.C. Supervision: A.B., A.W., B.C. and K.B.M.C. Funding acquisition: A.W., B.C. and K.B.M.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Lab Animal thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods: generation of the PE protein.
Supplementary Tables (download XLSX )
Supplementary Tables 1–8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vanhooydonck, M., De Neef, E., De Saffel, H. et al. Prime editing outperforms homology-directed repair as a tool for CRISPR-mediated variant knock-in in zebrafish. Lab Anim 54, 165–172 (2025). https://doi.org/10.1038/s41684-025-01560-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41684-025-01560-1
This article is cited by
-
Evaluating variants of uncertain significance in adult zebrafish via prime editing: a proof of concept with a COL1A2 variant
BMC Medical Genomics (2025)
-
O-GlcNAcylation in novel regulated cell death: ferroptosis, pyroptosis, and necroptosis
Cell Death Discovery (2025)
-
Morpholino Knockdown in Zebrafish: A Tool to Investigate the Functional Impact of Variants of Unknown Significance in Mitochondrial Diseases
NeuroMolecular Medicine (2025)