Abstract
This study developed and validated a deep learning network using baseline magnetic resonance imaging (MRI) to predict Ki-67 status in meningioma patients. A total of 1239 patients were retrospectively recruited from three hospitals between January 2010 and December 2023, forming training, internal validation, and two external validation cohorts. A representation learning framework was utilized for modeling, and performance was assessed against existing methods. Furthermore, Kaplan–Meier survival analysis was conducted to investigate whether the model could be used for tumor growth prediction. The model achieved superior results, with areas under the curve (AUCs) of 0.797 for internal testing and 0.808 for generalization, alongside 0.756 and 0.727 for 3- and 5-year tumor growth predictions, respectively. The prediction was significantly associated with the growth of asymptomatic small meningiomas. Overall, the model provides an effective tool for early prediction of Ki-67 and tumor volume growth, aiding in individualized patient management.
Similar content being viewed by others

Introduction
Meningiomas are the most common primary intracranial tumors with an annual incidence of 9.73 per 100,000 individuals1,2. Magnetic resonance imaging (MRI) is the cornerstone in patient management as it plays the most important role in tumor diagnosis, follow-up, and treatment planning3. For symptomatic meningioma, complete surgical resection with/without adjuvant radiotherapy is recommended as it can reach favorable prognosis in most cases4. In contrast, asymptomatic meningiomas of small size (maximal diameter less than 3 cm) are conservatively managed through radiological follow-up, with the decision for active intervention determined by development of clinical symptoms or radiological evidence of tumor growth5. Recognizing the importance in both prognostic risk stratification and therapeutic decision making, there is a growing emphasis on achieving individualized patient management and prognosis evaluation6.
Recent studies have provided accumulated evidence regarding the clinical significance of Ki-67 index in individualized patient care. As a classical bio-marker of cell proliferation, it can be used for identifying asymptomatic small meningioma patients who need early intervention. Additionally, it holds promise as prognostic indicator related to both increased recurrent risk after surgical resection7 and poor progression-free survival (PFS) after post-operative stereotactic radiosurgery8. The gold standard for Ki-67 assessment remains immunohistochemistry staining of tumor specimens obtained from biopsy or surgery9. Nevertheless, the invasiveness of these approaches imposes significantly increased physical burden and risks of complication. Hence, there is a critical need to identify an alternative approach for accurately assessing Ki-67 expression in meningioma cases based on information obtained from MRI.
Previous research has identified a series of radiological characteristics on routine MRI that independently correlated with Ki-67 expression10. Parameters obtained from advanced MR technology, such as the relative apparent diffusion coefficient (rADC), may also potentially serve as independent predictors11,12,13,14. Yet, accurate prediction by human being remains challenging as these indicators remain controversial, imprecise, relying on clinical expertize15, nor advanced MR technology is routinely performed in patient management.
Multi-modal representation learning models have drawn great attention in recent years. Presented as Transformer-based structure16,17, they have demonstrated a remarkable adaptability in medical-related tasks and offered distinctive advantages. By leveraging cross-attention module to interact and integrate features of multi-modal inputs, both performance and generalization can be enhanced with redundancy of information, therefore representing the logical next step18,19,20. Returning to this topic, radiological characteristics, radiomics features, and deep learning features are interconnected but distinct from the perspective of medical imaging analysis21. Nevertheless, to the best of knowledge, the potential of multi-modal representation learning in prediction of Ki-67 on MRI for meningiomas has not been investigated yet.
Thus, in this study, with a large multicenter dataset collected from three healthcare institutions, we aimed to developed and validated a multi-modal representation learning model to predict the Ki-67 index for meningiomas.
Results
Resected meningioma cohort
Table 1 provides a full overview of the patient demographics and baseline clinical characteristics. Generally, a total of 1008 patients with resected meningioma were selected in this study. The mean patient age was 55.2 year (range: 27–76), and 70.0% (n = 705) of the study cohort was female. A total of 296 patients presented with skull base meningioma, and the other 712 patients presented with non-skull base meningioma.
There were 791 patients with Ki-67 < 5% and 217 patients with Ki-67 of ≥ 5%. The univariate analysis of the relationship between radiological characteristics and Ki-67 index is provided in the Supplementary Table 3. As suggested, meningiomas with Ki-67 ≥ 5% were larger in volume compared to tumors with Ki-67 < 5% (mean 46.3 ± 48.2 and 27.0 ± 25.9 cm3; p < 0.001), Similarly, meningiomas with Ki-67 ≥ 5% had significantly larger EI compared to tumors with Ki-67 < 5% (p < 0.001), and also more likely to present intra-tumoral necrosis (p = 0.008). Whereas, there was no statistic difference regarding to CSF cleft surrounding tumor (p = 0.493) or absent capsular enhancement (p = 0.315) between two groups. As presented in the Supplementary Table 4, multivariate analysis suggested that intra-tumoral necrosis was the only characteristic independently associated with Ki-67 expression (Odds ratio (OR) = 4.048; 95CI: 2.878–5.694; p < 0.001).
Model performance and generalization
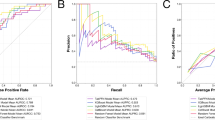
The results of ablation experiment to determine the suitable input are provided in the Supplementary Note 3. Table 2 presents the results of setting different combinations of modalities as input. The results suggested that setting combination of 3D MRIs, radiological characteristics, and Radiomics as input was the best one as it achieved the highest AUC as well as the MCC. More specifically, Output 1, utilizing MRI and radiological characteristics as input, exhibited relatively better performance. In the internal test, the model achieved an AUC of 0.797 (95CI: 0.758–0.837), accuracy of 0.748, sensitivity of 0.739, specificity of 0.760, F1 score of 0.746, and MCC of 0.495. The metrics demonstrated robustness in the external test as well, with an AUC of 0.808 (95CI: 0.726–0.895), accuracy of 0.769, sensitivity of 0.756, specificity of 0.771, F1 score of 0.755 and MCC of 0.537. The ROC curves of the model with closest performance to the average performance are illustrated in Fig. 1.
Receiver operating characteristics curves of different combinations of modalities as input with closest performance to the average performance of all 50 models in the internal validation cohort (A) and external validation cohort (B).
Supplementary Figure 2 presents representative Grad-CAM and cross-attention maps for predictions of both high Ki-67 and low Ki-67 cases. As suggested, the Grad-CAM analysis indicated that the MRI block highlighted the tumor region and extracted high-level features for prediction. Similarly, the cross-attention map demonstrated effective interaction between the blocks, wherein the model selectively attended to related MRI regions consistent with radiological characteristics during output generation. The weights of combined loss function in this sequential task were W mri = 0.5, W radiological characteristics = 0.4, W radiomics = 0.1, respectively.
Comparison with clinical model and the existing methods
The clinical model built with logistic regression presented performance with AUC of 0.664 (95CI: 0.624–0.710) in the internal test, and 0.506 (95CI: 0.439–0.587) in the external test. As comparing to the previously reported deep learning or machine learning methods, the best machine learning model achieved AUC of 0.746 (95CI: 0.717–0.780), accuracy of 0.655, sensitivity of 0.675, specificity of 0.629, F1 score of 0.675, and MCC of 0.299 in internal test, and AUC of 0.676 (95CI: 0.585–0.769), accuracy of 0.817, sensitivity of 0.314, specificity of 0.915 F1 score of 0.423, and MCC of 0.351 in external test. More detailed results were presented in the Supplementary Table 5 and Supplementary Table 6.
Prognostic value in tumor growth prediction
Significant tumor volume growth was observed in 22.5% (52 cases) within three years, and 33.8% (78 cases) within five years. Generally, as shown in Table 3, the predictive model (output 1) exhibited promising potential in anticipating tumor growth, achieving the following metrics for three-year tumor growth prediction (Fig. 2a): AUC of 0.756 (95CI: 0.722–0.793), accuracy of 0.753, sensitivity of 0.503, specificity of 0.825, F1 score of 0.641, and MCC of 0.321. For five-year tumor growth prediction, the model yielded an AUC of 0.727 (95CI: 0.699–0.756), accuracy of 0.721, sensitivity of 0.452, specificity of 0.859, F1 score of 0.588, and MCC of 0.305.
A ROC curves of the model in 3-years tumor growth prediction and 5-years tumor growth prediction; B Decision curves of Output 1 in predicting tumor growth of asymptomatic small meningiomas after radiological diagnosis in 5 years; C K–M curves of the low-risk group and high-risk group stratified by model; D the corresponding tumor growth status and time for each patient stratified by output 1; E Heatmaps of predictors according to multivariate Cox regression analyzes between high-risk and low-risk groups.
In survival analysis, the decision curves illustrated that the developed model was the more useful prediction model for clinical decision-making (Fig. 2b). Kaplan-Meier survival curves and log-rank analysis also demonstrated a significant statistical difference (p < 0.001) between the two groups categorized by the model (Fig. 2c). Subgroup analysis also suggested that our model can achieve satisfactory patient stratification, whether for skull-base meningiomas (p = 0.015) or non-skull-base meningiomas (p < 0.001) (Table 3 and Supplementary Figure 3). Furthermore, as presented in the Fig. 2d, the corresponding predicting probability along with their survival status and survival time for each patient in Dataset 3 is visualized. Supplementary Table 7 and Fig. 2e provided the results of univariate and multivariate Cox regression analyzes of predictors of tumor volume growth, suggesting that the prediction could be considered as an independent predictor (HR = 1.996; 95CI: 1.117–3.568; p = 0.020).
Discussion
This large-scale multicenter study proposed a multi-modal representation learning model to pre-surgically predict the Ki-67 index of meningiomas on MRI. The model achieved good performance along with generalization, and could be regarded as a useful tool to facilitate individualized management of asymptomatic meningioma patients.
Medical image analysis technology aims at capturing complicated patterns of tumor images that are imperceptible to naked eye22,23,24, previous studies applied voxel-level analysis and artificial intelligence (AI) algorithms for modeling. Both radiomic-based machine learning10,25,26 and deep learning27 approaches demonstrated the feasibility with their results, suggesting a non-invasive method of predicting the Ki-67 index in meningioma patients, as outlined in Supplementary Table 8. However, limited feasibility is still the primary concern from the perspective of clinical translation. It can be presented from three aspects. The first one is the sharp decline in model performance metrics when transitioning well-trained model from internal validation to independent external test. The objectives in medical images, especially referred to MRI, can be dramatically various in image pattern due to heterogeneous acquisition protocols, different institutions, or distinctive patient populations28,29. These domain shifts challenge model performance but must be overcome in clinical translation. The second one is the incapability in handling missing data. The standard protocols for radiomics-related research have not yet achieved consensus, and features selected for modeling were considerably variate. Moreover, the complicated but highly variant procedures of image pre-processing are also common and significantly lead to low reproducibility of feature extraction. Whereas, the real clinical landscape presents a far more complicated scenario. Given the highly fixed nature of the model’s input, the absence of any single feature or modality, a situation highly likely to be encountered in clinical settings, will render the model unusable28. The third one is that no study has demonstrated how can the model facilitate clinical decision-making yet. Although there is a consensus that predicting Ki-67 is of great clinical significance, it’s still unclear for clinicians that if they can make therapeutic decision consistent with the latest guidelines when using AI model. Therefore, our results can be interpreted as important as it provided critical information from following distinctive aspects.
The first one is that the developed model could be seamlessly integrated into workstations that commonly used in the healthcare institutions, and supported an automated, accurate, and stable clinical management workflow. It should be noted that the advantage of our model in flexibly accommodating various data scenarios. The combination of radiomics with machine learning is the most common method used in previous studies. However, these handicraft features necessitate delineating tumor segmentation, which is subjective, time-consuming, low-reproducing, and laborious. In contrast, MRI and radiological characteristics are much more easily obtained in routine diagnostic workflow. Compared with previous studies, the inputs of our model were more flexible as they were fed into the network separately rather than simultaneously, and the dropout modules and the auxiliary output module were introduced to obtain the intermediate outputs. So, the clinicians were able to obtain the prediction with/without radiomics input, and it is rational to say that our model is more adaptable to the practical use of clinicians.
The second perspective is that it provides valuable insights into a novel method of handling multi-modal data to the field of CAD research. Low generalization is a major concern of previous studies. For instance, a deep learning model might thrillingly achieve an AUC of 0.966 in internal test, while its performance would notably deteriorate to an AUC of 0.591 in external test27. Empirically, multi-modal models are anticipated to address this question owing to potential informative redundancy between modalities. Previous studies also indicated the superiority of clinical-radiomic models over single radiomic models10,25. However, directly setting multi-modal information as input is not mathematically rational, as it cannot capture complex relationships between the different modalities. Besides, it may technically lead to challenge in handling high-dimensional feature spaces which results in computational inefficiency, increased training time, and overfitting30. Therefore, we hypothesized that the multi-modal representation learning with cross-attention was a viable solution. More specifically, the design of our model was to use the radiological characteristics to guide the attention of deep learning block to the related regions of the MRI, and the deep learning block could consequently extract more enriched and informative representations of the input data. So, the predictions of the model were made by integrating information from multiple modalities into a cohesive representation. The results suggested the improvement of multi-modal learning in superior performance comparing to the existing methods, and indicated a new method to use multi-modal inputs to enhance both reproducibility and performance of deep learning model.
Thirdly, this research also provided concrete evidence showing the potential of AI model in aiding clinical decision-making. Regarding the prediction of Ki-67 expression in meningiomas, one of the most important clinical relevance is its application in predicting tumor growth of asymptomatic meningiomas. As reported by previous studies, the therapeutic decision of asymptomatic tumor was commonly determined by the location, as it was rather difficult for neurosurgeons to identify the easily growing one15. Comparatively, the results of our model suggested to be thrilling with AUC of 0.756 and 0.727 in prediction, as well as significant statistical difference in PFS between the stratified groups. Also, within these results, two key points warrant attention: 1) Follow-up outcomes indicated that the majority of patients’ tumors did not exhibit significant volume growth (three years: 77.5%; five years: 66.2%); 2) Similar to previous studies, although the model maintained rather balanced performance metrics during training, specificity was notably higher than sensitivity in the external test. These findings demonstrated that when using the model to make therapeutic decision for asymptomatic meningiomas, the decisions might lean towards conservatism as it encouraged clinicians to opt for follow-up observation rather than active intervention. To be more specific, for patients predicted to be at high risk, intervention should be performed actively as the high specificity; whereas, for patients predicted to be at low risk, regular follow-up should still be maintained to monitor tumor progression as the rather high false negative rate. It seems clinically rational as the therapeutical decisions based on this model aligns with the diagnostic and treatment recommendations outlined in the latest guidelines2,5. Whereas, further research is still warranted to explore methods for enhancing the model’s sensitivity in identifying such patients.
There is another contradicted point should be also worthy noted. As evidenced by the results, including radiomics in network training does indeed improve model performance, but during the inference phase, it negatively impacts the model’s decision-making. The most common reason for this is that this modality provides information redundancy and stabilizes the model optimization. But due to the redundancy or even noise it carries, removing it in inference may lead the model to focus more on other more valuable modalities, thereby improving performance. These findings suggested that the radiomics features might be treated as noise rather than useful features in this model, and it acted more like an implicit regularization effect rather than useful inputs. This could be explained from following perspectives: firstly, the CNN module is equally adept at extracting local features (such as texture and shape), partially replacing the informative role of radiomics; secondly, as the tabular data, radiomics are known for its high dimensionality, heterogeneity, and correlative relationships, but the neural networks often struggle to handle them effectively31. Therefore, considering both the model’s performance and clinical requirements, we concluded that output 1 was the most suitable one.
Our study possesses several limitations. Firstly, as retrospective research, the selection bias was inherent and unavoidable. Secondly, limited by the image quality of the dataset, we only included T1C images of MP-RAGE in this modeling. Future studies should investigate the method of integrating multi-sequences and advanced MR imaging to enhance the model’s performance. Thirdly, there is still deficiency from the perspective of clinical validation. Studies have shown that Ki-67 also holds significance in predicting patient prognosis (such as tumor recurrence after surgical resection and/or gamma knife treatment). Therefore, more research is required to investigate how the model can facilitate therapeutic decision-making for surgically resected meningioma patients in the future. Fourth, while Ki-67 is a valuable biomarker, WHO grade is also a well-established, clinically actionable parameter that significantly predicts recurrence and outcome. We do not include this parameter in our model because there have already been numerous studies that had successfully predicted WHO grades with promising results. Moreover, Transformer-based models are typically applied to large language datasets. Although our training set contains 578 patients, high-grade meningiomas account for only 25.3%, leading to significant imbalanced data distribution. Therefore, the comparison between two biomarkers in terms of prognostic value is still absent, and future study is still required to investigate which one is better for meningioma patients.
Methods
Study overview
Figure 3 illustrates the overview of the current research. Briefly, this study consisted of two steps: first, we collected surgically resected meningioma patients from two institutions to develop the multi-modal representation learning model, and evaluated its performance in both internal set and external set; then, to further demonstrate that how exactly our model could be used to aid in the clinical work, we collected a follow-up asymptomatic small meningioma from the third institution, and investigated that if the stratification by the developed model could show significant difference in patient survival.
The flowchart of model training, assessment, and clinical validation.
Input definition
Patients who underwent surgical resection for meningioma were recruited from West China Hospital Dataset A) and the Third People’s Hospital of Mianyang (Dataset B) between January 2017 and December 2023. The inclusion criteria were set as follows: (1) histopathological diagnosis of meningioma according to the 2021 WHO classification; (2) immunohistochemical record of Ki-67 index; (3) standardized MR scans within 2 weeks before surgery; (4) age ≥ 18 years old. The exclusion criteria included: (1) history of biopsy or radiotherapy; (2) recurrent meningiomas; (3) severe motion artifacts; (4) multiple meningiomas; (5) history of other intracranial diseases, such as subarachnoid hemorrhage. Based on above criteria, a total of 723 cases were identified from Center A, and 285 cases were identified from Center B, respectively. Ki-67 index was assessed by immunohistochemistry using an avidin-biotin-peroxidase complex method by using Aperio IHC image analysis software, and patients were stratified by a cutoff of 5%, a threshold deemed clinically relevant and used in most of previous studies26,32,33. The patient recruitment workflow of this research is shown in Fig. 4.
Patient recruitment workflow of this research.
Four MR sequences were acquired and exported from Picture Archive and Communication System (PACS), including T1-weighted imaging (T1WI), T2-weighted imaging (T2WI), fluid attenuated inversion recovery (Flair) and contrast-enhanced T1-weighted imaging (T1C). The scanning protocols of each dataset are provided in the Supplementary Note 1, which demonstrates that the MRIs were acquired from scanners of different vendors along with a wide variety of acquisition settings.
There is the noticeable difference in slice thickness between T1WI, T2WI, and FLAIR (5 mm) and T1C (1 mm), which means that the information they contain is quite limited. Therefore, quantitative image analysis is not suitable for these sequences and only T1C was set as input. Image preprocessing for each of the patients involved voxel resampling to 1 × 1 × 1 mm3, and N4 bias field correction for intensity nonuniformity adjustment. The intensity values for all image volumes were scaled in the range [0 255].
The evaluation of radiological characteristics was performed by two senior neuro-radiologists from two institutions (with more than 15 years’ experience in image reading) on pre-treatment MRI. The results were also independently reviewed by a neuro-radiologist with 20 years of experience from another Hospital. It should be noted that some radiological characteristics, like signal intensity and tumor size, can be quantified as radiomics features. Therefore, we focused on four “yes or no” characteristics and set them as inputs, including peritumoral edema (presenting as Edema Index, EI), intra-tumor necrosis, cerebral spinal fluid (CSF) cleft surrounding tumor, and capsular enhancement (Supplementary Fig. 1).
As the same reason described in the 2.2.1, T1WI, T2WI, and FLAIR images were excluded in quantitative radiomics extraction. Here, tumor segmentation was performed on T1C images using an automatic deep learning approach that the authors developed previously34. Afterwards, two senior neuro-radiologists with 15 and 20 years of experience in image reading, blinded to clinicopathological information, manually adjusted the preliminary segmentation using 3D Slicer software, and any discrepancies were resolved through consensus. Finally, a total number of 1218 radiomics features were retrieved from tumor segmentation by using PyRadiomics and standardized by mean removal and unit variance scaling (Supplementary Table 1 and 2).
Deep learning modeling
The predictive model was developed using the Python programming language (PyTorch 1.3.1) on a workstation equipped with four NVIDIA RTX3090 data center accelerators (RAM = 24GB). The cascaded multi-modal Transformer architecture consisted of three blocks sequentially integrating features from deep learning features of 3D-MRI, radiological characteristics, and radiomics. This architecture aimed to refine class prediction by integrating and interacting representations of individual modalities20. The class prediction process initiated with a learned Query vector Qi, progressing through successive blocks while assimilating multi-modal information to generate the final output. Within each block, modalities were transformed into vector embeddings represented as Key vectors Ke and Value vectors Ve, followed by a cross-attention module to calculate relationships between the modality and previous input sequences. The Dropout module probabilistically determined whether the backbone could receive information in each training iteration, indicating “not available.” Auxiliary output module was used to obtain intermediate output. More details, including input embedding, cascaded cross-attention calculation, dropout module, auxiliary output module, and hyperparameter settings, are provided in Supplementary Note 2.
Dataset A was randomly divided into a training group (n = 578) and an internal test group (n = 145) at an 8:2 ratio. Dataset B (n = 285) was used as the external test group to assess model generalizability. Here we conducted a series of ablation experiments were conducted to determine the best input of each block. Moreover, experiment of setting different combinations of modalities as input was also performed as it could provide valuable insights into which modality was necessary for prediction. Only the model with the best performance was set for subsequent analysis.
Model performance was evaluated using metrics including area under the curve (AUC), accuracy, sensitivity, specificity, F1 score, and Matthews correlation coefficient (MCC). The mean and 95CI (95% confidence interval) of each metrics were calculated based on 50-fold cross-validation. Gradient-weighted class activation mapping (Grad-CAM) was employed to generate heatmaps highlighting important regions based on target class score gradients, along with cross-attention maps to visualized and analyzed with the contributions of each modality to the class prediction.
Clinical validation
Follow-up asymptomatic small meningioma patients (Dataset C) were identified from the Shangjin Hospital between January, 2010 and December, 2017. The inclusion criteria were as follows: (1) radiographic diagnosis of meningioma by brain MRI5; (2) asymptomatic lesion; (3) maximal diameter less than 3 cm; (4) without venous sinus involvement; (5) age ≥ 18 years old; (6) with radiological follow-up for at least 5 years. The exclusion criteria were as follows: (1) occurrence of an intervention unrelated to tumor growth (including symptom development, unrelated meningioma-specific mortality, patient request); (2) MR scans with severe motion artifacts; (3) multiple meningiomas; (4) occurrence of irrelevant intracranial diseases. Based on above criteria, a total of 231 cases were accordingly collected.
Survival time was calculated from the date of MR scan to the date of the endpoint, which was defined as fifteen percentage increase in tumor volume in concordance with prior studies35,36,37. Then, tumor growth happened in both 3 years and 5 years were set as ground truth, and predictive metrics of the developed model selected from 2.3.2 were calculated accordingly. There was no other fine tuning or new modeling was performed.
The 231 cases were directly stratified into low-group and high-group based on the prediction of well-trained model. Kaplan–Meier (KM) analysis and log-rank test were used to examine if there was statistical difference between two groups. Moreover, multivariate Cox regression was conducted to investigate if the model prediction was independently associated with patient survival in 5 years. Sub-group analysis was performed to evaluate if the model could achieve similar accuracy in skull-base versus non-skull-base meningiomas.
Statistical analysis
Categorical variables were described using frequencies and percentages, while continuous variables were summarized using means and standard deviations/95CI. The chi-square test was employed for univariate analysis, and Logistic regression was used for multivariate analysis. P-values < 0.05 considered statistically significant.
Ethics approval and consent to participate
The study was conducted in accordance with the Declaration of Helsinki, and approved by the institutional review board of West China Hospital, Sichuan University (protocol code 2021-S-851). The informed consent was waived by the institutional review board.
Data availability
Data is provided within the manuscript or supplementary information files. Due to the privacy of patients, the data related to patients cannot be available for public access but can be obtained from the corresponding author on reasonable request approved by the institutional review board of all enrolled centers. Whereas, we’d like to share the deep learning model publicly once the paper is accepted.
References
Ostrom, Q. T. et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2016-2020. Neuro Oncol. 25), iv1–iv99 (2023).
Louis, D. N. et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 23, 1231–1251 (2021).
Huang, R. Y. et al. Imaging and diagnostic advances for intracranial meningiomas. Neuro Oncol. 21, i44–i61 (2019).
Goldbrunner, R. et al. EANO guideline on the diagnosis and management of meningiomas. Neuro Oncol. 23, 1821–1834 (2021).
Louis Burt Nabors, J.P., Joachim Baehring, Ankush Bhatia, Orin Bloch. NCCN Clinical Practice Guidelines in Oncology- Central Nervous System Cancers (Version 1.2023). 2023; https://www.nccn.org/.
Wang, J. Z. et al. Meningioma: International Consortium on Meningiomas (ICOM) consensus review on scientific advances & treatment paradigms for clinicians, researchers, and patients. Neuro Oncol. 2024.
Oya, S., Kawai, K., Nakatomi, H. & Saito, N. Significance of Simpson grading system in modern meningioma surgery: integration of the grade with MIB-1 labeling index as a key to predict the recurrence of WHO Grade I meningiomas. J. Neurosurg. 117, 121–128 (2012).
Shinya, Y. et al. Long-Term Outcomes of Stereotactic Radiosurgery for Postoperative World Health Organization Grade I Skull Base Meningioma: Utility of Ki-67 Labeling Index as a Prognostic Indicator. Neurosurgery. 2023.
Wen, P. Y. & Packer, R. J. The 2021 WHO classification of tumors of the central nervous system: clinical implications. Neuro Oncol. 23, 1215–1217 (2021).
Zhao, Y., Xu, J., Chen, B., Cao, L. & Chen, C. Efficient Prediction of Ki-67 Proliferation Index in Meningiomas on MRI: From Traditional Radiological Findings to a Machine Learning Approach. Cancers (Basel). 14 (2022).
Zhang, R. et al. The value of diffusion-weighted imaging in the natural history of meningiomas: a predictor of tumor growth. J. Neurosurg. 140, 377–385 (2024).
Li, Z. et al. Preoperative Subtyping of WHO Grade 1 Meningiomas Using a Single-Shot Ultrafast MR T2 Mapping. J Magn Reson Imaging. 2023.
Park, Y. W. et al. Magnetic resonance imaging-based 3-dimensional fractal dimension and lacunarity analyses may predict the meningioma grade. Eur. Radio. 30, 4615–4622 (2020).
Yu, H. et al. Can amide proton transfer-weighted imaging differentiate tumor grade and predict Ki-67 proliferation status of meningioma? Eur. Radio. 29, 5298–5306 (2019).
Lee, E. J., Park, J. H., Park, E. S. & Kim, J. H. Wait-and-See” strategies for newly diagnosed intracranial meningiomas based on the risk of future observation failure. World Neurosurg. 107, 604–611 (2017).
Dosovitskiy, A. et al. An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. 2020; abs/2010.11929.
Vaswani, A. et al. Attention is all you need. 2017; 30.
Zhou, H.Y. et al. A transformer-based representation-learning model with unified processing of multimodal input for clinical diagnostics. Nat. Biomed. Eng. 7, 743–755 (2023).
Han, Y. et al. Radiomics-guided global-local transformer for weakly supervised pathology localization in chest X-rays. IEEE Trans. Med. Imaging 42, 750–761 (2023).
Liu, L. et al. Cascaded multi-modal mixing transformers for Alzheimer’s disease classification with incomplete data. Neuroimage 277, 120267 (2023).
Xu, P., Zhu, X. & Clifton, D. A. Multimodal learning with transformers: a survey. IEEE Trans. Pattern Anal. Mach. Intell. 45, 12113–12132 (2023).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: images are more than pictures, they are data. Radiology 278, 563–577 (2016).
Bera, K., Braman, N., Gupta, A., Velcheti, V. & Madabhushi, A. Predicting cancer outcomes with radiomics and artificial intelligence in radiology. Nat. Rev. Clin. Oncol. 19, 132–146 (2022).
Roisman, L. C. et al. Radiological artificial intelligence - predicting personalized immunotherapy outcomes in lung cancer. NPJ Precis Oncol. 7, 125 (2023).
Moon, C. M. et al. Preoperative prediction of Ki-67 and p53 status in meningioma using a multiparametric MRI-based clinical-radiomic model. Front Oncol. 13, 1138069 (2023).
Khanna, O. et al. Machine learning using multiparametric magnetic resonance imaging radiomic feature analysis to predict Ki-67 in World Health Organization grade I meningiomas. Neurosurgery 89, 928–936 (2021).
Chen, J., et al. Predicting meningioma grades and pathologic marker expression via deep learning. Eur Radiol. 2023.
Huang, E. P. et al. Criteria for the translation of radiomics into clinically useful tests. Nat. Rev. Clin. Oncol. 20, 69–82 (2023).
Bloem, J. L., Reijnierse, M., Huizinga, T. W. J. & van der Helm-van Mil, A. H. M. MR signal intensity: staying on the bright side in MR image interpretation. RMD Open 4, e000728 (2018).
Ma, M., Ren, J., Zhao, L., Testuggine, D., Peng, X. Are multimodal transformers robust to missing modality? Paper presented at: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition 2022.
Du, L., et al. TabularNet: A neural network architecture for understanding semantic structures of tabular data. Paper presented at: Proceedings of the 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining 2021.
Nowak-Choi, K. et al. Resected WHO grade I meningioma and predictors of local control. J. Neurooncol 152, 145–151 (2021).
Haddad, A. F. et al. WHO grade I meningioma recurrence: identifying high risk patients using histopathological features and the MIB-1 index. Front Oncol. 10, 1522 (2020).
Chen, C. et al. Performance test of a well-trained model for meningioma segmentation in health care centers: secondary analysis based on four retrospective multicenter data sets. J. Med Internet Res 25, e44119 (2023).
Hashimoto, N. et al. Slower growth of skull base meningiomas compared with non-skull base meningiomas based on volumetric and biological studies. J. Neurosurg. 116, 574–580 (2012).
Hashiba, T. et al. Serial volumetric assessment of the natural history and growth pattern of incidentally discovered meningiomas. J. Neurosurg. 110, 675–684 (2009).
Behbahani, M. et al. A prospective study of the natural history of incidental meningioma-Hold your horses! Neurooncol Pr. 6, 438–450 (2019).
Acknowledgements
This work was funded by Postdoctoral Research Foundation of China (ID: 2022M712245, Chaoyue Chen), Science and Technology Department of Sichuan Province (ID: 2024NSFSC1886, Chaoyue Chen), and West China Hospital of Sichuan University (ZYAI24018, Jianguo Xu).
Author information
Authors and Affiliations
Contributions
Conceptualization: Jianfeng Xu, Xin Zan, Lei Zhang and Jianguo Xu; methodology: Chaoyue Chen, Haoze Jiang, Fumin Zhao, Yang Zhang, and Yanjie Zhao; investigation, Chaoyue Chen, Fumin Zhao, Linrui Cai, and Xiaolong Xu; data curation: Chaoyue Chen, Linrui Cai, Shuangyi Zhang, Haoze Jiang, Fumin Zhao, Yuen Teng, Junkai Zheng, Junkai Zheng, and Yanjie Zhao; validation: Linrui Cai, Yang Zhang, Zhouyang Huang and Fumin Zhao; funding acquisition: Chaoyue Chen; writing—original draft preparation: Chaoyue Chen; writing—review and editing: Xin Zan, Lei Zhang and Jianguo Xu. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, C., Zhao, Y., Cai, L. et al. A multi-modal deep learning model for prediction of Ki-67 for meningiomas using pretreatment MR images. npj Precis. Onc. 9, 21 (2025). https://doi.org/10.1038/s41698-025-00811-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-00811-1
This article is cited by
-
Research progress of deep learning based on magnetic resonance imaging in meningioma
Magnetic Resonance Materials in Physics, Biology and Medicine (2025)