Abstract
Head and neck squamous cell carcinoma (HNSCC) is the sixth most common cancer worldwide, significantly impacting patient survival and quality of life. The recent emergence of immunotherapy has provided new hope for HNSCC patients, improving survival rates; however, only 15%–20% of patients benefit, and side effects are inevitable. With advancements in omics technologies and the growing prevalence of bioinformatics research, the immune microenvironment of HNSCC has become increasingly well understood, and the molecular mechanisms underlying immunotherapy responses continue to be elucidated. In this review, we summarize commonly used omics techniques and their applications in the research of HNSCC immunotherapy, including predicting and enhancing efficacy, formulating personalized treatment plans, establishing robust preclinical research models, and identifying new immunotherapy targets. Finally, we explore future perspective in terms of sequencing samples, data integration analysis, emerging technologies, clinicopathological features, and interdisciplinary approaches.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC) is the sixth most common cancer worldwide, with 931,000 new cases and 467,000 deaths in 20201. In addition to early oral cancer (treated only by surgery) or laryngeal cancer (treated only by surgery or radiotherapy), the treatment of most HNSCC cases requires a multimodal approach and therefore requires multidisciplinary care2. Since 2016, anti-programmed death 1 (PD-1) inhibitors, such as nivolumab and pembrolizumab, have been approved for patients with recurrent or metastatic HNSCC who experience disease progression during or after platinum-based chemotherapy3,4. Despite these advances, patient prognosis remains poor, with no significant improvement in survival rates, and only a subset of patients responding to treatment. Therefore, there is a critical need to explore more effective and targeted immunotherapy strategies to enhance clinical outcomes for patients with HNSCC5. For example, the emerging neoadjuvant immunotherapy has shown significant promise, particularly when combined with chemotherapy, demonstrating a 91.2% event-free survival rate6.
Immunotherapy under the guidance of traditional pathological classification faces the challenge of response heterogeneity, which is mainly due to the high complexity of the tumor ecosystem. Omics technologies, including genomics, epigenomics, transcriptomics, and single-cell omics, have been widely applied in the field of immunotherapy for HNSCC7,8,9. These technologies help dissect the complex tumor microenvironment, explore mechanisms of immunotherapy, and identify new targets. However, single-omics data are typically analyzed at a singular level, which may overlook information from other dimensions and fail to reveal biological functions holistically. For instance, genomic analysis may fail to capture epigenetic regulation-induced PD-L1 expression heterogeneity (e.g., DNA methyltransferases inhibitor effects).
The interplay and complementarity of multi-omics data facilitate a comprehensive analysis of tumors, heralding a new era of precise, efficient, and personalized immunotherapy. For example, single-cell transcriptome can evaluate the functional status of different T cell subsets based on gene set enrichment scores. Proteomics makes up for the low sensitivity of the transcriptome to the identification of T cell expression markers, and has a higher degree of recognition for T cells with different functional states. Combined with spatial omics analysis, the potential cell interactions of T cells in specific functional states can be explored. By combining single-cell proteomics, single-cell transcriptomics, T cell receptors (TCR) sequencing and spatial omics techniques, Rahim et al. intuitively demonstrated the characteristics of T cell responses in uninvolved lymph nodes and emphasized the key role of lymph nodes in maintaining the efficacy of immunotherapy10,11. This integrated application of multi-omics not only enhances the depth and breadth of cancer research but also provides valuable data support and innovative insights for the development of new therapeutic methods and diagnostic tools.
This article reviews the application of omics technology in immunotherapy research for HNSCC, highlighting its critical role in various aspects of immunotherapy. Finally, we discuss the future perspectives for the application of omics technology in HNSCC immunotherapy research.
The study of HNSCC before the era of multi-omics
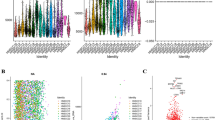
Before the advent of the era of multi-omics, researchers mainly relied on flow cytometry, immunohistochemistry and immunofluorescence techniques to explore the immune microenvironment in HNSCC. These traditional methods can qualitatively and quantitatively analyze the immune cell populations in tumors by detecting pre-selected immune markers. For example, flow cytometry uses its advantages of high-throughput and multi-parameter detection to finely distinguish different subsets of CD8+ T cells, regulatory T cells (Tregs), and macrophages. Study has shown that in HNSCC, tumor-infiltrating CD8+ T cells are closely related to the better prognosis of patients, mainly due to their key role in directly killing tumor cells and activating other anti-tumor effector cells12. On the contrary, Tregs and some phenotypes of tumor-associated macrophages often mediate immunosuppression and promote tumor progression, which is associated with poor prognosis13,14.
Although flow cytometry and immunohistochemistry/immunofluorescence provide us with valuable preliminary information, these techniques are usually limited to the detection of a single or a few indicators, and it is difficult to fully reflect the complexity of the tumor microenvironment and the dynamic interaction between cells. For example, due to sensitivity limitations, flow cytometry can only detect a small abundance of cell subsets. Moreover, flow cytometry provides only a single time point snapshot and cannot track the dynamic evolution of T cell clones.
The emergence of high-throughput technology has greatly expanded the research horizon in this field. For example, the typical immunohistochemical panel can only detect 3–5 markers at the same time, but the mass spectrometry flow technology can detect more than 50 parameters, which greatly improves the data throughput and resolution. Furthermore, by integrating genome, transcriptome, proteome and metabolome data, multi-omics methods can analyze the phenotype, functional status and molecular regulatory network of immune cells in a higher dimension, thus revealing the tumor heterogeneity and immune regulation mechanism that are difficult to capture by traditional methods15. This global data integration not only helps to identify new immune cell subsets, but also reveals their interactions and potential regulatory pathways in the tumor microenvironment, providing a theoretical basis for the development of accurate immunotherapy strategies16.
Omics approaches: a brief overview
Since American geneticist Thomas H. Roderick first introduced the concept of genomics in 1986, omics technologies have increasingly been applied in medical research. The advent of next-generation sequencing and single-cell sequencing technologies has significantly accelerated this progress, providing new scientific insights and research directions for tumor immunotherapy.
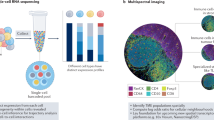
In general, multi-omics technologies are capable of extracting valuable biological information from saliva, blood, tumor tissue, and lymph node samples collected from HNSCC patients. Multi-omics joint analysis enables the construction of gene regulatory networks across multiple dimensions, such as DNA, RNA, and protein, allowing for a comprehensive analysis of tumorigenesis and development over time and spatial dimensions (Fig. 1).
BCR B cell receptors, FFPE formalin fixed paraffin embedded, HNSCC head and neck squamous cell carcinoma, TCR T cell receptors. Created with BioRender.com.
Genomics
High-throughput sequencing-based genomic methods can reveal individual tumor mutation burden (TMB), complex mutation characteristics, and tumor-specific antigens, providing valuable information for targeted therapies, immune checkpoint inhibitors (ICI), and personalized anti-cancer vaccines17. Notably, patients with higher TMB have an increased number of neoantigens produced by tumor cells, enhancing antigen presentation by immune cells and facilitating the recognition and elimination of these tumor cells. A meta-analysis indicated that ICI therapy yields a more pronounced response and clinical benefit in HNSCC patients with high TMB18.
In addition to TMB, in the context of high-throughput genome sequencing, more tumor neoantigens based on other source, such as exon retention events, frame shift mutations, and abnormal expression of human endogenous retroviruses have been continuously excavated19,20,21. The diversity and quality of these neoantigens are also key to the success of tumor immunotherapy. Additionally, mutations induced by carcinogens promote the enrichment of immunosuppressive M2 macrophages in the TIME, leading to resistance to ICI treatment22.
Epigenomics
Epigenomics involves the study of all chemical modifications outside genomic DNA in cells, such as DNA methylation, histone modification, non-coding RNA, and chromatin remodeling. These modifications can regulate gene expression to affect cell function, thereby affecting individual development and even causing diseases, such as malignant tumor23.
In recent years, the role of epigenetics in tumor immunotherapy has been continuously emphasized. Through epigenetic regulation, the normal silent genomic non-coding regions, such as long terminal repeats, long/short scattered nuclear elements, are restored to express, thereby enhancing the immunogenicity of cancer cells. Additionally, epigenetic characteristics can influence T cell exhaustion through multiple mechanisms, potentially diminishing the effectiveness of ICI treatment24. In studies of HNSCC immunotherapy, significant differences in methylation patterns have been observed between patients who respond to ICI treatment and those who do not, highlighting the potential of the HNSCC methylation spectrum in predicting treatment response25. A comprehensive understanding of epigenetic characteristics will aid in identifying patient responsiveness and drug resistance to immunotherapy.
Transcriptomics
Transcriptome sequencing enables a detailed analysis of target gene expression levels, the identification of new transcripts, single nucleotide polymorphisms, and splicing variants, as well as the provision of allele-specific gene expression. Transcriptomic data can elucidate the signaling pathways and key transcription factors involved in tumorigenesis and development26. Furthermore, various analytical methods can derive distinct biological insights from the original data. For instance, deconvolution methods can reveal the composition of TIME, differential analysis can identify gene signatures associated with different tumor phenotypes, and prognostic analysis can predict patient survival. It is worth noting that transcriptome analysis plays a key role in predicting the response of HNSCC patients to immunotherapy and screening immune checkpoint inhibitor adjuvant drugs. In particular, the integration of immune-related gene expression characteristics, such as those reflecting T cell infiltration and interferon-γ (IFN-γ) signaling pathways, can more accurately predict the patient’s response to ICI therapy27,28.
Single-cell RNA sequencing (scRNA-seq) technology has advanced TIME research to unprecedented levels of detail, significantly contributing to our understanding of TIME. ScRNA-seq enables non-targeted quantification of transcripts at the single-cell level, facilitating the identification of new immune cell subtypes, rare immune cell populations, and the mapping of immune cell status and development29. Additionally, scRNA-seq provides insights into cell interactions within TIME, allowing researchers to accurately identify ligand-receptor pairs involved in cell-cell interactions, thereby offering a more comprehensive view of immune cell functionality. In the context of immunotherapy, scRNA-seq applied to immune and stromal cells in TIME elucidates transcriptional states associated with therapeutic response and drug resistance. For instance, intratumoral CD103+ CD8+ T cells have been identified as predictors of response in patients with advanced HNSCC undergoing neoadjuvant chemotherapy immunotherapy30.
Immune repertoire profiling
Immunoreceptor repertoire sequencing provides new insights into the dynamics of tumor-immune interactions. By capturing the complete diversity of TCRs and B cell receptors (BCRs) within the TIME at single-cell resolution, this approach not only identifies the clonal composition of T/B cells but also quantifies clonal expansion events driven by tumor-specific antigens31,32.
Critically, the integration of scRNA-seq with paired single-cell TCR/BCR sequencing enables two key advances: precise tracking of clonally expanded T/B cell populations (e.g., tumor-reactive CD8+ T cell clones with high TCR clonality) and their functional phenotypes (e.g., activation, memory, exhaustion), and reconstruction of clonal lineage relationships to infer antigen-driven selection pressures during immunotherapy33. Zhou et al. demonstrated that clonal expansion of CD8+ T cells with cytotoxic phenotypes correlates with durable response to ICIs in HNSCC34. Moreover, study highlights that dominant TCR clones with high clonal frequency in pretreatment biopsies may serve as predictive biomarkers for ICI efficacy35.
Proteomics
Genome and transcriptome analyses offer insights into the characteristics and potential effects of genomic changes, while proteomics provides direct information on protein regulation and signal transduction in response to these changes. At present, mass spectrometry has become one of the most widely used techniques in high-throughput proteomics, which quantifies post-translational modifications through direct fragments or specific protein decomposition activities responsible for their formation. Mass spectrometry can be combined with a variety of separation and pre-fractionation techniques to identify target proteins/peptides and improve the accuracy and yield of recognition. In addition, commonly used high-throughput proteomics techniques include protein pathway array, next generation tissue microarrays, multiplex bead-or aptamer-based assays, proximity extension assay, and nanopore based single-molecule proteomics36.
Mass spectrometry-based proteomics can accurately reflect the functional status of tumors, distinguish between different immune subtypes, and inform the development of personalized immunotherapy strategies37. It is worth noting that proteogenomic has gradually demonstrated its advantages in the field of immunotherapy research. On the one hand, by combining mass spectrometry with whole exome sequencing, it can identify and verify neoantigens at the protein level, providing a new potential strategy for immunotherapy38. On the other hand, based on the results of proteomics analysis, the analysis of immune infiltration in different cohorts, combined with the results of whole genome sequencing, can help to determine the internal driving factors of low immune infiltration, so as to formulate personalized precise immunotherapy strategies for this population in the future39.
Single-cell proteomics provides distinct advantages in cell population annotation by enabling precise identification of cell surface receptor proteins, thereby facilitating intuitive visualization of cellular subgroup proportions. Current single-cell proteomics techniques are generally classified into two primary categories. The first involves targeted protein analysis, which employs antibody-based detection methods such as spectral flow cytometry (e.g., CyTOF) and antibody-coupled oligonucleotide technologies. The second category encompasses global proteome analysis, exemplified by methodologies like SCoPE-MS and nanoPOTS. These technologies enable in-depth phenotypic characterization of immune cells, facilitate dynamic modeling of signaling pathways, and reveal protein expression signatures associated with immunotherapy responsiveness. Notably, single-cell proteomics has been applied to classify circulating tumor cells (CTCs), enabling patient stratification based on CTC distribution patterns and immune checkpoint expression profiles. Such analyses show significant potential for guiding immunotherapeutic strategies40.
Metabolomics
Tumor-infiltrating immune cells often experience metabolic stress due to the metabolic dysregulation of tumor cells, leading to an impaired anti-tumor immune response. The reuse of anti-cancer drugs that target metabolism may synergistically enhance immunotherapy by reprogramming the tumor microenvironment (TME)41.
Metabolomics can analyze the metabolic landscape at various stages of tumor development through high-throughput methods, allowing for the identification of metabolites or metabolic pathways associated with immunotherapy resistance. For example, metabolomic analysis has shown that an increased ratio of kynurenine to tryptophan correlates with heightened resistance to ICI treatment in patients, indicating that therapeutic targets against kynurenine production are critical for improving immunotherapy efficacy42.
Microbiomics
Microbiota may play a crucial role in the onset and progression of HNSCC, treatment-related toxicity, disease recurrence, and the efficacy of immunotherapy. For instance, a correlation has been observed between the presence of bifidobacteria in melanoma lesions and the response to ICIs43. Recent advancements in tools such as 16S rDNA/RNA sequencing and metagenomic shotgun sequencing have facilitated the analysis of microbial composition in HNSCC and its impact on remodeling the TIME. Notably, 16S rDNA sequencing identified an enrichment of Peptostreptococcus in oral squamous cell carcinoma, with its upregulation enhancing the efficacy of ICI treatment44.
Spatial omics
Single-cell omics plays a crucial role in decoding TIME and immune cell interaction information. However, different subtypes and development stages of HNSCC exhibit distinct tissue architectures and hierarchical structures. Due to the lack of spatial information in single-cell omics, cell interaction analysis only relies on the expression of related genes, and lacks the verification of spatial dimension. Furthermore, external stimuli, such as immunotherapy, can trigger spatial reprogramming within tumors, leading to anti-tumor immune regeneration and stromal cell repositioning. Thus, analyzing the spatial structure of TIME is essential for developing new immunotherapy strategies.
A study based on spatial transcriptomics technology pointed out that there is a regulatory axis of epithelial cells-inflammatory cancer-associated fibroblasts (CAFs)-regulatory T cells (Treg) in the high metabolic region of oral squamous cell carcinoma, thus shaping the immunosuppressive microenvironment45. The application of spatial transcriptomics offers potential therapeutic targets for optimizing HNSCC immunotherapy. Additionally, spatial proteomics has been employed to evaluate the impact of tumor immune architecture in patients both at baseline and post-immunotherapy46. Notably, Chen et al. utilized spatially resolved metabolomics in combination with tumor-immune cell co-cultured spheroids to visualize the metabolic interactions between tumor and immune cells, developing a novel platform for screening and imaging metabolites that change during T cell anti-tumor responses. This approach provides new insights into the metabolic alterations associated with HNSCC immunotherapy47.
Multi-omics data integration strategies
Contemporary cancer research employs multi-omics integration strategies across three levels: early-stage data-level integration (e.g., principal component analysis, joint matrix factorization), mid-stage feature-level integration (e.g., multi-omics network analysis, Bayesian causal inference), and late-stage decision-level integration (e.g., hierarchical clustering combined with machine learning models). Researchers commonly use cross-omics pathway enrichment, multidimensional molecular subtyping, and machine learning frameworks (e.g., random forest, deep learning) to harmonize genomic, transcriptomic, epigenomic, proteomic, and metabolomic data. Such integration enables the identification of tumor driver genes, molecular interaction networks, and drug resistance mechanisms48,49.
Emerging spatial omics and single-cell omics approaches further resolve tumor microenvironment heterogeneity at subcellular resolution. Concurrently, systems biology approaches—including protein-protein interaction networks and regulatory circuit modeling—decode functional modules across omics layers. Together, these strategies systematically link molecular signatures to clinical phenotypes, identifying actionable therapeutic targets and biomarkers to advance precision oncology50.
Application of multi-omics in immunotherapy of head and neck squamous cell carcinoma
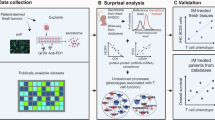
The rapid advancement of omics technology has generated new ideas and insights for both preclinical and clinical research in HNSCC immunotherapy, significantly contributing to the prediction of efficacy and the formulation of personalized treatment strategies (Fig. 2). Here, we summarize the biomarkers found in HNSCC immunotherapy studies by multi-omics techniques (Table 1).
First, multi-omics helps to understand and strengthen the role of immunotherapy, identify the role of tdLN and various immunosuppressive cells in HNSCC immunotherapy, and reveal the molecular mechanism of the good efficacy of multi-target immunotherapy. Secondly, multi-omics can predict the response of immunotherapy by revealing the immune microenvironment of patients and combining small molecules, specific cell subtypes, prediction models, and tertiary lymphoid structures. In addition, multi-omics can develop personalized treatment options by combining the patient’s gene mutation information, HPV infection, and anatomical sites, such as immunotherapy combined with chemotherapy and targeted therapy. At the same time, multi-omics helps to establish highly simulated preclinical research models, such as cell lines, organoids, and animal models. Finally, multi-omics can help to discover new therapeutic targets in different immunotherapy, including immune checkpoints, tumor vaccines, cell adoptive therapy, small molecule inhibitors, and therapeutic antibodies. CAFs cancer-associated fibroblasts, CAR-T chimeric antigen receptor T, CTLA-4 cytotoxic T-lymphocyte-associated protein 4, HNSCC head and neck squamous cell carcinoma, MDSCs myeloid-derived suppressor cells, 4-NQO 4-Nitroquinoline N-oxide, tdLN tumor-draining lymph node, PD-1 programmed death 1, PD-L1 programmed death ligand 1. Created with BioRender.com.
In recent years, the application of multi-omics research has increased significantly, with most studies combining single-cell resolution data with clinical cohort information from bulk analyses. These studies predominantly focus on predicting the efficacy of immunotherapy and identifying potential targets for enhancing immune responses. Notably, tertiary lymphoid structures and B cell characteristics have emerged as strong predictive biomarkers for immunotherapy outcomes. Additionally, there has been a growing number of studies utilizing multi-omics approaches for the subclassification of HNSCC, showcasing the advantages of integrating multiple layers of molecular data. This integrated approach is crucial for better guiding personalized immunotherapy strategies. Looking ahead, multi-omics holds immense potential in the field of HNSCC immunotherapy research.
Understand and enhance the efficacy of immunotherapy
PD-1 inhibitors (e.g., pembrolizumab) are approved as the first-line therapy for recurrent/metastatic HNSCC, either as monotherapy in PD-L1 positive tumors or combined with platinum-based chemotherapy. However, clinical trials demonstrate an overall response rate of only 15–20%51, highlighting the need to investigate additional cellular components within the TME that may influence ICI responsiveness, including potential contributions from specific cell subtypes or patient subgroups.
The response of CD8+ T cells is critical for anti-tumor immunity. Recent findings from single-cell transcriptome sequencing, TCR sequencing, and proteomics indicate that CD8+ T cells are activated in non-metastatic lymph nodes before migrating to the tumor, a process that appears to be disrupted in metastatic lymph nodes10. This underscores the significant role of tumor-draining lymph nodes (tdLNs) in HNSCC immunotherapy. Exploring targeted lymph node treatment strategies may present a promising avenue for the next generation of immunotherapies. Furthermore, the application of nanomedicine delivery systems holds great potential for lymph node targeting52. In a melanoma mouse model, a lymph node-targeting mRNA vaccine using lipid nanoparticles elicited a robust CD8+ T cell response while minimizing side effects, showcasing its therapeutic and protective efficacy53.
In addition, various immunosuppressive cell subtypes within tumors regulate the infiltration of CD8+ T cells, leading to immune dysfunction54. Recent studies utilizing single-cell and spatial transcriptomics have identified a subset of CAF that highly express MHC-I molecules and galectin-9, which are involved in restricting CD8+ T cell infiltration and promoting tumor growth55. Targeting CAF could yield synergistic effects when combined with anti-PD-1 therapy, thereby enhancing immunotherapy efficacy56.
To address the drug resistance associated with single-agent immunotherapy, clinical practice often employs combinations of different ICIs. A thorough investigation of the response mechanisms underlying combination therapy is essential for improving therapeutic outcomes. For instance, the immune response mechanism of anti-PD-L1 combined with cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) therapy has been analyzed using tumor biopsy samples from HNSCC patients pre- and post-immunotherapy. The activation of CD4+ T cells and the recruitment of tdLNs serve as markers of early response to HNSCC immunotherapy. Notably, the combination therapy enhances T cell activation along the CD4+ helper T cell trajectory compared to anti-PD-L1 alone. Furthermore, it appears that anti-CTLA-4, rather than anti-PD-L1, may have a direct impact on cells residing within the tdLNs. Following treatment with anti-PD-L1 and anti-CTLA-4, CD4+ T cells located in the tdLNs are subsequently transported to the tumor via the bloodstream57.
Predict the response to immunotherapy
Predicting the response to immunotherapy based on immune infiltration characteristics is a critical step in improving therapeutic outcomes. Tumors exhibiting an immune-inflamed phenotype are rich in immune cells, particularly T cells expressing CD4 and CD8, which are located in proximity to tumor cells. Clinical responses to anti-PD-L1/PD-1 treatment are most frequently observed in tumors with this inflammatory phenotype58. In contrast, tumors displaying an immune-excluded phenotype, where immune cells cluster at the tumor margin without infiltrating malignant cell nests, and those with an immune-desert phenotype, characterized by a lack of T cell infiltration, often exhibit reduced sensitivity to immunotherapy59.
The heterogeneous expression of immune checkpoint ligands in malignant cells correlates with distinct immune microenvironments. Utilizing single-cell transcriptome sequencing to assess the expression of immune checkpoint ligands has enabled the stratification of OSCC patients into three groups: “high immune checkpoint ligands/high IFN-γ,” “low immune checkpoint ligands/low IFN-γ,” and “low immune checkpoint ligands/low IFN-γ/high PD-L1.” Patients in the first group are predominantly associated with an immune-inflamed phenotype, while groups 2 and 3 are more commonly linked to an immune-desert phenotype60.
It is challenging to fully predict therapeutic outcomes based on a single biomarker. In some instances, multi-gene models may offer greater accuracy and predictive value than single-gene assessments, allowing for a more comprehensive evaluation of the complexities within the TME. One study identified eight genes closely linked to tumor progression and immune regulation by integrating single-cell and bulk RNA sequencing data, subsequently developing and validating an eight-gene risk model. This model revealed that low-risk groups exhibited higher infiltration rates of memory activated CD4+ T cells, CD8+ T cells, and plasma cells, as well as higher immune scores, suggesting they are more likely to benefit from immunotherapy compared to high-risk groups with greater infiltration of activated mast cells and M2 macrophages61.
Develop personalized strategies for immunotherapy
An in-depth study of tumor pathogenesis at the cellular and molecular levels allows omics technologies to better guide individualized treatments, improving clinical outcomes. The latest National Comprehensive Cancer Network guidelines for HNSCC recommend utilizing next-generation sequencing gene mapping to detect biomarkers that inform treatment choices (www.nccn.org/patients).
Human papillomavirus (HPV) is one of the important factors leading to head and neck cancer. HPV-positive patients usually show good immune cell infiltration and respond well to existing treatment options (including immune checkpoint inhibitors). Therefore, antiviral therapy or vaccination against HPV can be considered to improve clinical prognosis62. In contrast, although some HPV-negative patients can also achieve certain efficacy when receiving ICIs (such as anti-PD1), the overall response rate is significantly lower than that of HPV-positive patients, and it is difficult to develop individualized treatment strategies for HPV-negative patients. Multi-omics provides a direction for the exploration of treatment strategies for such patients. Based on the combined analysis of proteomics, genome and transcriptome in 108 HPV-negative HNSCC patients, the researchers integrated DNA, RNA, protein, and phosphopeptide data, divided the patients into three subtypes, and recommended corresponding precise treatment for each subtype, including cyclin-dependent kinases inhibitor, epidermal growth factor receptor (EGFR) antibody and ICI therapy. This study provides a new direction for the precise treatment of HPV-negative HNSCC, and also lays a foundation for further development of individualized immunotherapy strategies39.
Given that HNSCC encompasses a heterogeneous group of tumors across various anatomical sites—including the oral cavity, pharynx, and larynx—conducting omics studies specific to individual sites can help identify their susceptibility to different drug therapies. This enables the formulation of targeted combination treatment strategies to advance personalized care.
Moreover, HNSCC exhibits marked intratumoral heterogeneity, which results in substantial variability in patient responses to conventional anti-PD-1/PD-L1 therapies. Relying solely on classical biomarkers such as TMB, PD-L1 expression, and immune cell infiltration is increasingly insufficient to meet the growing demands of personalized immunotherapy. Recent studies employing multi-omics approaches for molecular subtyping of HNSCC have demonstrated considerable potential for patient stratification. For example, a recent study integrated miRNA, mRNA, methylation, mutation, and copy number variation data from HNSCC patients and identified three distinct molecular subtypes63. These subtypes showed significant differences in clinicopathological features, prognosis, TIME, and treatment vulnerabilities, underscoring their considerable clinical relevance.
Establish preclinical models for immunotherapy research
At present, a significant challenge in cancer treatment development is the discrepancy between existing preclinical models and the in vivo TME64. To enhance drug development and improve immunotherapy outcomes for HNSCC patients, it is crucial to create preclinical models that closely resemble primary tumors.
For instance, 4-nitroquinoline-1-oxide serves as an effective carcinogen for establishing experimental oral carcinogenesis models. Research utilizing genomics and transcriptomics has mapped genomic alterations and immune infiltration throughout the tumorigenesis process in a mouse model of tongue cancer induced by this carcinogen. Several mutated genes identified in this model are frequently observed in human HNSCC, suggesting that this mouse model effectively recapitulates human disease and facilitates the evaluation of various immunotherapy treatments65.
Multi-omics approaches are vital for assessing organoid models. Techniques such as transcriptome sequencing, whole exome sequencing, and whole genome sequencing help identify genetic characteristics of organoids and primary tissues, thereby effectively evaluating their functions in vitro66. Driehuis et al. successfully established tumor mimics from HNSCC patient cells, verifying their ability to identify or validate tumor biomarkers through transcriptome and gene sequencing. The genetic alterations and tumorigenic potential observed following xenotransplantation were documented, confirming that such organoids can serve as platforms for determining effective targeted therapies67. Integrating multi-omics with organoid technology has significantly advanced our understanding of disease mechanisms and therapeutic possibilities68. Exploring organoid-based therapies, along with the utilization and integration of multi-omics, holds promise for elucidating the synergy between these approaches and guiding personalized treatment strategies in the future.
Develop new targets for immunotherapy
Immune checkpoint inhibitors
At present, immune checkpoint inhibitors targeting PD-1/PD-L1 and CTLA-4 have been widely studied and clinically applied in a variety of solid tumors (including HNSCC), and have brought significant survival benefits to some patients. However, only a small number of patients can achieve long-term remission from such single-target therapy, mainly due to the complexity of the tumor immune microenvironment and the heterogeneity between patients. Therefore, it is urgent to explore other immune checkpoints other than PD-1/CTLA-4 in order to exert synergistic effects through joint intervention and improve the overall treatment response.
Lymphocyte-activating gene 3 (LAG-3) is a protein composed of four domains, mainly expressed on activated CD4+ T cells, CD8+ T cells and Tregs. LAG-3 binds to its canonical ligand, leading to exhaustion of immune cells and reduced cytokine secretion69. Based on the analysis of RNA-seq data in The Cancer Genome Atlas, it was found that high mutation load and the expression of exogenous viruses (such as EBV and HPV) and endogenous retroviruses (ERV3-2) are closely related to the high expression of LAG-3 in various cancers, which provides important clues for identifying the types of cancers that are most likely to benefit from LAG-3 blockade therapy, and emphasizes the clinical application prospects of targeting LAG-3 as a potential new target in cancer immunotherapy70. Currently, LAG-3 inhibitor relatlimab combined with nivolumab received FDA approval in 2022 for advanced melanoma, marking the first LAG-3/PD-1 dual checkpoint blockade regimen in clinical practice.
T-cell immunoglobulin and mucin-domain containing-3 (TIM-3) is expressed in tumor cells and various immune cells, such as natural killer (NK) cells, CD8+ T cells, Tregs, etc. The interaction of TIM-3 with its ligand has been shown to induce T cell suppression, and the up-regulation of TIM-3 is uniquely initiated by CD4+ and CD8+ T cells that produce IFN-γ71,72. A recent study used scRNA-seq to explore the phenotype and function of TIM-3+ NK cells in HNSCC patients73. The results showed that the interaction between TIM-3 and its ligand galectin-9 significantly inhibited NK cell-mediated cytotoxicity and proliferation. At the same time, elevated expression of TIM-3+ NK cell signatures in tumors of HNSCC patients are associated with worse prognosis. This indicates the potential of TIM-3 as an immunotherapy target for HNSCC.
The expression of T cell immunoreceptor with Ig and ITIM domains (TIGIT) is tightly restricted to lymphocytes and is mainly observed in natural killer cells and various T cell subsets. TIGIT can inhibit NK cell-mediated tumor killing, induce the formation of immunosuppressive dendritic cells, and hinder the function of CD8+ T cells. In HNSCC and malignant melanoma, TIGIT was significantly overexpressed on both CD4+ and CD8+ T cells, and this expression pattern was closely related to the levels of PD-1, TIM-3 and LAG-374,75. As a widely studied immune checkpoint gene, TIGIT antibody combined with PD-L1 antibody has shown good anti-tumor efficacy in clinical studies. Combined with scRNA-seq and proteomics techniques, researchers have found that combined therapy can activate myeloid cells, that is, TIGIT antibody activates tumor-associated macrophages and monocytes through Fcγ receptor, promotes CD8+ T cells from exhaustion to memory-like state, and reverses the immunosuppressive function of Tregs76. This finding lays a solid theoretical foundation for the development of a new generation of immunotherapy strategies.
Natural Killer Group 2 Member A (NKG2A) is an inhibitory receptor expressed on both T and NK cells, featuring intracytoplasmic tyrosine-based inhibitory motifs. Its binding to cognate ligands inhibits the effector functions of these immune cells77. The ligand of NKG2A is the unconventional MHC-I molecule HLA-E, which is expressed at a low level in normal tissues, but can be overexpressed in a variety of cancers78. Quantitative RNA sequencing analysis of lymphocytes from HNSCC patients shows that NKG2A is expressed by the majority of NK cells and selectively by CD8+ T cells within the tumor microenvironment. Furthermore, transcriptomic data indicate that high transcription levels of CD8 genes correlate with better prognosis; however, high co-expression of KLRC1 (which encodes NKG2A) neutralizes this benefit, suggesting a potentially detrimental role for NKG2A in HNSCC. Therefore, blocking NKG2A may enhance the immune therapy response in HNSCC patients79.
Tumor vaccines
Therapeutic cancer vaccines typically contain tumor-specific or tumor-associated antigens that activate the body’s immune response to produce anti-tumor effects80.
A study successfully combined genome, transcriptome, and proteome data to identify several oropharyngeal squamous cell carcinoma-specific tumor-associated peptides, which can serve as potential targets for immunotherapy. This approach not only aids in developing personalized cancer vaccines but also activates cytotoxic T cells to enhance anti-tumor immune responses81.
HPV, particularly HPV-16, is etiologically linked to approximately 70% of oropharyngeal squamous cell carcinomas, while accounting for less than 5% of non-oropharyngeal HNSCC. The viral oncoproteins E6 and E7 drive carcinogenesis through degradation of p53 and retinoblastoma (pRb) tumor suppressor proteins, respectively, while evading immune detection by downregulating MHC class I expression82.
Although prophylactic HPV vaccines have demonstrated 90–100% efficacy in preventing oral HPV-16/18 infections in clinical trials, their therapeutic potential in established HNSCC remains limited. In the study of HPV-16 vaccine combined with anti-PD-1 treatment, the objective remission rate of combined treatment is higher than that of patients treated with anti-PD-1 alone83.
Mechanistic insights from integrated scRNA-seq and TCR repertoire analysis reveal that HPV-specific CD8+ T cells were activated after HPV mRNA vaccine inoculation, and the effector memory and exhausted T cell subsets showed excessive expansion of TCR clonality84.
Future investigations should leverage multi-omics approaches (e.g., spatial transcriptomics, TCRβ chain immunophenotyping) to resolve the spatiotemporal dynamics of vaccine-primed T cell populations within the immunosuppressive tumor microenvironment (TME). This may inform rational combinations with novel agents targeting exhausted T cell reinvigoration (e.g., anti-TIGIT, IL-15 superagonists) to achieve durable antitumor immunity.
Cell therapy
Cell therapy refers to the transplantation of viable autologous or allogeneic cells into patients following in vitro manipulation, aimed at replacing diseased or damaged cells, regulating cell function, or assisting in the removal of pathogenic or dysfunctional cells85. In the context of HNSCC, while adoptive cell therapies such as tumor-infiltrating lymphocytes and CAR-T cells have shown promise in early-phase trials, their clinical translation remains hindered by the paucity of tumor-specific surface antigens with optimal therapeutic windows. Recent pan-cancer analyses reveal that HNSCC exhibits particularly low expression of currently targeted CAR-T antigens (e.g., EGFRvIII, HER2) compared to other solid tumors, with less than 30% of tumors expressing these targets at clinically actionable levels. This underscores the critical need for novel target discovery specific to HNSCC’s molecular landscape86.
Addressing this gap, Sanna Madan et al. employed single-cell transcriptomic and proteomic profiling to comprehensively map CAR targets across diverse cancer types and systematically identified 20 new cell surface targets that are either safer or more specific than current options. Five of these targets demonstrate excellent selectivity and safety scores, making them potential candidates for CAR therapy in HNSCC86.
Other immunotherapy targets
As our understanding of the molecular mechanisms involved in cancer development and progression deepens, small molecule inhibitors are increasingly becoming essential components of cancer treatment87. Recent studies utilizing single-cell transcriptome sequencing and transcriptomics have shown that HPV-negative tumors are more reliant on IL-6/IL-6R and CCL2/CCR2 signaling within the TME to evade NK cell immune attacks. Inhibiting IL-6 can enhance NK cell infiltration and proliferation, while the combined use of CCR2 chemokine receptor antagonists and IL-6 blockers may yield a more pronounced anti-tumor effect.
In addition to small molecule inhibitors, antibody therapeutics represent another essential component of cancer treatment. Currently, HNSCC lacks mature and highly specific antigens. Monoclonal antibodies targeting EGFR are among the earliest antibody-targeted therapies utilized clinically, yet this target is also widely expressed in normal cells88. Interleukin-10 (IL-10) is recognized as an anti-inflammatory mediator that inhibits antigen-presenting cells and shows promise in anti-tumor therapy. Spatial transcriptomics and transcriptome analyses have been employed to explore the correlation between IL-10 expression, IL-10 receptor alpha, and colony-stimulating factor 1 receptor (CSF1R) levels with CD8+ T cell and tumor-associated macrophage scores in HNSCC. A profile characterized by high IL-10 and low CSF1R expression correlates with an activated immune signature and better prognosis in HNSCC patients. A fusion protein combining IL-10 with an anti-CSF1R antibody has demonstrated the ability to reprogram the TME into an immunologically active state. Furthermore, combining this therapy with αPD-1 has been shown to enhance anti-tumor activity and promote the production of pro-inflammatory cytokines89.
Challenges and perspectives
Challenges
While multi-omics technology has significantly advanced the development of immunotherapy for HNSCC, it also encounters several challenges. Firstly, current sequencing technologies impose limitations on sequencing depth, which can impact the accuracy of transcriptomic data. This limitation may hinder the identification of low-frequency variations and result in the omission of crucial biological information. Additionally, significant batch effects often arise during cross-platform sequencing and multi-sample integration. There are substantial discrepancies in the standards for batch processing across various studies, which can reduce the reproducibility of bioinformatics analyses. Furthermore, conclusions drawn from the analysis of limited omics data frequently depend on in vitro and in vivo experiments. This reliance can diminish the credibility of the findings, and interpretations are often subjective, potentially leading to deviations from reality and even opposite conclusions. Finally, in the integration and analysis of multi-omics data, the sequencing samples are often from different sources, and there is a lack of multi-parameter sequencing analysis of the same sample. The conclusions obtained by simply combining the results of different omics methods may cover up important biological information due to sample heterogeneity. At present, there are methods to extract mRNA, protein and spatial information of the same sample for analysis (DBiT-seq)90, and a full-coverage spatial full-transcriptome sequencing technology (Patho-DBiT) for clinical archived formalin fixed paraffin embedded (FFPE) tissues has been developed91, which accurately decodes information such as mRNA, non-coding RNA expression, alternative splicing, genetic variation, microRNA regulation and RNA dynamic changes in FFPE complex tissues. The emergence of these technologies provides a new idea for the combined application of multi-omics.
Future perspectives
This section outlines five pivotal directions to advance immunotherapy research in HNSCC: (1) optimization of sequencing samples, (2) enhanced data integration and analysis, (3) application of emerging omics technologies, (4) incorporation of clinicopathological features, and (5) multidisciplinary convergence (Fig. 3).
Future multi-omics has better development potential in five aspects, including the selection and processing of sequencing samples, the integration and analysis of data, the use of emerging technologies, the analysis combined with clinicopathological features, and the use of artificial intelligence. Created with BioRender.com.
Current sequencing efforts predominantly focus on primary tumors, lymph nodes, peripheral blood, and liquid biopsies (e.g., saliva). However, emerging evidence highlights the untapped potential of alternative biospecimens. For instance, T cell-derived extracellular vesicles have been shown to induce systemic immunosuppression and predict immunotherapy responses in HNSCC92, while post-neck dissection drainage fluid may serve as a biomarker for lymph node metastasis93. To address sample heterogeneity and improve data reliability, multi-center collaborations and expanded sequencing cohorts are imperative. Standardization of sample processing protocols and batch management systems will further ensure reproducibility and cross-study comparability.
Public repositories such as GEO (www.ncbi.nlm.nih.gov/geo), TCGA (www.cancer.gov/ccg/research/genome-sequencing/tcga), and the Human Cell Atlas (HCA) (www.humancellatlas.org) provide extensive multi-omics datasets encompassing single-cell transcriptomics, epigenomics, and proteomics. Integrative analysis of these resources could refine drug efficacy predictions and identify immunotherapy-responsive subpopulations. Dedicated platforms like TIGER94 and ICBatlas95 enable direct interrogation of immune response markers and single-cell spatial distributions. Advanced computational tools (e.g., CIBERSORT, TIMER, xCell) should be systematically employed to deconvolute bulk sequencing data and quantify immune cell infiltration dynamics.
Cutting-edge spatial and single-cell omics are revolutionizing TME characterization. Single-cell Assay for Transposase-Accessible Chromatin with high throughput sequencing (ATAC-seq) enables chromatin accessibility mapping at cellular resolution, elucidating context-dependent gene regulatory networks96. Spatiotemporal omics platforms now permit dynamic monitoring of immunotherapy-induced TME remodeling, including shifts in immune cell neighborhoods associated with treatment response in melanoma models97. These technologies hold significant promise for resolving HNSCC-specific immune evasion mechanisms.
Patient heterogeneity—driven by anatomical subsite, HPV status, lesion classification, and metastatic patterns—critically impacts immunotherapy outcomes. Stratified sequencing based on these features could unravel resistance mechanisms and guide personalized therapeutic strategies. Concurrently, omics-driven identification of risk predictors for immune-related adverse events (e.g., pneumonia and myocarditis) is essential for optimizing patient selection and toxicity management98.
Artificial intelligence is emerging as a transformative tool for multi-omics integration. Deep learning models that synthesize genomic, molecular, and clinical data have demonstrated superior predictive accuracy for immunotherapy responses in non-small cell lung cancer and melanoma99,100. In HNSCC, such approaches could overcome the limitations of single-biomarker strategies by decoding complex interactions between tumor biology and host immunity.
Conclusion
In this review, we have synthesized the transformative role of multi-omics technologies in advancing immunotherapy research for head and neck squamous cell carcinoma (HNSCC). By dissecting tumor biology at genomic, transcriptomic, epigenomic, and proteomic levels, these tools have not only deepened our understanding of immune evasion mechanisms but also accelerated the translation of preclinical insights into clinical strategies. As a cornerstone of twenty-first-century biomedical research, multi-omics approaches are reshaping therapeutic paradigms—enabling biomarker discovery, patient stratification, and dynamic monitoring of treatment responses.
Despite these advancements, challenges persist in data integration, technical standardization, and clinical validation. Addressing these limitations through interdisciplinary collaboration and AI-driven analytics will be critical to unlocking the full potential of multi-omics in precision immunotherapy. Looking ahead, the convergence of emerging technologies (e.g., spatiotemporal omics, single-cell profiling) with robust clinical frameworks promises to refine personalized treatment regimens, mitigate immune-related toxicities, and ultimately improve survival outcomes for HNSCC patients.
Data availability
No datasets were generated or analysed during the current study.
Abbreviations
- BCR:
-
B cell receptor
- CAF:
-
cancer-associated fibroblast
- CAR:
-
chimeric antigen receptor
- CSF1R:
-
colony-stimulating factor 1 receptor
- CTCs:
-
circulating tumor cells
- CTLA-4:
-
cytotoxic T-lymphocyte-associated protein 4
- EGFR:
-
epidermal growth factor receptor
- FFPE:
-
formalin fixed paraffin embedded
- HNSCC:
-
head and neck squamous cell carcinoma
- HPV:
-
human papillomavirus
- ICI:
-
immune checkpoint inhibitor
- IFN-γ:
-
interferon-γ
- LAG-3:
-
lymphocyte-activating gene 3
- NK:
-
natural killer
- OSCC:
-
oral squamous cell carcinoma
- PD-1:
-
programmed death 1
- PD-L1:
-
programmed death ligand 1
- scRNA-seq:
-
single-cell RNA sequencing
- TCR:
-
T cell receptor
- tdLNs:
-
tumor-draining lymph nodes
- TIGIT:
-
T cell immunoreceptor with Ig and ITIM domains
- TIM-3:
-
T-cell immunoglobulin and mucin-domain containing-3
- TIME:
-
tumor immune microenvironment
- TMB:
-
tumor mutation burden
- TME:
-
tumor microenvironment
- Tregs:
-
regulatory T cells
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Chow, L. Q. M. Head and neck cancer. N. Engl. J. Med. 382, 60–72 (2020).
Bhatia, A. & Burtness, B. Treating head and neck cancer in the age of immunotherapy: a 2023 update. Drugs 83, 217–248 (2023).
Harrington, K. J. et al. Pembrolizumab with or without chemotherapy in recurrent or metastatic head and neck squamous cell carcinoma: updated results of the phase III KEYNOTE-048 study. J. Clin. Oncol. 41, 790–802 (2023).
Cramer, J. D., Burtness, B. & Ferris, R. L. Immunotherapy for head and neck cancer: recent advances and future directions. Oral. Oncol. 99, 104460 (2019).
Liu, H. M. et al. Neoadjuvant immunotherapy with or without chemotherapy in locally advanced oral squamous cell carcinoma: randomized, two-arm, phase 2 trial. Cell Rep. Med. 6, 101930 (2025).
Sepiashvili, L. et al. Novel insights into head and neck cancer using next-generation “omic” technologies. Cancer Res. 75, 480–486 (2015).
Baysoy, A. et al. The technological landscape and applications of single-cell multi-omics. Nat. Rev. Mol. Cell Biol. 24, 695–713 (2023).
Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582 (2015).
Rahim, M. K. et al. Dynamic CD8(+) T cell responses to cancer immunotherapy in human regional lymph nodes are disrupted in metastatic lymph nodes. Cell 186, 1127–1143.e18 (2023).
Zhong, N. N., Liu, B. & Bu, L. L. Neoadjuvant immunotherapy: new horizon for lymph node preservation. MedComm5, e577 (2024).
Badoual, C. et al. PD-1-expressing tumor-infiltrating T cells are a favorable prognostic biomarker in HPV-associated head and neck cancer. Cancer Res. 73, 128–138 (2013).
Sun, W. et al. A positive-feedback loop between tumour infiltrating activated Treg cells and type 2-skewed macrophages is essential for progression of laryngeal squamous cell carcinoma. Br. J. Cancer 117, 1631–1643 (2017).
Hadler-Olsen, E. & Wirsing, A. M. Tissue-infiltrating immune cells as prognostic markers in oral squamous cell carcinoma: a systematic review and meta-analysis. Br. J. Cancer 120, 714–727 (2019).
Cheng, D. et al. Molecular and transcriptional basis of bidirectional CD4(+) T cell exhaustion in oropharyngeal squamous cell carcinoma. MedComm5, e572 (2024).
Parra, E. R. et al. Multi-omics analysis reveals immune features associated with immunotherapy benefit in patients with squamous cell lung cancer from phase III Lung-MAP S1400I trial. Clin. Cancer Res. 30, 1655–1668 (2024).
Berger, M. F. & Mardis, E. R. The emerging clinical relevance of genomics in cancer medicine. Nat. Rev. Clin. Oncol. 15, 353–365 (2018).
Rodrigo, J. P. et al. Tumor mutational burden predictability in head and neck squamous cell carcinoma patients treated with immunotherapy: systematic review and meta-analysis. J. Transl. Med. 22, 135 (2024).
Peng, Q. et al. Impacts and mechanisms of alternative mRNA splicing in cancer metabolism, immune response, and therapeutics. Mol. Ther. 30, 1018–1035 (2022).
Griffith, O. L. et al. The prognostic effects of somatic mutations in ER-positive breast cancer. Nat. Commun. 9, 3476 (2018).
Wang, J. et al. Single-cell RNA sequencing highlights the functional role of human endogenous retroviruses in gallbladder cancer. EBioMedicine 85, 104319 (2022).
Shi, Y. et al. Mutant p53 drives an immune cold tumor immune microenvironment in oral squamous cell carcinoma. Commun. Biol. 5, 757 (2022).
Wang, K. C. & Chang, H. Y. Epigenomics: technologies and applications. Circ. Res. 122, 1191–1199 (2018).
Micevic, G., Bosenberg, M. W. & Yan, Q. The crossroads of cancer epigenetics and immune checkpoint therapy. Clin. Cancer Res. 29, 1173–1182 (2023).
Starzer, A. M. et al. DNA methylation profiles differ in responders versus non-responders to anti-PD-1 immune checkpoint inhibitors in patients with advanced and metastatic head and neck squamous cell carcinoma. J. Immunother. Cancer 10, e003420 (2022).
Byron, S. A. et al. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat. Rev. Genet. 17, 257–271 (2016).
Gavrielatou, N. et al. Biomarkers for immunotherapy response in head and neck cancer. Cancer Treat. Rev. 84, 101977 (2020).
Zhao, M. et al. T cell dynamics with neoadjuvant immunotherapy in head and neck cancer. Nat. Rev. Clin. Oncol. 22, 83–94 (2025).
Papalexi, E. & Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat. Rev. Immunol. 18, 35–45 (2018).
Ren, S. et al. Intratumoral CD103(+) CD8(+) T cells predict response to neoadjuvant chemoimmunotherapy in advanced head and neck squamous cell carcinoma. Cancer Commun.43, 1143–1163 (2023).
Zhang, S. et al. Antibody repertoire sequencing analysis. Acta Biochim. Biophys. Sin.54, 864–873 (2022).
Jiang, N., Schonnesen, A. A. & Ma, K. Y. Ushering in integrated T cell repertoire profiling in cancer. Trends Cancer 5, 85–94 (2019).
Luoma, A. M. et al. Tissue-resident memory and circulating T cells are early responders to pre-surgical cancer immunotherapy. Cell 185, 2918–2935.e29 (2022).
Zhou, L. et al. Checkpoint blockade-induced CD8+ T cell differentiation in head and neck cancer responders. J. Immunother Cancer 10, e004034 (2022).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med. 25, 1251–1259 (2019).
Cui, M., Cheng, C. & Zhang, L. High-throughput proteomics: a methodological mini-review. Lab Invest. 102, 1170–1181 (2022).
Monti, C. et al. Proteomics turns functional. J. Proteom. 198, 36–44 (2019).
Tretter, C. et al. Proteogenomic analysis reveals RNA as a source for tumor-agnostic neoantigen identification. Nat. Commun. 14, 4632 (2023).
Huang, C. et al. Proteogenomic insights into the biology and treatment of HPV-negative head and neck squamous cell carcinoma. Cancer Cell 39, 361–379.e16 (2021).
Payne, K. et al. Feasibility of mass cytometry proteomic characterisation of circulating tumour cells in head and neck squamous cell carcinoma for deep phenotyping. Br. J. Cancer 129, 1590–1598 (2023).
Cao, X. et al. Oral immunotherapy reshapes intestinal immunosuppression via metabolic reprogramming to enhance systemic anti-tumor immunity. Adv. Sci. 10, e2302910 (2023).
Li, H. et al. Metabolomic adaptations and correlates of survival to immune checkpoint blockade. Nat. Commun. 10, 4346 (2019).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
Zheng, D. W. et al. Biomaterial-mediated modulation of oral microbiota synergizes with PD-1 blockade in mice with oral squamous cell carcinoma. Nat. Biomed. Eng. 6, 32–43 (2022).
Liu, Z. et al. Spatial transcriptomics reveals that metabolic characteristics define the tumor immunosuppression microenvironment via iCAF transformation in oral squamous cell carcinoma. Int. J. Oral. Sci. 16, 9 (2024).
Taylor, M. H. et al. Safety and efficacy of pembrolizumab in combination with acalabrutinib in advanced head and neck squamous cell carcinoma: phase 2 proof-of-concept study. Clin. Cancer Res. 28, 903–914 (2022).
Chen, P. et al. Spatially resolved metabolomics combined with the 3D tumor-immune cell coculture spheroid highlights metabolic alterations during antitumor immune response. Anal. Chem. 95, 15153–15161 (2023).
Karczewski, K. J. & Snyder, M. P. Integrative omics for health and disease. Nat. Rev. Genet. 19, 299–310 (2018).
He, X. et al. Artificial intelligence-based multi-omics analysis fuels cancer precision medicine. Semin. Cancer Biol. 88, 187–200 (2023).
Vandereyken, K. et al. Methods and applications for single-cell and spatial multi-omics. Nat. Rev. Genet. 24, 494–515 (2023).
Wu, D. et al. Neoadjuvant chemo-immunotherapy with camrelizumab plus nab-paclitaxel and cisplatin in resectable locally advanced squamous cell carcinoma of the head and neck: a pilot phase II trial. Nat. Commun. 15, 2177 (2024).
Li, Z. Z. et al. Nanoparticles targeting lymph nodes for cancer immunotherapy: strategies and influencing factors. Small 20, e2308731 (2024).
Chen, J. et al. Lipid nanoparticle-mediated lymph node-targeting delivery of mRNA cancer vaccine elicits robust CD8(+) T cell response. Proc. Natl Acad. Sci. USA 119, e2207841119 (2022).
Lakins, M. A. et al. Cancer-associated fibroblasts induce antigen-specific deletion of CD8 (+) T Cells to protect tumour cells. Nat. Commun. 9, 948 (2018).
Li, C. et al. Spatial and single-cell transcriptomics reveal a cancer-associated fibroblast subset in HNSCC that restricts infiltration and antitumor activity of CD8+ T cells. Cancer Res. 84, 258–275 (2024).
Obradovic, A. et al. Immunostimulatory cancer-associated fibroblast subpopulations can predict immunotherapy response in head and neck cancer. Clin. Cancer Res. 28, 2094–2109 (2022).
Franken, A. et al. CD4(+) T cell activation distinguishes response to anti-PD-L1+anti-CTLA4 therapy from anti-PD-L1 monotherapy. Immunity 57, 541–558.e7 (2024).
Chen, D. S. & Mellman, I. Elements of cancer immunity and the cancer-immune set point. Nature 541, 321–330 (2017).
Galluzzi, L. et al. The hallmarks of successful anticancer immunotherapy. Sci. Transl. Med. 10, eaat7807 (2018).
Le Meitour, Y. et al. Uncovering immune checkpoint heterogeneity in oral squamous cell carcinoma using single cell RNA-sequencing data highlights three subgroups of patients with distinct immune phenotypes. Oral. Oncol. 149, 106680 (2024).
Zhang, S., Zhang, W. & Zhang, J. 8-Gene signature related to CD8(+) T cell infiltration by integrating single-cell and bulk RNA-sequencing in head and neck squamous cell carcinoma. Front. Genet. 13, 938611 (2022).
Lechner, M. et al. HPV-associated oropharyngeal cancer: epidemiology, molecular biology and clinical management. Nat. Rev. Clin. Oncol. 19, 306–327 (2022).
Diao, P. et al. Integrative multiomics analyses identify molecular subtypes of head and neck squamous cell carcinoma with distinct therapeutic vulnerabilities. Cancer Res. 84, 3101–3117 (2024).
Wen, L. & Tang, F. Organoid research on human early development and beyond. Med. Rev.2, 512–523 (2022).
Lee, Y. M. et al. Genomic and transcriptomic landscape of an oral squamous cell carcinoma mouse model for immunotherapy. Cancer Immunol. Res. 11, 1553–1567 (2023).
Kanton, S. et al. Organoid single-cell genomic atlas uncovers human-specific features of brain development. Nature 574, 418–422 (2019).
Driehuis, E. et al. Oral mucosal organoids as a potential platform for personalized cancer therapy. Cancer Discov. 9, 852–871 (2019).
Han, X. et al. Landscape of human organoids: ideal model in clinics and research. Innovation5, 100620 (2024).
Huard, B. et al. Lymphocyte-activation gene 3/major histocompatibility complex class II interaction modulates the antigenic response of CD4+ T lymphocytes. Eur. J. Immunol. 24, 3216–3221 (1994).
Panda, A. et al. Genomic and immunologic correlates of LAG-3 expression in cancer. Oncoimmunology 9, 1756116 (2020).
Sakuishi, K. et al. Targeting Tim-3 and PD-1 pathways to reverse T cell exhaustion and restore anti-tumor immunity. J. Exp. Med. 207, 2187–2194 (2010).
Gao, X. et al. TIM-3 expression characterizes regulatory T cells in tumor tissues and is associated with lung cancer progression. PLoS ONE 7, e30676 (2012).
Wang, J. et al. Differential impact of TIM-3 ligands on NK cell function. J. Immunother. Cancer 13, e010618 (2025).
Wu, L. et al. Blockade of TIGIT/CD155 signaling reverses T-cell exhaustion and enhances antitumor capability in head and neck squamous cell carcinoma. Cancer Immunol. Res. 7, 1700–1713 (2019).
Chauvin, J. M. et al. TIGIT and PD-1 impair tumor antigen-specific CD8(+) T cells in melanoma patients. J. Clin. Invest. 125, 2046–2058 (2015).
Guan, X. et al. Anti-TIGIT antibody improves PD-L1 blockade through myeloid and T(reg) cells. Nature 627, 646–655 (2024).
Andre, P. et al. Anti-NKG2A mAb is a checkpoint inhibitor that promotes anti-tumor immunity by unleashing both T and NK cells. Cell 175, 1731–1743.e13 (2018).
Borst, L., van der Burg, S. H. & van Hall, T. The NKG2A-HLA-E axis as a novel checkpoint in the tumor microenvironment. Clin. Cancer Res. 26, 5549–5556 (2020).
van Montfoort, N. et al. NKG2A blockade potentiates CD8 T cell immunity induced by cancer vaccines. Cell 175, 1744–1755.e15 (2018).
Fritsch, E. F. et al. HLA-binding properties of tumor neoepitopes in humans. Cancer Immunol. Res. 2, 522–529 (2014).
Abou Kors, T. et al. Multi-omics analysis of overexpressed tumor-associated proteins: gene expression, immunopeptide presentation, and antibody response in oropharyngeal squamous cell carcinoma, with a focus on cancer-testis antigens. Front. Immunol. 15, 1408173 (2024).
Aggarwal, C. et al. Safety and efficacy of MEDI0457 plus durvalumab in patients with human papillomavirus-associated recurrent/metastatic head and neck squamous cell carcinoma. Clin. Cancer Res. 29, 560–570 (2023).
Massarelli, E. et al. Combining immune checkpoint blockade and tumor-specific vaccine for patients with incurable human papillomavirus 16-related cancer: a phase 2 clinical trial. JAMA Oncol. 5, 67–73 (2019).
Qiu, K. et al. mRNA-LNP vaccination-based immunotherapy augments CD8(+) T cell responses against HPV-positive oropharyngeal cancer. NPJ Vaccines 8, 144 (2023)
El-Kadiry, A. E., Rafei, M. & Shammaa, R. Cell therapy: types, regulation, and clinical benefits. Front. Med.8, 756029 (2021).
Madan, S. et al. Pan-cancer analysis of patient tumor single-cell transcriptomes identifies promising selective and safe chimeric antigen receptor targets in head and neck cancer. Cancers 15, 4885 (2023).
Zhang, J. et al. If small molecules immunotherapy comes, can the prime be far behind?. Eur. J. Med. Chem. 218, 113356 (2021).
He, C. H. & Li, X. M. Research progress in immunotherapy of head and neck squamous cell carcinoma]. Zhonghua Er Bi Yan Hou Tou Jing Wai Ke Za Zhi 57, 82–88 (2022).
Chang, Y. W. et al. A CSF-1R-blocking antibody/IL-10 fusion protein increases anti-tumor immunity by effectuating tumor-resident CD8(+) T cells. Cell Rep. Med. 4, 101154 (2023).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681.e18 (2020).
Bai, Z. et al. Spatially exploring RNA biology in archival formalin-fixed paraffin-embedded tissues. Cell 187, 6760–6779.e24 (2024).
Zhang, L. Z. et al. PD-1/CD80(+) small extracellular vesicles from immunocytes induce cold tumours featured with enhanced adaptive immunosuppression. Nat. Commun. 15, 3884 (2024).
Wang, J. et al. Preliminary extracellular vesicle profiling in drainage fluid after neck dissection in OSCC. J. Dent. Res. 102, 178–186 (2023).
Chen, Z. et al. TIGER: a web portal of tumor immunotherapy gene expression resource. Genomics Proteom. Bioinforma. 21, 337–348 (2023).
Yang, M. et al. ICBatlas: a comprehensive resource for depicting immune checkpoint blockade therapy characteristics from transcriptome profiles. Cancer Immunol. Res. 10, 1398–1406 (2022).
Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39, 1578–1593.e8 (2021).
Hickey, J. W. et al. T cell-mediated curation and restructuring of tumor tissue coordinates an effective immune response. Cell Rep. 42, 113494 (2023).
Jing, Y. et al. Multi-omics prediction of immune-related adverse events during checkpoint immunotherapy. Nat. Commun. 11, 4946 (2020).
Vanguri, R. S. et al. Multimodal integration of radiology, pathology and genomics for prediction of response to PD-(L)1 blockade in patients with non-small cell lung cancer. Nat. Cancer 3, 1151–1164 (2022).
Chowell, D. et al. Improved prediction of immune checkpoint blockade efficacy across multiple cancer types. Nat. Biotechnol. 40, 499–506 (2022).
Song, A. et al. Tumor-intrinsic SIRPA drives pyroptosis evasion in head and neck cancer. J. Dent. Res. 0, 220345241305590 (2025).
Qin, H. et al. Disulfidptosis-related gene signatures as prognostic biomarkers and predictors of immunotherapy response in HNSCC. Front. Immunol. 15, 1456649 (2024).
Wu, C. S. et al. Integrated multi-omics analyses of oral squamous cell carcinoma reveal precision patient stratification and personalized treatment strategies. Cancer Lett. 614, 217482 (2025).
Chang, T. G. et al. Tumor and blood B-cell abundance outperforms established immune checkpoint blockade response prediction signatures in head and neck cancer. Ann. Oncol. 36, 309–320 (2025).
Ding, Q. et al. Inhibition of PNCK inflames tumor microenvironment and sensitizes head and neck squamous cell carcinoma to immune checkpoint inhibitors. J. Immunother. Cancer 12, e009893 (2024).
Cha, J. et al. Single-cell analysis reveals cellular and molecular factors counteracting HPV-positive oropharyngeal cancer immunotherapy outcomes. J. Immunother. Cancer 12, e008667 (2024).
Li, K. et al. Single cell analysis unveils B cell-dominated immune subtypes in HNSCC for enhanced prognostic and therapeutic stratification. Int. J. Oral. Sci. 16, 29 (2024).
Zhang, X. Y. et al. Metabolic landscape of head and neck squamous cell carcinoma informs a novel kynurenine/Siglec-15 axis in immune escape. Cancer Commun.44, 670–694 (2024).
Xu, Y. et al. A novel CAF-cancer cell crosstalk-related gene prognostic index based on machine learning: prognostic significance and prediction of therapeutic response in head and neck squamous cell carcinoma. J. Transl. Med. 22, 645 (2024).
Chen, Z. et al. TP63 transcriptionally regulates SLC7A5 to suppress ferroptosis in head and neck squamous cell carcinoma. Front. Immunol. 15, 1445472 (2024).
Cao, Y. et al. Inferring characteristics of the tumor immune microenvironment of patients with HNSCC from single-cell transcriptomics of peripheral blood. Cancer Res. Commun. 4, 2335–2348 (2024).
Yan, S. et al. Deciphering the interplay of HPV infection, MHC-II expression, and CXCL13(+) CD4(+) T cell activation in oropharyngeal cancer: implications for immunotherapy. Cancer Immunol. Immunother. 73, 206 (2024).
Zhang, J. W. et al. Integrated analysis of methylation and transcriptome identifies a novel risk model for diagnosis, prognosis, and immune characteristics in head and neck squamous cell carcinoma. Mol. Genet. Genomics 299, 71 (2024).
Cao, H. et al. FAT1 as a tumor mutation burden specific gene affects the immunotherapy effect in head and neck squamous cell cancer. Drug Resist. Updat. 76, 101095 (2024).
Li, Y., Wang, N. & Yang, G. Multi-omic analysis and validation reveal ZBP1 as a potential prognostic and immunotherapy-related biomarker in head and neck squamous cell carcinoma. J. Stomatol. Oral. Maxillofac. Surg. 125, 101901 (2024).
Ma, H. et al. Comprehensive investigation into the influence of glycosylation on head and neck squamous cell carcinoma and development of a prognostic model for risk assessment and anticipating immunotherapy. Front. Immunol. 15, 1364082 (2024).
Li, R. et al. DNA-methylome-derived epigenetic fingerprint as an immunophenotype indicator of durable clinical immunotherapeutic benefits in head and neck squamous cell carcinoma. Cell Oncol.47, 1129–1148 (2024).
Sadeghirad, H. et al. Spatial dynamics of tertiary lymphoid aggregates in head and neck cancer: insights into immunotherapy response. J. Transl. Med. 22, 677 (2024).
Lin, X. et al. CXC ligand 13 orchestrates an immunoactive microenvironment and enhances immunotherapy response in head and neck squamous cell carcinoma. Int. J. Immunopathol. Pharm. 38, 3946320241227312 (2024).
Quah, H. S. et al. Single cell analysis in head and neck cancer reveals potential immune evasion mechanisms during early metastasis. Nat. Commun. 14, 1680 (2023).
Li, Z. et al. A novel oxidative stress-related gene signature as an indicator of prognosis and immunotherapy responses in HNSCC. Aging15, 14957–14984 (2023).
Dai, Y. et al. Integrative single-cell and bulk transcriptomes analyses identify intrinsic HNSCC subtypes with distinct prognoses and therapeutic vulnerabilities. Clin. Cancer Res. 29, 2845–2858 (2023).
Cui, J. et al. Cancer germline antigen gene MAGEB2 promotes cell invasion and correlates with immune microenvironment and immunotherapeutic efficiency in laryngeal cancer. Clin. Immunol. 240, 109045 (2022).
Tanagala, K. K. K. et al. SP140 inhibits STAT1 signaling, induces IFN-gamma in tumor-associated macrophages, and is a predictive biomarker of immunotherapy response. J. Immunother. Cancer 10, e005088 (2022).
Wang, W. et al. Stress keratin 17 expression in head and neck cancer contributes to immune evasion and resistance to immune-checkpoint blockade. Clin. Cancer Res. 28, 2953–2968 (2022).
Zhao, X. et al. Somatic 9p24.1 alterations in HPV(-) head and neck squamous cancer dictate immune microenvironment and anti-PD-1 checkpoint inhibitor activity. Proc. Natl Acad. Sci. USA 119, e2213835119 (2022).
Woolaver, R. A. et al. Differences in TCR repertoire and T cell activation underlie the divergent outcomes of antitumor immune responses in tumor-eradicating versus tumor-progressing hosts. J. Immunother. Cancer 9, e001615 (2021).
Shi, C. et al. A TP53 mutation model for the prediction of prognosis and therapeutic responses in head and neck squamous cell carcinoma. BMC Cancer 21, 1035 (2021).
Lin, W. et al. Multi-omics data analyses identify B7-H3 as a novel prognostic biomarker and predict response to immune checkpoint blockade in head and neck squamous cell carcinoma. Front. Immunol. 12, 757047 (2021).
Feng, B. et al. Integrative analysis of multi-omics data identified EGFR and PTGS2 as key nodes in a gene regulatory network related to immune phenotypes in head and neck cancer. Clin. Cancer Res. 26, 3616–3628 (2020).
de Vos, L. et al. The landscape of CD28, CD80, CD86, CTLA4, and ICOS DNA methylation in head and neck squamous cell carcinomas. Epigenetics 15, 1195–1212 (2020).
Acknowledgements
This study was supported by Hubei Provincial Natural Science Foundation of China (2024AFD448) to Z.L., the Fundamental Research Funds for the Central Universities (Wuhan University, Clinical Medicine + X) (2042024YXB017), Hubei Province Chinese Medicine Research Project (ZY2023Q015), Natural Science Foundation of Hubei Province (2023AFB665), Medical Young Talents Program of Hubei Province, and Wuhan Young Medical Talents Training Project to L.-L.B.
Author information
Authors and Affiliations
Contributions
Xuan-Hao Liu: investigation, methodology, conceptualization, writing—original draft, visualization, writing—review & editing. Guang-Rui Wang: investigation, methodology, conceptualization, writing—original draft, writing—review & editing. Nian-Nian Zhong: investigation, methodology, conceptualization, writing—original draft, writing—review & editing. Wei-Yu Wang: investigation, methodology, writing—original draft. Bing Liu: conceptualization, supervision, writing—review & editing. Zheng Li: investigation, methodology, conceptualization, funding acquisition, writing—review & editing. Lin-Lin Bu: conceptualization, supervision, project administration, funding acquisition, writing—review & editing. All authors have reviewed and approved the final version of this manuscript for publication. Each author agrees to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, XH., Wang, GR., Zhong, NN. et al. Multi-omics in immunotherapy research for HNSCC: present situation and future perspectives. npj Precis. Onc. 9, 93 (2025). https://doi.org/10.1038/s41698-025-00886-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-00886-w
This article is cited by
-
Single-cell omics in tumor lymph node metastasis: mechanisms and therapeutic implications
Molecular Cancer (2026)
-
The impact of genomic ancestry on tumor genomics in head and neck squamous cell carcinoma
Cancer and Metastasis Reviews (2026)
-
Overcoming resistance to anti-PD-L1 immunotherapy: mechanisms, combination strategies, and future directions
Molecular Cancer (2025)
-
Interferon signaling and STING pathway in head and neck cancers: unlocking immune secrets and therapeutic frontiers
Cancer Cell International (2025)