Abstract
The clinical application of deep learning (DL)-based brain tumor segmentation remains limited by missing MRI sequences and cross-center data inconsistencies. Existing supervised generative models can synthesize missing sequences but rely on paired, fully sampled training data, which are often unavailable in routine practice. This study aims to assess the use of an unsupervised generative model to complete missing sequences and eliminate cross-center data inconsistency, and to verify whether using the generated images can enhance brain tumor segmentation. We retrospectively evaluated 921 glioblastoma (GBM) patients from BraTS, UCSF-PDGM, and our institutional datasets, together with 1000 meningioma cases from BraTS-MEN cohort. We developed an unsupervised multi-center multi-sequence generative adversarial transformer (UMMGAT) to generate MRI sequences from incomplete datasets. Key features of UMMGAT include a sequence encoder that disentangles and encodes modality-specific characteristics, and a lesion-aware module (LAM) that enhances the generation of tumor regions, all trained via multi-task learning for generating multi-modal images. Validation on GBM and meningioma segmentation task demonstrated that generated MRI sequences significantly improved segmentation performance across various missing-sequence scenarios. The enhancement in segmentation performance when T1ce was missing is an improvement that previous methods have not achieved. Further validation on an external local dataset confirmed that UMMGAT effectively adapts to cross-center data variations. With minimal training data requirements and the ability to generate multi-sequence MRI across multiple centers, UMMGAT provides a practical solution for handling incomplete and heterogeneous MRI data, facilitating more consistent and accurate brain tumor segmentation in diverse clinical contexts.
Similar content being viewed by others
Introduction
Brain tumors severely threaten human health, accounting for over 250,000 reported cases annually1,2. Among malignant forms, glioblastoma emerges as a primary contributor to morbidity and mortality among adult brain tumors, exhibiting an alarming 6.9% 5-year survival rate and contributing to 10,000 annual deaths in the US3. Meningiomas are the most common primary brain tumors, accounting for roughly 30–38% of all cases, and the vast majority are benign4. Despite generally favorable prognoses with observation or surgery, cases with higher grade histology or complex anatomical involvement present ongoing challenges. The significant global burden of brain tumors and their poor survival rates highlight the need for improved diagnostic and therapeutic strategies.
Segmenting brain tumors from multiple MRI sequences is crucial for better diagnosis, treatment planning, monitoring, and clinical trials5. Deep learning (DL)-based models can automatically segment brain tumors on multiple MRI sequences, saving tedious manual work and avoiding user subjectivity6,7,8. However, the widespread adoption of multi-sequence brain tumor segmentation models in clinical practice encounters two major stumbling blocks. First, some MRI sequences are often unavailable due to limited scan time, image artifacts, scan corruption, incorrect machine settings, and allergies to contrast agents9. Most DL-based segmentation models could not handle missing inputs that lead to failure in sequence-missing situations10,11. The second stumbling block is that the MR images acquired at different centers may differ in their characteristics due to differences in manufacturers, acquisition parameters, site procedures, and scanner configurations12,13. A well-trained model may fail when performed on images from novel centers.
To address the sequence missing problem, a common approach is to use the most correlated available sequence to replace the missing one, as also reported in our comparative analysis14. To address the cross-center inconsistency problem, a typical way is to register all the brains to a common brain or a standard space, which is computationally intensive and time-consuming15,16. There are also risks of introducing biases during the registration process, as the chosen standard template may not be equally representative of all subject populations17. Image generation can serve as a unified solution for all the abovementioned problems. Using AI-generated images to substitute missing sequences or simulate the testing images to have consistent shape and distribution with the training images is an intuitive way to enhance the generalizability of the model without altering its structure or retraining its parameters. Generative adversarial networks (GANs) are widely used for medical image synthesis18,19,20. GANs are trained using two neural networks—a generator and a discriminator. The generator learns to create data that resemble examples contained within the training data set, and the discriminator learns to distinguish real examples from the ones created by the generator. The two networks are trained together until the generated examples are indistinguishable from the real examples.
Existing works have explored using GANs as a possible solution for brain MRI image generation21,22,23. However, these methods usually require an amount of aligned and paired data for training, and only synthesize specific types of sequences. This strictly applicable scene limits their use in real clinical settings, where complete multiple MRI sequences are often difficult to obtain. Our original intention was to address the issue of missing data through image generation. However, the paradox lies in the fact that image generation models themselves require complete data for training, which contradicts the practical scenarios of real-world applications. Moreover, in clinical practice, it is often uncertain which sequences are missing or available, leading to complex data gaps involving various missing sequences. Current one-to-one or multiple-to-one image generation models can only handle fixed missing sequences, further limiting their utility.
The novel image generation method developed in this work aims to address the aforementioned issues by incorporating two key techniques: unsupervised learning and multi-task learning. The former enables the image generation model to be trained on incomplete data, while the latter allows for flexible transformation between any sequences. Additionally, to better preserve the lesion region information, we introduced a lesion-aware module (LAM) that enhances the generation of these regions, which often exhibit different features from the rest of the image. Furthermore, while previous studies have largely relied on objective metrics and subjective evaluations by physicians to assess the quality of generated images, there has been insufficient evaluation of their potential for use in DL-based models.
In this study, we realistically simulated the complex multi-center inconsistencies and sequence-missing scenarios found in clinical practice. Under these conditions, we developed an unpaired multi-center multi-sequence generative adversarial transformer (UMMGAT) for image generation, which can be effectively trained when each patient has only one sequence, simulating the most challenging data-missing scenarios in clinical practice. We then used the generated images to complete the missing sequences and align cross-center multi-sequence MRI data. These cross-center consistent and complete multi-sequence data were subsequently used as input for a brain tumor segmentation model (overall pipeline can be seen in Fig. 1). We validated that the effectiveness of generated images across both glioblastomas and meningiomas cohorts, demonstrating consistent improvements in segmentation performance under various sequence-missing and cross-center scenarios. These results demonstrates the proposed pipeline’s robustness, versatility, and applicability in complex clinical settings.
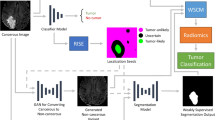
a Schematic representation of using image generation to enhance brain tumor segmentation in real-world scenarios. While segmentation models perform well on complete multi-sequence datasets, their performance deteriorates when faced with incomplete and inconsistent multi-center inputs. To address this, an image generation model synthesizes missing sequences and standardizes multi-center data as input for the segmentation model, thereby improving its performance. The main challenges in designing such an image generation model include training with incomplete data, where supervised image generation fails, necessitating an unsupervised generative AI approach, and multi-sequence image generation, where one-to-one models are inefficient, requiring a multi-task strategy. b Application of our proposed UMMGAT in real-world scenarios for enhancing brain tumor segmentation. In the training dataset for UMMGAT, each patient has exactly one MR image from a single sequence. During each training epoch, UMMGAT is randomly trained on different sequences from different patients. Images are encoded by the ‘Encoder’ into sequence codes (S), and the sequence number (Seq. num.) is mapped to sequence codes (S) by the ‘Mapper.’ The ‘Generator’ then utilizes these sequence codes to generate images, with lesion regions separately encoded to enhance lesion area generation. Testing scenarios involve various missing sequence and cross-center conditions to evaluate UMMGAT’s ability to improve brain tumor segmentation performance. UMMGAT leverages the available sequence image and the missing or different center’s sequence number to generate missing images and align cross-center multi-sequence MRI data. The resulting multi-center consistent and complete multi-sequence data are subsequently used as input for a well-trained brain tumor segmentation model. c Structure of the Generator. The generator is based on the Swin-Unet architecture, which effectively captures multi-scale features through its U-Net structure while leveraging a transformer-based architecture to model long-range dependencies and global context. Adaptive Instance Normalization (AdaIN) is incorporated to integrate the sequence code into the model.
Results
Patient characteristics
The BraTS2019 dataset comprises multi-institutional pre-operative MRI scans from 335 patients, and the BraTS2023-MEN dataset includes 1000 patients. Both datasets provide four MRI sequences: T1-weighted (T1), contrast-enhanced T1-weighted (T1ce), T2-weighted (T2), and T2-weighted fluid-attenuated inversion recovery (FLAIR). The local dataset consists of T1- and T2-weighted sequences from 91 patients with a median age of 54 years (range: 24 to 83 years), with 49 male (53.85%) and 42 female patients (46.15%). The UCSF-PDGM dataset consists of preoperative MRI scans from 494 patients, each containing the aforementioned four sequences in addition to susceptibility-weighted imaging (SWI), diffusion-weighted imaging (DWI), 3D arterial spin labeling (ASL) perfusion, and 2D 55-direction high angular resolution diffusion imaging (HARDI).
UMMGAT’s capability to encode sequence features from unpaired datasets
The key to UMMGAT’s ability to train using an unpaired dataset lies in its use of a sequence encoder to extract sequence codes, which ensure disentangled and significant encoding of modality-specific characteristics in the absence of supervision. Figure 2 shows the UMAP visualization of the extracted sequence codes. As observed, the sequence codes can well distinguish different sequences, while they show no clear differentiation between the generated images and the original images. Moreover, the style codes of generated images do not cluster by source sequence, indicating that the sequence encoder effectively performs cross-modality style transformation. In addition, the sequence encoder independently captures lesion-specific features, which are further emphasized by the LAM to enhance lesion synthesis.
Each point represents a style code of an MRI image in the UCSF-PDGM test cohort. a Colored by target sequence. b Colored by code type (blue = real, red = generated). c Colored by source sequence. d UMAP of target sequence codes including lesion region sequence codes.
Multi-sequence multi-center image generation results
UMMGAT can generate synthetic MR images respect to a specified target sequence by inputting an original image and the target sequence number. Figure 3 shows the generated images of each MRI sequence. The generated MR images demonstrate high overall quality and faithfully preserve tumor-related features, with lesion boundaries, enhancement patterns, and peritumoral edema closely matching real scans. Incorporation of the LAM (Supplementary Fig. 2 shows example generated images with and without LAM) further improved lesion synthesis, particularly enhancing the depiction of peritumoral edema and capturing tumor heterogeneity. Supplementary Fig. 6 shows axial, coronal, and sagittal stacks of generated MRI sequences for a single case, illustrating that the generated sequences retain the contour and anatomical structure of the brain. Quantitative evaluation using FID (Fig. 4) further confirmed the effectiveness of UMMGAT. Baseline FID values between original modalities reflected inherent inter-sequence variation (mean = 542.21 ± 310.09). Incorporating LAM (mean = 258.21 ± 129.04) consistently reduced FID scores across modalities compared with the without-LAM setting (mean = 310.96 ± 166.15). Qualitative assessment by multiple experienced clinicians supported these findings: most generated images clearly displayed tumor heterogeneity and provided sharp demarcation between tumor regions and normal brain tissue. In particular, when generating sequences from those in which edema is less visible (e.g., T1 and T1ce), the synthesized images revealed the perilesional region more clearly and even highlighted vascular patterns around the lesion (red boxes in Supplementary Fig. 2), providing additional diagnostic information. Nevertheless, occasional limitations were observed, including false enhancement or absence of expected enhancement within the tumor core (blue boxes in Supplementary Fig. 2).
The first columns display the input images while the remaining columns show the images generated by our proposed UMMGAT. Each row corresponds to images generated from the leftmost input image into the target sequence indicated on the vertical axis.
Shown are comparisons of (left) real-to-real comparison, (middle) real images versus generated images without the lesion-aware module (LAM), and (right) real images versus generated images with LAM. Lower FID values indicate higher similarity. Lower FID values indicate higher similarity. The optimal FID score is 0.0, signifying that the two sets of images are identical (the values along the diagonal from the bottom left to the top right are 0). Friedman test indicated significant differences among the three FID datasets (χ² = 120.82, p < 0.001). Dunn’s post hoc tests with Bonferroni correction showed significant pairwise differences: without LAM vs. real (−231.25, Cohen’s d = −1.07, p < 0.001), with LAM vs. real (−283.99, Cohen’s d = −1.31, p < 0.001), and with LAM vs. without LAM ( − 52.74, Cohen’s d = −0.24, p < 0.001).
Quantitative evaluation of segmentations under various sequence-missing and cross-center scenarios
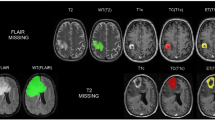
We validate the segmentation results of using generated and copied images to replace the missing ones for brain tumor segmentation. We first evaluated glioblastoma segmentation in the BraTS dataset. As shown in Fig. 5, visualized segmentation masks indicate that replacing missing sequences with generated images results in more accurate tumor segmentations. Specifically, using copied images often overestimates the extent of the whole tumor (as seen in the scenarios of missing T2, missing Flair, missing (T2 and Flair), and missing (Flair, T1ce, and T2)), whereas using generated images provides more accurate segmentation of the whole tumor. Additionally, using generated images better identifies heterogeneous tumor components compared to copied images. For example, when T2 is missing, using copied images tends to classify all regions as the necrotic tumor core (red), and when (T1 and T1ce) are missing, using copied images tends to classify all regions as the enhancing tumor (blue). From Table 1 and the scatter plots in Supplementary Fig. 4, the median DSCs are significantly improved by using generated images compared with copied images in most scenarios. Specifically, for single sequence missing, generating missing T1 and Flair from T2, or generating missing T2 and T1ce from T1 significantly improves the DSCs of WT, TC, and ET, compared with using copied T2 or T1. Generating T1, T2 from each other achieves comparable segmentation results in WT, TC, and ET with using complete sequences (missing T1: 0.905, 0.822, 0.797; missing T2: 0.865, 0.759, 0.781; complete data: 0.895, 0.811, 0.790). Generating Flair from T2 restores the segmentation of TC and ET to 0.683 and 0.775, respectively. Also, generating T1ce from T1 achieves a WT segmentation performance of 0.894, almost identical to complete data. When two or more sequences are missing, the copying strategy fails to restore the decreased segmentation performance. However, using generated images for WT segmentation in scenarios of missing (T1 and T1ce), (T1 and T2), (T2 and T1ce), and (T1, T1ce, and T2) achieves comparable results with complete data (0.897, 0.850, 0.854, 0.844). Using generated images for TC and ET segmentation in cases of missing (T2 and Flair), (T1 and T2), (T1 and Flair, and T1), (T2, and Flair) still yields acceptable results (0.659 and 0.718, 0.796 and 0.76, 0.673 and 0.769, 0.63 and 0.667).
The red area indicates the necrotic tumor core (NCR); the green area indicates the peritumoral edematous/invaded tissue (ED); the blue area indicates the enhancing tumor (ET); the whole tumor region (WT) includes all tumor areas (NCR + ED + ET); the tumor core region (TC) includes NCR and ET.
We further assessed the impact of synthetic images in single-modality-missing scenarios using the UCSF-PDGM dataset, which includes additional perfusion and diffusion sequences (Table 2). Incorporating of these modalities improved the model’s adaptability to missing inputs, highlighting the complementary value of multi-sequence information. However, segmentation performance remained difficult to recover when T1ce was missing, indicating the critical role of this modality. Among the newly included sequences, the absence of HARDI had a pronounced negative effect on glioblastoma segmentation, indicating that it conveys unique and indispensable tumor-related information.
To further evaluate the generalizability of our pipeline across tumor types, we applied the trained UMMGAT to meningioma segmentation using the BraTS-MEN dataset (Fig. 6 and Table 3). Similar improvements were observed under sequence-missing conditions, confirming that the generated images effectively supported segmentation even for this distinct tumor entity. Interestingly, although meningioma segmentation is generally considered computationally simpler with high Dice scores when all sequences are available—it was more susceptible to missing modalities. This likely reflects the lower-grade tumors less distinctive features in individual sequences, thereby relying more heavily on the complementary information provided by multiple sequences.
Color coding is the same as in Fig. 5: red = NCR, green = ED, blue = ET, WT = NCR + ED + ET, TC = NCR + ET.
Discussion
Generative AI has garnered significant enthusiasm, yet its application in medical imaging necessitates careful consideration and comprehensive evaluation, particularly for patient-facing tasks24. Currently, the assessment of generated images is often based on subjective evaluations by physicians25. These studies aim to provide a pattern which we term “Generative - Doctor”, where the generative method aims to provide images directly to doctors for diagnostic purposes. However, translating such laboratory research into clinical practice poses significant challenges because of the potential risks associated with inevitable errors in the generated images as inacceptable in clinical settings. In contrast, we explore a “Generative - AI - Doctor” pattern, where generative methods support other AI models, which are already integral to clinical diagnostics but often face obstacles due to high data completeness and consistency demands. Our work demonstrates generative AI can address the issues of data missing and inconsistency, thereby enhancing the performance and generalizability of AI-empowered models in real clinical scenarios.
We propose a unified pipeline to extend multi-sequence brain tumor segmentation models to “imperfect” datasets characterized by sequence missing and inter-center inconsistencies. The key development is the proposed image generation model, UMMGAT, which can be trained on unpaired, incomplete multi-center multi-sequence MRI data to generate images for any center and any sequence. Our method showed to improve the robustness and applicability of AI-empowered models in clinical practice by leveraging generative AI to overcome the limitations posed by incomplete and inconsistent data. This provides a promising avenue for reducing the need for multiple scans in clinical practice.
Compared with previous works, we have significantly expanded both the application scenarios and the technological approaches. In terms of scenario settings, our primary contribution lies in using a unified image generation solution to simultaneously address sequence missing and cross-center data inconsistency. Conte et al. attempted to enhance brain tumor segmentation models by replacing missing sequences with generated ones, but their study was limited to scenarios with missing T1 and FLAIR sequences22. Technically, they employed a one-to-one synthesis approach. Therefore, it requires to train n2-1 models to achieve mutual generation among n sequences. To avoid this model complexity, our method employs a multi-task learning strategy, enabling mutual generation among any number of sequences with a unified model. Recently, Sharma et al. employed a unified model to generate any missing sequence, but they still relied on paired data for training23. They acknowledged the limitations of their work, such as the need for image registration for multi-contrast inputs, which is time-consuming, and the necessity for a multi-center evaluation to assess the model’s generalizability across different sites, scanners, and clinical settings. Our work effectively overcomes the limitations mentioned in their study.
Our results suggest that in AI-assisted clinical settings, reducing the scanning of several sequences for brain tumor diagnosis can significantly save time and cost without compromising diagnostic accuracy. T1 and T2 are fundamental MRI sequences with distinct clinical values. Our results concluded that T1 and T2 can be synthesized from other contrasts, which is consistent with Lee et al.‘s work26. Moreover, Flair and T1ce sequences are known to provide clearer ROI information but also require more time and resources, which may not be available in some hospitals. In our experiments, model with absence of advanced sequence, such as T1ce and Flair sequences resulted in a significant performance decrease. However, we discovered that Flair can be well-synthesized from other contrasts with maintainable performance. This finding has potential clinical implications, as it suggests that T2 sequences may be used as a substitute for Flair in situations where time, resources, or equipment are limited, and thus can help mitigate the loss of diagnostic information caused by the absence of Flair in brain tumor imaging. Previous studies have shown that the missing T1ce sequence greatly impacts the segmentation effect, and none of the previous studies succeeded in improving segmentation by generating images22,26. This is likely because the contrast agent used in T1ce sequences provides unique information that is challenging to replicate through synthetic methods alone. However, in our study, generating missing T1ce sequence from T1 also improves the segmentation performance of WT. This suggests the feasibility of utilizing low-dose imaging. We further extended our framework to generate advanced functional and structural MRI sequences, including arterial spin labeling (ASL), diffusion-weighted imaging (DWI), high angular resolution diffusion imaging (HARDI), and susceptibility-weighted imaging (SWI), all of which yielded promising results. Each sequence provide unique clinical insights: ASL provides quantitative information on cerebral blood flow, DWI and HARDI capture microstructural and diffusion anisotropy features that inform tumor infiltration and cellularity, and SWI highlights venous structures and microhemorrhages. Notably, the absence of HARDI had a pronounced negative effect on glioblastoma segmentation, suggesting its indispensable role in conveying tumor-related information; our model was able to partially mitigate this deficit. Collectively, these findings highlight the potential of UMMGAT not only to overcome the limitations of incomplete clinical datasets but also to expand access to advanced imaging biomarkers in centers constrained by scanning time, equipment availability, or patient tolerance.
Data from a single center is vulnerable to biases due to small sample sizes, which result in conflicting or inconclusive conclusions and are insufficient for training an effective DL model. Transferring a model trained on large datasets to local center applications is a practical and effective solution. While the trained DL-based methods are expected to work on data with identical or similar distributions, image generation techniques can adapt the input data to the same distribution. Yan et al. evaluated the generalizability of a DL-based cardiac segmentation model to MRI data from scanners of different manufacturers and trained a GAN to adapt the input image to improve the segmentation model13. Here, we expanded the above work by treating images from different centers as different sequences, thereby unifying cross-sequence and cross-center MRI image generation.
We focused solely on adapting the input data to handle missing data and cross center data inconsistencies, without changing the model architecture or retrain the model, which may potentially yield better results. However, the original training data for the well-trained model is unavailable in local centers, which motivated us to develop the method. Although not the primary focus of this work, we additionally evaluated several more recent generator architectures, including ConvNeXt-Unet27 and Attention-Gated U-Net28 (Supplementary Table 2 and Supplementary Table 3), which achieved FID scores comparable to or even better than Swin-Unet, suggesting that our framework can readily benefit from advances in backbone design. Given the modular design of our framework, future backbones with superior representational capacity can be seamlessly integrated into the generator. Moreover, with the scalability of UMMGAT to additional sequences, future work could extend image generation to additional sequences, such as DTI, DSA, and CTA. We also plan to explore the value of using generated images in other DL models, such as those for diagnosing IDH1 and MGMT.
In conclusion, we validated that using generated images can enhance brain tumor segmentation models in sequence-missing and cross-center scenarios. We proposed a novel unsupervised image generation model, namely UMMGAT, which can be trained with unpaired, incomplete data, making it highly applicable in real clinical settings. Future research should explore integrating UMMGAT with other DL models to further improve their robustness and accuracy.
Methods
Data setup and preprocessing
For this retrospective study, we used both publicly available MRI scans and self-collected MRI scans from our institution. For glioblastoma (GBM) dataset, the inclusion criteria were a histologic diagnosis of GBM. The public data set was the Multimodal Brain Tumor Image Segmentation Benchmark (BRATS) 2019 data set, which comprises 335 subjects from 19 institutions29,30,31. Each subject includes T1-weighted (T1), contrast-enhanced T1-weighted (T1ce), T2-weighted (T2), and T2w–fluid-attenuated-inversion-recovery (FLAIR) images obtained at multiple institutions. In addition, we used the UCSF-PDGM dataset32, which initially included 501 subjects; after excluding incomplete cases, 494 subjects were retained. Each subject includes T1, T1ce, T2, FLAIR, SWI, DWI, ASL, and HARDI. The local dataset included 92 patients who were admitted to Nanjing Drum Tower Hospital. All patients were scanned with T1-weighted and T2-weighted MRI sequences. The whole tumor was manually annotated by two experienced radiologists. All procedures performed were in accordance with the ethical standards of the Declaration of Helsinki. The use of local data was reviewed and approved by the Institutional Ethics Committee of Nanjing Drum Tower Hospital. Furthermore, informed consent to participate in the study was obtained from all individual participants. For meningioma, we employed the Brain Tumor Segmentation (BraTS) 2023 Meningioma Challenge training dataset33, which contains 1000 patients, each with T1, T1ce, T2, and FLAIR sequences. Cases were identified either by histopathological confirmation following resection or biopsy or by formal clinical and radiographic diagnosis of meningioma, commonly classified under the International Classification of Diseases, 10th Revision (ICD-10) code D32.9 (“benign neoplasm of the meninges”). For BraTS, BraTS-MEN and UCSF-PDGM dataset, all scans are resampled to 1 mm3 isotropic resolution using a linear interpolator, skull stripped, and co-registered with a single anatomical template using a rigid registration model with mutual information similarity metric. All the imaging datasets have been segmented manually by experienced neuro-radiologists. Annotations comprise the enhancing tumor (ET - label 4), the peritumoral edematous/invaded tissue (ED - label 2), and the necrotic tumor core (NCR - label 1). The whole tumor region (WT) includes all tumor areas (label1 + label2 + label4). The tumor core region (TC) is a combination of NCR (label1) and ET (label4). We normalize the values of all series to (0, 255) and crop them to (155, 176, 176) to crop out the brain region from each sequence.
Overall pipeline: leveraging unsupervised generative AI to bridge real-world data gaps in segmentation
In real clinical settings, data distribution often consists of multi-center, inconsistent, and incomplete datasets. Many deep learning (DL)-based models fail to perform effectively under these conditions. To address this, we propose a pipeline that leverages an unsupervised generative model to transform the multi-center inconsistent and incomplete dataset into a consistent and complete dataset, which can then be seamlessly utilized by downstream segmentation models (Fig. 1).
Training and validation of UMMGAT
We developed an UMMGAT to generate MRI sequences. UMMGAT can be trained using these multi-center inconsistent and incomplete datasets through an unsupervised learning strategy and enables the simultaneous generation of various MRI sequences using a multi-task learning strategy. We simulated a challenging multi-center, incomplete, and inconsistent dataset distribution, where each patient contributed exactly one MR image from a single sequence. We trained UMMGAT for image generation using the BraTS 2019 dataset and our local dataset, covering six MRI sequences (T1, T2, FLAIR, and T1ce from BraTS2019, together with T1 and T2 from the local dataset). For internal validation, the combined dataset was randomly split into training and testing subsets at a 4:1 ratio, with the results primarily presented in the Supplementary Materials. Specifically, Supplementary Fig. 3 illustrates the UMAP visualization of sequence codes, Supplementary Fig. 4 reports the FID scores, and Supplementary Figs. 5, 6 provide representative generated images. External evaluation was performed on the BraTS-MEN dataset. In a separate experiment, we trained UMMGAT with the UCSF-PDGM and local datasets across ten MRI sequences, including advanced modalities such as ASL, DWI, SWI, and HARDI. Internal validation again followed a 4:1 train–test split. Given the greater modality diversity of this dataset, the corresponding generation results are presented in the main text, including the UMAP visualization of sequence codes (Fig. 2), representative generated images (Fig. 3), and FID scores (Fig. 4). Thus, results from the six-sequence setting are reported in the Supplementary Materials to provide methodological completeness, while results from the more diverse ten-sequence setting are highlighted in the main text to better demonstrate the robustness and generalizability of UMMGAT.
UMMGAT unsupervised multi-task learning strategies
UMMGAT employed a multi-task learning strategy to simultaneously learn image generation between any two sequences. In each training epoch, UMMGAT was randomly trained on different sequences from different patients. After multiple epochs, UMMGAT successfully generated images for any given sequence. The Sequence Encoder (E), Mapper (M), and Discriminator (D) each contain multiple branches, allowing a single model to generate images across various sequences from different centers. More structural details are available in Supplementary Fig. 1. Inspired by StarGAN-v2, sequence codes were encoded for unpaired unsupervised learning19. The lesion regions were separately encoded to enhance the generation of the lesion region. For visualizing sequence codes, we employed Uniform Manifold Approximation and Projection (UMAP) to create a low-dimensional representation of their distribution34.
We use multiple loss functions to train our framework, ensuring the generated image not only matches the style of the target MRI sequence but also retains the original image’s content. The adversarial loss (Loss_adv) guides the generator to create images resembling MRI images while the discriminator distinguishes them from real ones. The style reconstruction loss (Loss_sty) enables the sequence encoder and mapping network to extract representative codes. The cycle consistency loss (Loss_cyc) ensures the generated image preserves the domain-invariant characteristics of the input. The generator was built on Swin-Unet35. Its U-net structure effectively captures multi-scale features, while the transformer-based architecture captures long-range dependencies and global context. Adaptive Instance Normalization (AdaIN) was used to insert the sequence code36.
The batch size is set to 8 for all experiments, and the model is trained for 20,000 iterations, which cost about half a day on a single Tesla V100 GPU with our implementation in PyTorch. We adopt the non-saturating adversarial loss with R1 regularization using γ = 1. All models are trained using Adam with β1 = 0, β2 = 0.99, and weight-decay = 10−4. The learning rates are set to 10−4. For data augmentation, we flip the images horizontally with a probability of 0.5. For evaluation, we employ exponential moving averages over the parameters of all modules except D. We initialize the weights of all modules using He initialization and set all biases to zero.
Lesion-aware module (LAM)
During training, lesion masks were provided to delineate the lesion regions from both the reference and generated images. In our framework, the sequence encoder treats the lesion region as an additional modality (e.g., in a 10-sequence UMMGAT, FLAIR is considered domain 1, while FLAIR_lesion is considered domain 11). This design allows the sequence encoder to extract lesion-specific codes through a dedicated style reconstruction loss for lesions (Loss_sty_lesion). Importantly, this loss not only enforces the encoder to disentangle discriminative lesion features but also provides supervision for the generator, ensuring that the synthesized lesion regions conform to the modality-specific lesion characteristics. Notably, we did not adopt a naive approach of feeding the lesion and non-lesion regions separately into the generator and then fusing them, as this resulted in overfitting to tumor areas and produced unrealistic, sharp boundaries between lesions and surrounding tissue. Instead, the generator always receives the full-brain image as input, while lesion regions are extracted only for additional encoding in the sequence encoder. If the lesion region of the generated image is inconsistent with the expected modality-specific pattern, the sequence encoder penalizes the generator through the loss function, thereby guiding it to iteratively improve lesion synthesis.
The effectiveness of LAM was further validated in comparative experiments with and without the lesion-specific branch. As illustrated in Supplementary Fig. 2, the incorporation of LAM enhanced lesion representation, with red arrows highlighting examples of improved lesion boundaries and heterogeneity.
Evaluating the Impact of Generated Images on Segmentation
We next applied the generated images to a brain tumor segmentation model to assess their impact under then following realistic conditions: (a). Various sequence-missing scenarios, including fixed missing sequences of one (4 scenarios), two (6 scenarios), or three sequences (4 scenarios), as well as randomly missing sequences. For random sequence missing, we ensured that each patient had at least one kind of MRI sequence and at most three kinds of MRI sequences, resulting in one, two, or three MRI sequences being missing for each patient. The specific distribution of randomly deleted sequences is detailed in Supplementary Table 1b). A cross-center scenario, where a brain tumor segmentation model was applied to local T2 images for whole tumor (WT) segmentation. UMMGAT uses the available sequence image and the missing or cross-center sequence number to generate the missing images and align multi-center multi-sequence MRI data. These resulting consistent and complete multicenter multi-sequence data were then used as input for a well-trained brain tumor segmentation model. This strategy was compared with a method of copying the most correlated images to assess its effectiveness in enhancing the model’s performance. Considering the easy accessibility of T1 and T2 in clinical practice, these sequences were preferentially used to synthesize missing modalities. The framework was validated across multiple brain tumor types, including segmentation of glioblastomas in the BraTS dataset and meningiomas in the BraTS-MEN dataset, demonstrating its generalizability across distinct tumor entities.
Metrics
We evaluated UMMGAT-generated images using Fréchet Inception Distance (FID), which measures the distance between feature vectors of real and generated images, extracted using a trained Inception v3 model. A FID of 0.0 indicates identical image sets.
We used the dice similarity coefficient (DSC) as the metric of the segmentation effect. \({Y}_{\mathrm{gt}}\) indicate manual annotation and \({Y}_{\mathrm{pred}}\) indicate the prediction of the segmentation model in the scenarios where we simulated missing MRI sequences. The DSC ranged from 0 (no overlap) to 1(perfect overlap).
Statistics were computed with GraphPad Prism 10 software. All analyses employed nonparametric tests. The Friedman test (>3 groups) or Wilcoxon signed-rank test (2 groups) was applied for comparisons, with Dunn’s post hoc test used for multiple comparison correction. P < 0.05 indicated a statistically significant difference.
Data availability
The system was developed using standard libraries and scripts available in PyTorch. The full code, including the training code, test code, and local data used for training, are available from the corresponding author upon reasonable request. The BraTS2019 dataset can be downloaded from https://www.med.upenn.edu/cbica/brats-2019/. The UCSF-PDGM dataset can be obtained from https://www.cancerimagingarchive.net/collection/ucsf-pdgm/. The BraTS2023-MEN dataset is available from https://www.synapse.org/Synapse:syn51514106.
References
Khalighi, S. et al. Burden and trends of brain and central nervous system cancer from 1990 to 2019 at the global, regional, and country levels. Arch. Public Health 80, 209 (2022).
Kuang, Z. et al. Global Disease Burden, Trends, and Inequalities of Brain and Central Nervous System Cancers, 1990–2021: a Population-Based Study with Projections to 2036. World Neurosurg. 198, 123970 (2025).
Fekete, B. et al. What predicts survival in glioblastoma? A population-based study of changes in clinical management and outcome. Front. Surg. 10, 1249366 (2022).
Ogasawara, C., Philbrick, B. D. & Adamson, D. C. Meningioma: a review of epidemiology, pathology, diagnosis, treatment, and future directions. Biomedicines 9, 319 (2021).
Huang, Z., Lin, L., Cheng, P., Peng, L. & Tang, X. Automated brain tumor segmentation using multimodal brain scans: a survey based on models submitted to the BraTS 2012–2018 challenges. IEEE Rev. Biomed. Eng. 13, 156–168 (2020).
Kamnitsas, K. et al. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 36, 61–78 (2017).
Pereira, S., Pinto, A., Alves, V. & Silva, C. A. Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans. Med. imaging 35, 1240–1251 (2016).
Havaei, M. et al. Brain tumor segmentation with deep neural networks. Med. Image Anal. 35, 18–31 (2017).
Zhou, T. et al. A literature survey of MR-based brain tumor segmentation with missing modalities. Comput. Med. Imaging Graph. 104, 102167 (2023).
Zhou, T., Canu, S., Vera, P. & Ruan, S. Latent correlation representation learning for brain tumor segmentation with missing MRI Modalities. IEEE Trans. Image Process 30, 4263–4274 (2021).
Yang, Q., Guo, X., Chen, Z., Woo, P. Y. M. & Yuan, Y. D2-Net: Dual disentanglement network for brain tumor segmentation with missing modalities. IEEE Trans. Med. Imaging 41, 2953–2964 (2022).
Dewey, B. E. et al. A disentangled latent space for cross-site MRI Harmonization. in Medical Image Computing and Computer Assisted Intervention – MICCAI 2020 (eds Martel A. L., Abolmaesumi P., Stoyanov D., et al.) (Springer International Publishing, 2020).
Yan, W. et al. MRI manufacturer shift and adaptation: increasing the generalizability of deep learning segmentation for MR images acquired with different scanners. Radiol. Artif. Intell. 2, e190195 (2020).
Zhou, T., Canu, S., Vera, P., Ruan, S. Brain tumor segmentation with missing modalities via latent multi-source correlation representation. in Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference (Springer, 2020).
Gholipour, A., Kehtarnavaz, N., Briggs, R., Devous, M. & Gopinath, K. Brain functional localization: a survey of image registration techniques. NeuroImage 54, 313–327 (2011).
Evans, A. C., Janke, A. L., Collins, D. L. & Baillet, S. Brain templates and atlases. NeuroImage 54, 313–327 (2011).
Trottet, C. et al. The problem of functional localization in the human brain. NeuroImage 3, 313–327 (2011).
Zhu, J.-Y., Park, T., Isola, P., Efros, A. A. Unpaired Image-to-Image Translation using Cycle-Consistent Adversarial Networks. 2020; published online Aug 24. http://arxiv.org/abs/1703.10593 (accessed 10 January 2023).
Choi, Y., Uh, Y., Yoo, J., Ha, J.-W. StarGAN v2: diverse Image synthesis for multiple domains. 2020; published online April 26. http://arxiv.org/abs/1912.01865 (accessed Jan 10, 2023).
Xie G., et al. Cross-Modality Neuroimage Synthesis: A Survey. 2022; published online Dec 15. http://arxiv.org/abs/2202.06997 (accessed 9 April 2023).
Tavse, S., Varadarajan, V., Bachute, M., Gite, S. & Kotecha, K. A systematic literature review on applications of GAN-Synthesized images for brain MRI. Future Internet 14, 351 (2022).
Conte, G. M. et al. Generative adversarial networks to synthesize missing T1 and FLAIR MRI sequences for use in a multisequence brain tumor segmentation model. Radiology 299, 313–323 (2021).
Liu, J. et al. One model to synthesize them all: multi-contrast multi-scale transformer for missing data imputation. IEEE Transac. Med. Imaging 42, 2577–2591 (2023).
Wachter, R. M. & Brynjolfsson, E. Will generative artificial intelligence deliver on its promise in health care?. JAMA 331, 65 (2024).
Jeong, J. J. et al. Systematic Review of Generative Adversarial Networks (GANs) for medical image classification and segmentation. J. Digit. Imaging 35, 137–152 (2022).
Lee, D., Moon, W.-J., Ye, J. C. Which contrast does matter? towards a deep understanding of MR contrast using collaborative GAN. ArXiv 2019; published online May 10. https://www.semanticscholar.org/paper/Which-Contrast-Does-Matter-Towards-a-Deep-of-MR-GAN-Lee-Moon/96f826aec079bd283a93ae5aa1cec942d0ef2697 (accessed 10 June 2024).
Han, Z., Jian, M. & Wang, G.-G. ConvUNeXt: an efficient convolution neural network for medical image segmentation. Knowl.Based Syst. 253, 109512 (2022).
Masse-Gignac, N., Flórez-Jiménez, S., Mac-Thiong, J. & Duong, L. Attention-gated U-Net networks for simultaneous axial/sagittal planes segmentation of injured spinal cords. J. Appl. Clin. Med. Phys. 24, e14123 (2023).
Bakas, S. et al. Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. Preprint at https://doi.org/10.48550/arXiv.1811.02629 (2018).
Bakas, S. et al. Advancing the cancer genome atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci. Data 4, 1–13 (2017).
Menze, B. H. et al. The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. imaging 34, 1993–2024 (2014).
Calabrese, E. et al. The University of California San Francisco Preoperative Diffuse Glioma MRI (UCSF-PDGM). https://doi.org/10.7937/tcia.bdgf-8v37.
LaBella, D. et al. The ASNR-MICCAI Brain Tumor Segmentation (BraTS) Challenge 2023: Intracranial Meningioma. 2023; published online May 12. https://doi.org/10.48550/arXiv.2305.07642.
McInnes, L., Healy, J., Melville, J. Umap: Uniform manifold approximation and projection for dimension reduction. Preprint athttps://doi.org/10.48550/arXiv.1802.03426 (2018).
Cao, H. et al. Swin-Unet: Unet-Like Pure Transformer for Medical Image Segmentation. in Lecture Notes in Computer Science (eds Karlinsky L., Michaeli T., Nishino K) (Springer Nature Switzerland, 2023).
Huang, X. & Belongie, S. Arbitrary style transfer in real-time with adaptive instance normalization. In Proc. IEEE international conference on computer vision 1501–1510 (IEEE, 2017).
Acknowledgements
This work was supported in part by the National Nature Science Foundation of China under Grant No. 62476122,62106101. This work was also supported in part by the Natural Science Foundation of Jiangsu Province under Grant No. BK20210180. This work is also partly supported by the AI \& AI for Science Project of Nanjing University.
Author information
Authors and Affiliations
Contributions
Z.L., K.H., and C.H. designed the research. W.L., B.Z., and X.Z. collected and annotated the data. Z.L. and K.H. developed and tested the Unsupervised Generative AI. Z.L. and K.H. co-wrote the manuscript. K.H., W.L., T.Z., B.Z., and C.H. critically revised the manuscript, and all authors discussed the results and provided feedback on the manuscript. All authors had final responsibility for the decision to submit for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Z., Zhou, T., Zhang, B. et al. Unsupervised generative AI for enhancing brain tumor segmentation in multi-center, incomplete real-world data scenarios. npj Precis. Onc. 9, 384 (2025). https://doi.org/10.1038/s41698-025-01173-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01173-4