Abstract
Multiple myeloma (MM) is a hematologic malignancy characterized by monoclonal proliferation of plasma cells. As most patients with MM relapse after multiple lines of therapy, seeking new treatment strategies is an urgent task for current research. We found succinate dehydrogenase subunit A (SDHA) was closely related to prognosis of MM patients. SDHA promoted the tricarboxylic acid (TCA) cycle and inhibited glycolysis. It subsequently repressed the proliferation and invasion of MM both in vitro and in vivo. Endothelial pAS domain protein1 (EPAS1), also known as hypoxia-inducible factor-2α (HIF-2α), is a key transcription factor in response to hypoxia. We found that patients with high EPAS1 expression had a better prognosis. EPAS1 promoted the TCA cycle, suppressed glycolysis, and inhibited the proliferation and invasion of MM both in vitro and in vivo. EPAS1 enhanced SDHA expression by inhibiting HDAC2 mRNA expression and increasing acetylation at the SDHA histone H3K27 site. In conclusion, activating the EPAS1-HDAC2-SDHA axis could restore the TCA cycle as the dominant glucose metabolism pathway in order to inhibit proliferation of MM cells.
Similar content being viewed by others
Introduction
Multiple myeloma (MM) is the most common plasma cell malignant tumor. It is the second most common malignancy in hematological diseases, accounting for about 10% of all hematological malignant tumors1. MM patients often present with the following clinical symptoms, hypercalcemia, renal insufficiency, anemia, and bone destruction or pathological fractures. With the continuous emergence of new drugs, the prognosis of newly diagnosed MM patients has been significantly improved2. However, most MM patients will relapse even after completing autologous hematopoietic stem cell transplantation3. Existing chemotherapy drugs will fail, and patients will eventually die due to tumor progression or treatment-related complications. Therefore, seeking new effective treatment strategies is the top priority for MM research.
Succinate dehydrogenase (SDH) is a critical enzyme in the tricarboxylic acid (TCA) cycle, linking oxidative phosphorylation to the electron transport chain (ETC)4. SDH is a complex composed of four subunits: SDHA, SDHB, SDHC, and SDHD, with SDHA serving as the catalytic core5. The SDH gene is the first identified nuclear-encoded mitochondrial tumor suppressor, and its deletion mutation is implicated in the onset and progression of various tumors, including hereditary paraganglioma/pheochromocytoma syndrome, gastrointestinal stromal tumors, thyroid tumors, testicular seminoma, neuroblastoma, ovarian cancer, pituitary adenoma, and breast cancer6,7,8,9. SDH deficiency disrupts the TCA cycle, leading to creating a pseudohypoxic environment within cells10. Under hypoxic conditions, hypoxia inducible factor 1α (HIF1α) is inadequately hydroxylated and degraded, resulting in the accumulation of reactive oxygen species (ROS), which ultimately promotes tumor proliferation and metastasis11. Our previous research identified low expression of SDHA in MM patients, with high SDHA expression serving as an independent favorable prognostic factor12. Furthermore, we discovered that chidamide, an effective histone deacetylase (HDAC) inhibitor, regulated the proliferation and invasion of MM cells by upregulating SDHA expression13. However, specific mechanisms of SDHA in the pathogenesis and progression of MM remain unclear.
In normal physiology, Endothelial pAS domain protein1 (EPAS1, also known as HIF-2α) orchestrates systemic oxygen homeostasis14: it drives hypoxia-inducible erythropoietin expression to regulate erythropoiesis15, supports carotid-body oxygen sensing16, and contributes to vascular and sympathoadrenal development17. In the hematopoietic system, EPAS1 primarily acts through the bone-marrow microenvironment to sustain effective hematopoiesis rather than being intrinsically required for long-term HSC maintenance18. These physiological roles, together with human genetic evidence linking EPAS1 to high-altitude hemoglobin phenotypes, situate EPAS1 as a canonical regulator of oxygen-dependent tissue function in normal cells and niches.
Proliferation of tumor cells is highly dependent on the tumor microenvironment. Tumor microenvironment of MM protects MM cells from being killed by chemotherapy drugs through direct contact, secretion of cytokines, and production of different metabolites19. Metabolic changes of tumor cells are one of the most important characteristics of the tumor microenvironment. Many tumor cells rely on glycolysis instead of the TCA cycle for cellular energy under normal oxygen conditions, which is a metabolic phenomenon known as aerobic glycolysis or Warburg effect20. The dominant glycolysis increases the efficiency of glucose uptake and substances production for tumor cells, which causes rapid proliferation of tumor cells. This study aimed to discover the impact of EPAS1 and SDHA deficiency on MM metabolism and explore its underlying mechanisms.
Results
Enhancing expression of SDHA in MM cells restored the TCA cycle and reduced level of glycolysis in vitro and in vivo
To explore the effect of SDHA expression on MM cells, we first established SDHA overexpression cell lines. Relative expression of SDHA was shown in Supplementary Fig. 1A, we chose two cell lines with the lowest SDHA expression, H929 and AMO-1 to construct stable transgenic MM cell lines overexpressing SDHA (SDHA-OE, Supplementary Fig. 1B, C).
Compared with normal control, SDHA overexpression inhibited cell proliferation in H929 (Fig. 1A, left panel, P = 0.0537 and P = 0.0021, respectively, cultured after 24 h and 48 h) and AMO-1 (Fig. 1A, right panel, P = 0.0398 and P = 0.0008 respectively cultured after 24 h and 48 h) cell lines. As revealed by transwell invasion assay, compared with ncH929 cells, SDHA overexpression cells had a lower invasion ability (Supplementary Fig. 1D, B P = 0.0340 and P = 0.0004, respectively for 24 h and 48 h). In AMO-1 cell line, compared with normal control group, overexpression of SDHA showed a decreasing trend in cell invasion (Fig. 1C, P = 0.0808). Above results revealed that SDHA overexpression inhibited proliferation and invasion of MM cells.
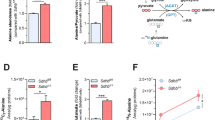
A Left panel showed H929 cell proliferation curves of SDHA-NC and SDHA-OE. After 24 h and 48 h, cell counts of SDHA-OE were significantly lower than SDHA-NC; Right panel showed AMO-1 cell proliferation curves of SDHA-NC and SDHA-OE. After 48 h, cell counts of SDHA-OE were significantly lower than SDHA-NC. B Cells protruding from the outer side of the basement membrane were observed under a microscope. Cell counts observed of SDHA-OE were significantly lower than SDHA-NC in H929 cells. C Cell counts observed of SDHA-OE were lower than SDHA-NC, but no statistical difference in AMO-1 cells. D OCR and ECAR curves of H929 cell lines. Compared with SDHA-NC, SDHA-OE had higher OCR and lower ECAR. E OCR and ECAR curves of AMO-1 cell lines. Compared with SDHA-NC, SDHA-OE had higher OCR and lower ECAR. F Energy metabolite hierarchical clustering diagram by targeted metabolomics of SDHA-NC and SDHA-OE cells. G Tumor volume was measured until the 24th day after subcutaneous cell inoculation in mice, growth curves of subcutaneous tumors in SDHA-NC and SDHA-OE groups of mice. Tumor volume of SDHA-OE group was significantly lower than SDHA-NC group (P = 0.0350, P = 0.0008, P < 0.0001, and P < 0.0001, respectively, on the day 18, 20, 22, and 24 after subcutaneous cell inoculation). H Statistical results of subcutaneous tumor weight. I Left panel showed micro PET/CT scanning imaging of NC and SDHA-OE group mice; Right panel showed statistical results of subcutaneous tumor SUVmax in NC and SDHA-OE groups of mice. J Energy metabolite hierarchical clustering diagram by targeted metabolomics of B-NDG mice xenograft tumor tissue. (SDHA-NC stands for normal control cells or mice tumor models by subcutaneous injection of normal control H929 cells. SDHA-OE stands for SDHA overexpression cells or mice tumor models by subcutaneous injection of H929 cells overexpressing SDHA. *P < 0.05, **P < 0.01,***P < 0.001,****P < 0.001; ns stands for no statistically significant difference between the two groups; Oligo stands for oligomycin; FCCP stands for mitochondrial oxidative phosphorylation uncoupling agent; Ant/Lot stands for antifungal/rotenone; 2-DG stands for 2-deoxy-D-glucose).
We evaluated the effect of overexpression of SDHA on mitochondrial aerobic respiratory function in H929 cells and AMO-1 cells by detecting mitochondrial OCR through Seahorse energy metabolism experiments. Results showed that compared with normal control group (SDHA-NC), SDHA overexpression group (SDHA-OE) had significantly higher OCR, indicating overexpression of SDHA promoted mitochondrial aerobic respiratory function in MM cells (Figs. 1D, E, left panels). The effect of overexpression of SDHA on the glycolysis of H929 cells and AMO-1 cells was evaluated by detecting ECAR. Results indicated that overexpression of SDHA inhibited glycolysis in MM cells (Figs. 1D, E, right panels). We conducted whole metabolomics analysis and pathway analysis on SDHA-NC and SDHA-OE H929 cells. Results showed that TCA cycle ranked at the top of enrichment terms (Supplementary Fig. 1A).
In order to investigate the effects of overexpression of SDHA on TCA cycle and glycolysis in MM cells, we performed hierarchical clustering analysis on energy metabolites in SDHA-OE group and SDHA-NC group H929 cells. TCA cycle metabolites of SDHA-OE cells were significantly increased compared to SDHA-NC. In contrast, glycolytic products were significantly reduced (Fig. 1F). We further selected important metabolites in TCA cycle and glycolysis for quantitative analysis. Concentrations of glycolytic metabolites such as D-glucose 6-phosphate (Supplementary Fig. 2B, P = 0.0006), Beta-D-Fructose 6-phosphate (Supplementary Fig. 2C, P = 0.0228), lactate (Supplementary Fig. 2D, P = 0.0011) in SDHA-OE cells were significantly lower than those in SDHA-NC cells. Concentrations of metabolites of SDHA-OE cells in TCA cycle, such as isocitrate (Supplementary Fig. 2F, P = 0.0018), alpha-ketoglutarate (Supplementary Fig. 2H, P = 0.0004), fumarate (Supplementary Fig. 2I, P = 0.0426), succinate (Supplementary Fig. 2J, P < 0.0001), oxaloacetate (Supplementary Fig. 2K, P = 0.0205), were significantly increased compared to SDHA-NC group. Concentrations of phosphoeno pyruvate and L-malic acid showed no significant difference in statistically, but there was still a clear trend (Supplementary Fig. 2E, G). The results indicated that high expression of SDHA promoted TCA cycle, and while glycolysis was inhibited in vitro. The roles of 32 metabolites in glycolysis and the TCA cycle were shown in Supplementary Table 2.
To explore the effects of overexpression of SDHA on proliferation of MM tumors in vivo, we established a mice MM subcutaneous tumor model by subcutaneous injection of H929 cells. Growth curve of the subcutaneous tumor in mice was plotted (Fig. 1G). Results showed that the tumor growth rate of SDHA-OE mice was slower than that of SDHA-NC mice. On the 24th day after subcutaneous cell inoculation in mice, the average tumor volumes in SDHA-NC group and SDHA-OE group were 1728.3 ± 147.1 mm3, and 1130.4 ± 168.3 mm3 respectively (Supplementary Fig. 3A and B, P < 0.0001). The tumor weights of SDHA-NC group and SDHA-OE group were 1.58 g and 0.78 g, respectively (Fig. 1H, P = 0.0022). The results of realtime RT-PCR were shown in Supplementary Fig. 3C. Compared with SDHA-NC group, SDHA-OE group showed a significant decrease in tumor invasion markers (MMP-2, MMP-9, CXCR4, MKI67) and increase in anti-invasion molecule (CDH1) expression level in tumor tissue. The results of immunohistochemistry (IHC) were shown in Supplementary Fig. 3D. Compared with SDHA-NC group, SDHA-OE group showed a significant increase in SDHA expression level in tumor tissue (P = 0.0150). Compared with SDHA-NC group, SDHA-OE group showed a decrease in Ki-67 and MMP2 expression (Supplementary Fig. 3C, P = 0.0406 and P = 0.0129, respectively). The IHC staining experiment results confirmed that overexpression of SDHA inhibited proliferation and invasion of MM tumors.
Micro PET/CT scans were performed to further explore the effects of overexpressing SDHA on glucose metabolism of subcutaneous tumors. As shown in Fig. 1I, the SUVmax of SDHA-OE group (1.8, 2.0, and 2.1 for three mice, respectively) was significantly lower than that of SDHA-NC group (3.7, 3.9, and 4.4, respectively, P = 0.0008). The decrease in 18F-FDG uptake in MM subcutaneous tumors of SDHA-OE mice suggested that overexpression of SDHA inhibited glucose metabolism in mice xenograft tumors. 18F-FDG is a glucose analogue, and tumor uptake of 18F-FDG is based on enhanced glycolysis21,22. Therefore, overexpression of SDHA in tumor tissues with decreased SUVmax suggested its inhibition of glycolysis.
In order to investigate the effects of overexpression of SDHA on TCA cycle and glycolysis in tumor tissues, we performed hierarchical clustering analysis on energy metabolites in SDHA-OE group and SDHA-NC group mice tumor tissues. TCA cycle metabolites of SDHA-OE cells were significantly increased compared to SDHA-NC. By contrast, glycolytic products were significantly reduced (Fig. 1J). We further selected important metabolites in TCA cycle and glycolysis for quantitative analysis. Concentrations of glycolytic metabolites such as D-Fructose 1,6-bisphosphate (Supplementary Fig. 4A, P = 0.0032), beta-D-Fructose 6-phosphate (Supplementary Fig. 4D, P = 0.0122) in SDHA-OE tumors were significantly lower than those in SDHA-NC tumors. On the contrary, concentrations of metabolites of SDHA-OE tumors in TCA cycle such as L-Malic acid (Supplementary Fig. 4F, P = 0.0036), fumarate (Supplementary Fig. 4G, P < 0.0001), isocitrate (Supplementary Fig. 4H, P = 0.0410), were significantly increased compared to SDHA-NC group. Concentrations of lactate, 3-phospho-D-glyceratate, and α-ketoglutarate were no significant difference in statistically, but there was still a strong trend (Supplementary Fig. 4B, C, E). The results indicated that high expression of SDHA promoted TCA cycle, and while glycolysis was inhibited in vivo.
EPAS1 was screened as a favorable prognostic factor for MM
We previously found the level of histone acetylation of SDHA was low in MM cells12, but the underlying mechanism was not clear. In order to explore regulatory molecules targeting SDHA in MM, we conducted multi-directional exploration. We analyzed GSE24080 dataset in the Gene Expression Omnibus (GEO) database to explore genes closely related to MM prognosis. The GSE24080 dataset contains 20183 gene information and clinical prognosis data from 340 MM patients who received the same treatment regimen. Twenty-nine genes related to overall survival (OS) of patients were screened through LASSO regression analysis (Supplementary Fig. 5A, B), including EPAS1. To predict biological function of these genes in MM, we conducted Gene Set Enrichment Analysis (GSEA). Results showed that compared to MM patients with low EPAS1 expression, MM patients with high EPAS1 expression showed significant activation of oxidative phosphorylation and hypoxia pathways (Supplementary Fig. 5C). Only EPAS1 showed strongly relationship to energy metabolism among the 29 genes. In order to explore the relationship between expression level of EPAS1 and prognosis of MM patients, patients in the GSE24080 dataset were divided into three groups of low, medium, and high expression of EPAS1 by X-tile software, and a K-M curve was plotted (Supplementary Fig. 5D). Patients with high expression of EPAS1 had a better prognosis than groups with medium and low expression of EPAS1, and the OS difference between the three groups was statistically significant (P < 0.0001).
In order to further confirm if the expression of EPAS1 is related to prognosis of our patients, real-time RT-PCR was used to determine the expression of EPAS1 in 138 MM patients. Based on X-tile software23, these patients were divided into three groups, including EPAS1 low expression group (n = 64), EPAS1 medium expression group (n = 36), and EPAS1 high expression group (n = 38). Table 1 showed the baseline characteristics of these 138 MM patients. The information of treatment received by the patients was shown in Supplementary Table 1. As shown in Fig. 2A, expression of SDHA and EPAS1 did not change during their courses of disease. We then analyzed OS and progression-free survival (PFS) of three groups of patients. As shown in Fig. 2B, C, there were statistically significant differences in OS and PFS among the three groups with low, medium, and high expression of EPAS1 (P = 0.041 and 0.0077, respectively).
A Four bone marrow CD138+ cells samples were randomly before and after MM treatment. It was found that there was no significant difference in the expression of SDHA and EPAS1 in patients before and after treatment (4 patients were treated with VTD, VCD, VCD, and RD regimens, respectively). B OS in three groups of patients with low, medium, and high expression of EPAS1. C PFS in three groups of patients with low, medium, and high expression of EPAS1. D Nomogram diagram shows two variables of the best prognosis prediction model of OS in Cox regression, including expression of SDHA, and R-ISS stage. E Nomogram diagram shows two variables of the best prognosis prediction model of PFS in Cox regression, including expression of SDHA, and R-ISS stage. The integral weight of each variable was shown in the figure. This diagram also predicted the relationship between this model and 1-, 2-, and 4-year OS and PFS of MM patients.
We included expression of EPAS1 and all the variables shown in Table 1 into Cox multivariate regression analysis. MM patients with high SDHA expression had significantly longer OS. Nomogram graphs (Fig. 2D, E) were constructed to quantify specific survival probability of a MM patient. Based on the statistically significant variables in the multivariate stepwise Cox regression model, nomogram graphs could predict 1-, 2-, and 4-year OS and PFS.
Enhancing expression of EPAS1 in MM cells restored the TCA cycle and reduced level of glycolysis in vitro and in vivo
To explore the reason that the expression of EPAS1 decreased in MM cells, we first detected the K48 ubiquitination level of EPAS1 protein in H929 cells. As shown in Supplementary Fig. 6A, after deubiquitination by DUB, a K48-linkage-specific deubiquitinase, the signal of the target protein (EPAS1) was enhanced, indicating that EPAS1 was ubiquitinated in H929 cells; thus, the expression was reduced. To explore the effects of EPAS1 expression on MM cells, we established EPAS1 overexpression and knockdown cell lines. The relative expression of EPAS1 of MM cell lines was shown in Supplementary Fig. 6B. We chose AMO1 and H929 to construct EPAS1 overexpression cell lines (Supplementary Fig. 6D, E). 8226 and H929 were chosen to establish transient transfection cells with EPAS1 knockdown MM cell lines by siRNA (EPAS1-siRNA, Supplementary Fig. 6C).
H929 and AMO-1 cells were cultured with an original concentration of 0.5×105/ml to detect proliferation of MM cells. Compared with the normal control, EPAS1 overexpression inhibited cell proliferation in H929 (Fig. 3A, left panel, P = 0.1469 and P = 0.0061, respectively, cultured after 24 h and 48 h) and AMO-1 (Fig. 3A, right panel, P = 0.0116 and P = 0.0006, respectively, cultured after 24 h and 48 h) cell lines. As revealed by the transwell invasion assay, compared with ncH929 cells, EPAS1 overexpression cells had a lower invasion ability (Supplementary Fig. 6F and Fig. 3B, P = 0.0276 and P = 0.0004, respectively for 24 h and 48 h). In the AMO-1 cell line, compared with the normal control group, overexpression of EPAS1 showed a decreasing trend in cell invasion (Fig. 3C, P = 0.1404). The above results revealed that EPAS1 overexpression inhibited proliferation and invasion of MM cells. Seahorse results showed that, compared with the normal control group (EPAS1-NC), the EPAS1 overexpression group (EPAS1-OE) had significantly higher OCR (Fig. 3D, E, left panels) and lower ECAR (Fig. 3D, E, right panels), indicating that overexpression of EPAS1 promoted mitochondrial aerobic respiratory function and inhibited glycolysis in MM cells.
A Left panel showed H929 cell proliferation curves of EPAS1-NC and EPAS1-OE. After 24 h and 48 h, cell counts of EPAS1-OE were significantly lower than EPAS1-NC; Right panel showed AMO-1 cell proliferation curves of EPAS1-NC and EPAS1-OE. After 48 h, cell counts of EPAS1-OE were significantly lower than EPAS1-NC. B Cell counts observed of EPAS1-OE were significantly lower than EPAS1-NC in H929 cells. C Cell counts observed of EPAS1-OE were lower than EPAS1-NC but no statistical difference in AMO-1 cells. D OCR and ECAR curves of H929 cell lines. Compared with EPAS1-NC, EPAS1-OE had higher OCR and lower ECAR. E OCR and ECAR curves of AMO-1 cell lines. Compared with EPAS1-NC, EPAS1-OE had higher OCR and lower ECAR. F Energy metabolite hierarchical clustering diagram by targeted metabolomics of EPAS1-NC and EPAS1-OE cells. (EPAS1-NC stands for normal control cells. EPAS1-OE stands for EPAS1 overexpression cells. G Tumor volume was measured until the 24th day after subcutaneous cell inoculation in mice, growth curves of subcutaneous tumors in EPAS1-NC and EPAS1-OE groups of mice. Tumor volume of EPAS1-OE group was significantly lower than EPAS1-NC group (P = 0.0088, P = 0.0045, P = 0.0003, and P < 0.0001, respectively, on the day 18, 20, 22, and 24 after subcutaneous cell inoculation). H Statistical results of subcutaneous tumor weight. I Left panel showed micro PET/CT scanning imaging of EPAS1-NC and EPAS1-OE group mice; Right panel showed statistical results of subcutaneous tumor SUVmax in NC (the same with Figue 1I NC group) and EPAS1-OE groups of mice. J Energy metabolite hierarchical clustering diagram by targeted metabolomics of B-NDG mice xenograft tumor tissue. (EPAS1-NC stands for normal control cells or mice tumor models by subcutaneous injection of normal control H929 cells. EPAS1-OE stands for SDHA overexpression cells or mice tumor models by subcutaneous injection of H929 cells overexpressing EPAS1.*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; ns stands for no statistically significant difference between the two groups; Oligo stands for oligomycin; FCCP stands for mitochondrial oxidative phosphorylation uncoupling agent; Ant/Lot stands for antifungal/rotenone; 2-DG stands for 2-deoxy-D-glucose).
In order to investigate the effects of overexpression of EPAS1 on TCA cycle and glycolysis in MM cells, we performed hierarchical clustering analysis on energy metabolites in EPAS1-OE group and EPAS1-NC group H929 cells. TCA cycle metabolites of EPAS1-OE cells were significantly increased compared to EPAS1-NC, while glycolytic products were significantly reduced, indicating that overexpression of EPAS1 promoted TCA cycle of MM cells and inhibited glycolysis (Fig. 3F). Concentrations of glycolytic metabolites such as lactate (Supplementary Fig. 7A, P = 0.0006), 3-phospho-D-glyceratate (Supplementary Fig. 7B, P = 0.0029), D-glucose 6-phosphate (Supplementary Fig. 7D, P = 0.0013) in EPAS1-OE cells were significantly lower than those in EPAS1-NC cells. On the contrary, concentrations of important metabolites of EPAS1-OE cells in the TCA cycle, such asisocitrate (Supplementary Fig. 7H, P = 0.0003), citrate (Supplementary Fig. 7I, P < 0.0001), cis-aconitate (Supplementary Fig. 7J, P < 0.0001) were significantly increased compared to the EPAS1-NC group. Concentrations of β-D-Fructose 6-phosphate, thiamine pyrophosphate, succinate, and α-ketoglutarate were no significant difference in statistically, but there was still a strong trend (Supplementary Fig. 7C, E–G). The results indicated that high expression of EPAS1 promoted TCA cycle, and while glycolysis was inhibited in vitro.
In vivo, the growth curve of the subcutaneous tumor in mice was plotted (Fig. 3G). Tumor growth rate of EPAS1-OE mice was slower than that of EPAS1-NC mice. On the 24th day after subcutaneous cell inoculation in mice, the average tumor volumes in the EPAS1-NC group and EPAS1-OE group were 1728.3 ± 147.1 mm³ and 964.6 ± 293.5 mm3, respectively (Supplementary Fig. 3A, B, P < 0.0001). Tumor weights of EPAS1-NC group and EPAS1-OE group were 1.58 g and 0.88 g, respectively (Fig. 3H, P = 0.0378). The results of realtime RT-PCR were shown in Supplementary Fig. 8C. Compared with EPAS1-NC group, EPAS1-OE group showed a significant decrease in tumor invasion markers (MMP-2, MMP-9, CXCR4, MKI67) and increase in anti-invasion molecule (CDH1) expression level in tumor tissue. The results of IHC were shown in Supplementary Fig. 8D. Compared with EPAS1-NC group, EPAS1-OE group showed a significant increase in EPAS1 expression level in tumor tissue (P = 0.0317) and a decrease in Ki-67 and MMP2 expression (Supplementary Fig. 8D, P = 0.0390 and P = 0.0363, respectively). As shown in Fig. 3I, through micro PET/CT, the SUVmax of EPAS1-OE group (2.1, 2.5, and 3.0 for three mice, respectively) was significantly lower than that of NC group (3.7, 3.9, and 4.4, respectively, P = 0.0177).
TCA cycle metabolites of EPAS1-OE cells were significantly increased compared to EPAS1-NC. While on the other hand, glycolytic products were significantly reduced (Fig. 3J). Concentrations of glycolytic metabolites such as dihydroxyacetone phosphate (Supplementary Fig. 9A, P = 0.0011), D-glucose 6-phosphate (Supplementary Fig. 9B, P < 0.0001), lactate (Supplementary Fig. 9D, P < 0.0001), D-fructose 1,6-bisphosphate (Supplementary Fig. 9E, P = 0.0474) in EPAS1-OE tumors were significantly lower than those in EPAS1-NC tumors. On the contrary, concentrations of important metabolites of EPAS1-OE tumors in the TCA cycle, such as isocitrate (Supplementary Fig. 9H, P = 0.0018), citrate (Supplementary Fig. 9J, P = 0.0027), succinate (Supplementary Fig. 9K, P < 0.0001), cis-aconitate (Supplementary Fig. 9L, P = 0.0009), were significantly increased compared to EPAS1-NC group. Concentrations of 3-phospho-D-glycerate, β-D-Fructose 6-phosphate, α-ketoglutarate, and oxaloacetate were no significant difference in statistically, but there was still a strong trend (Supplementary Fig. 9C, F, G, I). The results indicated that high expression of EPAS1 promoted TCA cycle, and while glycolysis was inhibited in vivo.
EPAS1 inhibited HDAC2 and activated transcription of SDHA
In order to discover the relationship between EPAS1 and SDHA, Realtime RT-PCR and Western blot were performed. Expression levels of SDHA mRNA and protein were significantly decreased in H929 cells when EPAS1 was knocked down (Fig. 4A, B). Upregulation of EPAS1 increased expression level of SDHA in H929 cells (Fig. 4C, D). However, there was no significant change in expression of EPAS1 mRNA after overexpression or knockdown of SDHA (Fig. 4E). Moreover, compared with SDHA-NC group, there was no significant change in expression of EPAS1 protein in SDHA-OE and SDHA-KD groups (Fig. 4F). Based on the above results, EPAS1 regulated the expression of SDHA.
A Left panel showed relative expression level of EPAS1 mRNA after knocking down EPAS1 (P = 0.0158). Right panel showed relative expression level of SDHA mRNA after knocking down EPAS1 (P = 0.0393). B Knocking down of EPAS1 (P = 0.0328) resulted in decreased expression of SDHA protein (P = 0.0478); C Left panel showed relative expression level of EPAS1 mRNA after overexpressing EPAS1 (P = 0.0001). Right panel showed relative expression level of SDHA mRNA after overexpressing EPAS1 (P = 0.0044). D Overexpressing of EPAS1 (P = 0.0475) resulted in increased expression of SDHA protein (P = 0.0419); E Left panel showed relative expression level of SDHA mRNA after knocking down and overexpressing SDHA (P = 0.0006 and P = 0.0087, respectively). Right panel showed relative expression level of EPAS1 mRNA after knocking down and overexpressing SDHA (P = 0.3721 and P = 0.7328, respectively). F Neither overexpressing nor knocking down of SDHA (P = 0.0024 and P = 0.0333, respectively) resulted in changes of EPAS1 protein (P = 0.0829 and P = 0.3817, respectively); G Distribution heatmap of Reads in gene regions reflects the enrichment of Reads in the upstream and downstream 2 kb regions of TSS and TES in the entire genome of genes. The signal enrichment of H3K27 acetylation site in the EPAS1-OE cells was significantly more active than that in the EPAS1-NC cells. H ChIP seq sequencing reads were significantly enriched in the SDHA gene region, and the area and number of enriched peaks in the EPAS1-OE cells were significantly higher than those in the EPAS1-NC cells, indicating that overexpression of EPAS1 increcased the acetylation level of SDHA histone H3K27. I Relative expression levels of multiple HDACs mRNA in H929 cells of EPAS1-NC and EPAS1-OE (P = 0.0010, P = 0.1446, P = 0.0017, P = 0.0197, P = 0.0022, P = 0.0024, P = 0.3182, P = 0.2046, P = 0.8931, P = 0.0002, P = 0.0107, P = 0.2196, P = 0.0044, P = 0.0068, P = 0.0002, P < 0.0001, P = 0.8404 relatively for EPAS1, HDAC1, HDAC2, HDAC3, HDAC4, HDAC5, HDAC6, HDAC7, HDAC8, HDAC9, HDAC10, HDAC11, SirT1, SirT2, SirT3, SirT4 and SirT7). J Enrichment comparison map of Reads in human whole gene regions. Upper panel showed HDAC2 gene region sequence alignment and nether panel showed SirT4 gene region sequence alignment. K ChIP qPCR showed EPAS1 protein bound to HDAC2 gene promoter (P = 0.0081). (EPAS1-NC stands for normal control cells. EPAS1-OE stands for EPAS1 overexpression cells. EPAS1-siRNA stands for EPAS1 knocking down cells. SDHA-NC stands for normal control cells. SDHA-OE stands for SDHA overexpression cells. SDHA-siRNA stands for SDHA knocking down cells.***P < 0.001; ****P < 0.0001; ns stands for no statistically significant difference between the two groups.
Our previous studies have confirmed that SDHA expression was low in H929 cells due to its low histone H3K27 site acetylation level12. We detected the effect of EPAS1 on acetylation level of histone H3K27 acetylation site. Based on the heat map, it was seen that concentration of Reads upstream of transcription starting site (TSS) in EPAS1-OE cells was significantly higher than that in EPAS1-NC cells, indicating that the acetylation level of histone H3K27 acetylation site in EPAS1-OE cells was significantly higher than that in EPAS1-NC (Fig. 4G). What’s more, compared with EPAS1-NC cells, the peak area and peak number of SDHA binding to histone H3K27 acetylation site in EPAS1-OE cells were significantly higher, indicating that overexpression of EPAS1 increased acetylation level of histone H3K27 in SDHA (Fig. 4H). As EPAS1-overexpression enhanced the histone acetylation intensity of H929 cells, we hoped to screen out which histone deacetylase expression decreased when EPAS1 was overexpressed. In other words, we tried to find an HDAC affected downstream of EPAS1. We detected mRNA expression levels of 18 kinds of HDACs (HDAC1-11 and SirT1-7) in EPAS1-OE cells and EPAS1-NC cells. Compared with EPAS1-NC group, HDACs such as HDAC2, HDAC3, HDAC4, HDAC5, HDAC10 and SirT2 were significantly reduced in EPAS1-OE group, while expression levels of HDAC9, SirT1, SirT3, and SirT4 were significantly increased (Fig. 4I). We screened the above HDAC genes to find HDACs binding to EPAS1 protein by ChIP-seq. By visualizing sequencing reads data and aligning them with the entire human genome gene sequence, we found HDAC2 and SirT4 genes binding to EPAS1 (Fig. 4J), while realtime RT-PCR results indicated that overexpression of EPAS1 reduced expression of HDAC2 and increased expression of SirT4. Therefore, we intersected the above results that HDAC2 might be the HDAC interacting with EPAS1. To confirm the result of ChIP-seq, we measured EPAS1 level in HDAC2 promoter region by ChIP qPCR experiment. The results are shown in Fig. 4K, indicating that EPAS1 bound to the promoter of HDAC2.
To confirm the effects of EPAS1 on HDAC2 and histone H3K27 acetylation, we detected expression of HDAC2 and H3K27 acetylation protein (H3K27ac) after overexpressing and knocking down EPAS1 in H929 cells, respectively. Compared with EPAS1-NC cells, knocking down EPAS1 (EPAS1-KD) in cells significantly increased expression of HDAC2 mRNA and protein, while expression of H3K27ac decreased. By contrast, overexpression of EPAS1 resulted in a significant decrease in HDAC2 mRNA and protein expression, while H3K27ac expression increased (Fig. 5A-C). In order to explore the effects of HDAC2 on EPAS1 and SDHA, highly selective HDAC2 inhibitor Santacruzamate A (HDAC2i), was applied to treat H929 cells. Results showed that SDHA and H3K27ac were significantly increased, while expression of EPAS1 did not show significant changes (Figs. 5D, E). Figure 5F–H showed that after overexpression or knocking down SDHA had no significant effect on mRNA and protein expression of HDAC2. To further confirm that the regulation of histone H3K27 acetylation by EPAS1 is mediated by HDAC2, we applied HDAC2i to treat EPAS1 knockdown H929 cells, which inhibited activity of HDAC2 without affecting protein expression, and detecting expression levels of H3K27ac and SDHA. Results showed that knocking down EPAS1 inhibited HDAC2, and expression levels of histone H3K27ac and SDHA proteins were significantly increased (Fig. 5I). After inhibiting HDAC2, the regulatory effect of EPAS1 on H3K27 acetylation and SDHA disappeared, indicating that the regulation of histone H3K27 acetylation level by EPAS1 depends on HDAC2.
A Left panel showed relative expression level of EPAS1 mRNA after overexpression and knocking down EPAS1 (P = 0.0005 and P = 0.0008, respectively). Right panel showed relative expression level of SDHA mRNA after overexpression and knocking down EPAS1 (P = 0.0112 and P = 0.0056, respectively). B Changes in the expression levels of HDAC2 and H3K27ac proteins (P = 0.0170 and P < 0.0001, respectively) after knocking down EPAS1 (P = 0.0064); C Changes in the expression levels of HDAC2 and H3K27ac proteins (P = 0.0043 and P = 0.0002, respectively) after overexpressing EPAS1 (P = 0.0020). D Left panel showed relative expression level of SDHA mRNA after adding 10 μM HDAC2i to H929 cells for 24 h (P = 0.0104). Right panel showed relative expression level of EPAS1 mRNA after adding 10uM HDAC2i to H929 cells for 24 h (P = 0.2501). E Changes in the expression levels of EPAS1, SDHA, HDAC2, and H3K27ac proteins (P = 0.4326, P = 0.0106, P = 0.9638, and P = 0.0022, respectively) after adding 10 μM HDAC2i to H929 cells for 24 h. F Changes in relative expression levels of SDHA mRNA after overexpression and knockdown of SDHA; Left panel showed changes in relative expression levels of SDHA mRNA after overexpression and knocking down SDHA (P = 0.0025 and P = 0.0167, respectively). Right panel showed changes in HDAC2 mRNA expression after overexpression and knocking down SDHA (P = 0.2606 and P = 0.2010, respectively). G Changes in SDHA and HDAC2 protein expression after knocking down SDHA (P = 0.0190 and P = 0.6895, respectively). H Changes in SDHA and HDAC2 protein expression after overexpressing SDHA (P = 0.0018 and P = 0.6946, respectively). I Knocking down EPAS1 and inhibiting HDAC2 resulted in increased expression levels of histone H3k27ac and SDHA proteins (P = 0.1878, P = 0.0012, P = 0.7507 and P = 0.0022, respectively for EPAS1, SDHA, HDAC2 and H3K27ac). J Standard curve of the relationship between ATP concentration and RLU. The curve equation is Y = 0.6371X + 5.012. Y stands for log(RLU), and X stands for log(concentration of ATP). K Changes of ATP production after overexpressing and knocking down SDHA (P = 0.0209 and P = 0.0040, respectively). L Changes of ATP production after overexpressing and knocking down EPAS1 (P = 0.0105 and P = 0.0038, respectively). M Change of ATP production after adding HDACi into MM cells (P = 0.0057). N Pattern diagram showed EPAS1-HDAC2-SDHA-glycolysis/TCA cycle axis in normal cells. O Pattern diagram showed EPAS1-HDAC2-SDHA-glycolysis/TCA cycle axis in MM cells. *P < 0.05; **P < 0.01.
Finally, we detected the concentration of intracellular ATP to intuitively reflect the role of EPAS1-HDAC2-SDHA axis on cellular energy metabolism. The standard curve was drawn according to the ATP concentration and the corresponding RLU value (Fig. 5J). After overexpression of EPAS1 or SDHA (Fig. 5L, K) and inhibiting HDAC2 by Santacruzamate A (Fig. 5M), the production of intracellular ATP in H929 cells was significantly decreased. On the contrary, after EPAS1 or SDHA was knocked down, the production of intracellular ATP in H929 cells was significantly increased (Fig. 5L, K). After 10 μM Santacruzamate A was added to cells for 24 h, TCA cycle metabolites of H929 cells were significantly increased compared to control H929 cells. While on the other hand, glycolytic products were significantly reduced (Fig. 5N).
Discussion
In conclusion, as shown in Fig. 5O, low expression of EPAS1 in MM cells resulted in high expression of HDAC2. This phenomenon hinders acetylation of histone of SDHA, leading to a decrease in expression of SDHA in MM. Low level of SDHA blocks TCA cycle in MM cells, and glycolysis dominates metabolism of glucose, releasing a large amount of essential substances and promoting rapid cell proliferation.
We discovered that EPAS1 was downregulated in MM cells; however, higher EPAS1 expression is associated with better prognosis in patients. FISH results were shown in Table 1. The results showed that expression of SDHA was not related to regular cytogenetic abnormalities in MM patients. Speaking of the samples, we selected bone marrow samples with follow-up information from the myeloma sample bank randomly. Newly diagnosed and treated patients were both included. We regarded expression of EPAS1-SDHA axis as a unique cellular biological characteristic of an MM patient. This was just like expression of c-MYC and BCL-2 in DLBCL patients24. It might not change during their courses of disease unless they accepted special treatment like HDACi. To confirm our hypothesis, we have performed some experiments before. As shown in Fig. 2A, there was no difference in the expression of SDHA and EPAS1 in MM cell line H929 and patients before and after treatment. What’s more, we also analyzed initial patients and treated patients respectively. As shown in Supplementary Fig. 10, both newly diagnosed and treated patients showed a correlation between EPAS1 expression and prognosis.
EPAS1 is a hypoxia-inducible factor and also a key transcription factor for cells to respond to hypoxia25. EPAS1 plays an important role in the occurrence and development of various tumors. Raval et al. reported that EPAS1 promoted proliferation in renal cell carcinoma with VHL deficiency while HIF1α acted as a tumor suppressor. They also showed in renal tumor cells, the enhanced expression of EPAS1 could suppress HIF1α26. Another study reported that EPAS1 promoted tumor proliferation and migration in renal cancer, but inhibited proliferation and migration of breast cancer cells27. Ebeid et al. reported that the expression of EPAS1 was significantly upregulated in breast cancer tissues. Moreover, it was significantly positively correlated with Her-2 expression and β-catenin levels in breast cancer patients28. A study showed that overexpression of EPAS1 in rat glioma tumors enhanced angiogenesis but reduced the growth of tumors by increasing tumor cell apoptosis. Moreover, siRNA knockdown of EPAS1 reduced apoptosis in hypoxic human malignant glioblastoma cells. Furthermore, EPAS1 deficiency in teratomas reduced vascularization but accelerated the growth of these tumor types29. Westerlund et al. reported that EPAS1 expression in neuroblastoma was associated with a better patient outcome and characteristics of low-risk tumors. In neuroblastoma tumors, genes co-expressed with EPAS1 strongly predict favorable patient outcomes and features of low-risk tumors30.
We also found low levels of EPAS1 binding to HDAC2 resulted in low expression of SDHA in MM cells and finally promoted proliferation of tumor cells by blocking the TCA cycle. HDAC2, a member of the class I HDACs, has been reported to play important roles in cellular proliferation, cell signaling, cancer initiation and progression, inflammation, and gene expression regulation31. C-MYC was reported to increase the expression of HDAC2 by binding to the HDAC2 gene directly32. Yin Yang 1 (YY1) bound to HDAC2 and regulated the sensitivity of tyrosine kinase inhibitors in clear cell renal cell carcinoma33. HDAC2 was overexpressed in lung cancer tissues andmetastatic human colorectal cancer34. High levels of HDAC2 could predict the poor prognosis of OS osteosarcoma, hepatocellular carcinoma, and colorectal cancer patients35,36,37. A higher level of HDAC2 was found in oral squamous cell carcinoma than in controls and was associated with the tumor stages of patients38. The downregulation of HDAC2 induced the expression of E3 ubiquitin-protein ligase c-Cbl in lung cancer cells. The ectopic expression of c-Cbl decreased EGFR expression and inhibited the growth of lung cancer39. What’s more, HDAC2-specific inhibitors are now developing as anti-cancer drugs. As specific HDAC1/2 inhibitors, romidepsin and givinostat suppressed the proliferation of urothelial carcinoma cells by inducing non-apoptotic cell death40. Treatment of dedifferentiated liposarcoma cell lines with romidepsin reduced tumor growth and decreased mouse double minute 2 (MDM2) expression41. Valproic acid (VPA) induced apoptosis by inhibiting Akt signaling and inducing autophagy in GCs42. As an HDAC2 inhibitor, VPA increased acetyl-histone H3 level by upregulating p27, caspase 3, and caspase 9 in gastric cancer cells43. The binding of VPA to HDAC2 also enhanced the sensitivity of melanoma cells to radiation44.
However, a previous study revealed that MM cell survival and growth depended on the expression of HDAC3 and HDAC1, but not of HDAC245. We also checked the HDAC3 gene region sequence alignment from ChIP-seq (Supplementary Fig. 11A) and performed ChIP qPCR (Supplementary Fig. 11B). The results showed no significant enrichment of EPAS1 was detected at the HDAC3 promoter. The expression of HDAC3 did not changed after knocking down EPAS1 in H929 cells (Supplementary Fig. 11C). When 100 nM BRD3308, a HDAC3 inhibitor, was added into MM cells, the expressions of SDHA were not changed (Supplementary Fig. 11D). What’s more, the inhibitor of HDAC2 inhibited proliferation of MM cells while the inhibitor of HDAC3 did not (Supplementary Fig. 11E). In addition, Patients with high expression of HDAC2 had a shorter OS and PFS than those with low expression. While although there was also a trend that high expression of HDAC3 had a shorter OS and PFS in MM patients, but the differences were not significant. So, our data showed HDAC2 appears to play a more critical role than HDAC3 in EPAS1-SDHA axis of MM cells.
Metabolic reorganization is an important marker of malignant tumors46. Abnormal cellular metabolism often occurs before tumors47. In 1927, Otto Warburg first observed an abnormal feature of energy metabolism in tumor cells48. Even in the presence of oxygen, tumor cells can focus their energy metabolism mainly on glycolysis, thereby reprogramming their glucose metabolism to produce energy. The main reasons for this are as follows. First of all, metabolites produced by glycolytic pathway can provide necessary substances for the synthesis and metabolism of rapidly proliferating tumor cells49. For example, the pyruvate produced by glycolysis can be converted into substances such as cholesterol and fatty acids, while the TCA cycle mainly produces ATP and lacks this ability50. Secondly, rapid proliferation of tumor cells requires production of a large amount of energy in a short period of time, and the glycolytic pathway is more efficient and adaptable than the TCA cycle51. Therefore, glycolytic pathway meets the substance requirements for the rapid proliferation of tumor cells. Although this effect is commonly observed in various tumors, it may not act as a dominant mechanism in tumorigenesis52,53.
Multiple myeloma is a hematological malignancy that has evolved from antibody-secreting B lymphocytes. Like other types of cancers, myeloma cells have acquired functional capabilities which are referred to as “Hallmarks of Cancer”, and one of their most important features is the metabolic disorders54. Metabolic changes not only induced aberrant proliferation of tumor cells, but also led to metabolic wastes accumulation55. Moreover, metabolites could regulate gene and protein expression that would influence the behavior of tumor cells, normal cells, and tumor microenvironment. These cells would in turn produce more metabolites and wastes impacting the regulation of genes and proteins that affected disease progression56.
The metabolic abnormalities of MM are mainly manifested as severe dependence on glycolysis57,58. MM cells also exhibit abnormally active glycolysis, leading to uptake of more glucose by cells, enabling them not only to quickly obtain energy from aerobic and anaerobic conditions, but also provide intermediate metabolites for other synthetic metabolic pathways59. There are also a few studies on inhibiting the Warburg effect of MM. Pyruvatede hydrogenase kinase (PDK), a key regulator of glycolysis and oxidative phosphorylation, was activated by HIF-1α under hypoxic conditions. As a pyruvate mimetic, dichloroacetate (DCA) reactivated pyruvate dehydrogenaseby targeting PDK, restored the supply of acetyl-CoA to the TCA cycle, finally leading to apoptotic cell death60. In MM, DCA was reported to alter the metabolic phenotype by inhibiting the Warburg effect and increasing sensitivity to bortezomib61. FOXM1 was also reported as a promising target in MM. Targeting FOXM1 with its specific inhibitor NB73 could enhance the proteasomal degradation of FOXM1, and finally inhibited MM cell growth62. The International Myeloma Working Group (IMWG), European Myeloma Network, and the European Society for Medical Oncology have recommended clinicians use 18F-FDG PET/CT scanning to localize and quantify myeloma load in patients, and for the assessment of extramedullary disease as well as response to therapies63. Although the Warburg effect plays an important role in MM, PET/CT seems to be its only application, and the treatment strategy to counteract the Warburg effect has not yet appeared in clinical practice.
In addition to glucose metabolism, lipid metabolism and amino acid metabolism also have a significant impact on MM. Excessive lipids and cholesterolare now considered hallmarks of cancer aggressiveness and chemoresistance64. MM cells need exogenous cholesterol for survival, and low-density lipoprotein (LDL) cholesterol was reported to be an anti-apoptotic factor for MM cells65. As the frontline medicine for lipid lowering treatment, statins were reported to reduce mortality in MM patients by inducing cell apoptosis and cell growth arrest66,67. MM cells were reported to induce lipolysis in BM adipocytes, and free fatty acids (FFAs) were then absorbed by MM cells, leading to an increased proliferation of MM cells at lower concentrations of FFA68. MM cells also rely on metabolism of amino acids for their survival and proliferation. As the most abundant amino acid in the body, glutamine was needed to meet urgent nutrient demands of MM cells, and glutamine served both as a component of proteins and as a supplement to the TCA cycle, which sustained the metabolic status of MM cells69. Moreover, MM cells absorbed glycine via solute carrier family 6 member 9 (SLC6A9) to synthesize glutathione and purines. The knockdown of SLC6A9 could inhibit proliferation of MM cells and enhance the effect of bortezomib70.
We screened that EPAS1 and SDHA were both good prognostic factors, and they were involved in glucose metabolism. This phenomenon also inferred that the Warburg effect might play an important role in MM. As there is treatment for MM with the strategy of anti-Warburg effect applied in clinical settings, potential therapeutic implications are now our new concern, and we are conducting the relative research. Actually, SDHA was found by a kind of HDACi, chidamide. We found that chidamide could inhibit the proliferation of MM cells in vitro, and SDHA was screened by RNA-seq after treating primary MM cells in vitro. Our goal is to apply the key metabolic pathways of MM we found, and then search for more suitable treatment methods. Now we have screened several FDA approved drugs that act on the EPAS1-SDHA axis, and further studies are currently underway. We are looking forward to more related treatment methods to enrich the treatment options for MM.
Methods
Patients
The histological diagnosis of MM was established according to IMWG 2014 criteria71. The study was approved by Shanghai Zhongshan Hospital Review Board (Y-2021085). Informed consent was obtained from patients in accordance with the Declaration of Helsinki. Bone marrow-derived mononuclear cells (BMMCs) were isolated from the bone marrow samples of MM patients by Ficoll-isopaque centrifugation. CD138+ cells were isolated by CD138 magnetic microbeads (130-051-301, Miltenyi Biotec, Cologne, Germany).
Cells and reagents
BMMCs and MM cell lines were cultured in RPMI-1640 medium with 10% heat-inactivated fetal bovine serum (FBS) in a humidified atmosphere of 95% air and 5% CO2 at 37 °C. MM cell lines AMO-1, MM1S, NCI-H929, RPMI8226, and U266 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). Santacruzamate A (CAY10683) and BRD3308 (S8962) were purchased from Selleck (Houston, TX, USA).
Detection of genes
Total RNA was extracted from BMMCs or cell lines. Gene expression was analyzed by real-time RT-PCR using 7500HT Fast Real-time PCR system (Applied Biosystem, Foster City, CA, USA). Primers used were listed in Supplementary Table 3. Relative expressions were calculated by the method of ΔΔCT.
Western blot
Western blot was performed as described previously72. Antibodies against SDHA (14865-1-AP) and EPAS1 (26422-1-AP) were from Proteintech (Wuhan, Hubei, China). Antibody against HDAC2 was from Santa Cruz Biotechnology (sc-81599) and H3K27ac was from Cell Signaling Technology (8173 T). Actin (Beyotime, Shanghai, China) was used to ensure equivalent protein loading. The full and uncropped western blots were uploaded as ‘Supplemental Material’.
Clinical prognosis analysis
138 BMMC samples were randomly selected to extract RNA with high quality at Zhongshan Hospital. Real-time RT-PCR was used to determine expression of EPAS1 in these patients. We collected clinical and laboratory data of these 138 patients. The consecutive variables were analyzed by t test. Categorical outcomes were compared by Chi-square test. The association between EPAS1 and outcomes was shown via Kaplan-Meier (K-M) curves. All statistical tests were two-sided, and the analysis was made by SPSS 21 software and R software, version 3.6.0. P < 0.05 was regarded as statistical significance.
Cell transfection
Lenti-virus with SDHA, EPAS1, or control vector was synthesized by Genomeditech (Shanghai, China). H929 and AMO-1 cells were infected by lenti-virus with cell-virus ratio of 1:100. The transfected clones were detected after transfection for 72 h. Small interfering RNA (siRNA) was synthesized by Biotend (Shanghai, China). Cells were transfected by siRNA with an ultimate concentration of 100 nM. The transfected clones were detected after transfection for 48 h.
Cell proliferation assay
Cells were seeded at a density of 0.5×105/ml cells per well in 6 well plates and incubated at 37 °C. Cell proliferation was measured after 24 h and 48 h, respectively, by cell counting.
Cell invasion assay
Cell invasion was tested by Matrigel Invasion Chamber (BD Pharmingen, Franklin Lakes, NJ, USA), which was composed of the upper and lower compartment separated by membranes with matrigel and pores (8μm pore size). About 6 × 104 cells were incubated with RPMI-1640 (FBS-free, 200 μl) for 24 h and then added into the upper compartment, while RPMI-1640 with 10% FBS (500 μl) was added to the lower compartment. Cells were cultured for 24 h, and then invasive ability of cells was tested as described previously12. The membrane was stained by 0.1% crystalline violet, and the invading cells were observed.
Ubiquitination detection
The K48 Linkage-Specific UbiTest-Magnetic TUBE Elution Kit was purchased from LifeSensors (UM411M, Malvern, PA, USA). Magnetic beads were added into the centrifuge tube and let it stand on the magnetic frame for 2 min, and the supernatant was carefully discarded. 1 ×;107 cells was lysed by 100ul RIPA solution, and then was added into the centrifuge tube containing magnetic beads and incubated at room temperature for 1 h. K48 link specific DUB was added and incubated for 1 h after the magnetic beads were fully washed on the magnetic frame. After adding elution buffer, the supernatant was transferred to a new centrifuge tube. Finally, neutralization buffer and β-mercaptoethanol were added and mixed well for Western blot detection.
Establishment of a mice subcutaneous tumor model
The study was approved by Shanghai Zhongshan Hospital Review Board (2021-268). The ethics committee review board number was SYXK2021-0022. B-NDG mice with severe immunodeficiency at 4 weeks of age were selected and were fed in room with no specific pathogen free (SPF) level. 2.5 × 106 H929 lentivirus stable transgenic cells were injected into the subcutaneous area of the right shoulder of the mice. Long and short diameters of tumors were measured by a caliper. Tumor volume = 0.5 × long diameter × short diameter2. Animals were monitored twice daily once any trigger was approached, and immediate euthanasia (cervical dislocation) was performed upon reaching an endpoint. All experiments were performed in accordance with the approved protocol and other relevant guidelines and regulations. All experiments were performed in accordance with the approved protocol and other relevant guidelines and regulations. The maximal tumor size did not exceed the limit permitted by the ethics committee.
Immunohistochemistry
Immunohistochemical analyses were performed with an indirect immunoperoxidase method by antibodies against Ki67 (ab15580, Abcam, Cambridge, MA, USA), MMP-2 (ab97779, Abcam), SDHA (14865-1-AP, Proteintech), and EPAS1 (abs122932, Absin, Shanghai, China). Anti-rabbit-IgG antibody was (Invitrogen, Carlsbad, CA, USA) secondary antibodies. Nuclei were stained by DAPI. To quantify the immunohistochemistry, the semi-quantitative scoring method was applied. The percentage of counting positive cells in the total cells was recorded as A (1–25% was 1 point, 26–50% was 2 points, 51–75% was 3 points, 75–100% was 4 points), and the staining intensity was recorded as B (pale yellow 1 point, brown yellow 2 points, brown brown 3 points), and A × B was recorded as the final quantification of immunohistochemistry.
Seahorse cell energy metabolism experiment
4 × 104 cells were used to perform this examination per test. Drugs such as Oligomycin, mitochondrial oxidative phosphorylation uncoupler (FCCP), and Rotenone/Antimycin A were added sequentially to detect cellular oxygen consumption rate (OCR). Drugs such as glucose, oligomycin, and 2-DG were added to cells in sequence to detect extracellular acidification rate (ECAR). Seahorse XFe96 analyzer (Agilent) was used for detection.
Metabolomics analysis
1 × 107 cells or 200 mg or more tissue samples were prepared and stored at −80 °C. After grinding and ultrasonic extraction, samples were stored at −20 °C for 30 min. The extract was centrifuged at 4 °C (13,000 rpm) for 10 min. 300 μL of supernatant in a glass vial was dried in a freeze concentration centrifugal dryer. 300 μL mixture of methanol and water (volume ratio was 1:4) were added to each sample, samples vortexed for 30 s, extracted by ultrasonic for 3 min, then placed at −20 °C for 2 h. Samples were centrifuged at 4 °C (13,000 rpm) for 10 min. The supernatants (150 μL) from each tube were collected by crystal syringes, filtered through 0.22 μm microfilters and transferred to LC vials. The vials were stored at −80 °C until LC-MS analysis.
The metabolomic data analysis was performed by Shanghai Luming Biological Technology Co., Ltd. (Shanghai, China). An ACQUITY UPLC I-Class plus (Waters Corporation, Milford, USA) fitted with Q-Exactive mass spectrometer equipped with heated electrospray ionization (ESI) source (Thermo Fisher Scientific, Waltham, MA, USA) was used to analyze the metabolic profiling in both ESI positive and ESI negative ion modes.
The original LC-MS data were processed by software Progenesis QI V2.3 (Nonlinear, Dynamics, Newcastle, UK) for baseline filtering, peak identification, integral, retention time correction, peak alignment, and normalization. Differential metabolites were selected with variable importance of projection values greater than 1.0 and p values less than 0.05. Differential metabolites were further used to for KEGG pathway enrichment analysis.
Mice micro PET/CT scanning
Micro PET/CT scanning was performed when tumor in mice grows to 1.0–1.2 cm. Measure the weight and the radioactive activity of the radioactive tracer before and after injection of each mouse. F-18 fluorodeoxyglucose (18F-FDG) is injected into caudal vein as a radioactive tracer. Maximum standard uptake value (SUVmax) was measured by Syngo MI Applications on the Multi Modality workstation.
Chromatin immunoprecipitation (ChIP) sequencing
Cells were crosslinked with 1% formaldehyde at room temperature for 10 min. After chromatin shearing by sonication, lysates were incubated overnight at 4 °C with protein A Dynabeads coupled with 5 μg antibody. DNA was eluted and crosslinks reverted at 65°C for 4 h and then purified with QIAGEN Kit. For ChIP-Seq, 5 ng of purified ChIP DNA were used to generate the sequencing library using a NEB kit and sequenced with the Illumina HiSeq X Ten.
ChIP PCR
CHIP Assay kit was purchased from Beyotime Biotechnology, Inc. (P2078). Cells were fixed with 1% formaldehyde for 10 min at 37 °C. Protein was extracted from cells using lysis buffer (Beyotime Institute of Biotechnology) and an ultrasonic cell crusher (Sonics & Materials, Inc.). The supernatant was incubated for 2 h at room temperature with EPAS1 antibodies (Proteintech, Wuhan, Hubei, China) or an isotype control (IgG). Precipitated DNA was extracted from the beads by placing them in 1.1 M NaHCO3 solution (Sangon Biotech Co., Ltd.) and 1% SDS solution at 65˚C for 6 h. The quality of the PCR products was verified by a gel, and qPCR was used to quantify DNAs affiliated to the target protein.
Intracellular ATP detection
Intracellular ATP detection assay was purchased from Beyotime (S0027, Shanghai, China). Standard solution was diluted to a volume of 100 μl with ATP concentrations of 0.01, 0.03, 0.1, 0.3, and 1 μM. And then 100 μl prepared detection working solution was added into the prepared standard solution. The mixed solution was placed at room temperature for 5 min, and then the value of relative light unit (RLU) value was detected by a luminometer. A standard curve was drawn according to the ATP concentration and the corresponding RLU value. About 1 × 104 cells were lysed by 100 μl lysate. And then 100 μl prepared detection working solution was added into the prepared standard solution. The mixed solution was placed at room temperature for 5 min, and then the value RLU value was detected by a luminometer. The corresponding ATP concentration was calculated according to the RLU value.
Statistical analysis of experiments
Each experiment was repeated three times and experimental results obtained from three independent experiments and determined by t-test to compare variance. One-way ANOVA was applied for ≥3 groups, and two-way ANOVA for time course data. P < 0.05 was considered statistically significant.
Data availability
The data that support the findings of this study are available on request to the corresponding author.
References
van de Donk, N., Pawlyn, C. & Yong, K. L. Multiple myeloma. Lancet 397, 410–427 (2021).
Lucca, L. E. Multiple myeloma treatment: one bridge closer. Blood 142, 1763–1764 (2023).
Rajkumar, S. V. & Harousseau, J. L. Next-generation multiple myeloma treatment: a pharmacoeconomic perspective. Blood 128, 2757–2764 (2016).
Huang, S. & Millar, A. H. Succinate dehydrogenase: the complex roles of a simple enzyme. Curr. Opin. Plant Biol. 16, 344–349 (2013).
Bannon, A. E. et al. Biochemical, molecular, and clinical characterization of succinate dehydrogenase subunit a variants of unknown significance. Clin. Cancer Res. 23, 6733–6743 (2017).
Gill, A. J. Succinate dehydrogenase (SDH) and mitochondrial driven neoplasia. Pathology 44, 285–292 (2012).
Wang, L. et al. Upregulation of Succinate Dehydrogenase (SDHA) contributes to enhanced bioenergetics of ovarian cancer cells and higher sensitivity to anti-metabolic agent shikonin. Cancers 14, https://doi.org/10.3390/cancers14205097 (2022).
Evenepoel, L. et al. Toward an improved definition of the genetic and tumor spectrum associated with SDH germ-line mutations. Genet. Med. 17, 610–620 (2015).
K Aggarwal. R. et al. Functional succinate dehydrogenase deficiency is a common adverse feature of clear cell renal cancer. Proc. Natl. Acad. Sci. USA 118, https://doi.org/10.1073/pnas.2106947118 (2021).
Eniafe, J. & Jiang, S. The functional roles of TCA cycle metabolites in cancer. Oncogene 40, 3351–3363 (2021).
Bardella, C., Pollard, P. J. & Tomlinson, I. SDH mutations in cancer. Biochim. Biophys. Acta 1807, 1432–1443 (2011).
Sun, Y. et al. High expression of succinate dehydrogenase subunit a which is regulated by histone acetylation, acts as a good prognostic factor of multiple myeloma patients. Front. Oncol. 10, 563666 (2020).
Sun, Y. et al. Chidamide, a novel histone deacetylase inhibitor, inhibits multiple myeloma cells proliferation through succinate dehydrogenase subunit A. Am. J. Cancer Res. 9, 574–584 (2019).
Haase, V. H. Regulation of erythropoiesis by hypoxia-inducible factors. Blood Rev. 27, 41–53 (2013).
Scortegagna, M., Morris, M. A., Oktay, Y., Bennett, M. & Garcia, J. A. The HIF family member EPAS1/HIF-2alpha is required for normal hematopoiesis in mice. Blood 102, 1634–1640 (2003).
Moreno-Domínguez, A. & Ortega-Sáenz, P. Acute O(2) sensing through HIF2α-dependent expression of atypical cytochrome oxidase subunits in arterial chemoreceptors. 13, https://doi.org/10.1126/scisignal.aay9452 (2020).
Castrop, H. & Kurtz, A. Functional evidence confirmed by histological localization: overlapping expression of erythropoietin and HIF-2alpha in interstitial fibroblasts of the renal cortex. Kidney Int. 77, 269–271 (2010).
Guitart, A. V. et al. Hif-2α is not essential for cell-autonomous hematopoietic stem cell maintenance. Blood 122, 1741–1745 (2013).
Kawano, Y. et al. Targeting the bone marrow microenvironment in multiple myeloma. Immunol. Rev. 263, 160–172 (2015).
Koppenol, W. H., Bounds, P. L. & Dang, C. V. Otto Warburg’s contributions to current concepts of cancer metabolism. Nat. Rev. Cancer 11, 325–337 (2011).
Waki, A. et al. Reassessment of FDG uptake in tumor cells: high FDG uptake as a reflection of oxygen-independent glycolysis dominant energy production. Nucl. Med. Biol. 24, 665–670 (1997).
Chen, X. et al. Fibroblast activation protein and glycolysis in lymphoma diagnosis: comparison of (68)Ga-FAPI PET/CT and (18)F-FDG PET/CT. J. Nucl. Med. 64, 1399–1405 (2023).
Camp, R. L., Dolled-Filhart, M. & Rimm, D. L. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin. Cancer Res. 10, 7252–7259 (2004).
Karube, K. & Campo, E. MYC alterations in diffuse large B-cell lymphomas. Semin. Hematol. 52, 97–106 (2015).
Tian, H., McKnight, S. L. & Russell, D. W. Endothelial PAS domain protein 1 (EPAS1), a transcription factor selectively expressed in endothelial cells. Genes Dev. 11, 72–82 (1997).
Raval, R. R. et al. Contrasting properties of hypoxia-inducible factor 1 (HIF-1) and HIF-2 in von Hippel-Lindau-associated renal cell carcinoma. Mol. Cell. Biol. 25, 5675–5686 (2005).
Sowter, H. M., Raval, R. R., Moore, J. W., Ratcliffe, P. J. & Harris, A. L. Predominant role of hypoxia-inducible transcription factor (Hif)-1alpha versus Hif-2alpha in regulation of the transcriptional response to hypoxia. Cancer Res. 63, 6130–6134 (2003).
Ebeid, S. A., Abd El Moneim, N. A., El-Benhawy, S. A., Ramadan, R. & Ismail, S. E. Znhit1 and HIF-2α are correlated with cancer stem cell markers in breast cancer patients. Sci. Rep. 12, 13918 (2022).
Acker, T. et al. Genetic evidence for a tumor suppressor role of HIF-2alpha. Cancer Cell 8, 131–141 (2005).
Westerlund, I., Shi, Y. & Holmberg, J. EPAS1/HIF2α correlates with features of low-risk neuroblastoma and with adrenal chromaffin cell differentiation during sympathoadrenal development. Biochem. Biophys. Res. Commun. 508, 1233–1239 (2019).
Jo, H., Shim, K., Kim, H. U., Jung, H. S. & Jeoung, D. HDAC2 as a target for developing anti-cancer drugs. Comput. Struct. Biotechnol. J. 21, 2048–2057 (2023).
Hu, L. et al. The anti-angiogenesis role of FBXW7 in diabetic retinopathy by facilitating the ubiquitination degradation of c-Myc to orchestrate the HDAC2. J. Cell Mol. Med. 25, 2190–2202 (2021).
Li, W. et al. YTHDC1 is downregulated by the YY1/HDAC2 complex and controls the sensitivity of ccRCC to sunitinib by targeting the ANXA1-MAPK pathway. J. Exp. Clin. Cancer Res. 41, 250 (2022).
Stypula-Cyrus, Y. et al. HDAC up-regulation in early colon field carcinogenesis is involved in cell tumorigenicity through regulation of chromatin structure. PLoS ONE 8, e64600 (2013).
Li, J. et al. HDAC2-mediated upregulation of IL-6 triggers the migration of osteosarcoma cells. Cell Biol. Toxicol. 35, 423–433 (2019).
Noh, J. H. et al. HDAC2 provides a critical support to malignant progression of hepatocellular carcinoma through feedback control of mTORC1 and AKT. Cancer Res. 74, 1728–1738 (2014).
Qi, Z. P. et al. HDAC2 promotes the EMT of colorectal cancer cells and via the modular scaffold function of ENSG00000274093.1. J. Cell Mol. Med. 25, 1190–1197 (2021).
Krishna, A. et al. Upregulated histone deacetylase 2 gene correlates with the progression of oral squamous cell carcinoma. Cancer Biomark. 29, 543–552 (2020).
Wei, T. T. et al. Induction of c-Cbl contributes to anti-cancer effects of HDAC inhibitor in lung cancer. Oncotarget 6, 12481–12492 (2015).
Pinkerneil, M. et al. Inhibition of class I histone deacetylases 1 and 2 promotes urothelial carcinoma cell death by various mechanisms. Mol. Cancer Ther. 15, 299–312 (2016).
Seligson, N. D. et al. Inhibition of histone deacetylase 2 reduces MDM2 expression and reduces tumor growth in dedifferentiated liposarcoma. Oncotarget 10, 5671–5679 (2019).
Sun, J. et al. Valproic acid targets HDAC1/2 and HDAC1/PTEN/Akt signalling to inhibit cell proliferation via the induction of autophagy in gastric cancer. FEBS J. 287, 2118–2133 (2020).
Yagi, Y. et al. Effects of valproic acid on the cell cycle and apoptosis through acetylation of histone and tubulin in a scirrhous gastric cancer cell line. J. Exp. Clin. Cancer Res. 29, 149 (2010).
Kalal, B. S., Pai, V. R. & Behera, S. K. HDAC2 inhibitor valproic acid increases radiation sensitivity of drug-resistant melanoma cells. 7 https://doi.org/10.3390/medsci7030051 (2019).
Minami, J. et al. Histone deacetylase 3 as a novel therapeutic target in multiple myeloma. Leukemia 28, 680–689 (2014).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Pavlova, N. N. & Thompson, C. B. The emerging hallmarks of cancer metabolism. Cell Metab. 23, 27–47 (2016).
Carroll, V. A. & Ashcroft, M. Role of hypoxia-inducible factor (HIF)-1alpha versus HIF-2alpha in the regulation of HIF target genes in response to hypoxia, insulin-like growth factor-I, or loss of von Hippel-Lindau function: implications for targeting the HIF pathway. Cancer Res. 66, 6264–6270 (2006).
Hanse, E. A. et al. Cytosolic malate dehydrogenase activity helps support glycolysis in actively proliferating cells and cancer. Oncogene 36, 3915–3924 (2017).
Ganapathy-Kanniappan, S. & Geschwind, J. F. Tumor glycolysis as a target for cancer therapy: progress and prospects. Mol. Cancer 12, 152 (2013).
Lunt, S. Y. & Vander Heiden, M. G. Aerobic glycolysis: meeting the metabolic requirements of cell proliferation. Annu. Rev. Cell Dev. Biol. 27, 441–464 (2011).
Cohen, I. J. et al. Increased tumor glycolysis is associated with decreased immune infiltration across human solid tumors. Front. Immunol. 13, 880959 (2022).
Li, W. et al. Aerobic glycolysis controls myeloid-derived suppressor cells and tumor immunity via a specific CEBPB isoform in triple-negative breast cancer. Cell Metab. 28, 87–103.e106 (2018).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Pavlova, N. N., Zhu, J. & Thompson, C. B. The hallmarks of cancer metabolism: still emerging. Cell Metab. 34, 355–377 (2022).
Zhou, X., He, R., Hu, W. X., Luo, S. & Hu, J. Targeting myeloma metabolism: How abnormal metabolism contributes to multiple myeloma progression and resistance to proteasome inhibitors. Neoplasia 50, 100974 (2024).
Gu, Z. et al. NEK2 promotes aerobic glycolysis in multiple myeloma through regulating splicing of pyruvate kinase. J. Hematol. Oncol. 10, 17 (2017).
Zhang, B. et al. A novel glycolysis-related gene signature for predicting the prognosis of multiple myeloma. Front. Cell Dev. Biol. 11, 1198949 (2023).
Cheng, Y. et al. FOXM1 regulates glycolysis and energy production in multiple myeloma. Oncogene 41, 3899–3911 (2022).
Michelakis, E. D., Webster, L. & Mackey, J. R. Dichloroacetate (DCA) as a potential metabolic-targeting therapy for cancer. Br. J. Cancer 99, 989–994 (2008).
Sanchez, W. Y. et al. Dichloroacetate inhibits aerobic glycolysis in multiple myeloma cells and increases sensitivity to bortezomib. Br. J. Cancer 108, 1624–1633 (2013).
Cheng, Y. & Sun, F. FOXM1 regulates glycolysis and energy production in multiple myeloma. Oncogene 41, 3899–3911 (2022).
Zamagni, E., Tacchetti, P. & Cavo, M. Imaging in multiple myeloma: How? When?. Blood 133, 644–651 (2019).
Qin, J. et al. Fatty acids in cancer chemoresistance. Cancer Lett. 572, 216352 (2023).
Tirado-Vélez, J. M., Benítez-Rondán, A., Cózar-Castellano, I., Medina, F. & Perdomo, G. Low-density lipoprotein cholesterol suppresses apoptosis in human multiple myeloma cells. Ann. Hematol. 91, 83–88 (2012).
Afzal, A., Fiala, M. A., Gage, B. F., Wildes, T. M. & Sanfilippo, K. Statins reduce mortality in multiple myeloma: a population-based US study. Clin. Lymphoma Myeloma Leuk. 20, e937–e943 (2020).
Dai, Y. et al. Statins synergistically potentiate 7-hydroxystaurosporine (UCN-01) lethality in human leukemia and myeloma cells by disrupting Ras farnesylation and activation. Blood 109, 4415–4423 (2007).
Panaroni, C., Fulzele, K., Mori, T., Siu, K. T. & Onyewadume, C. Multiple myeloma cells induce lipolysis in adipocytes and uptake fatty acids through fatty acid transporter proteins. Blood 139, 876–888 (2022).
Tabe, Y. & Lorenzi, P. L. Amino acid metabolism in hematologic malignancies and the era of targeted therapy. Blood 134, 1014–1023 (2019).
Xia, J. et al. Blocking glycine utilization inhibits multiple myeloma progression by disrupting glutathione balance. Nat. Commun. 13, 4007 (2022).
Chng, W. J. et al. IMWG consensus on risk stratification in multiple myeloma. Leukemia 28, 269–277 (2014).
Xu, P. P. et al. JAM-A overexpression is related to disease progression in diffuse large B-cell lymphoma and downregulated by lenalidomide. Sci. Rep. 7, 7433 (2017).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82100216) and Shanghai Science and Technology Commission Yangfan Program Project (21YF1406200). The fundings belong to the first author Y.S.
Author information
Authors and Affiliations
Contributions
Y.S., A.M. and P.L. designed the experiments. Y.S., A.M., Y.W., J.J., H.Z., C.P. and Y.P. performed the experiments. Y.S., A.M., Z.X. and L.T. analyzed the data. YS, AM, and PL provided professional writing services and materials and wrote the manuscript. All authors met the criteria for authorship and approved the final manuscript for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, Y., Maihemaiti, A., Xu, Z. et al. EPAS1 increases SDHA to inhibit proliferation of multiple myeloma cells by restoring TCA Cycle. npj Precis. Onc. 9, 406 (2025). https://doi.org/10.1038/s41698-025-01194-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-025-01194-z