Abstract
Mesenchymal-epithelial transition exon 14 skipping mutations (METex14) define a targetable molecular subset of non-small cell lung cancer (NSCLC), but optimal circulating tumor DNA (ctDNA) biomarkers to guide MET tyrosine kinase inhibitor therapy remain undefined. Within the EMBRACE trial of ensartinib, we conducted a prospective biomarker analysis with plasma collected at baseline (V0) and 4 weeks after treatment initiation (V1). ctDNA was profiled using a hybrid-capture next-generation sequencing panel covering 229 cancer-related genes, and mutations were classified into three monitoring paradigms: MET-specific alterations, canonical alterations (tier I/II), and pan-alterations (any variant in any gene on the panel). Baseline ctDNA positivity rates were 48.3%, 48.3%, and 75.9% for MET-specific, canonical, and pan-alterations, respectively, and declined across all categories at V1. Conversion to ctDNA negativity in any paradigm was associated with a tendency toward higher objective response rate (ORR) and longer progression-free survival (PFS). Moreover, clearance of MET-specific alterations conferred the greatest clinical benefit, with an ORR of 75.0% versus 16.7% and a median PFS of 9.3 months versus 2.2 months (p = 0.005). Among the three paradigms, MET-specific monitoring provided the most favorable diagnostic performance to identify long-term responders, with a specificity of 90% and a positive predictive value of 80%. These data demonstrate that early MET-specific ctDNA clearance is a robust on-treatment biomarker for ensartinib benefit in METex14 NSCLC, while broader ctDNA profiling remains valuable for uncovering emerging resistance mechanisms.
Similar content being viewed by others
Introduction
Mesenchymal-epithelial transition exon 14 skipping mutations (METex14) occur in 2 – 4% of non-small cell lung cancer (NSCLC) cases and represent actionable targets for type 1b MET tyrosine kinase inhibitors (TKIs)1. Current approved type Ib selective MET inhibitors demonstrated an objective response rate (ORR) ranging from 40.5 to 70.6% in later-line therapy, with a median progression-free survival (mPFS) of approximately 5.5 to 11 months2. Ensartinib, a multitargeted kinase inhibitor, has been approved for the first-line treatment of ALK-rearranged NSCLC in both China and the United States. In our phase II trial evaluating type Ia MET inhibitor, ensartinib as a second-line therapy for METex14 NSCLC3,4, the agent demonstrated a comparable ORR of 53.3% and mPFS of 6.0 months. We further conducted an exploratory assessment of predictive biomarkers to identify patients most likely to derive clinical benefit.
Circulating tumor DNA (ctDNA), a fraction of cell-free DNA shed by tumors via apoptosis, necrosis, or NETosis, enables non-invasive monitoring of tumor burden and the emergence of resistance5. The International Association for the Study of Lung Cancer (IASLC) has formally endorsed ctDNA profiling as a standard-of-care component for molecular stratification in advanced NSCLC6. Landmark FLAURA and AURA3 studies demonstrated that longitudinal ctDNA monitoring, using assays targeting both common (Ex19del, L858R) and acquired resistance (T790M, C797S) epidermal growth factor receptor mutations (EGFR), could detect progressive disease (PD) prior to radiographic evidence7. Within the MET domain, the tepotinib VISION trial and savolitinib phase 2 study demonstrated that undetectable baseline METex14 ctDNA or post-treatment clearance predicts favorable clinical outcomes8,9. However, the optimal monitoring paradigm, whether to focus on the driver mutation, canonical drivers, or broader genomic alterations, remains undefined.
To date, no study has systematically compared different ctDNA monitoring strategies in METex14 NSCLC treated with MET TKIs. Leveraging the EMBRACE trial of ensartinib, we performed a post-hoc analysis of a prospective study to evaluate three ctDNA paradigms: MET-specific alterations, canonical alterations (tier I/II mutations), and pan-alterations. We hypothesized that early ctDNA clearance, particularly MET-specific alterations, would constitute the most robust predictor of therapeutic efficacy and survival in METex14 NSCLC. Unlike prior MET TKI studies such as VISION, which largely defined ctDNA positivity based on broad pan-gene profiling, our analysis directly compares three distinct ctDNA monitoring paradigms and demonstrates that MET-specific alterations offer superior predictive specificity and simplicity for early response assessment.
Results
Patient characteristics
Among the 31 patients enrolled in the Phase II trial, 30 were evaluable for efficacy assessment. As of March 1, 2025, the median follow-up duration was 9.8 months (95% confidence interval, 95% CI: 7.8–16.3). The updated ORR was 53.3% (95% CI: 35.5–71.2%), disease control rate (DCR) was 90.0% (95% CI: 73.5–97.9%), the mPFS was 6.0 months (95% CI: 3.0–8.8), and the mOS (median overall survival) was 16.3 months (95% CI: 8.2-NA) (Fig. 1A).
A Kaplan-Meier curves for progression free survival (PFS) and overall survival (OS) for the entire evaluable cohort (n = 30). B Waterfall plot depicting the distribution of somatic alterations detected in 29 patients. Genes are ordered by mutation frequency. Variant classification is indicated by color. C Correlation between tumor burden as calculated by the sum diameters of target lesion and the baseline variant allele frequency (VAF) of MET-specific alterations in MET ctDNA positive patients. D Correlation between tumor burden as calculated by the sum diameters of target lesion and the baseline mean VAF of pan-alterations.
Baseline plasma samples were available for ctDNA analysis in 29 patients. Somatic mutations were detected in 24 patients (82.8%), while 5 patients (17.2%) showed no detectable mutations. We identified 102 somatic mutations, which included 68 missense, 12 splice-site, 11 nonsense, 8 frameshift, and 3 in-frame variants. TP53 (21.0%) and RET (14.0%) were the most frequently mutated genes. (Fig. 1B, Supplementary Table 1).
Patient demographics are detailed in Table 1. Across all three profiling strategies, ctDNA negative cohorts exhibited only numerically lower tumor burdens (defined by sum of target lesion diameters) than ctDNA positive groups, with no significant difference detected (MET-specific alterations profiling, 40.3 mm vs. 45.5 mm, p = 0.631; pan-alterations profiling, 33.3 mm vs. 46.5 mm, p = 0.176). In MET-specific alterations ctDNA positive patients, target lesion size demonstrated a marginal positive correlation with variant allele frequency (VAF) (R = 0.49, p = 0.075; Fig. 1C). Conversely, no significant correlation was observed between lesion size and VAF in pan-alterations profiling (R = 0.18, p = 0.35; Fig. 1D). Moreover, metastatic organ involvement showed no association with ctDNA detection rates or VAF across all mutation classification paradigms (MET alterations profiling, p = 0.540; pan-alterations profiling, p = 1) (Table 1) (Supplementary Fig. 1).
The impacts of ctDNA at baseline on treatment efficacy
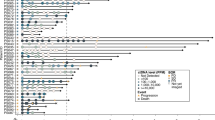
MET alterations and canonical alterations profiling were both detected in the same 14 patients (48.3%), while 15 (51.7%) were negative for both at baseline (V0), with their profiles precisely mirroring each other and no additional oncogenic drivers observed. For pan-alterations profiling, 22 patients (75.9%) showed ctDNA positivity, with 7 (24.1%) were negative (Fig. 2A). The swimmer plot was pooled to illustrate the dynamics of the ctDNA status and tumor response by radiographic RECIST criteria (Fig. 2B).
A The venn diagram showing the number of patients with ctDNA positive results detected by three profiling paradigms at V0. B Swimmer plot showing treatment duration, best of response (BOR), and MET mutation status at V0 and 4 weeks after the initiation of treatment (V1) for each patient. C Kaplan-Meier curves for progression-free survival (PFS) stratified by baseline MET ctDNA status. D Kaplan-Meier curves for PFS stratified by baseline ctDNA status of pan-alterations. E Bar plot comparing the sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) of the three profiling paradigms for predicting treatment response at V0. F Kaplan-Meier curves for PFS stratified by baseline ctDNA status of TP53 or ATRX.
At baseline, the ORR was 60.0% (95% CI: 35.2–84.8%) in the MET-specific alterations negative group and 50.0% (95% CI: 23.0–77.0%) in the positive group (p = 0.715). The mPFS was 7.5 months (95% CI: 6.0–20.2) for the negative group, compared to 4.9 months (95% CI: 2.2-NR) for the positive group (Hazard ratio, HR = 0.78, 95% CI: 0.31–1.96; p = 0.545). Although ORR and mPFS were numerically higher in the baseline MET-specific alterations negative, these differences did not reach statistical significance, indicating limited prognostic value of baseline ctDNA alone. (Fig. 2C; Table 2).
For pan-alterations profiling, the ORR in ctDNA negative patients (71.4%; 95% CI: 29.0–96.3%) was associated with a better tendency but no significant difference when compared to those with ctDNA positive detection (50.0%; 95% CI: 28.2–71.8%; p = 0.410). Similarly, mPFS was numerically longer in the negative cohort (7.7 months; 95% CI: 6.0-NA) versus the positive cohort (6.0 months; 95% CI: 3.0–9.2; p = 0.350, HR = 0.61, 95% CI: 0.22–1.72) (Fig. 2D; Table 2). Although the pan-alterations negative subgroup consistently showed numerically superior outcomes across all efficacy endpoints, these differences did not reach statistical significance in Kaplan-Meier analysis. Of note, at the V0 timepoint, the pan-alterations analysis demonstrated a higher sensitivity (85%) for predicting PR, although its specificity was limited (31%) (Fig. 2E). Besides, patients harboring baseline ATRX and/or TP53 variants showed significantly shorter mPFS (2.9 vs. 7.5 months, p = 0.049; HR = 0.32, 95% CI: 0.13–0.79; p = 0.014; Fig. 2F).
Overall, these primary findings suggested that baseline ctDNA status alone was insufficient as a robust prognostic biomarker in this cohort, whereas early on-treatment dynamics at 4 weeks provided stronger predictive signals.
The impacts of ctDNA at 4 weeks post-treatment initiation on treatment efficacy
There were 20 patients with available plasma samples at 4 weeks after the initiation of treatment (V1) (Supplementary Table 2). The number of detectable variants decreased significantly (Fig. 3A), accompanied by reduced positive rates across all three monitoring paradigms. 6 patients (30.0%) tested positive for MET-specific alterations profiling, while 5 (25.0%) of them were also positive for canonical alterations profiling. Notably, the remaining patient had a newly acquired non-canonical MET alteration. Thirteen patients (65.0%) maintained detectable pan-alterations, substantially higher than MET-specific alterations and canonical alterations detection rates. Additionally, novel acquired missense mutations in ALK (c.2698 G > A, c.1486 C > T, c.2257 C > T) and NTRK1 (c.2083 C > T) were detected in 15.0% (3/20) of patients (Fig. 3B).
A The venn diagram showing the number of patients with ctDNA positive results detected by the three profiling paradigms at V1. B Longitudinal alterations of ctDNA status during treatment. Kaplan-Meier analysis of progression-free survival (PFS) stratified by ctDNA status for (C) MET-specific alterations, D canonical alterations, and E pan-alterations at V1. F Bar plot comparing the sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) of the three profiling paradigms for predicting treatment response at V1.
For MET-specific alterations profiling, striking clinical differences emerged between these groups. The negative cohort showed the superior outcomes with markedly prolonged mPFS (negative vs. positive; 7.9 months, 95% CI: 6.0-NA vs. 2.2 months, 95% CI: 2.2-NA; HR = 0.04, 95% CI: 0.004–0.35; p = 0.004; Fig. 3C), though only a numerically better ORR was observed (64.3%, 95% CI: 35.1–87.2% vs. 16.7%, 95% CI: 4.0–64.1%; p = 0.141; Table 2, Supplementary Fig. 2).
Patients achieving ctDNA negativity of canonical alterations showed a trend toward superior outcomes, with a numerically threefold higher ORR but no significant difference when compared to those with ctDNA positivity (60.0%, 95% CI: 35.2–84.8% vs. 20.0%, 95% CI: 5.0–71.6%, p = 0.303). The mPFS of canonical alterations mirrored the prognostic patterns observed with MET-specific monitoring, with prolonged survival benefits in the ctDNA negative cohort (Fig. 3D; Table 2).
Patients with pan-alterations ctDNA positivity also had poorer outcomes, with the ORR of 71.4% (95% CI: 29.0–96.3%) in ctDNA negative subgroup versus 38.5% (95% CI: 13.9–68.4%) in ctDNA positive subgroup (p = 0.350). The mPFS was 8.2 months (95% CI: 4.1-NA) in the ctDNA negative subgroup compared to 6.0 months (95% CI: 3.0-NA) in the ctDNA positive subgroup (HR = 0.73, 95% CI: 0.24–2.23; p = 0.542) (Fig. 3E; Table 2).
At the V1 timepoint, pan-alterations analysis continued to show the highest sensitivity (80%) for predicting treatment response, and its specificity increased compared to V0. However, the specificity and positive predictive value (PPV) remained modest, at 50% and 62%, respectively. Sub-clonal low-VAF variants may reduce specificity of pan-alterations strategy. In contrast, MET-specific alterations analysis demonstrated markedly improved specificity (90%) and PPV (83%) at V1 (Fig. 3F).
The impacts of dynamic changes of ctDNA on treatment efficacy
Longitudinal ctDNA trajectories further emphasized the prognostic values of molecular response. After 4-week treatment, 10 patients maintained persistent MET negativity (ctDNA negativity: negative at V0-negative at V1), 4 achieved MET clearance (ctDNA clearance: positive at V0-negative at V1), 5 exhibited sustained MET positivity and 1 acquired new MET positivity (ctDNA residual: positive at V0-positive at V1 and negative at V0-positive at V1) (Fig. 4A). Specifically, patients with MET-specific ctDNA residual exhibited markedly inferior survival, with a mPFS of just 2.2 months (95% CI: 2.2-NA) and an ORR of 16.7% (95% CI: 4.0–70.5%). In contrast, those achieving MET ctDNA clearance showed substantially prolonged mPFS (9.3 months; 95% CI: 6.0-NA) and best ORR (75.0%, 95% CI: 19.4–99.4%), while persistent MET-specific ctDNA negative patients had intermediate outcomes (mPFS: 6.8 months, 95% CI: 5.9-NA; ORR: 60.0%, 95% CI: 26.2–87.8%). Patients achieving MET-specific alterations clearance showed significantly prolonged mPFS compared to those with residual (mPFS: 9.3 vs. 2.2 months, p = 0.005). Similarly, the persistently negative group had significantly better outcomes than the residual group (mPFS: 6.8 vs. 2.2 months, p < 0.001). Although a numerical advantage was observed for the clearance group over the persistently negative group, this difference did not reach statistical significance (mPFS: 9.3 vs. 6.8 months, p = 0.291). (Fig. 4B-C; Table 2).
Sankey plots illustrating the evolution of ctDNA status and its association with the best of response (BOR) of the three profiling paradigms: A MET-specific alterations, D Canonical (tier I/II) alterations, and G Pan-alterations. Bar charts depicting the distribution of BOR stratified by ctDNA dynamic categories (persistent negativity, clearance, or residual) for (B) MET-specific, E Canonical, and H Pan-alterations, profiling. Kaplan-Meier curves for progression-free survival (PFS) based on dynamic ctDNA status for (C) MET-specific, F Canonical, and I Pan-alterations monitoring.
Post-treatment longitudinal profiling of canonical alterations closely mirrored MET-specific alterations (Fig. 4D), with only two discrepancies observed: one acquired EGFR T790M mutation (tier I/II driver variant) and one acquired non-canonical MET alteration. This minimal molecular divergence resulted in nearly identical predictive performance between canonical alterations profiling and MET-specific monitoring. 11 patients achieving persistent ctDNA negativity of canonical alterations demonstrated superior outcomes with an ORR of 54.5% (95% CI: 23.4–83.2%). This contrasted sharply with the 20.0% ORR (95% CI: 5.0–71.6%) observed in 5 patients with residual ctDNA. Notably, four patients achieving canonical alterations clearance attained the highest ORR (75.0%, 95% CI: 19.4–99.4%). The mPFS of canonical alterations mirrored the prognostic patterns observed with MET monitoring, with prolonged survival benefits in the ctDNA negative cohort and ctDNA clearance (Fig. 4E, F; Table 2).
Longitudinal pan-alterations dynamics also demonstrated prognostic value for mPFS. Among 20 evaluable patients, 2 maintained persistent pan-alterations ctDNA negativity, 5 achieved ctDNA clearance and 13 exhibited ctDNA residual (10 exhibited sustained pan-alterations positivity, and 3 developed new ctDNA positivity) (Fig. 4G). Patients with residual pan-alterations showed markedly inferior outcomes (mPFS: 5.9 months, 95% CI: 3.0–6.0; ORR: 38.5%, 95% CI: 14.7–65.2%). Conversely, those achieving ctDNA clearance demonstrate substantially prolonged survival (mPFS: 8.5 months, 95% CI: 4.1-NA; ORR: 60.0%, 95% CI: 14.7–94.7%), while persistently negative patients maintain intermediate outcomes (mPFS: 7.1 months, 95% CI: 6.0-NA; ORR: 100.0%, 95% CI: 15.8–100%). In the pairwise comparisons of mPFS, while numerical differences were observed between the groups, none reached statistical significance. The clearance group showed similar mPFS compared to the residual group (mPFS: 8.5 vs. 5.9 months, p = 0.862). Similarly, the persistently negative group demonstrated a non-significant survival advantage over the residual group (mPFS: 7.1 vs. 5.9 months, p = 0.780). The difference between the persistently negative and clearance groups was also not statistically significant (mPFS: 7.1 vs. 8.5 months, p = 0.429). (Fig. 4H–I).
Critically, MET-specific ctDNA clearance subgroup achieved the best outcome, and dynamic ctDNA monitoring demonstrated superior predictive value over single-timepoint analysis. MET-specific clearance emerged as the optimal indicator for treatment response, providing exceptional specificity.
Discussion
Our findings are consistent with prior work showing that ctDNA clearance predicts improved outcomes under targeted therapy, but further extend the field by directly contrasting three monitoring paradigms in MET exon 14 skipping NSCLC. This biomarker study of the EMBRACE trial provides supporting evidence for the clinical utility of dynamic ctDNA monitoring in patients with METex14 skipping alterations NSCLC receiving MET-TKI therapy.
In baseline plasma samples detection, nearly half of the patients with previously tissue-confirmed METex14 skipping alterations showed negative results for MET-specific alterations. This rate of ctDNA detection appears quite comparable to that reported in VISION trial, where only 74 out of 180 tissue-positive patients (41.1%) had detectable METex14 skipping in matched liquid biopsies. This observation likely reflects a combination of biological and analytical factors. Biologically, limited ctDNA shedding, often associated with low tumor burden, indolent metastatic behavior, or reduced tumor vascularity, can substantially diminish the detectability of known oncogenic drivers in plasma, as demonstrated in prior studies10,11. Analytically, differences between tissue-based and plasma-based assays, including methodological platforms (PCR versus NGS), input DNA requirements, and variant-type, dependent detection efficiencies, may also contribute to discordant findings12,13. In our cohort, tissue plasma concordance reached 100% among the eight patients with known METex14 junctions (Supplementary Table 3), indicating that assay sensitivity is unlikely to be a major driver of discordance when ctDNA shedding is adequate. Crucially, we show that ctDNA assessment at 4 weeks post-treatment initiation constitutes a clinically actionable strategy for early response evaluation. This aligns with accumulating evidence that early ctDNA dynamics can effectively complement RECIST-based imaging. In scenarios where radiologic findings are delayed, subtle, or equivocal, ctDNA provides an orthogonal molecular readout that may reveal treatment response or emerging progression earlier than conventional imaging14. Such early molecular signals hold particular value for refining on-treatment decision-making and may support timely therapeutic adjustments in clinical practice15,16.
Our comparative analysis of three monitoring paradigms, MET-specific alterations, canonical alterations, and pan-alterations, offers critical insights into the strengths and limitations of different approaches. MET-specific monitoring achieved the highest specificity, accurately identifying patients most likely to benefit from ensartinib. These findings suggest that when the expected prevalence of MET alterations is relatively high and the therapeutic pathway is clearly defined, selective MET single-gene testing offers distinct economic advantages. Compared with large multi-gene NGS panels, single-gene assays involve a simpler workflow, lower analytical costs, and more straightforward interpretation, making them particularly valuable in settings with limited sample availability or constrained diagnostic budgets17. Moreover, single-gene testing avoids the burden of “low-yield findings” frequently encountered in broad panels, namely, numerous low-frequency variants without actionable relevance, thereby improving the cost-to-benefit efficiency of molecular testing18. Collectively, these features render MET-specific monitoring both cost-saving and clinically practical. Canonical alterations profiling yielded comparable trends and allowed detection of emergent resistance variants, such as EGFR T790M, underscoring its utility in preemptive treatment adaptation. Notably, although EGFR T790M is a well-established mechanism of resistance to EGFR-TKIs, it is not considered a typical resistance mechanism to MET inhibitors. Given that all patients in our study were TKI-naïve, the emergence of T790M is more likely attributable to clonal evolution or expansion of a pre-existing subclone rather than representing true on-target resistance to ensartinib19. Isolated reports have also described the appearance of EGFR L858R/T790M or isolated T790M after treatment with non-EGFR targeted agents, highlighting that spatiotemporal tumor heterogeneity and therapy-induced selective pressures can give rise to new genomic events20. Thus, while canonical profiling can reveal such unexpected alterations, these findings should be interpreted cautiously in this context. By contrast, pan-alterations profiling provided the highest sensitivity for capturing molecular events but was limited by low specificity and reduced predictive accuracy for therapeutic response. Within this paradigm, only two patients remained persistently ctDNA-negative (both achieving PR; mPFS 7.1 months), and five achieved on-treatment ctDNA clearance (three PR and two SD; mPFS 8.5 months). Although both groups exhibited numerically better outcomes than patients without clearance (5/13 PR; mPFS 5.9 months), the differences were not statistically significant. This pattern is most likely attributable to the very small subgroup sizes and resulting limited statistical power. Biological factors may also contribute. Patients achieving broader pan-alterations clearance may harbor lower-burden or more genomically homogeneous disease, whereas others may retain diverse sub-clonal alterations. Such sub-clonal variants, typically present at low allele frequencies and confined to a minority of tumor cells, may not be uniformly eliminated even when dominant clones regress21. As a result, their dynamics may fail to fully reflect global tumor response, diminishing the prognostic resolution of pan-alteration-based clearance. Together, these findings suggest that an integrated strategy may be most informative: MET-specific monitoring for early efficacy evaluation, supplemented by broader profiling to detect resistance mechanisms.
Consistent with prior MET TKI studies, including the savolitinib8 and tepotinib9 trials, early MET clearance was associated with improved outcomes, reinforcing its validity as an effective biomarker. However, distinct mutation profiles emerged between the cohorts: the savolitinib-treated cohort was characterized by predominant DNMT3A mutations (11.4%), whereas our ensartinib-treated cohort exhibited enrichment in MUC16, HGF, ATRX, and NOTCH family alterations (10.0%). Notably, these non-MET mutations were uniformly cleared during the initial treatment response. This dynamic pattern suggests that ensartinib may exert its biological effect through simultaneous targeting of multiple pathways: direct suppression of MET signaling; modulation of collateral pathways, particularly NOTCH signaling22; epigenetic remodeling via ATRX-associated chromatin regulation23; immune microenvironment regulation24.
Paired molecular profiling of 11 plasma and 4 tissue samples revealed enriched resistance-associated alterations. Tissue analysis identified persistent MET mutations in 50% (2/4) and emergent TP53 mutations in 75% (3/4) of specimens. Plasma ctDNA profiling demonstrated recurrent genomic events, most frequently involving ARID1B (3/11, 27.3%), followed by HGF (2/11, 18.2%), persistent MET (2/11, 18.2%), and TP53 (2/11, 18.2%) alterations. Clinically, patients with detectable MET mutations at progression consistently exhibited primary lesion enlargement, whereas pan-mutation positive cases demonstrated heterogeneous progression patterns, including new brain metastases, central nervous system (CNS) progression, or systemic progression such as primary lesion enlargement. Longitudinal analysis further confirmed enrichment of specific resistance variants: recurrent TP53 missense mutations (c.473 G > A, c.839 G > T, c.701 A > G in 2/7 progression samples), ARID1B missense mutations (c.6365 G > A, c.1720 C > T, c.4697 G > A in 2/7), and HGF missense mutations (c.1187 G > A in 2/7), collectively suggesting distinct resistance trajectories (Supplementary Fig. 3).
Nevertheless, several limitations should be acknowledged. The sample size was modest, limiting statistical power and generalizability. The single-arm design precludes direct comparison with alternative therapeutic approaches, and overall survival data remain immature. Furthermore, this study was limited to ctDNA-based targeted sequencing. Future investigations could be enhanced by incorporating complementary analytical modalities, such as methylation profiling or fragmentomics, and by the addition of circulating tumor RNA assessment, which may improve the detection yield of MET exon 14 skipping alterations that can be missed by ctDNA testing alone. Finally, although the 4-week ctDNA assessment aligned with early radiological responses (with 85% and 75% of patients achieving PR at 4 weeks in our early phase data and the EMBRACE study), the optimal timing for molecular monitoring requires further prospective validation. We hypothesize that a 4-week ctDNA assessment is more informative for early efficacy evaluation than later timepoints. Future study designs will validate this by incorporating serial ctDNA measurements at 6 and 8 weeks, benchmarking them against long-term clinical outcomes to define the optimal monitoring schedule.
In conclusion, dynamic ctDNA monitoring provides an early indicator of treatment response in MET exon 14 skipping NSCLC receiving ensartinib in this exploratory cohort, with MET-specific profiling showing the highest predictive value, and broader profiling helping uncover resistance mechanisms. These findings warrant validation in larger prospective studies before implementation in routine clinical practice.
Methods
Study design
This multicenter, single-arm, open-label phase II trial (ChiCTR2100048767) assessed ensartinib (225 mg once daily) in 31 patients with histologically confirmed advanced or metastatic METex14 skipping mutation-positive NSCLC. METex14 alterations were confirmed in tumor tissue by reverse transcription-polymerase chain reaction or NGS. Patients were enrolled across six Chinese centers and were required to be MET TKI-naïve with documented disease progression after first-line chemotherapy and/or immunotherapy. Primary endpoint was investigator-assessed ORR, and the secondary endpoints included DCR, progression-free survival (PFS), duration of response (DoR) and safety profiles. The complete study design and efficacy endpoint results were previously reported3. Clinical follow-up and efficacy evaluation would be conducted for the enrolled patients at weeks 4, 8, and 16 after the initiation of the treatment, and then every 8 weeks thereafter. Each follow-up included safety assessments (physical examination, vital signs, Eastern Cooperative Oncology Group Performance Status, hematology, biochemistry, coagulation, and 12-lead electrocardiogram) and tumor evaluations. Imaging assessments consisted of chest computed tomography, abdominal ultrasonography, brain magnetic resonance imaging, superficial lymph node ultrasonography. In addition, contrast-enhanced computed tomography of the abdomen would be conducted every 16 weeks, and abdominal ultrasonography was not repeated during those visits. Tumor response was evaluated per RECIST 1.1 criteria, and disease staging followed the 8th edition of the AJCC/UICC TNM staging system. This study was approved by the Ethics Committees of the Second Affiliated Hospital of Zhejiang University School of Medicine, West China Hospital of Sichuan University, Peking University Cancer Hospital, Union Hospital of Tongji Medical College, Handan Central Hospital, and Hunan Cancer Hospital (protocol No. 2021-0683). The study complied with ethical regulations, including the Declaration of Helsinki. Written informed consent was obtained from all participants.
Sample collection and sequencing
Plasma samples were prospectively collected at V0 and V1. V0 samples were obtained after enrollment but prior to study drug initiation. V1 samples coincided with the completion of the first treatment cycle.
Ten-milliliter blood samples were collected in Cell-Free DNA (cfDNA) BCT Streck and centrifuged at 1600 × g to isolate plasma (upper layer) and white blood cells (WBC, middle layer). The supernatant was carefully transferred to a new tube and clarified by secondary centrifugation. Clarified plasma aliquots and WBC were stored at –80 °C. Subsequent cfDNA extraction, library preparation, sequencing, and data analysis were performed according to established protocols25. Briefly, cfDNA was extracted using the NucleoSnap cfDNA kit (Macherey-Nagel, Hilden, Germany). Matched genomic DNA from WBCs was simultaneously extracted and used as a germline and clonal hematopoiesis of indeterminate potential (CHIP) control to filter out non-tumor-derived variants. Quality control confirmed ≥10 ng of input DNA per sample for sequencing. The DNA samples were prepared for DNA library construction using the Hieff NGS® DNA Library Prep Kit (YEASEN, Shanghai, China) and captured by SureSelect XT-HS Target Enrichment System (Agilent Technologies, Santa Clara, USA). A 229-gene panel (Astrocyte Technology, Hangzhou, China) was utilized, and sequencing was performed using the Illumina NovaSeq-6000 platform (Illumina, San Diego, USA) at ~20,000× mean depth. Variant calling entailed the use of VarScan2 and Mutect2 for the detection of somatic mutations and copy number variations, respectively, in candidate genes. METex14 positivity was defined as ≥ 3 variant-supporting reads. Molecular responses were calculated based on the mean VAF.
The MET mutation burden was quantified as the total count of nonsynonymous MET mutations identified in ctDNA per patient. To minimize technical variability, the VAFs of MET alterations were normalized using the PyClone2 software. Longitudinal VAF fluctuations within individual patients were evaluated using the Wilcoxon signed-rank tests.
The stratification system for mutation analysis incorporated several distinct categories to provide a comprehensive genetic evaluation: (1) MET-specific alterations: MET-specific alterations include gene fusions (with breakpoints in intron 1, intron 13, and intron 14), amplification, single nucleotide variant (SNV), insertion-deletion (Indel), exon 14 skipping. (2) Canonical alterations: stratified by clinical significance: Tier I-Variants with strong clinical significance (Level A/B evidence); Tier II-Variants of potential clinical significance (Level C/D evidence)26. (3) Pan-alterations: all detectable alterations from the 229-gene panel were included to capture a broad spectrum of genetic alterations potentially influencing disease phenotypes or resistance mechanisms. Decision trees and specific targets for the three analytical strategies are detailed in the supplementary material. (Supplementary Figs. 4, 5)
Statistical analysis
Survival analysis was conducted using the Kaplan-Meier method to estimate PFS and OS. Differences in survival curves between groups using log-rank (Mantel-Cox) tests. Fisher's exact test was employed to compare the frequency data between the two groups. Continuous data were compared between groups using the t test. The relationship between continuous variables was assessed using Pearson correlation analysis. The association between prognostic factors and survival outcomes was primarily evaluated using univariable Cox regression analysis, which was the main statistical approach for this study. Multivariable Cox regression was also applied to explore and assess prognostic factors. HRs and 95% CIs were derived from these models. Proportional hazards assumptions were verified using Schoenfeld residual testing. A two-sided p < 0.05 was considered statistically significant. The ORR was calculated with exact 95% CIs using Clopper-Pearson method. PFS estimates and corresponding 95% CIs were generated using the Brookmeyer-Crowley approach. All statistical analyses were performed with SPSS 22.0 and R software (version 4.3.1), with statistical significance defined at a two-sided α level of 0.05.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Reis, H. et al. MET expression in advanced non-small-cell lung cancer: Effect on clinical outcomes of chemotherapy, targeted therapy, and immunotherapy. Clin. Lung Cancer 19, e441–e463, https://doi.org/10.1016/j.cllc.2018.03.010 (2018).
Han, Y. et al. Targeting MET in NSCLC: An ever-expanding territory. JTO Clin. Res Rep. 5, 100630. https://doi.org/10.1016/j.jtocrr.2023.100630 (2024).
Xia, Y. et al. Ensartinib for advanced or metastatic non-small-cell lung cancer with MET exon 14 skipping mutations (EMBRACE): A multi-center, single-arm, phase 2 trial. EClinicalMedicine 81, 103099. https://doi.org/10.1016/j.eclinm.2025.103099 (2025).
Xia, Y. et al. Potent antitumor activity of ensartinib in MET exon 14 skipping-mutated non-small cell lung cancer. Cancer Lett. 561, 216140. https://doi.org/10.1016/j.canlet.2023.216140 (2023).
Stejskal, P. et al. Circulating tumor nucleic acids: Biology, release mechanisms, and clinical relevance. Mol. Cancer 22, 15. https://doi.org/10.1186/s12943-022-01710-w (2023).
Rolfo, C. et al. Liquid biopsy for advanced NSCLC: A consensus statement from the international association for the study of lung cancer. J. Thorac. Oncol. 16, 1647–1662, https://doi.org/10.1016/j.jtho.2021.06.017 (2021).
Gray, J. E. et al. Longitudinal analyses of circulating tumor DNA for the detection of EGFR mutation-positive advanced NSCLC progression during treatment: Data From FLAURA and AURA3. J. Thorac. Oncol. 19, 1525–1538, https://doi.org/10.1016/j.jtho.2024.07.008 (2024).
Yu, Y. et al. Circulating tumour DNA biomarkers in savolitinib-treated patients with non-small cell lung cancer harbouring MET exon 14 skipping alterations: a post hoc analysis of a pivotal phase 2 study. Ther. Adv. Med. Oncol. 14, 17588359221133546. https://doi.org/10.1177/17588359221133546 (2022).
Rolfo, C. et al. Liquid and tissue biopsies for identifying MET exon 14 skipping NSCLC: Analyses from the phase II VISION study of tepotinib. Clin. Cancer Res. 31, 2675–2684, https://doi.org/10.1158/1078-0432.Ccr-24-4097 (2025).
Abbosh, C. et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 545, 446–451, https://doi.org/10.1038/nature22364 (2017).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 6, 224ra224, https://doi.org/10.1126/scitranslmed.3007094 (2014).
Kemper, M. et al. Liquid biopsies in lung cancer. Cancers (Basel) 15, https://doi.org/10.3390/cancers15051430 (2023).
Ko, E. Y. et al. The diagnostic accuracy of next-generation sequencing in advanced NSCLC. J. Liq. Biopsy 9, 100325. https://doi.org/10.1016/j.jlb.2025.100325 (2025).
Frank, M. S. et al. Circulating tumor DNA monitoring reveals molecular progression before radiologic progression in a real-life cohort of patients with advanced non-small cell lung cancer. Cancer Res. Commun. 2, 1174–1187, https://doi.org/10.1158/2767-9764.Crc-22-0258 (2022).
Assaf, Z. J. F. et al. A longitudinal circulating tumor DNA-based model associated with survival in metastatic non-small-cell lung cancer. Nat. Med. 29, 859–868, https://doi.org/10.1038/s41591-023-02226-6 (2023).
Dong, S. et al. Circulating tumor DNA-guided de-escalation targeted therapy for advanced non-small cell lung cancer: A nonrandomized controlled trial. JAMA Oncol. 10, 932–940, https://doi.org/10.1001/jamaoncol.2024.1779 (2024).
Loong, H. H. et al. Clinical and economic impact of upfront next-generation sequencing for metastatic NSCLC in East Asia. JTO Clin. Res. Rep. 3, 100290. https://doi.org/10.1016/j.jtocrr.2022.100290 (2022).
Brancato, D. et al. NGS approaches in clinical diagnostics: From workflow to disease-specific applications. Int. J. Mol. Sci. 26, https://doi.org/10.3390/ijms26199597 (2025).
Bean, J. et al. MET amplification occurs with or without T790M mutations in EGFR mutant lung tumors with acquired resistance to gefitinib or erlotinib. Proc. Natl. Acad. Sci. USA 104, 20932–20937, https://doi.org/10.1073/pnas.0710370104 (2007).
Kim, S. Y. et al. Metachronous development of L858R and T790M EGFR mutations following ALK inhibitor therapy in stage IV lung adenocarcinoma: A case report. Transl. Lung Cancer Res. 14, 1021–1031, https://doi.org/10.21037/tlcr-2024-1071 (2025).
McGranahan, N. & Swanton, C. Clonal heterogeneity and tumor evolution: Past, present, and the future. Cell 168, 613–628, https://doi.org/10.1016/j.cell.2017.01.018 (2017).
Sun, J., Dong, M., Xiang, X., Zhang, S. & Wen, D. Notch signaling and targeted therapy in non-small cell lung cancer. Cancer Lett. 585, 216647. https://doi.org/10.1016/j.canlet.2024.216647 (2024).
Hou, T. et al. Alpha thalassemia/intellectual disability X-linked deficiency sensitizes non-small cell lung cancer to immune checkpoint inhibitors. Front Oncol. 10, 608300. https://doi.org/10.3389/fonc.2020.608300 (2020).
Aithal, A. et al. MUC16 as a novel target for cancer therapy. Expert Opin. Ther. Targets 22, 675–686, https://doi.org/10.1080/14728222.2018.1498845 (2018).
Yang, Y. et al. Decoding the evolutionary response to ensartinib in patients with ALK-positive NSCLC by dynamic circulating tumor DNA sequencing. J. Thorac. Oncol. 16, 827–839, https://doi.org/10.1016/j.jtho.2021.01.1615 (2021).
Li, M. M. et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: A Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J. Mol. Diagn. 19, 4–23, https://doi.org/10.1016/j.jmoldx.2016.10.002 (2017).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grant Nos. 82370028, 82422001, and 82403366), and Beijing Xisike Clinical Oncology Research (Y-2024AZ(NSCLC)MS-0200).
Author information
Authors and Affiliations
Contributions
Y.X. conceived and designed the research, analyzed and interpreted the patient data. M.Z. and Y.Z. acquired the data, performed formal analysis, and were major contributors to writing the manuscript. X.Q.S. analyzed and interpreted the patient data and revised the manuscript for important intellectual content. Y.H. and R.J. acquired the data and conducted statistical analysis. P.W.T., J.Z., Y.J., Z.Y.G., and Y.C.Z. acquired and assembled the data. Y.R.M. and F.L. performed statistical analysis. W.L. secured funding and provided supervision. X.N.L. designed the research. H.F.S. analyzed and interpreted the patient data, revised the manuscript for important intellectual content, and contributed to the methodology. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, M., Zhu, Y., Sun, X. et al. Comparative analysis of ctDNA monitoring strategies in advanced NSCLC with MET exon 14 skipping mutations treated with ensartinib. npj Precis. Onc. 10, 110 (2026). https://doi.org/10.1038/s41698-026-01284-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01284-6