Abstract
T-cell receptor (TCR) repertoires are central to antitumor immunity, yet their dynamics in digestive system cancers remain poorly defined. We profiled TCR repertoires from 415 tumors in 145 patients with colorectal cancer (CRC, n = 96), gastric cancer (GC, n = 47), and hepatocellular carcinoma (LIHC, n = 2), integrating clinical and pathological features. Distinct repertoire architectures emerged: CRC was characterized by abundant TRB V–J combinations (e.g., TRBV10-2*00–TRBJ2-4*00), whereas GC showed higher abundance of TRG/TRD pairings (e.g., TRGV5P*00–TRGJP1*00, TRDV3*00–TRDJ1*00), reflecting tumor-specific immune surveillance. Conserved motifs (“CATWD,” “YKKLF”) across cancers indicate shared selective pressures, while antigen mapping revealed both common (KRAS, SF3B1, and BST2) and tumor-specific targets (MAGEA10, WT1 in CRC; PABPC1 in GC). In CRC, repertoire dynamics were tightly coupled to disease stage. Metastatic tumors (MT) displayed larger size, vascular invasion, and elevated serum markers, whereas primary tumors (PT) exhibited stronger immune infiltration with lymphocyte- and myeloid-driven responses. Tumor size was significantly and positively correlated with the number of TRD/TRG clonotypes shared between PT and MT. Shared clones were further classified into three categories, including stable, contracted, and expanded. Among these, expanded MT clones were dominated by the “NYGYTF” motif within the TRB chain (e.g., TRBV7-9*00–TRBJ1-2*00). The most abundant “NYGYTF”-containing clones recognized MLANA, a tumor-associated antigen linked to prognosis and therapeutic responsiveness, underscoring its potential role in CRC progression. Collectively, these findings delineate cancer- and stage-specific TCR repertoire alterations and antigen specificities, highlighting novel biomarkers and therapeutic targets to inform TCR-based diagnostics and personalized immunotherapies in CRC and GC.
Similar content being viewed by others
Introduction
The remarkable efficacy of immune checkpoint inhibitors (ICIs) across various tumors has highlighted the pivotal role of T cells in antitumor immunity1,2. Both the abundance and functional state of tumor-infiltrating lymphocytes (TILs) are closely associated with patient prognosis and response to immunotherapy3,4,5. Through their specific receptors, T cells recognize tumor antigens—a process essential for immune surveillance and tumor elimination6. Advancements in immune repertoire sequencing have enabled systematic analysis of the T-cell receptor (TCR) repertoire, offering new insight into the complex tumor immune microenvironment7.
In gastrointestinal cancers, T-cell-mediated immunity has gained increasing attention. For instance, in colorectal cancer (CRC), Galon et al. demonstrated that the “Immunoscore” is a strong predictor of patient prognosis8. In gastric cancer (GC), Zhao et al. showed that patients with mismatch repair deficiency (dMMR) respond favorably to immunotherapy, correlating with high TIL densities9. These findings highlight the need to better understand T-cell characteristics in gastrointestinal tumors. Additionally, the immune microenvironment in these cancers is shaped by sustained exposure to exogenous antigens, diverse microbiomes10,11, and abundant tissue-resident immune cells, all of which influence local T-cell composition and function.
Recent studies have further explored the role of the TCR in tumor immunity. Porciello et al. found that TCR clone diversity in CRC is associated with immunotherapy outcomes12, while Li et al. identified prognostic TCR clonotypes in liver hepatocellular carcinoma (LIHC)13. Borràs et al. reported higher TCR diversity in microsatellite instability (MSI) CRC tumors14. However, these studies have primarily focused on the TCR-β chain, providing valuable but incomplete insights into tumor-immune interactions15. Notably, most analyses center on αβ T cells, with limited investigation of γδ T cells, despite their important roles in gastrointestinal immunity16. Moreover, comparative analyses of TCR features across different gastrointestinal cancer types remain limited, even though tumors in close anatomical regions can exhibit markedly distinct immune landscapes17. In the realm of metastatic disease, researches have focused largely on primary tumors, leaving the dynamics of TCR repertoire differences during metastasis poorly understood. Mlecnik et al. identified significant immune differences between primary and metastatic CRC patients18,19, yet detailed investigations at the TCR level are lacking. Given that metastasis is a key determinant of patient outcomes, understanding immune changes in the metastatic setting is essential.
To address these gaps, we conducted comprehensive TCR sequencing on 145 patients with gastrointestinal cancers, profiling the β, γ, and δ TCR chains across CRC, GC, and LIHC, including both primary and metastatic CRC lesions. A subset of samples also included data for the α chain. In addition, we collected extensive clinical data for each patient (see Table S1). By integrating clinical characteristics and survival outcomes, this study aimed to: (1) systematically characterize TCR repertoire features across different gastrointestinal tumor types and identify potential antigens; (2) elucidate the clinical characteristics and changes of the TCR repertoire during CRC metastasis; and (3) explore the associations between TCR features, clinical outcomes, and antigens for shared clones in primary and metastatic CRC patients. These investigations may serve as a valuable source for guiding future efforts to advance precision immunotherapy in gastrointestinal cancers.
Results
Clonotypic specialization and diversity of T-cell receptor (TCR) chains in cancer patients
We profiled 145 cancer patients, generating 415 samples using the TCR-VDJ sequencing platform. The cohort included individuals with colorectal cancer (CRC, 96 cases), gastric cancer (GC, 47 cases), and liver hepatocellular carcinoma (LIHC, 2 cases), accompanied by comprehensive clinical data such as tumor type, tumor node metastasis (TNM) stage, vascular invasion status, distant metastasis status, and platelet levels (Fig. 1A, B; see Materials and methods; Table S1). TCR sequencing targeted V, D, and J segments of TCR α (TRA), β (TRB), δ (TRD) and γ (TRG) chains in T cells. These chains define two T-cell populations: αβ T cells, expressing TRA and TRB receptors, undergo thymic selection to recognize peptide-MHC complexes, driving adaptive immunity, while γδ T cells, expressing TRD and TRG receptors, mature differently to enable rapid, broad antigen responses that bridge innate and adaptive immunity20. We examined chain-specific clonotypic specialization patterns and their potential implications for tumor immunity. Initial analyses revealed distinct differences in clonal size distribution across TCR chains. The clone sizes of TRA and TRB were broadly distributed, primarily in the range of 1–1000, while those of TRD and TRG were mostly restricted to the range of 1–10. This pattern suggests that αβ T cells (TRA/TRB) are more prone to extensive clonal expansion, consistent with their role in adaptive immune responses, whereas γδ T cells (TRD/TRG) tend to undergo more limited expansion, possibly reflecting their involvement in specialized or innate-like immune functions (Fig. 1C). Complementarity-determining region 3 (CDR3) sequence length analysis further revealed chain-specific characteristics: TRD sequences peaked at 20 amino acids, whereas the other chains peaked around 15 amino acids, indicating a distinct structural feature of TRD that may underlie its specialized antigen recognition (Fig. 1D).
A Workflow of the study. B Circular plot illustrating sample grouping and TCR sequencing information. This study includes data from 145 clinical samples collected from patients with colorectal cancer (CRC), gastric cancer (GC), and liver hepatocellular carcinoma (LIHC). TCR-VDJ bulk sequencing was performed on these samples. The outermost circle represents tumor type, and the inner layers correspond to TCR α (TRA), TCR β (TRB), TCR δ (TRD), and TCR γ (TRG) chains, respectively. C Bar plot showing the clone size distribution of clonotypes detected in the TRA, TRB, TRD, and TRG receptor chains. D Bar plots showing the distribution of complementary determining region 3 (CDR3) lengths across TRA, TRB, TRD, and TRG chains. E Violin plots combined with box plots showing the Shannon and Simpson diversity indices for TRA, TRB, TRD, and TRG chains. P values for two-group comparisons were calculated using the two-sided Dunn’s test with Benjamini–Hochberg (BH) adjustment, and multi-group p value by the Kruskal–Wallis test. F Violin plots combined with box plots present the number of unique clonotypes, unique V genes, and unique J genes detected in TRA, TRB, TRD, and TRG chains. P values for two-group comparisons were obtained using the two-sided Dunn’s test with BH adjustment, and multi-group p value by the Kruskal–Wallis test. G Violin plots with box plots show TCR clonotype Shannon diversity across tumor node metastasis (TNM) stages. Two-group p values were obtained by two-sided Dunn’s test with BH adjustment, and multi-group p value by the Kruskal–Wallis test. H–L Violin plots combined with box plots showing the distribution of TCR clonotype Shannon diversity across various clinical subgroups, including metastatic state (H), vascular invasion (I), cancer types (J), preoperative platelet levels (K), and postoperative platelet levels (L). For subgroup comparisons, p values were calculated using two-sided t-tests (H, I, K, L) or Wilcoxon test (J).
Regarding clonotypic diversity, the TRB chain exhibited the trend of highest diversity index, followed by TRA, while the TRD chain showed the lowest diversity (Fig. 1E and Table S2; see Materials and methods). This pattern was consistent across unique clonotypes and V/J gene distributions, indicating that TRA and TRB chains could contribute more broadly to antigen recognition, whereas TRD and TRG chains might instead serve more specialized functions (Fig. 1E, F). Association with clinical data suggested that tumor progression (TNM stage) may influence TCR diversity between stage I and stage IV, with advanced-stage patients showing more complex clonotypic features (Fig. 1G; adjusted p value <0.05). For distant metastasis, patients with metastasis exhibited significantly higher diversity (Fig. 1H; p value <0.05). In the case of vascular invasion, patients with invasion showed significantly higher diversity (Fig. 1I; p value <0.05). No significant differences in TCR diversity were observed across cancer types, implying that clinical characteristics rather than tumor type may play a more prominent role (Fig. 1J). Platelet levels demonstrated a dynamic impact on TCR diversity. As shown in Fig. 1K, L, the Shannon diversity index differed significantly between patients with high and low platelet levels in both preoperative and postoperative settings (Fig. 1K, L; p value <0.05). Taken together, these observations point to chain-specific and clinically associated patterns of TCR repertoires in digestive system cancers.
Distinct patterns and cancer-specific insights into T-cell receptor (TCR) V-J gene combinations
To elucidate the characteristics of TCR gene combinations across cancer types, we analyzed V-J gene pairing patterns across TCR chains. Hierarchical clustering heatmaps revealed distinct V-J pairing differences within each chain (Fig. 2A). In the TRB chain, genes such as TRBV20-1, TRBV28, and TRBV19 frequently paired with TRBJ2-2 and TRBJ1-2. For the γδ TCR chains (TRG and TRD), TRGV8 and TRGV4 were commonly paired with TRGJ2 and TRGJP1, while TRDV3, TRDV2, and TRDV1 paired with their respective J genes. These non-random V-J combinations suggest that TCR combinations is governed by specific molecular mechanisms rather than occurring stochastically, further validating classical immunological principles and previous studies21,22 on TCR recombination biases. Analysis of V-J combination diversity further revealed chain- and cancer type-specific patterns (Fig. 2B). In the TRB chain, CRC exhibited a tendency toward higher diversity relative to GC and LIHC, but this difference did not reach statistical significance (p value >0.05). Likewise, diversity in the TRG chain appeared higher in GC and LIHC compared with CRC, yet no significant difference was observed. By contrast, the Shannon diversity index for TRD was significantly elevated in GC compared with CRC (p <0.05), suggesting a potential association between TRD V–J usage and the tumor immune microenvironment in GC. Notably, these findings should be interpreted with caution, as limited statistical power.
A Heatmap showing the frequency of V-J combinations across all samples. Color intensity reflects the log2-transformed number of clones detected for each V-J gene combination. Rows and columns are hierarchically clustered using the complete linkage method. B Violin plots combined with box plots showing the distribution of Simpson and Shannon diversity indices for V-J combinations across different tumor types. Panels represent: (left) T-cell receptor (TCR) β (TRB), (middle) TCR δ (TRD), and (right) TCR γ (TRG). P values were derived from two-sided Wilcoxon tests. C Bubble plots showing the abundances of V* (top) and J* (bottom) genes across cancer types. Red indicates enrichment, and green indicates depletion in specific cancer types, following the methodology of Zhang et al. (see Materials and Methods)39. D Density plot showing the specificity scores of V-J combinations across cancer types (see Materials and methods). The green line indicates the threshold used to define cancer-type-specific V-J combinations. E Bubble plot showing cancer-type-specific V-J combinations. Red indicates enrichment, and green indicates depletion, consistent with panel (C).
A detailed examination of V-J gene usage differences across cancer types revealed clear tissue-specific patterns (excluding LIHC hereafter due to limited sample size). The TRBV families exhibited higher frequencies in CRC, while TRDV and TRGV were more abundant in GC (Fig. 2C; top panel). Corresponding J gene distribution patterns mirrored these trends, emphasizing cancer-specific differences (Fig. 2C; bottom panel). To further identify cancer-specific V–J combinations, we applied a specificity score threshold of 2 (Fig. 2D; see Materials and methods) and identified several combinations strongly associated with specific cancer types (Fig. 2E and Table S3). Comparative analysis of Fig. 2C, E revealed that although the TRBV family showed broad representation in CRC, only certain combinations—such as TRBV10-2*00_TRBJ2-4*00 and TRBV12-1*00_TRBJ2-7*00—were specifically enriched, suggesting their potential roles in tumor immune responses. Similarly, in GC, although the TRDV and TRGV families were broadly expressed, specific combinations like TRDV2*00_TRDJ4*00 and TRGV5P*00_TRGJP1*00 were enriched, indicating their possible involvement in recognizing tumor-specific antigens. Importantly, we identified novel cancer-specific V–J combinations. For instance, TRGV5P*00_TRGJP1*00 and TRDV2*00_TRDJ4*00 showed dominant pairing in GC (Fig. 2E). Notably, although individual V or J genes in these combinations did not exhibit significant selection differences in Fig. 2C, their specific pairings were markedly enriched in a cancer-specific manner. This highlights the importance of V–J pairing patterns as more informative indicators of TCR-mediated antitumor immune responses than single gene preferences alone.
Collectively, these findings reveal distinct patterns of TCR gene combinations across cancer types and offer potential molecular targets for TCR-based diagnostics and immunotherapies.
Distinct complementarity-determining region 3 (CDR3) sequence characteristics and T-cell receptor (TCR) repertoire patterns across cancer types
We conducted a comprehensive analysis of CDR3 sequence characteristics across different cancer types, focusing on sequence features such as length distribution, amino acid composition, and conserved motifs. The density distribution of CDR3 lengths revealed similar patterns among TRB, TRD, and TRG chains (Fig. 3A; Kolmogorov–Smirnov (ks) test, p value >0.05); however, some chain-specific features were also evident. For TRB chains, peak density occurred at lengths between 15 and 20 in both CRC and GC. TRD chains demonstrated peak density at length 20. For TRG chains, CRC and GC showed high density distributions at lengths ranging from 15 to 30. Quantitative analysis of CDR3 length differences revealed specific enrichment patterns (Fig. 3B). Short sequences (5–13 amino acids) were more abundant in GC, which also demonstrated greater enrichment for lengths between 27 and 32, while CRC exhibited a distinctive abundance for sequences longer than 34 amino acids. These cancer-specific length differences may reflect distinct antigen recognition requirements in different tumor types. Analysis of amino acid usage patterns revealed some differences among cancer types (Fig. 3C). Amino acids such as A, R, N, Q, E, G, H, M, F, P, S, and Y were frequently used in CRC, whereas the remaining amino acids, including D, C, I, L, K, T, W, and V, were enriched in GC. However, the overall mean abundance of amino acids did not differ significantly between CRC and GC (Fig. 3D). Notably, previous studies23,24,25 have shown that the composition and distribution of amino acids within the CDR3 region are critical for determining TCR binding specificity and affinity. Although the differences we observed between CRC and GC were limited, they may still hint at the potential functional relevance of amino acid variability in antigen recognition.
A Density plots showing the distribution of CDR3 sequence lengths for T-cell receptor (TCR) β (TRB, left), TCR δ (TRD, middle), and TCR γ (TRG, right) chains. Colors represent different cancer types. Differences between distributions were assessed using the Kolmogorov–Smirnov (ks) test. B Bubble plot showing the cancer-type-specific differences for CDR3 lengths. Red indicates enrichment, and green indicates depletion, as described in Zhang et al. (see Materials and methods)39. C Grouped bar plot showing the average frequency of amino acids across cancer types. Frequency differences between cancers were assessed using the two-sided t-test. D Bubble plot showing cancer-type-specific amino acid distributions. Red indicates enrichment, and green indicates depletion. E Violin combined with box plot showing the distribution of conservation indices across cancer types (see Materials and methods). P value was calculated using a two-sided t-test. F Multiple sequence alignment of the top k most abundant CDR3 sequences in each cancer type. Gaps are denoted by “—”, and “X” indicates unidentified amino acids. G CDR3 sequence logo showing amino acid frequencies at each position (x-axis: positions 1–16; y-axis: information content in bits). Amino acids were depicted as letters, with their size proportional to frequency. Colors indicate chemical properties: acidic (red), basic (blue), hydrophobic (black), neutral (purple), and polar (green). H Heatmap showing cancer-specific CDR3 motifs in colorectal cancer (CRC) and gastric cancer (GC) patients. Redder shades indicate higher motif abundance. Rows and columns were hierarchically clustered using the “complete” method. I Heatmap showing the starting positions of cancer-specific motifs within CDR3 sequences. Redder shades indicate higher relative frequencies. Tau values represent the conservation index of motifs (see Materials and methods). J Violin combined with box plots depicting the clonal fraction distributions of cancer-specific motifs in CRC and GC. P values were calculated using two-sided Wilcoxon tests. K Bubble plot showing cancer-type-specific differences for conserved motif-associated V-J combinations. Red indicates enrichment, and green indicates depletion. L Venn diagram showing unique and shared antigen genes in CRC (purple) and GC (pink), based on VDJdb28 entries with vdjdb.score > 1. M Bar plot comparing log₂-transformed frequencies of antigen genes in CRC and GC from VDJdb (vdjdb.score >2). Bars are dodged. N, O Heatmaps showing the frequency of antigen genes versus TCR V–J combinations in (N) GC and (O) CRC from VDJdb (vdjdb.score >2). Rows represent antigen genes; columns represent V–J pairs. Frequencies are row-scaled and hierarchically clustered. P Network plot illustrating interactions among shared antigen genes (CRC and GC), major histocompatibility complex (MHC) alleles (mhc.a), and TCR V–J combinations from VDJdb (vdjdb.score >2). Nodes represent genes, MHC alleles, and V–J pairs; edge width reflects interaction frequency, and node size indicates connectivity.
Conservation analysis, which reflects the level of diversity or consistency of CDR3 sequences within the immune receptor repertoires, revealed that GC samples exhibited significantly higher (p value <0.05) sequence conservation compared to CRC (Fig. 3E and Table S4; see Materials and methods). This elevated conservation in GC suggests a more focused immune response, possibly indicating convergent evolution toward recognition of specific tumor antigens. Multiple sequence alignment of the most abundant CDR3 sequences identified highly conserved motifs “CATWD” at positions 1–5 and “YKKLF” at positions 11-15 across all cancer types (Fig. 3F, G; see Materials and methods). The preservation of these motifs across cancers underscores their fundamental role in TCR structure and function, consistent with previous studies26,27 highlighting the importance of these positions in antigen recognition. Analysis of cancer-specific motifs between GC and CRC revealed distinct patterns (Fig. 3H; see Materials and methods). Position differences analysis showed the most motifs occurred frequently at positions 5-10, while motifs beginning with “CA” were often appeared at position 1 (Fig. 3I). This positioning pattern aligns with the known structural requirements of TCR-antigen recognition, where certain positions maintain TCR folding integrity and others directly mediate antigen contact. Clonal analysis of cancer-specific motifs revealed significant differences in clone fractions between CRC and GC for motifs “ASSSK,” “DRPPY,” “QHSRR,” and “HSRRP” (Fig. 3J; t-test; p value <0.05). Furthermore, V-J gene usage analysis showed a higher abundance of TRB chain V-J combinations in CRC, whereas GC exhibited a greater abundance of TRG chain V-J pairings (Fig. 3K). These distinct V-J patterns suggest cancer-specific TCR repertoire development, possibly reflecting different immunological selection pressures in cancer types.
Using the VDJdb database28, we analyzed the antigen specificity associated with V-J gene combinations identified in CRC and GC (Fig. 3K; see Materials and methods). Applying a threshold of vdjdb.score >1, we detected 48 unique antigen genes shared between CRC and GC, with 132 antigens identified in CRC and 58 in GC (Fig. 3L). Under a more stringent criterion (vdjdb.score >2), 23 antigen genes were consistently shared between CRC and GC. These included well-characterized oncogenic and immunogenic targets such as KRAS, SF3B1, BST2, 5T4, NY-ESO-1, CDK4, BCL2L1, and IGF2BP2, reflecting conserved oncogenic pathways and immune recognition across these tumor types (Fig. 3M). CRC-specific antigens (n = 44) encompassed molecules such as MAGEA10, MUC1, MMP9, WT1, and the fusion gene DNAJB1-PRKACA. The presence of cancer-testis antigens (CTAs) and fusion genes highlights their value as highly specific tumor-associated antigens for immunotherapeutic targeting. Notably, PABPC1 was uniquely detected in GC but absent in CRC, implicating a GC-specific role potentially linked to mRNA stability regulation and identifying it as a novel candidate antigen warranting further validation for diagnostic and therapeutic applications. Distinct antigen interaction profiles were observed among specific V-J gene combinations: TRBV7-TRBJ2 and TRBV5-TRBJ2 demonstrated multiple antigen interactions predominantly in CRC, whereas TRBV9-TRBJ2, TRBV9-TRBJ1, TRBV27-TRBJ1, and TRBV10-TRBJ1 exhibited widespread antigen associations in GC (Fig. 3N, O). Furthermore, we constructed interaction networks integrating shared antigen genes, MHC molecules, and V-J gene pairs (Fig. 3P). These networks provide critical insights into the shared antigen recognition landscape in CRC and GC and may facilitate the identification of cross-cancer immunotherapeutic targets.
Clinical and immunological characteristics of colorectal cancer (CRC) metastasis
To investigate the clinical and immunological characteristics associated with CRC metastasis, we compared metastatic (MT, from primary sites with distant metastasis) and primary (PT, from primary sites without distant metastasis) CRC samples. Clinical analysis revealed a significantly higher proportion of vascular invasion in MT patients (Fig. 4A; Fisher’s exact test, p < 0.05) and a greater prevalence of tumors larger than 5 cm in diameter (Fig. 4A; Fisher’s exact test, p < 0.05). These findings align with prior studies identifying vascular invasion as a critical risk factor for CRC metastasis. In contrast, no significant difference in neural invasion was observed between MT and PT groups (Fig. 4A; Fisher’s exact test, p value >0.05), suggesting a weaker link between neural invasion and distant spread. Tumor size analysis further demonstrated significantly larger tumor volumes in the MT group (Fig. 4B; p value <0.05). TNM staging confirmed that all MT cases were classified as stage IV, whereas PT cases were predominantly staging I–III, consistent with clinical staging criteria (Fig. 4C). Microsatellite instability (MSI) analysis revealed a higher proportion of MSI-high (MSI-H) cases in the PT group, whereas all MT cases were microsatellite stable (MSS), corroborating previous findings29,30 that MSI-H CRCs are less likely to metastasis (Fig. 4D). Perioperative clinical indices showed several notable differences (Fig. 4E). Leukocyte counts were consistently higher in the PT group both pre- and postoperatively, with levels increasing after surgery in both groups. Neutrophil counts differed significantly between groups postoperatively, while lymphocyte counts differed preoperatively. Monocyte and platelet counts showed no significant differences. These observations underscore changes in the immune microenvironment during metastasis. Analysis of tumor markers showed significantly elevated levels of CEA, CA19-9, and CA72-4 in the MT group, aligning with their recognized prognostic roles in metastasis prediction (Fig. 4E).
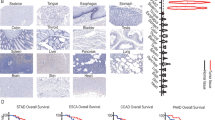
A Stacked bar plots showing the percentage distribution of metastatic tumors (MT, from primary sites with distant metastasis) and primary tumors (PT, from primary sites without distant metastasis) across different clinical phenotypes: vascular invasion (left), nerve invasion (middle), and tumor longest diameter (right). B Violin combined with box plot showing tumor size distribution between metastatic CRC and primary CRC. P values were calculated using the Wilcoxon test. C Stacked bar plot showing the distribution of tumor node metastasis (TNM) stages (stage 1 to 4) in metastatic CRC and primary CRC. D Stacked bar plot showing the distribution of microsatellite instability (MSI) stages in metastatic and primary CRC. MSI stages include MSI-high (MSI-H) and microsatellite stable (MSS). E Violin plots combined with box plots illustrating the distribution of different clinical characteristics in metastatic CRC and primary CRC. P values were calculated using the two-sided Wilcoxon tests. F Violin plots combined with box plots showing Shannon diversity between PT and MT in CRC patients (left), stratified by T-cell receptor (TCR) chains (right). P values were calculated using the t-tests. G Venn diagram showing the overlap and uniqueness of TCR clonotypes detected in PT and MT. H Heatmap showing the relative abundance of V-J gene combinations in PT (right) and MT (left). Color intensity represents the proportion of each combination relative to the total. I Heatmap comparing the distribution of V-J gene combinations between PT and MT. J Bubble plot illustrating the differences of V-J genes between PT and MT. Red indicates enrichment and green indicates depletion, based on the method of Zhang et al. (see Materials and methods). K Bubble plot showing the distribution of selected specific V-J gene combinations in the “The Cancer Genome Atlas Colorectal Adenocarcinoma” (TCGA-COAD) dataset, comparing signature scores between primary tumors and solid normal tissues. Redder colors indicate higher activity in a specific group. L Violin plots combined with box plots showing the distribution of signature scores for selected V-J combinations in primary tumors versus solid normal tissues in the TCGA-COAD dataset. M–Q Kaplan–Meier survival plots showing the association between selected V-J gene activity scores and patient prognosis. P values were calculated using log-rank tests.
TCR repertoire revealed metastasis-associated adaptive immune alterations (Fig. 4F; p < 0.05). The MT group exhibited greater clonal diversity (Shannon index), primarily driven by TRB chains, with a similar trend observed in TRG chains (Fig. 4F; p value = 0.059). This suggests enhanced antigen recognition complexity during metastasis, leading broader T-cell clonal expansion. V-J gene combination analysis indicated that approximately 92% of combinations were shared between MT and PT groups, reflecting a relatively stable TCR framework (Fig. 4G). However, TRB V-J combinations displayed higher abundances in the MT group, while TRG chain V-J combinations were more abundant in the PT group (Fig. 4H–J). To explore the expression pattern of PT-specific V-J combinations in normal tissues and their potential role in the immune environment, 19 PT-specific V-J combinations were identified and validated in the TCGA-COAD dataset, exhibiting higher activity in normal tissues (Fig. 4K, L; see Materials and methods). Among these, TRGV8_TRGJP2, TRGV8_TRGJP1, TRGV5_TRGJP1, TRGV3_TRGJP2, and TRGV3_TRGJ1 were significantly associated with favorable prognosis, with elevated expression correlating with improved patient outcomes (Fig. 4M–Q; see Materials and methods). Collectively, these findings suggest that specific V-J combinations may contribute to antitumor immunity and serve as potential prognostic biomarkers for CRC.
Alterations of T-cell clonality and T-cell receptor (TCR) repertoires in colorectal cancer (CRC) metastasis
Shared clone types—defined as clones with identical amino acid CDR3 sequences (aaSeqCDR3), V gene, and J gene—were analyzed between PT and MT to characterize T-cell clonal variations during CRC metastasis. The analysis revealed that TRG clones were predominant, followed by TRB clones, together accounting for over 96% of the total, while TRD clones comprised the smallest fraction (Fig. 5A). This distribution is consistent with prior reports31,32,33, emphasizing the role of γδ T cells within the colorectal cancer microenvironment. Clonal abundance correlation analysis demonstrated that β chains from αβ T cells (TRB clones) and δ chains from γδ T cells (TRD clones) exhibited significant positive correlations between PT and MT (Pearson correlation, R ≥ 0.26; two-sided t-test, p value <0.01), whereas γ chains from γδ T cells (TRG clones) showed no significant correlation (Fig. 5B). These findings suggest relatively stable clonal expansion among αβ T cells, contrasted with more dynamic remodeling of γδ T-cell clones during metastasis. Further evaluation of clonal frequency changes (log2-transformed) corroborated these observations: TRB clones demonstrated frequency changes predominantly within the range of –10 to 10, while TRD and TRG clones exhibited broader variations from –20 to 20. Notably, TRG clones displayed a multimodal distribution, indicative of pronounced fluctuations in abundance pre- and post-metastasis (Fig. 5C).
A Pie chart showing the distribution of shared clonotypes across TCR repertoires. Each color represents a different TCR chain. B Scatter plot illustrating the consistency of clone fractions for shared clonotypes between metastatic (MT, from primary sites with distant metastasis) and primary (PT, from primary sites without distant metastasis) CRC patients. Each dot represents an individual clonotype. “R” indicates the Pearson correlation coefficient, and p values were derived from two-sided t-tests. C Bar plot showing the distribution of clone frequency changes by TCR chain. The x-axis represents the log2 fold change in clone frequency between MT and PT samples. D, E Bubble plots showing differences in V (D) and J (E) gene usages between MT and PT patients. The y-axis represents the log2 fold change in gene representation. TCR chains are color-coded. F Heatmap showing V–J combination patterns in MT versus PT patients: (left) TCR β (TRB), (middle) TCR δ (TRD), and (right) TCR γ (TRG). Red indicates higher usage in MT. G Scatter plot showing the correlation between shared clone count and tumor size, categorized by TCR type. Each point represents an individual sample. “R” represents Pearson correlation, with p values obtained by t-tests. H Volcano plot highlighting differences in shared clonotypes between MT and PT patients. Clonotypes are categorized based on log2 fold change (|log2(FC)| >2) into stable, contracted, or expanded. “Stable” indicates no significant change, “contracted” denotes higher abundance in PT, and “expanded” indicates greater prevalence in MT. The stacked bar plot on the right shows the relative distribution of these categories. I Density plot showing the distribution of contracted, expanded, and stable clonotypes by log2 fold change. J Stacked bar plot showing the relative distribution of stable, contracted, and expanded clonotypes across different TCR chains (TRB, TRD, TRG). K Density plot illustrating the distribution of clone frequency changes between MT and PT patients along the log2 fold change axis. L Bar plot showing the top ten clonotypes with the highest abundance in the contracted and expanded categories. M Scatter plot showing the relationship between the clone sizes of expanded clonotypes in MT and PT patients. Each dot represents an individual expanded clonotype. “R” indicates the Pearson correlation coefficient, with p values obtained by t-test.
Analysis of V and J gene patterns revealed tissue-specific differences. TRB-related V and J genes were more abundant in MT, whereas TRD- and TRG-related genes were more frequent in PT (Fig. 5D, E; see Materials and methods). Notably, V-J combination analysis demonstrated strong enrichment of TRBV14*00_TRBJ1-5*00/TRBJ2-4*00, TRBV29-1*00_TRBJ1-6*00, and TRBV5-3*00_TRBJ1-2*00 combinations in MT (Fig. 5F). In contrast, TRD V-J combinations were more abundant in PT, while for TRG, only TRGV11*00_TRGJP*00 and TRGV7*00_TRGJP2*00 exhibited higher abundance in MT (Fig. 5F). An important observation was the significant positive correlation between tumor size and the number of shared TRD and TRG clones (Pearson correlation, R > 0.25; two-sided t-test, p < 0.01) (Fig. 5G). This correlation was consistent across metastatic sites, suggesting that γδ T-cell recruitment may be directly associated with tumor burden.
To further elucidate clonal variation patterns, we classified shared clones based on abundance changes (log2FC) between PT and MT into three categories: stable (−4≤ log2FC ≤4), contracted (log2FC <−4), and expanded (log2FC >4). Stable clones accounted for 40.4%, expanded clones for 25.2%, and contracted clones for 34.4%, reflecting clonal alterations during metastasis (Fig. 5H–K and Table S5; see Materials and methods). Representative sequences identified included CACDTVGGTADKLIF_TRDV2*00_TRDJ1*00 for contracted clones and CACDPLGDKRSDKLIF_TRDV2*00_TRDJ1*00 for expanded clones (Fig. 5L). Of particular interest, we observed a significant negative correlation between the abundance of TRD and TRG clones in PT and their degree of expansion during metastasis, whereas TRB clones showed no comparable correlation (Fig. 5M). This finding suggests that TRD and TRG clones with lower initial abundance in PT preferentially undergo selective expansion during metastasis, complementing the tissue-specific T-cell clonal expansion model.
Overall, these results provide insights into the alterations of T-cell clones during CRC metastasis and offer a deeper understanding of the distinct roles of T-cell subsets in tumor progression.
Investigation of complementarity-determining region 3 (CDR3) characteristics and different patterns in T-cell receptor (TCR) clones
We conducted an exploration of the CDR3 region of TCR clones to uncover the molecular features of expanded and contracted clones. Examination of CDR3 length distribution revealed a consistent peak density at ~15 amino acids across clone types, consistent with previously reported34,35 functional TCR CDR3 length distributions (Fig. 6A). This conserved length likely reflects spatial conformational constraints required for TCR binding to the pMHC complex. Amino acid composition analysis revealed chain-specific patterns (Fig. 6B). For the TRB chain, expanded and contracted clones exhibited similar amino acid usage frequencies. However, in the TRD and TRG chains, contracted clones demonstrated higher amino acid usage frequencies (Fig. 6B). These differences may indicate distinct functional requirements of T-cell subsets during immune responses. Hydrophobicity analysis further highlighted chain-specific differences: contracted clones in the TRD chain showed greater hydrophobicity, while some opposite trend was observed in the TRG chain (Fig. 6C; see Materials and methods; p value <0.05). These variations in hydrophobicity could influence TCR-antigen binding affinity, affecting clone fate.
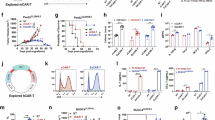
A Density plot showing the distribution of CDR3 lengths in expanded, contracted, and stable clones. B Bar plot showing amino acid usage frequency in expanded and contracted clones. Differences were evaluated using the two-sided t-test. C Density plot showing the distribution of CDR3 hydrophobicity in expanded and contracted clones. Differences were assessed using the Kolmogorov–Smirnov (ks) test. D SeqLogo showing the conserved motif distributions in expanded and contracted clones. E Bar plot highlighting the top 11 motifs in expanded clones with the highest log2 fold changes compared to contracted clones. F Bar plot showing the distribution of clones containing the NYGYTF motif, identified as the motif with the highest enrichment ratio. G Density plot showing the positional distribution of the NYGYTF motif within the CDR3 region of expanded and contracted clones. Differences were assessed using the Kolmogorov–Smirnov (ks) test. H Bar plot displaying the top 20 V − J gene combinations in clones containing the NYGYTF motif. I Kaplan–Meier survival curves showing the association between selected V − J gene activity scores and patient prognosis. P values were calculated using the log-rank test. J Sankey plot illustrating the flow from antigen genes (antigen.gene) to epitopes (antigen.epitope), major histocompatibility complex (MHC) alleles (mhc.a), and TCR V − J pairs in colorectal cancer (CRC), based on high-confidence Homo sapiens data from VDJdb28. Flow widths represent frequencies; data were filtered for specific V − J pairs. K Kaplan–Meier survival plot showing the association between MLANA expression and patient prognosis in the TCGA-COAD cohort. P value was calculated using the log-rank test. L Box plot showing the distribution of MLANA expression in tumor and normal samples from the “The Cancer Genome Atlas Colorectal Adenocarcinoma” (TCGA-COAD) cohort. (M) Violin plot showing the distribution of MLANA expression in tumor and normal samples from the GSE110224 cohort36. P value was obtained using the two-sided t-test.
Motif analysis identified a critical conserved sequence pattern (see Materials and Methods). While the first two positions of CDR3 sequences were generally conserved (dominated by “CA”), subsequent amino acids exhibited substantial variability (Fig. 6D). Notably, the motif NYGYTF was significantly enriched in expanded clones (Fig. 6E). Further investigation revealed positional differences within the CDR3 sequence: expanded clones showed a higher frequency of this motif at positions 6–8, whereas contracted clones predominated at positions 10–12 (Fig. 6F, G; p value <0.05). This positional specificity may reflect the role of TCR conformational changes in antigen recognition. V-J gene combination analysis revealed an intriguing pattern. Among clones containing the NYGYTF motif, the J genes were highly conserved as TRBJ1-2*00, despite diverse V genes (Fig. 6H and Table S6). This suggests a pivotal role for the J region in TCR function. The TRBV19*00-TRBJ1-2*00 combination was more common in expanded clones, while the TRBV28*00-TRBJ1-2*00 combination was prevalent in contracted clones (Fig. 6H). Survival analysis using the TCGA-COAD dataset demonstrated a correlation between specific V-J combinations and patient prognosis. Certain V-J combinations (TRBV9*00_TRBJ1-2*00, TRBV7-9*00_TRBJ1-2*00, TRBV5-1*00_TRBJ1-2*00, TRBV3-1*00_TRBJ1-2*00, TRBV28*00_TRBJ1-2*00) were associated with improved survival outcomes, underscoring the clinical relevance of these combinations (Fig. 6I; see Materials and methods).
Using the VDJdb database28, we mapped the antigen specificity of V-J pairs identified in Fig. 6H (see Materials and methods). Among these, eleven V-J pairs corresponded to antigen genes including BST2, IGF2BP2, MLANA, PMEL, and WT1 (Fig. 6J). BST2 encodes a transmembrane protein involved in antiviral responses and immune regulation, while IGF2BP2 is an RNA-binding protein implicated in mRNA stability and translational control, influencing cell proliferation and metabolism. MLANA (also known as MART-1) and PMEL are melanocyte differentiation antigens; MLANA is broadly recognized by HLA-A*02, with the highest frequency observed for the HLA-A*02–MLANA combination. These antigens play critical roles in melanoma immunobiology and are targets in cancer immunotherapy. WT1 encodes a transcription factor involved in cell growth and differentiation, often aberrantly expressed in various cancers, and considered a tumor-associated antigen. Given the immunological relevance of MLANA, we further investigated its prognostic significance using the TCGA-COAD cohort. High MLANA expression was significantly associated with improved overall survival in CRC patients (Fig. 6K). Consistently, MLANA expression was elevated in tumor tissues compared to adjacent normal tissues, a pattern validated in the independent GSE110224 dataset36 (Fig. 6L, M). The increased expression of MLANA within tumors may reflect enhanced immune recognition and antitumor activity, suggesting MLANA as a potential favorable prognostic biomarker in CRC.
Discussion
In this study, we investigated the clonotypic specialization, diversity, and gene rearrangement patterns of TCR chains in cancer patients across various tumor types, with a focus on CRC, GC, and LIHC. We observed differences in clonotypic expansion among TCR chains across the cancer cohort. The TRA and TRB chain displayed broad clonal expansion, consistent with its role in recognizing a wide range of antigens. In contrast, TRD and TRG chains exhibited more restricted expansion, primarily within small clones. These patterns are consistent with previous reports6,18,34 and reflect distinct functional roles of different TCR chains: while TRA and TRB are broadly involved in adaptive immune responses, TRD and TRG may mediate more specialized or tissue-resident immune functions. Notably, the lower diversity of clones in TRD chain compared to TRA and TRB supports its involvement in limited antigen recognition within specific immune contexts. We also observed a clear association between tumor progression and increased TCR diversity. Advanced-stage cancers, particularly those with metastasis or vascular invasion, exhibited broader TCR repertoires. This trend, consistent across cancer types, suggests that tumor-induced immune modulation—rather than tumor type—is a key driver of TCR repertoire evolution. Clinical parameters such as platelet levels further supported this dynamic; elevated platelet counts correlated with increased TCR diversity, implying that systemic factors like platelet activation contribute to immune plasticity during tumor progression and treatment.
Our analysis of V-J gene combinations revealed cancer-specific patterns. In CRC, TRB combinations—such as TRBV2*0-1 paired with TRBJ2-2—were more frequently expanded, suggesting their involvement in immune responses against colorectal tumors. In GC and LIHC, distinct TRGV and TRDV pairings (e.g., TRGV5P*00_TRGJP1*00 and TRDV2*00_TRDJ4*00) were enriched, indicating the potential of these gene combinations to mediate recognition of tumor-specific antigens. These findings support the notion that the TCR repertoire is shaped by selective pressures within the tumor microenvironment, rather than by random recombination events. Further characterization of CDR3 sequence features revealed cancer-specific patterns in both length distribution and amino acid composition. In GC, we observed enrichment of specific amino acids (e.g., D, C, I, L, K, T, W, V) and conserved motifs such as “CATWD” and “YKKLF” across multiple cancer types, suggesting structural and functional constraints in TCR-antigen binding. Importantly, our identification of shared tumor antigens—including KRAS, SF3B1, BST2, 5T4, NY-ESO-1, CDK4, BCL2L1, and IGF2BP2—in both GC and CRC underscores the translational value of cross-tumor antigenic overlap. These shared antigens provide promising targets for pan-cancer immunotherapies. Notably, we also identified PABPC1 as a novel, GC-enriched candidate antigen. Given its involvement in mRNA metabolism and previous associations with tumor proliferation and immune evasion, PABPC1 may represent a previously unrecognized immunogenic target in GC. Its tumor specificity and functional relevance position it as a compelling candidate for further investigation in the context of antigen discovery and vaccine-based or adoptive T-cell therapies.
In the context of CRC metastasis, we identified distinct immunological and clinical differences between PT and MT. MT exhibited greater clonal diversity, particularly in the TRB chains, suggesting a more complex antigenic landscape likely shaped by tumor heterogeneity, immune editing, or escape mechanisms. The increased presence of γδ T cells in MT, along with their correlation with tumor burden, reinforces their proposed role in metastatic immune surveillance. Notably, we observed prognostically relevant V-J combinations (e.g., TRGV8_TRGJP2) and enriched tumor-associated antigens such as MLANA. MLANA (Melan-A/MART1) is traditionally associated with melanocytic tumors, but its ectopic expression in gastrointestinal malignancies, including CRC metastases, highlights its broader immunogenic potential. The presence of MLANA in metastatic CRC suggests a convergence toward shared antigenic targets across distinct tumor types and disease stages. This ectopic expression may reflect either lineage plasticity or adaptive transcriptional reprogramming during metastasis, both of which could enhance immune visibility.
Despite these insights, several limitations remain. Limited sample sizes, particularly for LIHC and TRA chains, constrain generalizability. The absence of paired tissues (normal, primary tumor, and metastatic sites) precludes direct comparisons of tumor immune repertoires and hinders the capture of their evolutionary dynamics. Furthermore, the lack of paired TRA-TRB sequencing and functional validation of TCR clones and their antigen targets restricts interpretation of their antitumor activity. Future work should address these gaps through expanded cohorts, paired sequencing, and functional assays to enhance understanding of TCR clonal variations in the tumor immune microenvironment. Overall, our study provides a comprehensive characterization of TCR features in gastrointestinal cancers, delineates their differences during tumor progression and metastasis, and lays a foundation for the development of innovative immunotherapeutic strategies.
Methods
Study participants
From 2018 and 2024, 145 fresh tumor tissue samples were collected from patients undergoing surgical resection at the Third Affiliated Hospital of Shandong First Medical University (Affiliated Hospital of Shandong Academy of Medical Sciences). The cohort comprised patients with LIHC (2 cases), GC (47 cases), and CRC (96 cases). Among these, 13 CRC patients and 1 GC patient presented with distant metastasis. Importantly, paired tumor samples collected from the same patients before and after the development of distant metastasis were not available for analysis. (see Table S1 for details). Following surgical excision, tumor specimens were immediately rinsed with ice-cold saline following excision and promptly cryopreserved in liquid nitrogen.
Inclusion criteria for the study were: (1) histopathological confirmation of primary colon, liver, or gastric cancer and (2) availability of comprehensive clinical data. Exclusion criteria were: (1) receipt of radiotherapy, chemotherapy, immunotherapy, or targeted therapy prior to surgery; (2) a history of other malignant tumors; or (3) the presence of autoimmune diseases or chronic conditions. Detailed clinical information is provided in Table S1. This study was conducted in accordance with the Declaration of Helsinki and was approved by the Ethics Committee of the Third Affiliated Hospital of Shandong First Medical University (Approval No. FY2021018). Written informed consent was obtained from all participants prior to enrollment.
RNA extraction, cDNA synthesis, and T-cell receptor (TCR) library construction from tumor tissues
Tumor tissue specimens (at least 2 mL) were collected in EDTA vacutainer tubes to preserve RNA integrity. Total RNA was isolated using the RNAsimple Total RNA Kit (DP419, Tiangen Biotech, Beijing, China) according to the manufacturer’s instructions. RNA concentrations were quantified using a NanoDrop ND-2000 spectrophotometer (Thermo Scientific, UK). Complementary DNA (cDNA) was synthesized, and multiplex Polymerase Chain Reaction (PCR) was conducted to amplify rearranged TCR α, β, δ, and γ chain sequences, employing Immune Repertoire Library Preparation Kits (Geneway, Jinan, China) as described in a previously established method37. TCR libraries were sequenced on the DNBSEQ-T7 platform (MGI, Shenzhen, China), producing paired-end reads of 150 bp.
Preprocessing T-cell receptor (TCR) sequencing data
FASTQ files served as the initial format for sequencing data, with sample-specific index primers guiding the demultiplexing process. Following stringent quality control to eliminate sequences of low quality, the retained reads were mapped to the V, D, and J segments of TCR α (TRA), β (TRB), δ (TRD), and γ (TRG) chains (Table S1). This analysis utilized MiXCR (version 4.3.2)38, operating with its standard settings for both alignment and clonotype identification. The IMGT database (http://www.imgt.org/vquest/refseqh.html) provided the reference sequences for TCR gene annotation. Notably, αβ and γδ T cells follow distinct differentiation pathways: αβ T (expressing and TRB receptors) cells mature through positive and negative selection in the thymus, primarily recognizing peptide antigens presented by major histocompatibility complex (MHC) molecules and mediating specific adaptive immune responses; γδ T cells (expressing TRD and TRG receptors) mature along a unique pathway, enabling rapid responses to a broad range of antigens and bridging innate and adaptive immunity for immediate defense.
For each sample’s MiXCR output alignment file, we used R program (version 4.3.2) to read the data and generate a list, where each element is a data frame containing all the columns from the MiXCR output. Clones with CDR3 sequence lengths shorter than 6 amino acids were removed. The resulting filtered list was then used for downstream analyses.
Assessing T-cell receptor (TCR) clonotype and V-J gene combination diversity
To evaluate TCR clonotype or V-J combination diversity, Simpson’s and Shannon diversity metrics were computed, providing insights into the distribution of clonotypes and V-J combinations.
Simpson’s diversity index (D)
This metric quantifies the probability that two randomly selected TCR sequences or V-J combinations belong to different clonotypes or V-J combinations. It is defined as:
Shannon diversity index (H)
Shannon’s index captures the uncertainty or entropy in clonotype or V-J combination distributions. It is calculated as:
where \({p}_{i}\) is the proportion of the ith clonotype or V-J combination.
Differences analysis across different groups
To quantify the differences of elements across different groups, the number of elements observed and those expected (calculated using the Chi-square test) within each group were compared39. An element was considered enriched in a given group if the observed-to-expected ratio (Ro/e) was above 1.
“Observed” refers to the actual count of elements across various groups, while “Expected” denotes the frequencies anticipated if there were no relationship between the elements and groups. Expected frequencies were calculated by multiplying the respective row and column totals and dividing by the total number of elements.
Calculation of specificity scores among cancer types or V-J gene combinations
The specificity score was computed for each V-J gene combination (or amino acid sequence (AA) position, as shown in Fig. 3I) as the sum of the normalized frequencies across all cancer types:
Where N is the total number of cancer types within the group, fi is the frequency of expression for cancer i. fmax is the maximum frequency observed in the group. This sum of specificities is normalized by N − 1, which accounts for the total number of cancers minus one. The value S quantifies the relative specificity of V-J combinations (AA positions) across cancers, with higher specificity indicating more concentrated combinations or positions in fewer cancers. Notably, a specificity score (S) greater than 2 for a V-J combination (or AA position) is considered indicative of specificity.
Complementarity-determining region 3 (CDR3) sequences conservation analysis
The conservation analysis of CDR3 sequences was conducted using the Levenshtein distance metric. For each cancer type, 500 CDR3 sequences were randomly sampled, and pairwise distances were calculated using the stringdistmatrix function from “stringdist” package (version 0.9.12)40. A conservation score was obtained by calculating the mean of the upper triangular values of the resulting distance matrix. To ensure robustness, this sampling and computation process was repeated 10 times for each cancer type. The resulting conservation scores were compiled along with their corresponding cancer type labels for subsequent statistical analysis.
Analysis of dominant complementarity-determining region 3 (CDR3) patterns across cancer types
To identify and analyze dominant CDR3 patterns across cancer types, we implemented a comprehensive sequence analysis pipeline. First, V-J gene combinations were constructed by concatenating the best-matched V and J gene segments. For each cancer type, CDR3 sequences were grouped, and their frequencies were calculated. Sequence counts were then normalized to obtain relative frequencies within each cancer group. The top ten most abundant CDR3 sequences from each cancer type were selected based on their relative frequencies. These sequences were aligned using ClustalW via the “msa” R package (version 1.36.1)41. Prior to alignment, underscore characters were replaced with “X” to ensure compatibility with the alignment algorithm. The aligned sequences were visualized using the “ggmsa” package (version 1.10.0)42, with sequences ordered numerically by cancer type (top k per cancer type)”.
Motif analysis and cancer-type specificity assessment
To investigate CDR3 sequence motifs and their cancer-type specificity, we performed a two-step analysis consisting of motif extraction and specificity quantification. For motif extraction, a sliding window approach was employed to systematically identify all possible subsequences of length five (motif length) within each CDR3 sequence. To ensure computational efficiency, datasets containing more than 10,000 sequences were randomly down-sampled to 10,000 sequences for analysis. The frequency of each unique motif was recorded using a counting algorithm.
Motif specificity across cancer types was quantified using the previously defined specificity score, which ranges from 0 to 1, with a score of 1 indicating complete specificity to a particular cancer type. Frequencies were normalized against the maximum observed frequency, and safeguards were implemented to prevent division-by-zero errors. This analysis was conducted independently for each sample, yielding a comprehensive list of motif frequencies along with their corresponding cancer-type specificity scores.
Survival analysis
Overall survival analysis was performed using the R packages “survminer” (version 0.4.9)43 and “survival” (version 3.7-0)44. CRC Patient data from “The Cancer Genome Atlas Colorectal Adenocarcinoma” (TCGA-COAD) were stratified into high- and low-risk groups using an optimal cut-point determined by the surv_cutpoint function. Kaplan–Meier survival curves were generated using the ggsurvplot function, with statistical significance assessed by log-rank test. Risk tables were included to display the number of patients at risk during different time points. Survival analyses were performed across different cluster classifications based on vital status and survival time (i.e., days to death or last follow-up).
Signature scoring for V-J combination genes
TCGA-COAD transcriptomic and clinical data were organized using the Seurat object format. Signature scores for specific V-J combination genes were calculated using the AddModuleScore function from the “Seurat” R package (version 4.3.0)45.
Classification of clonal patterns in colorectal cancer (CRC) patients
To characterize clonal patterns between primary (from primary sites without distant metastasis) and metastatic (from primary sites with distant metastasis) CRC samples, we calculated the fold change in clone frequencies. For each shared clone, the log2 fold change was computed by comparing its frequency in metastatic versus primary samples. To avoid undefined values from zero frequencies, a small pseudo-count (1e-10) was added to both frequencies. Based on their log2 fold change values, clones were then categorized as follows: clones with log2 fold change >4 were classified as “Expanded” (representing >16-fold increase), those with log2 fold change <–4 were classified as “Contracted” (representing >16-fold decrease), and all others as “Stable”.
Identification of motifs in contracted/expanded clonotypes
To identify motifs associated with contracted or expanded clonotypes, CDR3 sequences corresponding to each group were exported into separate FASTA files using the writeXStringSet function from the “Biostrings” package (version 2.72.1)46. These FASTA files were then analyzed using the MEME tool (version 5.4.1)47 to detect potential sequence motifs. The motif length was set to 3–8 amino acids, with all other parameters left at their default settings.
Investigation of antigen genes associated with T-cell receptor (TCR) V–J combinations
To investigate antigen genes associated with T-cell receptor (TCR) V-J combinations, we analyzed TCR-antigen interaction data from the VDJdb database (version 2025-02, accessed via https://vdjdb.cdr3.net)28. The dataset, containing TCR sequences and their antigen specificities, was filtered to include only human-derived entries (antigen.species = Homo sapiens). A predefined set of V-J gene combination was used to select TCRs. A TCR V-J identifier was generated by concatenating variable (v.segm) and joining (j.segm) gene segments for each TCR. The filtered data were aggregated by grouping entries by antigen gene (antigen.gene), epitope (antigen.epitope), MHC allele (mhc.a), and TRVJ, calculating the frequency of each combination. Only TCRs matching the predefined V-J combinations were included to focus on interactions.
Statistical analysis
All statistical analyses were conducted in R (version 4.3.2). Two-sided Student’s t-test, Fisher’s exact test, and Wilcoxon rank-sum test were used to assess differences between two groups. For comparisons across multiple groups, the Kruskal–Wallis test was applied, followed by pairwise Dunn’s tests with p values adjusted using the Benjamini–Hochberg (BH) method to control the false discovery rate. Distributional differences were evaluated with the two-sided Kolmogorov–Smirnov (KS) test. Pearson correlation was used to examine associations between selected variables, and survival differences were analyzed using the log-rank test.
Data availability
The raw data generated in this study has been deposited at https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1205408. Publicly available data utilized in this research were obtained from the TCGA-COAD transcriptomic expression dataset and associated clinical data. In addition, the GSE110224 dataset36 was obtained from the GEO database and comprises 17 normal tissue samples and 17 colorectal cancer (CRC) tumor samples.
Code availability
All the code used for the analysis can be accessed by the corresponding author on reasonable request.
References
Naimi, A. et al. Tumor immunotherapies by immune checkpoint inhibitors (ICIs); the pros and cons. Cell Commun. Signal. 20, 44 (2022).
Vafaei, S. et al. Combination therapy with immune checkpoint inhibitors (ICIs); a new frontier. Cancer Cell Int. 22, 1–27 (2022).
Bassez, A. et al. A single-cell map of intratumoral changes during anti-PD1 treatment of patients with breast cancer. Nat. Med. 27, 820–832 (2021).
Du, K. et al. Pathway signatures derived from on-treatment tumor specimens predict response to anti-PD1 blockade in metastatic melanoma. Nat. Commun. 12, 1–16 (2021).
Hammerl, D. et al. Spatial immunophenotypes predict response to anti-PD1 treatment and capture distinct paths of T cell evasion in triple negative breast cancer. Nat. Commun. 12, 1–13 (2021).
Joyce, J. A. & Fearon, D. T. T cell exclusion, immune privilege, and the tumor microenvironment. Science 348, 74–80 (2015).
Valkiers, S. et al. Recent advances in T-cell receptor repertoire analysis: bridging the gap with multimodal single-cell RNA sequencing. Immunoinformatics 5, 100009 (2022).
Galon, J. et al. Towards the introduction of the ‘Immunoscore’in the classification of malignant tumours. J. Pathol. 232, 199–209 (2014).
Zhao, P., Li, L., Jiang, X. & Li, Q. Mismatch repair deficiency/microsatellite instability-high as a predictor for anti-PD-1/PD-L1 immunotherapy efficacy. J. Hematol. Oncol. 12, 1–14 (2019).
Yarahmadi, A. & Afkhami, H. The role of microbiomes in gastrointestinal cancers: new insights. Front. Oncol. 13, 1344328 (2024).
Zhao, L.-Y. et al. Role of the gut microbiota in anticancer therapy: from molecular mechanisms to clinical applications. Signal Transduct. Target. Ther. 8, 201 (2023).
Porciello, N., Franzese, O., D’Ambrosio, L., Palermo, B. & Nisticò, P. T-cell repertoire diversity: friend or foe for protective antitumor response? J. Exp. Clin. Cancer Res. 41, 356 (2022).
Li, R. et al. T-cell receptor sequencing reveals hepatocellular carcinoma immune characteristics according to Barcelona Clinic liver cancer stages within liver tissue and peripheral blood. Cancer Sci. 115, 94–108 (2024).
Borràs, D. M. et al. Single cell dynamics of tumor specificity vs bystander activity in CD8+ T cells define the diverse immune landscapes in colorectal cancer. Cell Discov. 9, 114 (2023).
Wang, H. et al. Characterization of the T-cell receptor repertoire associated with lymph node metastasis in colorectal cancer. Front. Oncol. 14, 1354533 (2024).
Ma, R., Yuan, D., Guo, Y., Yan, R. & Li, K. Immune effects of γδ T cells in colorectal cancer: a review. Front. Immunol. 11, 1600 (2020).
Conway. J. W. et al Unveiling the tumor immune microenvironment of organ-specific melanoma metastatic sites. J. Immunother. Cancer 10, e004884(2022),
Bruni, D., Angell, H. K. & Galon, J. The immune contexture and immunoscore in cancer prognosis and therapeutic efficacy. Nat. Rev. Cancer 20, 662–680 (2020).
Mlecnik, B. et al. The tumor microenvironment and Immunoscore are critical determinants of dissemination to distant metastasis. Sci. Transl. Med. 8, 327ra326–327ra326 (2016).
Li, Y. et al. Unraveling the spatial organization and development of human thymocytes through integration of spatial transcriptomics and single-cell multi-omics profiling. Nat. Commun. 15, 7784 (2024).
Tanno, H. et al. Determinants governing T cell receptor α/β-chain pairing in repertoire formation of identical twins. Proc. Natl. Acad. Sci. USA 117, 532–540 (2020).
George, A. J., Stark, J. & Chan, C. Understanding specificity and sensitivity of T-cell recognition. Trends Immunol. 26, 653–659 (2005).
Vujovic, M. et al. T cell receptor sequence clustering and antigen specificity. Comput. Struct. Biotechnol. J. 18, 2166–2173 (2020).
Wong, W. K., Leem, J. & Deane, C. M. Comparative analysis of the CDR loops of antigen receptors. Front. Immunol. 10, 2454 (2019).
Mayer-Blackwell, K. et al. TCR meta-clonotypes for biomarker discovery with tcrdist3 enabled identification of public, HLA-restricted clusters of SARS-CoV-2 TCRs. Elife 10, e68605 (2021).
Joglekar, A. V. & Li, G. T cell antigen discovery. Nat. Methods 18, 873–880 (2021).
Xu, X., Li, H. & Xu, C. Structural understanding of T cell receptor triggering. Cell. Mol. Immunol. 17, 193–202 (2020).
Goncharov, M. et al. VDJdb in the pandemic era: a compendium of T cell receptors specific for SARS-CoV-2. Nat. Methods 19, 1017–1019 (2022).
Jin, Z. et al. Outcome of mismatch repair-deficient metastatic colorectal cancer: the Mayo Clinic experience. Oncologist 23, 1083–1091 (2018).
Laplante, P. et al. Effect of mismatch repair deficiency on metastasis occurrence in a syngeneic mouse model. Neoplasia 62, 101145 (2025).
Yu, L. et al. Tumor-infiltrating gamma delta T-cells reveal exhausted subsets with remarkable heterogeneity in colorectal cancer. Int. J. Cancer 153, 1684–1697 (2023).
Pan, L. et al. Progress of research on γδ T cells in colorectal cancer. Oncol. Rep. 52, 160 (2024).
Wang, Y., Xu, Y., Chen, H., Zhang, J. & He, W. Novel insights based on the plasticity of T cells in the tumor microenvironment. Explor. Immunol. 2, 98–132 (2022).
Stadinski, B. D. et al. Hydrophobic CDR3 residues promote the development of self-reactive T cells. Nat. Immunol. 17, 946–955 (2016).
Stern, J. N. et al. Peptide 15-mers of defined sequence that substitute for random amino acid copolymers in amelioration of experimental autoimmune encephalomyelitis. Proc. Natl. Acad. Sci. USA 102, 1620–1625 (2005).
Vlachavas, E. I. et al. Radiogenomic analysis of F-18-fluorodeoxyglucose positron emission tomography and gene expression data elucidates the epidemiological complexity of colorectal cancer landscape. Computat. Struct. Biotechnol. J. 17, 177–185 (2019).
Ye, X. et al. High-throughput sequencing-based analysis of T cell repertoire in lupus nephritis. Front. Immunol. 11, 1618 (2020).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Liu, Y. et al. Immune phenotypic linkage between colorectal cancer and liver metastasis. Cancer Cell 40, 424–437.e5 (2022).
Van der Loo, M. P. The stringdist package for approximate string matching. R. J. 6, 111 (2014).
Bodenhofer, U., Bonatesta, E., Horejš-Kainrath, C. & Hochreiter, S. msa: an R package for multiple sequence alignment. Bioinformatics 31, 3997–3999 (2015).
Zhou, L. et al. ggmsa: a visual exploration tool for multiple sequence alignment and associated data. Brief. Bioinform. 23, bbac222 (2022).
Kassambara, A., Kosinski, M., Biecek, P. & Fabian, S. Package ‘survminer’. Drawing survival curves using ‘ggplot2’(R package version 0.3.1) 2017.
Brown, S. J., Goetzmann, W. N. & Ross, S. A. Survival. J. Financ. 50, 853–873 (1995).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Lifschitz, S. et al. Bio-strings: a relational database data-type for dealing with large biosequences. BioTech 11, 31 (2022).
Bailey, T. L., Johnson, J., Grant, C. E. & Noble, W. S. The MEME suite. Nucleic Acids Res. 43, W39–W49 (2015).
Acknowledgements
This work was supported by Shandong Provincial Natural Science Foundation (No. ZR2025MS1517).
Author information
Authors and Affiliations
Contributions
Y.n.Z conceived the study, applied for ethical approval, provided administrative support, and finalized the revised manuscript. L.L. conducted data analysis and drafted the manuscript, while J.L. organized clinical data, performed statistical analyses, and prepared the figures. F.W. collected and processed specimens and contributed to manuscript drafting, and R.J. carried out data analysis and figure preparation. H.W. was responsible for specimen processing, quality control, and guiding the writing and revision of the manuscript. X.L. performed data analysis and contributed to revising the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, L., Li, J., Wang, F. et al. T-cell receptor clonotypic diversity and specialization in digestive system cancers. npj Precis. Onc. 10, 91 (2026). https://doi.org/10.1038/s41698-026-01294-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01294-4