Abstract
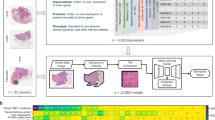
Molecular subtypes in prostate cancer significantly influence disease characteristics and treatment outcomes, yet obtaining this information requires specialized molecular testing. In this study, we develop and validate artificial intelligence models that can predict PAM50 and Prostate Subtyping Classifier (PSC) molecular classifications directly from standard hematoxylin and eosin (H&E)-stained biopsy slides. Using a cohort of 903 biopsy slides from 424 patients with matched molecular data, we demonstrate that our novel UNIv2-MIL framework, which fine-tunes a pre-trained pathology foundational model (UNIv2) using a multiple instance learning (MIL) strategy, achieves AUCs of 0.863 and 0.81 for PAM50 and PSC subtyping, respectively. Through computational clustering of high-attention regions, we identify histological patterns associated with molecular subtypes. Furthermore, in an independent validation cohort of 131 patients, we observed that patients with UNIv2-MIL predicted luminal subtypes tended to be more responsive to hormone therapy (HT) (p < 0.03). We also examined another independent cohort of 122 patients who transitioned from active surveillance to radical prostatectomy (RP) and observed that model-predicted PAM50 luminal B and PSC luminal proliferating scores showed significant correlation with adverse pathologic features (APFs) (p < 0.001 and p = 0.003, respectively). In conclusion, while further validation is needed, we have developed an AI-driven approach based on routine histopathology that has the potential to inform prostate cancer risk stratification and treatment response.
Similar content being viewed by others
Data availability
The datasets analyzed in the current study are not publicly available as they consist of electronic health records, and institutional policies require a data-sharing agreement for access.
Code availability
Source code is available via GitHub at https://github.com/RaminNateghi/PCaSub-MB-sMIL.
References
Wong, M. C. et al. Global incidence and mortality for prostate cancer: analysis of temporal patterns and trends in 36 countries. Eur. Urol. 70, 862–874 (2016).
Zhang, W. et al. Global burden of prostate cancer and association with socioeconomic status, 1990-2019: a systematic analysis from the global burden of disease study. J. Epidemiol. Glob. Health 13, 407–421 (2023).
Wikstrom, P. et al. Molecular subtypes provide possibilities for precision medicine in a advanced prostate cancer. Lakartidningen 121, 23179 (2024).
Feng, F. Y. et al. Association of molecular subtypes with differential outcome to apalutamide treatment in nonmetastatic castration-resistant prostate cancer. JAMA Oncol. 7, 1005–1014 (2021).
Zheng, K. et al. Integrative multi-omics analysis unveils stemness-associated molecular subtypes in prostate cancer and pan-cancer: prognostic and therapeutic significance. J. Transl. Med 21, 789 (2023).
Lee, D. J., Mahal, B. A. & Punnen, S. Validation of PAM50 for predicting progression in active surveillance: Results from the Miami MAST prospective clinical trial. J. Clin. Oncol 43, 238 (2025).
Kaffenberger, S. D. & Barbieri, C. E. Molecular subtyping of prostate cancer. Curr. Opin. Urol. 26, 213–218 (2016).
Coleman, I. M. et al. Therapeutic implications for intrinsic phenotype classification of metastatic castration-resistant prostate cancer. Clin. Cancer Res. 28, 3127–3140 (2022).
Zhao, S. G. et al. Associations of luminal and basal subtyping of prostate cancer with prognosis and response to androgen deprivation therapy. JAMA Oncol. 3, 1663–1672 (2017).
Ross, A. E. et al. Transcriptome-based prognostic and predictive biomarker analysis of ENACT: a randomized controlled trial of enzalutamide in men undergoing active surveillance. JCO Precis Oncol. 8, e2300603 (2024).
Patel, K. R. et al. Biopsy-based Basal-luminal Subtyping Classifi er in High-risk Prostate Cancer: A Combined Analysis of the NRG Oncology/RTOG 9202, 9413, and 9902 Phase 3 Trials. Eur. Urol. Oncol. S2588-9311(24)00246-3. Advance online publication. https://doi.org/10.1016/j.euo.2024.10.017.
Weiner, A. B. et al. A novel prostate cancer subtyping classifier based on luminal and basal phenotypes. Cancer 129, 2169–2178 (2023).
You, S. et al. Integrated classification of prostate cancer reveals a novel luminal subtype with poor outcome. Cancer Res. 76, 4948–4958 (2016).
Ben-Salem, S. et al. Diversity in androgen receptor action among treatment-naive prostate cancers is reflected in treatment response predictions and molecular subtypes. Eur. Urol. Open Sci. 22, 34–44 (2020).
Phillips, R. et al. Basal-luminal subtyping of localized high-risk prostate cancer and benefit of adding docetaxel to definitive radiotherapy with androgen suppression in the NRG Oncology/RTOG 0521 phase III trial. J. Clin. Oncol. 41, 5094 (2023).
Aisner, D. L. & Sams, S. B. The role of cytology specimens in molecular testing of solid tumors: techniques, limitations, and opportunities. Diagn. Cytopathol. 40, 511–524 (2012).
Erber, R. et al. Molecular subtyping of invasive breast cancer using a PAM50-based multigene expression test-comparison with molecular-like subtyping by tumor grade/immunohistochemistry and influence on oncologist's decision on systemic therapy in a real-world setting. Int. J. Mol. Sci. 23, 8716 (2022).
Picornell, A.C. et al. S.G. Breast cancer PAM50 signature: correlation and concordance between RNA-Seq and digital multiplexed gene expression technologies in a triple negative breast cancer series. BMC Genomics 20, 452 (2019).
Zhen, J. T. et al. Genetic testing for hereditary prostate cancer: current status and limitations. Cancer 124, 3105–3117 (2018).
Erak, E. et al. Predicting prostate cancer molecular subtype with deep learning on histopathologic images. Mod. Pathol. 36, 100247 (2023).
Mondol, R. K. et al. hist2RNA: an efficient deep learning architecture to predict gene expression from breast cancer histopathology images. Cancers 15, 2569 (2023).
Sirinukunwattana, K. et al. Image-based consensus molecular subtype (imCMS) classification of colorectal cancer using deep learning. Gut 70, 544–554 (2021).
Jaber, M. I. et al. A deep learning image-based intrinsic molecular subtype classifier of breast tumors reveals tumor heterogeneity that may affect survival. Breast Cancer Res. 22, 12 (2020).
Rawat, R. R. et al. Deep learned tissue “fingerprints” classify breast cancers by ER/PR/Her2 status from H&E images. Sci. Rep. 10, 7275 (2020).
Gao, F. et al. DeepCC: a novel deep learning-based framework for cancer molecular subtype classification. Oncogenesis 8, 44 (2019).
Fountzilas, E. et al. Convergence of evolving artificial intelligence and machine learning techniques in precision oncology. NPJ Digit Med 8, 75 (2025).
Wei, Z. et al. Deep learning-based multi-omics integration robustly predicts relapse in prostate cancer. Front. Oncol. 12, 893424 (2022).
Sarjezh, R. N. et al. Predicting PAM50 subtypes from whole slide images of prostate cancer biopsies. J. Clin. Oncol. 42, e17001 (2024).
Guramare, M. et al. PO3-07-04: prediction of PAM50 molecular subtypes from H&E-stained breast cancer specimens using tumor microenvironment features and additive multiple instance learning models. Cancer Res. 84, PO3-07-04-PO3-07-04 (2024).
Spratt, D.E. et al. A double-blinded placebo-controlled biomarker-stratified randomized trial of apalutamide (APA) and radiotherapy for recurrent prostate cancer (NRG GU006, BALANCE trial). Int. J Radiation Oncol. Bio. Phys. 123, 1197–1198 (2025).
McKenney, J. K. et al. Histologic grading of prostatic adenocarcinoma can be further optimized: analysis of the relative prognostic strength of individual architectural patterns in 1275 patients from the canary retrospective cohort. Am. J. Surg. Pathol. 40, 1439–1456 (2016).
Feng, F. Y.-C. et al. Luminal and basal subtyping of prostate cancer. J Clin. Oncol.: Am. Soc. Clin. Oncol. 35, 3 (2017).
Roy, S. et al. Early prostate-specific antigen response by 6 months is predictive of treatment effect in metastatic hormone sensitive prostate cancer: an exploratory analysis of the TITAN trial. J. Urol. 212, 672–681 (2024).
Chowdhury, S. et al. Deep, rapid, and durable prostate-specific antigen decline with apalutamide plus androgen deprivation therapy is associated with longer survival and improved clinical outcomes in TITAN patients with metastatic castration-sensitive prostate cancer. Ann. Oncol. 34, 477–485 (2023).
Harshman, L. C. et al. Seven-month prostate-specific antigen is prognostic in metastatic hormone-sensitive prostate cancer treated with androgen deprivation with or without docetaxel. J. Clin. Oncol. 36, 376–382 (2018).
Vaswani, A. et al. Attention is all you need. In Advances in Neural Information Processing Systems, 30 (NIPS, 2017).
Chen, R. J. et al. Towards a general-purpose foundation model for computational pathology. Nat. Med. 30, 850–862 (2024).
Acknowledgements
R.N. is supported by the Polsky Urologic Cancer Institute Award.
Author information
Authors and Affiliations
Contributions
R.N. led the conceptualization, methodology, analysis, visualization, validation, and manuscript drafting and editing. A.S. and H.D. implemented and validated the baseline method and contributed to writing the manuscript. N.H., M.S., J.M., M.Sa., J.W.J., and N.C. curated study cohorts, reviewed the manuscript, and provided clinical input. S.K. conducted statistical analysis, and E.M.S. oversaw project administration. H.D.P. contributed to the review of the manuscript. B.G.N. and X.J.Y. reviewed and validated pathology findings. L.A.D.C. provided resources and contributed to manuscript editing. A.E.R. supported the project by supervising, providing funding, guiding the concept, and reviewing and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nateghi, R., Sun, A., Dang, H. et al. Prediction of molecular subtypes from histology: AI-driven analysis of prostate cancer morphological patterns and therapeutic implications. npj Precis. Onc. (2026). https://doi.org/10.1038/s41698-026-01335-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-026-01335-y