Abstract
We report a child with an antenatally detected brain tumor that progressed over three years’ time despite surgery, chemo- and proton therapy. Retrospective whole-genome and transcriptome sequencing with methylation analysis of primary tumor tissue led to the molecular diagnosis infant-type hemispheric glioma, and identified a novel SNRNP70::ALK fusion, providing a therapeutic target for compassionate-use precision treatment with the ALK tyrosine kinase inhibitor lorlatinib. Functional studies confirmed the fusion protein to be expressed and active in the patient’s tumor. After two years of therapy, the child has sustained partial tumor regression on MRI and no new neurological symptoms. We conclude that comprehensive multi-omics analyses are required for correct molecular diagnosis in childhood CNS tumors and can radically impact patient outcome by identifying molecular targets for precision treatment.
Similar content being viewed by others
Introduction
Infant-type hemispheric glioma (ITHG) was introduced as a new tumor type in the fifth edition of the 2021 WHO Classification of Tumors of the Central Nervous System (CNS) 11. Since then, only a few new cases have been reported. The tumors are high-grade astrocytomas that develop in early childhood and commonly appear as large masses in the supratentorial compartment. They typically harbor receptor tyrosine kinase (RTK) fusion genes (NTRK1/2/3, ROS1, ALK or MET) that drive tumorigenesis via signaling through PI3K and/or MAPK pathways2,3. The presence of RTK fusions provides the opportunity to treat patients with small molecule tyrosine kinase inhibitors (TKIs) with improved outcome4. Furthermore, fusion sequences may serve as highly specific molecular markers for measurable residual disease detection in cell-free DNA (cfDNA) in liquid biopsies5.
Results
Case presentation
In a woman in gestational week 40, an abdominal ultrasonography was performed due to decreased fetal movements. A lesion was seen in the left cerebral hemisphere of the fetus, initially interpreted as a small hemorrhage. After an emergency C-section, the child stabilized with respiratory support. He had normal weight and length, no congenital malformations and no family history indicating a childhood cancer predisposition syndrome. Brain magnetic resonance imaging (MRI) showed a tumor measuring 7x7x6.5 cm with intra-tumoral bleedings (Fig. 1a).
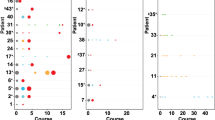
Brain imaging (MRI) with arrows indicating tumor: top row T2 images; bottom row post-gadolinium T1-images. a Imaging on the day of birth with left hemispheric tumor, (no contrast used at this time point). b 2 month post-operative control with primary tumor residue. c Control at 3 years of age, after SIOPE INFANT HIT-01 protocol and temozolomide, showing primary tumor residue and d spread into right ventricle/thalamus. e Control at 3 years and 8 months of age, post proton therapy, showing, lesion in right ventricle/thalamus (with some regression) and f new pontocerebellar metastasis. g Control at four years of age, 4 months into lorlatinib treatment, showing lesion in right ventricle/ thalamus, h pontocerebellar metastasis and i primary tumor residue. GW gestational week, d day, m month, y year, IVD intraventricular device, SIOP European Society for Pediatric Oncology, VCS ventriculocisternostomy, VPS ventriculoperitoneal shunt. Created with BioRender.com.
One day postpartum, the child developed seizures requiring treatment with levetiracetam and phenobarbital. The following day, he underwent neurosurgery and post-operative MRI indicated a possible complete resection of the tumor. However, MRI at two months of age showed a residual tumor (Fig. 1b). The final neuropathological diagnosis was a high-grade glial tumor, IDH wild-type (WHO grade IV) according to the WHO 2016 classification6 based on standard-of-care investigations (Supplementary appendix). At two weeks of age, the patient became lethargic due to hydrocephalus and received an intraventricular device (IVD). One week later, adjuvant chemotherapy was initiated according to the European Society for Pediatric Oncology (SIOPE) INFANT HIT-01 protocol. Treatment continued until the child was one year old. MRI indicated regression of the primary tumor lesion. At that time point, he had right-sided weakness of his arm and leg.
Sixteen months after completion of treatment, MRI showed new areas with intraventricular/right thalamic contrast enhancement. The child had no new symptoms related to the tumor and had no seizures. He had a developmental delay, but started babbling and crawling. After contact with experts within the SIOPE network, the treating oncologists requested analysis of ALK, NTRK and ROS1 gene fusions on the primary tumor material. Amplicon-based next-generation sequencing (NGS) with the targeted OncomineTM Childhood Cancer Research Assay (Thermo Fisher Scientific Waltham, MA, USA) showed no mutations or fusions. Immunohistochemistry (IHC) using D5F5 and ALK1 antibodies showed variable ALK reactivity (Suppl Fig. 1), while ROS and NTRK were negative. Following recommendations from the local multidisciplinary team, temozolomide, rather than irradiation, was chosen as the next treatment. Temozolomide had to be discontinued six months later due to persistent myelotoxicity and signs of bone marrow failure. One month after treatment termination, brain MRI revealed progression of the intraventricular/right thalamic contrast enhancement (Fig. 1c, d) and focal proton therapy (1.8 Gy relative biological effectiveness x33) was given to the whole tumor area including the intraventricular/thalamic lesion soon after the child’s third birthday.
A month after completion of proton therapy, neuroimaging again showed suspected progression and neurosurgeons were consulted to gain new tumor material for molecular analyses. Since a total resection of the tumor was not possible, the parents were reluctant to let their child undergo renewed surgery due to the risk of complications. The patient did not have progression of symptoms but had right-sided weakness, abnormal eye motility, right-sided blindness and poor vision in his left eye. Brain MRI two months later instead showed regression of the intraventricular/right thalamic lesion and during the coming months, the child was energetic and was able to attend preschool using his walker and stroller.
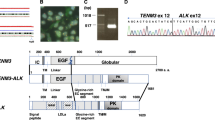
In a research setting, retrospective whole genome sequencing of DNA/RNA from the diagnostic tissue biopsy and a paired normal sample revealed an unbalanced translocation between chromosomes 2 and 19 and a predicted SNRNP70(e4)::ALK(e20) fusion in the tumor (Fig. 2a–c). Whole transcriptome sequencing confirmed an in-frame fusion with the kinase domain retained in the transcript (Fig. 2d, e). The copy-number profile showed a terminal duplication of 2p, including the ALKAL2 and MYCN genes, but not ALK itself, and a terminal deletion of 19q (Fig. 2f). Additionally, methylation array profiling showed a 1.0 match to the methylation class infantile hemispheric glioma (Fig. 2g, methods in Supplementary Appendix). No other genetic aberrations affecting regions known to be recurrently involved in childhood cancer were seen.
a Translocation between chromosomes 2 and 19 [t(2;19)(p23.2;q13.33)] involving exons 1-4/10::20-29/2, resulting in SNRNP70::ALK fusion, with genomic breakpoints annotated in figure (GRCh38, STAR-Fusion). b Break point junctions for ALK on chromosome 2 (purple) and SNRNP70 on chromosome 19 (pink), supported by 29 split reads and 27 paired reads, coinciding with a clear change of the read depth, indicating an unbalanced translocation. Variant allele frequency (VAF) was 24% and 26% based on split and paired reads, respectively (GRCh38, IGV). c Circus plot illustrating t(2;19)(p23.2;q13.33) (STAR-Fusion). d Predicted protein domains in the SNRNP70::ALK fusion (STAR-Fusion). e SNRNP70::ALK seen in-frame (FusionInspector). f Copy-number profile based on WGS data (Control-FREEC). g Copy-number profile based on methylation array analysis (brain tumor methylation classifier, DKFZ, German Cancer Research Center, https://www.molecularneuropathology.org/mnp/).
The finding of a novel ALK-fusion in the tumor was reported back to the treating clinicians as a potential therapeutic target. Fluorescent in situ hybridization (FISH) analysis on primary tumor material confirmed an ALK rearrangement with a split pattern in more than 50% of tumor cells (Supplementary Fig. 1c). The NGS panel failed to identify the SNRNP70::ALK fusion as the amplicons missed the ALK breakpoint by one base pair (communication with ThermoFisherTM). A license for compassionate use of the third-generation ALK inhibitor lorlatinib was granted by the Swedish Medical Agency.
After a three-month period of watchful waiting—at three years and eight months of age—a pontocerebellar metastasis was detected (Fig. 1e, f) and lorlatinib treatment was initiated with a total daily dose of 75 mg (divided into three daily doses, administered orally). Three months later, slight improvements of motor symptoms were described by the parents. MRI after four months of targeted therapy showed partial regression of the intraventricular tumor component as well as of the pontocerebellar metastasis (Fig. 1g, h) which had decreased in size from 13×17 mm to 11×7 mm, and from 14×17 mm to 7×10 mm, respectively. The tumor residue in the midline, occipital left had not confidently changed in size (Fig. 1i). Since then, consecutive MRIs at three-month intervals have shown stable disease. Due to weight gain, elevated cholesterol, constipation and anemia—all interpreted as side effects of treatment—the lorlatinib dose was lowered to 50 mg once daily. In parallel, a referral was sent to an obesity unit, and physiotherapy was initiated. The boy, now on ALK TKI therapy for two years, has improved in his general condition, has better sleep and is more alert.
Functional study results
A number of functional studies were performed to evaluate the expression of the fusion protein and its downstream effects (methods in Supplementary Appendix). Immunoblotting analyses of resected primary tumor material identified SNRNP70::ALK fusion expression with both anti-ALK (D5F3) and anti-SNRNP70 antibodies (Fig. 3A, B). Expression of the full length ALK RTK was also detected in the tumor lysate, as well as in control lysates from CLB-GAR and NB1 neuroblastoma cell lines (Fig. 3A). Further, phosphospecific ALK antibodies confirmed expression and activation of the ALK kinase domain of the SNRNP70::ALK fusion and full length ALK as well as activation of downstream targets, ERK and AKT (Fig. 3A).
a Anti-ALK immunoblotting detects a SNRNP70::ALK fusion protein with a size of approximately 85 kDa (arrow) in the patient tumor. In addition, endogenous full-length ALK proteins (indicated with arrowheads) was detected in the tumor sample. Lysates from SK-N-AS, SK-N-BE(2), CLB-GAR (ALK-R1275Q) and NB1 (ALK-amplified) neuroblastoma cells lines29 were loaded as controls (SK-N-AS and SK-N-BE(2) are ALK negative, while CLB-GAR and NB1 are ALK positive). Anti-pALK immunoblotting detects activation of SNRNP70-ALK fusion protein, and anti-pERK1/2 and pAkt confirm downstream AKT and ERK1/2 activity in the patient tumor. Tubulin was used as loading control. b Both anti-ALK and anti-SNRNP70 detect the approximately 85 kDa SNRNP70::ALK fusion protein in the patient tumor. As controls, SK-N-AS cells expressing either vector control, SNRNP7::ALK, SNRNP70::ALK (I1250T)-3xFLAG, or EML4::ALK V3a constructs were employed. Anti-SNRNP70 immunoblotting also detected endogenous SNRNP70 proteins (indicated with *) in all samples. Arrowheads indicate full-length and cleaved ALK. Arrow indicates the SNRNP70::ALK fusion protein. Note: due to a 3x FLAG tag, the molecular weight of the I1250T mutant SNRNP70::ALK fusion protein is slightly bigger than that of the wildtype fusion one. Original immunoblot figures are shown as Fig. S5. c Detection of active receptor tyrosine kinases by Human RTK array. A proteome Profiler Human Phospho-RTK Array Kit (ARY001B, R&D Systems) was used to detect active receptor tyrosine kinases (RTKs) in the tumor lysate following the protocol provided by the manufacturer. Approximately 300 µg clarified lysate was applied to one array. The layout of different RTKs (purple texts) in the array is provided on top. Boxed dots indicate pALK.
In addition, RTK array analysis of tumor lysate identified active ALK as well as several additional activated RTKs (Fig. 3c), such as PDGFRb, TYRO3 (Dtk) and ROR2.
A PC12 cell neurite outgrowth assay was employed to investigate if the SNRNP::ALK fusion protein was functional in a controlled system. ALK activation triggers differentiation of PC12 cells into sympathetic like neurons, a process characterized by extension of neurites7,8,9. The SNRNP70::ALK fusion was tested, as well as EML4::ALK variant 3a, which was employed as a positive control. An additional control, SNRPNP70::ALK harboring an ALK(I1250T) kinase dead mutation was also included10. Expression of either SNRNP70::ALK or EML4::ALK fusions resulted in neurite outgrowth within 48 h (Fig. S2a and Fig. S3), although the EML4::ALK, variant 3a is more potent. Both ALK fusions were phosphorylated on expression in PC12 cells and led to the activation of downstream targets such as AKT, ERK and S6K (Fig. S2b). Importantly, expression of the SNRNP70::ALK(I1250T) dead kinase fusion did not result in a significant increase in neurite outgrowth or downstream signaling activity (Fig. S2a, b and Fig. S3).
Taken together, these results indicate that the SNRNP70::ALK fusion protein is functional and able to drive progression of the disease in a similar manner as other ALK fusion proteins.
Liquid biopsy results
At three months of age, the patient underwent ventriculocisternostomy (VCS) and 16 ml of cerebrospinal fluid (CSF) was collected for a liquid biopsy study. cfDNA from CSF as well as genomic DNA (gDNA) from the CSF cell pellet, were analyzed as previously described5 using a customized ddPCR assay targeting the SNRNP70::ALK junction sequence (Supplementary Appendix). Both samples were positive for the fusion at very low levels; 7 and 12 copies in cfDNA and gDNA, respectively (Fig. S4). In comparison, three CSF samples taken between the age of two weeks and two months (6.5, 0.8 and 3.0 mL, respectively) were negative for malignant cells by morphologic examination of cytospin preparations. CfDNA from plasma collected prior to initiation of lorlatinib and during follow-up was negative.
Discussion
In this child with a large congenital brain tumor, multi-omics analysis led to a correct clinical diagnosis of ITHG and to the discovery of a novel gain-of-function SNRNP70::ALK fusion that was not identified by standard clinical investigation. These findings underscore the clinical importance of multi-layered omics analyses at diagnosis, as is now clinical routine in all children with cancer in Sweden11. The novel ALK fusion was confirmed to be active in patient tumor material and was amenable to targeted therapy. The child, who had suffered tumor progression despite surgery, chemo and photon therapy, displays continued radiological response to lorlatinib two years into treatment and is clinically improved.
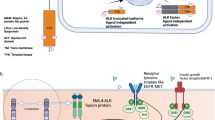
To the best of our knowledge, SNRNP70::ALK fusions have not been reported in cancer previously. The SNRNP70 gene encodes a ubiquitously expressed protein that forms a key component of the spliceosome (small nuclear ribonucleoprotein U1 subunit 70, alias U1-70K). The N-terminal part of the protein contains a coiled-coil domain encoded by exons 3 and 4 that likely mediates dimerization12,13,14. SNRNP70 is not sensitive to haploinsufficiency (pLoF 0.33 according to gnomAD v4.1.0 based on MANE Select transcript, ENST00000598441.6). Two publications have previously described SNRNP70 as a 5’ partner gene of an oncogenic fusion with MET or NTRK314,15. In the cBioPortal database, SNRNP70-fusions have not previously been reported in CNS tumors, but have been observed in several extracranial solid tumors, indicating that SNRNP70 is a recombination-prone gene, with partner genes on several chromosomes16. The variant observed in our patient is an in-frame fusion of the N-terminus of SNRNP70 (exons 1-4) to the C-terminus of ALK (exons 20-29), generating a fusion protein with ALK gain-of-function kinase activity, which was able to drive downstream signaling and neurite outgrowth in a PC12 cell differentiation system.
Interestingly, we also noted chromosomal 2p gain including the ALKAL2 (encoding an ALK receptor ligand) and the MYCN oncogene. The tumor clearly also expresses the full-length ALK RTK which is phosphorylated and active. It is known that neuroblastomas harboring 2p gain/amplification have a worse prognosis and are regarded as high-risk disease17,18. Furthermore, the combination of MYCN gain and ALK activation is known to increase oncogenic transformation19. It is therefore plausible that in addition to the SNRNP70::ALK fusion described here, the gain of other genes on chromosome arm 2p (such as ALKAL2 and the oncogene MYCN) enhance endogenous ALK signaling activity in the tumor20.
The SNRNP70::ALK target was detectable at low levels in cfDNA and gDNA from CSF using ddPCR, suggesting that this method outperforms the sensitivity of CSF cytology, which was negative for malignant cells on repeated occasions prior to the ddPCR analysis, and that this molecular approach may be feasible to measure residual disease in children with ITHG.
The strengths of our report include the comprehensive multi-omics methods (whole genome sequencing, whole transcriptome sequencing as well as methylation analysis) used to analyze the tumor; the robust functional assays performed that prove that the novel ALK-fusion is an active tumor driver and the clinical outcome of the licensed treatment. The report is, of course, limited by the fact that this genetic alteration is likely unique and thus based on only one individual.
In conclusion, a multi-omics approach was necessary to correctly classify ITHG, identify a target for precision treatment and enable personalized MRD detection in liquid biopsies.
Methods
Standard of care neuropathological assessment at time of diagnosis
Neuropathological assessment of fresh frozen tumor tissue showed a diffuse glioma with brisk mitotic activity, microvascular proliferation and palisading necrosis. Immunohistochemistry was positive for glial markers (INI1 and BRG1 retained, H3-K27M negative, and loss of H3-K27-me3). Genetic characterization, including Sanger sequencing of TP53 exons 4-8, IDH1 exon 4, IDH2 exon 4 and hTERT promoter positions -228 and -250, and real-time PCR of BRAF V600 mutation and the KIAA1549::BRAF fusion, did not reveal aberrations. Pyrosequencing showed no methylation of four CpG sites in the MGMT promoter.
Nucleic acid extraction from tumor
Nucleic acids from primary tumor tissue (DNA and RNA) and blood (DNA) were isolated by The Swedish Childhood Tumor Biobank, using the AllPrep DNA/RNA/Protein Mini kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions.
Whole genome/transcriptome sequencing and methylation profiling
WGS and methylation data were generated and processed as previously described21,22. An RNA library was prepared from total RNA using the Illumina TruSeq Stranded total RNA protocol, Illumina RiboZero GOLD (Illumina, SanDiego, CA, USA). The sample was sequenced on NovaSeq6000 (NovaSeq Control Software 1.7.5/RTA v3.4.4) with a 151nt(Read1)-10nt(Index1)-10nt(Index2)-151nt(Read2) setup using ‘NovaSeqXp’ workflow in ‘S4’ mode flowcell. Library preparation and sequencing are ISO-accredited applications, performed at the Genomic Production Center, SciLifeLab, Stockholm, Sweden. The Bcl to FastQ conversion was performed using bcl2fastq_v2.20.0.422 from the CASAVA software suite. The quality scale used was Sanger / phred33 / Illumina 1.8 + . Raw sequencing data is demultiplexed and converted to FastQ on site before being transferred securely to UPPMAX for delivery. Paired-end RNA-Seq FASTQ files with a read length of 150 bp were aligned to the GRCh38 human reference (hg38) using the STAR aligner (version 2.7.10b)23. Fusion calling was subsequently performed using STAR-Fusion (v1.12) and Arriba (v2.4.0) with default parameters24,25.
Liquid biopsy processing
CSF was collected in Cell-Free DNA BCT tubes (10 mL) (STRECK, La Vista, NE, USA) and sterile tubes. Blood samples were collected in the former tube type. Samples underwent cell removal by double centrifugation (10 min at 4 ◦C 1600× g and 16,000× g) within 5 days. Supernatants and cell pellets were frozen at −80 ◦C. Cell-free supernatants were then thawed and cfDNA was isolated from 16 mL of CSF and 3.8-5 mL of blood plasma (six consecutive samples) using the QIAamp Circulating Nucleic Acid Kit on the QIAvac24 Plus vacuum manifold (Qiagen, Manchester, UK). cfDNA was eluted in 40 μL of AVE buffer and stored at RT. Plasma from blood donors was processed in the same manner and used as negative control (NC). gDNA from CSF cell pellets was extracted using Puregene Blood Kit (Qiagen) and eluted in 100 μL rehydration buffer.
Droplet digital PCR
ddPCR assay design was based on WGS reads spanning the SNRNP70::ALK junction sequence (supplementary table 1) and optimized as previously described21. ddPCR experiments were run on the QX200 AutoDG Droplet Digital PCR System/QX200 Droplet Reader (BioRad, Hercules, CA, USA) according to the manufacturer’s instructions26,27. Amplicon size was 89 bp. Primer/probe sequences are available upon request. A reference assay (ABCC9, dHsaCP2506567, BioRad, USA) was used in all reactions to assess DNA concentration and amplifiability. Triplicate ddPCR reactions on non-amplified cfDNA from CSF and plasma were run on the QX200 AutoDG Droplet Digital PCR System/QX200 Droplet Reader (BioRad, Hercules, CA, USA) according to the manufacturer’s instructions using the SNRNP70::ALK fusion assay. Reactions contained 10 μL 2x ddPCR Supermix for probes (No dUTP) (BioRad, Hercules, CA, USA), 1 μL 20x SNRNP70::ALK fusion assay and 11 μL of cfDNA eluate. Nine ddPCR reactions on gDNA from CSF cell pellets were run in the same manner. Triplicate wells of no template controls (nuclease free water) and positive controls (genomic DNA from primary tumor), and 12 wells of NC cfDNA from donor plasma, were run on the same plate. Thermal cycling conditions were: 1 cycle at 95 ◦C for 10 min, 40 cycles at 94 ◦C for 30 s and 56 ◦C for 1 min, 1 cycle at 98 ◦C for 10 min, and 1 cycle at 4 ◦C ∞ at ramp rates of 2 ◦C/s. Results were visually reviewed applying Poisson and total error models with a 95% confidence interval using the QX ManagerTM Software Standard Edition Version 1.2 (BioRad, Hercules, CA, USA).
Preparation of patient tumor lysate
Snap-frozen tumor tissue was obtained from The Swedish Tumor Biobank under the permit BbK2337. Tumor tissue was homogenized with TissueLyser II (Qiagen) in RIPA buffer (ThermoScientific) with complete protease inhibitor cocktail (Roche). Crude lysate was then clarified by a 10-minute centrifugation at 14,000 rpm at 4 °C. Protein concentration of the clarified lysate was measured with the Pierce BCA Protein Assay kit (ThermoScientific). Prior to immunoblotting analysis, the lysate was mixed with 4 x SDS sample buffer (BioRad) to make 1x sample and boiled at 90 °C for 10 minutes.
Generation of the human SNRNP70::ALK construct
Based on sequence analysis of the fusion gene cDNA, a 342-bp DNA fragment beginning with the Kozak sequence of SNRNP70 and ending with the BlpI site of ALK, was synthesized by Invitrogen GeneArt Gene Synthesis services (ThermoScientific). A HindIII site was added to the 5’ end to facilitate subcloning. The pcDNA3-SNRNP70-ALK construct was generated by substituting the HindIII-BlpI portion of full-length ALK in pcDNA3 vector8 with the synthesized fragment. In addition, a construct expressing this fusion protein with ALK I1250T mutation (number refers to full-length ALK) was generated by substituting the HindIII-BlpI fragment of the full length ALK from the pcDNA3-ALK (I1250T)-3xFLAG construct10.
Cell culture, transfection, lysis and immunoblotting
PC12 cells were cultured in RPMI 1640 medium supplemented with 7% heat-inactivated horse serum and 3% fetal bovine serum (FBS) and a mixture of 1% penicillin/streptomycin. SK-N-AS, SK-N-BE(2), CLB-GAR and NB1 cells were cultured in RPMI 1640 medium supplemented with 10% FBS and a mixture of 1% penicillin/streptomycin. All cells were cultured under 37 °C, 95% humidity and 5% CO2 conditions as previously described28. PC12 cells (2 × 106) were transfected by electroporation in an Amaxa electroporator (Lonza, Basel, Switzerland) with 0.75 μg of either pcDNA3 vector, pcDNA3-SNRNP70-ALK, SNRNP70-ALK (I1250T) or pcDNA3-EML4-ALK V3a plasmid (constructed previously) and were cultured in complete medium for 24 hours. Cells were then starved in serum-free medium for 24 h prior to lysis. SK-N-AS cells were transfected with Lipofectamine 3000 (Invitrogen, Carlsbad. CA) following the manufacturers’ instructions. Cells were lysed in RIPA buffer with protease inhibitor cocktail. Lysates were clarified by centrifugation at 14,000 rpm for 10 min at 4 °C. After protein concentration measurement, lysates were mixed with 4x SDS sample buffer, boiled and analyzed by immunoblotting as previously described29. The intensity of different protein bands was quantified with Image Studio Lite 5.2 software. Ratios of phosphorylated proteins and total proteins are shown normalized. Primary antibodies used for immunoblotting: anti-pan-ERK (#610124) mouse monoclonal antibody (mAb) from BD Transduction Laboratories (Franklin Lakes, NJ); anti-pALK (Y1278) (D59G10) rabbit mAb (#6941), anti-ALK (D5F3) rabbit mAb (#3633), anti-pERK1/2 (T202/Y204) (D13.14.4E) rabbit mAb (#4370), anti-pAKT (S473) (D9E) rabbit mAb (#4060), and anti-AKT rabbit Ab (#9272) from Cell Signaling Technology (Danvers, MA); anti-SNRNP70 rabbit Ab (PA5-41729) and anti-β-Tubulin mouse mAb (MA5-16308) from ThermoScientific (Waltham, MA). Horseradish-peroxidase-conjugated secondary antibodies goat anti-rabbit IgG and goat anti-mouse IgG were from Thermo Scientific (Waltham, MA).
Neurite outgrowth assay
PC12 cells were transfected by electroporation in an Amaxa electroporator (Lonza, Basel, Switzerland) with 0.75 μg of either empty pcDNA3 vector, wildtype (WT) or I1250T mutant SNRNP70-ALK construct or EML4-ALK V3a construct as indicated, together with 0.5 μg of pEGFP-C1 in 100 μl of Ingenio electroporation solution (Mirus Bio LCC, Madison, WI). After electroporation, cells were transferred to RPMI 1640 medium supplemented with 7% horse serum and 3% FBS and seeded into 24-well plates. Two days after transfection, the percentage of both GFP-positive and neurite-carrying cells versus GFP-positive cells was calculated under a Zeiss Axiovert 40 CFL microscope. To be judged as a neurite-carrying cell, neurites had to reach at least twice the length of the cell body. Representative light microscope images showing the neurite outgrowth were taken with Olympus DP-12 microscope camera. Experiments were performed in triplicate and each sample within an experiment was assayed in duplicate. The bar chart was generated in GraphPad Prism 10 with bars representing mean percentage ± SD. P-values were calculated with two-tailed paired student’s t-test.
Ethical statement
The study was conducted in accordance with the Declaration of Helsinki and approved by the Regional Ethical Review Board/Swedish Ethical Review Authority. The patients’ parents provided written informed consent for inclusion in this study and for publication of the results. We followed the CARE-reporting guidelines30.
Data availability
Sequence data that support the findings of this study are available via the Swedish Childhood Tumor Biobank after an application: https://www.barntumorbanken.se/for-forskare/.
References
Louis, D. N. et al. The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 23, 1231–1251 (2021).
Guerreiro Stucklin, A. S. et al. Alterations in ALK/ROS1/NTRK/MET drive a group of infantile hemispheric gliomas. Nat. Commun. 10, 4343 (2019).
Clarke, M. et al. Infant high-grade gliomas comprise multiple subgroups characterized by novel targetable gene fusions and favorable outcomes. Cancer Discov. 10, 942–963 (2020).
Bagchi, A. et al. Lorlatinib in a child with ALK-fusion-positive high-grade glioma. N. Engl. J. Med. 385, 761–763 (2021).
Arthur C., et al. Simultaneous ultra-sensitive detection of structural and single nucleotide variants using multiplex droplet digital PCR in liquid biopsies from children with medulloblastoma. Cancers 15, 15071902 (2023);
Louis, D. N. et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 131, 803–820 (2016).
Guan J., et al. Clinical response of the novel activating ALK-I1171T mutation in neuroblastoma to the ALK inhibitor ceritinib. Cold Spring Harb. Mol. Case Stud. 4, 002500 2018).
Martinsson, T. et al. Appearance of the novel activating F1174S ALK mutation in neuroblastoma correlates with aggressive tumor progression and unresponsiveness to therapy. Cancer Res. 71, 98–105 (2011).
Motegi, A., Fujimoto, J., Kotani, M., Sakuraba, H. & Yamamoto, T. ALK receptor tyrosine kinase promotes cell growth and neurite outgrowth. J. Cell Sci. 117, 3319–3329 (2004).
Schönherr, C. et al. The neuroblastoma ALK(I1250T) mutation is a kinase-dead RTK in vitro and in vivo. Transl. Oncol. 4, 258–265 (2011).
Wadensten E., et al. Diagnostic yield from a nationwide implementation of precision medicine for all children with cancer. JCO Precis Oncol. 7, e2300039 (2023)
Pomeranz Krummel, D. A., Oubridge, C., Leung, A. K., Li, J. & Nagai, K. Crystal structure of human spliceosomal U1 snRNP at 5.5 A resolution. Nature 458, 475–480 (2009).
Kondo Y., Oubridge C., van Roon A. M., Nagai K. Crystal structure of human U1 snRNP, a small nuclear ribonucleoprotein particle, reveals the mechanism of 5’ splice site recognition. Elife. 4 https://doi.org/10.7554/eLife.04986 (2015).
Kobayashi, H. et al. Dramatic response to entrectinib in a patient with malignant peripheral nerve sheath tumor harboring novel SNRNP70-NTRK3 fusion gene. Genes Chromosomes Cancer 62, 47–51 (2023).
Wang, J. et al. Comprehensive analysis of oncogenic fusions in mismatch repair deficient colorectal carcinomas by sequential DNA and RNA next generation sequencing. J. Transl. Med. 19, 433 (2021).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Jeison, M. et al. 2p24 Gain region harboring MYCN gene compared with MYCN amplified and nonamplified neuroblastoma: biological and clinical characteristics. Am. J. Pathol. 176, 2616–2625 (2010).
Javanmardi, N. et al. Analysis of ALK, MYCN, and the ALK ligand ALKAL2 (FAM150B/AUGα) in neuroblastoma patient samples with chromosome arm 2p rearrangements. Genes Chromosomes Cancer 59, 50–57 (2020).
Zhu, S. et al. Activated ALK collaborates with MYCN in neuroblastoma pathogenesis. Cancer Cell 21, 362–373 (2012).
Guan J., Hallberg B., Palmer R. H. Chromosome imbalances in neuroblastoma-recent molecular insight into chromosome 1p-deletion, 2p-gain, and 11q-deletion identifies new friends and foes for the future. Cancers 13 https://doi.org/10.3390/cancers13235897 (2021).
Arthur C., et al. Simultaneous ultra-sensitive detection of structural and single nucleotide variants using multiplex droplet digital PCR in liquid biopsies from children with medulloblastoma. Cancers 15 https://doi.org/10.3390/cancers15071972 (2023).
Capper, D. et al. DNA methylation-based classification of central nervous system tumours. Nature 555, 469–474 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Haas, B. J. et al. Accuracy assessment of fusion transcript detection via read-mapping and de novo fusion transcript assembly-based methods. Genome Biol. 20, 213 (2019).
Uhrig, S. et al. Accurate and efficient detection of gene fusions from RNA sequencing data. Genome Res 31, 448–460 (2021).
Rare Mutation Detection Best Practices Guidelines Droplet Digital™ PCR. Bio-Rad Laboratories, Inc. https://www.bio-rad.com/webroot/web/pdf/lsr/literature/Bulletin_6628.pdf.
Droplet Digital™ PCR Applications Guide. Bio-Rad Laboratories, Inc. https://www.bio-rad.com/webroot/web/pdf/lsr/literature/Bulletin_6407.pdf.
Guan, J., Chuang, T. P., Vikstrom, A., Palmer, R. H. & Hallberg, B. ALK F1174S mutation impairs ALK kinase activity in EML4-ALK variant 1 and sensitizes EML4-ALK variant 3 to crizotinib. Front. Oncol. 13, 1281510 (2023).
Guan, J. et al. IGF1R Contributes to Cell Proliferation in ALK-Mutated Neuroblastoma with Preference for Activating the PI3K-AKT Signaling Pathway. Cancers 15, 4252 (2023).
Gagnier J. J., Kienle G., Altman D. G., Moher D., Sox H, Riley D; the CARE Group. The CARE guidelines: consensus-based clinical case reporting guideline development. Glob. Adv. Health Med. 2, 38–43 (2013).
Acknowledgements
The authors acknowledge support from Science for Life Laboratory, the Knut and Alice Wallenberg Foundation, the National Genomics Infrastructure funded by the Swedish Research Council, as well as the Uppsala Multidisciplinary Center for Advanced Computational Science for assistance with massively parallel sequencing and access to the UPPMAX computational infrastructure. Methylation profiling was performed by the SNP&SEQ Technology Platform in Uppsala (www.genotyping.se). The facility is part of the National Genomics Infrastructure (NGI) Sweden and Science for Life Laboratory. The SNP&SEQ Platform is also supported by the Swedish Research Council and the Knut and Alice Wallenberg Foundation. The authors acknowledge The Swedish Childhood Tumor Biobank, supported by The Swedish Childhood Cancer Fund, for access to sequencing and methylation data as well as to tumor DNA. This work was funded by Cancerfonden, Vetenskapsrådet and Barncancerfonden.
Funding
Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Contributions
C.A., G.B., D.H., E.T., B.H., and R.P. conceived the study. C.A., K.G., T.D.d.S., J.G., B.O., C.J., C.I., J.S.a., J.S.v., J.E., and U.S. collected and generated data - (i.e., methodology, clinical data, formal analysis, visualization). C.A., G.B., R.R., D.H., E.T., B.H., and R.P. reviewed the data and supervised the analysis as well as the writing of the manuscript. E.T. and G.H. funded the work. C.A., J.G., E.T., B.H., and R.P. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arthur, C., Georgantzi, K., de Ståhl, T.D. et al. Novel activating SNRNP70-ALK fusion in congenital infant-type hemispheric glioma displays clinical response to lorlatinib: a case-report. npj Precis. Onc. 10, 113 (2026). https://doi.org/10.1038/s41698-026-01336-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41698-026-01336-x