Abstract
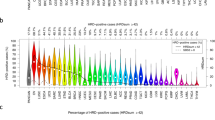
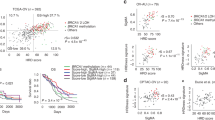
While homologous recombination deficiency (HRD) presents therapeutic opportunities in endometrial cancer (EC), its molecular determinants and clinical implications remain poorly characterized. Through genomic analysis of 688 cancer-related genes combined with genomic scar assessment, we investigated HRD molecular features and clinical relevance of HRD across three cohorts: an EC cohort from Sun Yat-sen University Cancer Center (SYSUCC, n = 114), the Cancer Genome Atlas EC cohort (n = 500), and a high-grade serous ovarian cancer (HGSOC) cohort (n = 118). HRD was identified in 23.7% of SYSUCC EC cases, and HRD tumors paradoxically had fewer short-nucleotide variations in HRR genes than proficient (HRP) tumors (18.52% vs. 48.28%, P = 0.007). Mechanistic analysis revealed large-scale transition (LST) losses as the potential predominant HRD driver in EC, occurring significantly more frequently in HRD versus HRP tumors (74.1% vs 5.7%; P < 0.001). Comparative genomics demonstrated enrichment of HRR gene LST losses was EC-specific, contrasting with HGSOC where LST distribution was HRD-independent. Clinically, elevated HRD scores predicted reduced progression-free survival (HR 1.74, 95% CI 1.03-2.94; P = 0.04) yet enhanced platinum sensitivity (HR 0.41, 95% CI 0.18–0.94; P = 0.034). Our findings indicate that the HRD phenotype in EC, driven primarily by LST losses rather than short-nucleotide variations, serves as both a prognostic and predictive biomarker.
Similar content being viewed by others
Data availability
Human DNA data of the TCGA cohort (EC) are available via UCSC XENA (cohort name: UCEC; dataset name: MuTect2 Variant Aggregation and Masking; https://xenabrowser.net/datapages/) for interactive use. The HRD scores of the TCGA cohort (EC) analyzed during this study are included in ref. 35 and its supplementary information files. The data of the SYSUCC cohort are not publicly available but are available from the corresponding author on reasonable request. The data of the FUDAN cohort have been deposited into the CNGB Sequence Archive(CNSA) of China National GeneBank DataBase (CNGBdb) with accession numberCNP0001456.
Code availability
The underlying code for this study is not publicly available but may be made available to qualified researchers on reasonable request from the corresponding author.
References
Brooks, R. A. et al. Current recommendations and recent progress in endometrial cancer. CA Cancer J. Clin. 69, 258–279 (2019).
Lee, E. K. et al. Uterine serous carcinoma: molecular features, clinical management, and new and future therapies. Gynecol. Oncol. 160, 322–332 (2021).
Muggia, F. M. et al. Phase II trial of the pegylated liposomal doxorubicin in previously treated metastatic endometrial cancer: a Gynecologic Oncology Group study. J. Clin. Oncol. 20, 2360–2364 (2002).
Lincoln, S. et al. Activity of paclitaxel as second-line chemotherapy in endometrial carcinoma: a Gynecologic Oncology Group study. Gynecol. Oncol. 88, 277–281 (2003).
Slomovitz, B. M. et al. A phase II study of the oral mammalian target of rapamycin inhibitor, everolimus, in patients with recurrent endometrial carcinoma. Cancer 116, 5415–5419 (2010).
Heeke, A. L. et al. Prevalence of homologous recombination-related gene mutations across multiple cancer types. JCO Precis. Oncol. 2, 1–13 (2018).
Wiegand, K. C. et al. Loss of BAF250a (ARID1A) is frequent in high-grade endometrial carcinomas. J. Pathol. 224, 328–333 (2011).
Kandoth, C. et al. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Jönsson, J.-M. et al. Homologous recombination repair mechanisms in serous endometrial cancer. Cancers 13, 254 (2021).
Mills, G. B. et al. Homologous recombination deficiency score shows superior association with outcome compared with its individual score components in platinum-treated serous ovarian cancer. Gynecol. Oncol. 141, 2–3 (2016).
Principe, D. R. et al. Frequency and prognostic value of mutations associated with the homologous recombination DNA repair pathway in a large pan cancer cohort. Sci. Rep. 10, 20223 (2020).
Post, C. C. B. et al. Efficacy and safety of durvalumab with olaparib in metastatic or recurrent endometrial cancer (phase II DOMEC trial). Gynecol. Oncol. 165, 223–229 (2022).
Madariaga, A. et al. Phase II trial assessing niraparib with or without dostarlimab (anti-PD-1) in recurrent endometrial carcinoma. J. Clin. Oncol. 39, 5574–5574 (2021).
Westin, S. N. et al. DUO-E/GOG-3041/ENGOT-EN10: a randomized phase III trial of first-line carboplatin and paclitaxel in combination with durvalumab, followed by maintenance durva with or without olaparib, in patients with newly diagnosed advanced or recurrent endometrial cancer. J. Clin. Oncol. 38, 283–299 (2020).
De Jonge, M. M. et al. Frequent homologous recombination deficiency in high-grade endometrial carcinomas. Clin. Cancer Res. 25, 1087–1097 (2019).
Patch, A.-M. et al. Whole–genome characterization of chemoresistant ovarian cancer. Nature 521, 489–494 (2015).
Elze, L. et al. Genomic instability in non-breast or ovarian malignancies of individuals with germline pathogenic variants in BRCA1/2. J. Natl. Cancer Inst. 116, 1904–1913 (2024).
Rempel, E. et al. Pan-cancer analysis of genomic scar patterns caused by homologous repair deficiency (HRD). npj Precis. Onc. 6, 36 (2022).
Ewing, A. et al. Structural variants at the BRCA1/2 loci are a common source of homologous repair deficiency in high-grade serous ovarian carcinoma. Clin. Cancer Res. 27, 3201–3214 (2021).
Smith, E. S. et al. Endometrial cancers in BRCA1 or BRCA2 germline mutation carriers: assessment of homologous recombination DNA repair defects. JCO Precis. Oncol. 3, PO.19.00103 (2019).
Huang, Y. et al. SMYD3 promotes endometrial cancer through epigenetic regulation of LIG4/XRCC4/XLF complex in non-homologous end joining repair. Oncogenesis 13, 3 (2024).
Wang, Z. C. et al. Profiles of genomic instability in high-grade serous ovarian cancer predict treatment outcome. Clin. Cancer Res. 18, 5806–5815 (2012).
Elvin, J. A. et al. Comprehensive genomic profiling with loss of heterozygosity to identify therapeutically relevant subsets of ovarian cancer. J. Clin. Oncol. 35, 5512 (2017).
Dillon, L. W.et al. DNA instability at chromosomal fragile sites in cancer. In Keeping up with Genome Sequence and Expression (ed. Neri, C. & Papaioannou, V.) 161–188 (Bentham Science Publishers, 2014).
Thys, R. G. et al. DNA secondary structure at chromosomal fragile sites in human disease. Curr. Genom. 16, 60–70 (2015).
Park, W. et al. Clinico-genomic characterization of ATM and HRD in pancreas cancer: application for practice. Clin. Cancer Res. 28, 4782–4792 (2022).
Karakostis, K. et al. The DNA damage sensor ATM kinase interacts with the p53 mRNA and guides the DNA damage response pathway. Mol. Cancer 23, 21 (2024).
Pandey, S. N. et al. ATM and p53 in aging and cancer: a double-edged sword in genomic integrity. Biogerontology 26, 102 (2025).
Do, Q.-T. et al. Genome-wide mapping and quantification of DNA damage induced by catechol estrogens using Click-Probe-Seq and LC-MS. Commun. Biol. 8, 357 (2025).
Bostan, I.-S. et al. Landscape of endometrial cancer: molecular mechanisms, biomarkers, and target therapy. Cancers 16, 2027 (2024).
Goel, A. et al. Characterization of sporadic colon cancer by patterns of genomic instability. Cancer Res. 63, 1608–1614 (2003).
León-Castillo, A. et al. Clinicopathological and molecular characterisation of ‘multiple-classifier’ endometrial carcinomas. J. Pathol. 250, 312–322 (2020).
Zhang, Y. et al. ATM mutations as an independent prognostic factor and a potential biomarker for immune checkpoint therapy in endometrial cancer. Pathol. Res. Pract. 216, 153032 (2020).
Makker, V. et al. Lenvatinib plus pembrolizumab for advanced endometrial cancer. N. Engl. J. Med. 386, 437–448 (2022).
Knijnenburg, T. A. et al. Genomic and molecular landscape of DNA damage repair deficiency across The Cancer Genome Atlas. Cell Rep. 23, 239–254 (2018).
Feng, Z. et al. Homologous recombination deficiency status predicts response to platinum-based chemotherapy in Chinese patients with high-grade serous ovarian carcinoma. J. Ovarian Res. 16, 53 (2023).
Acknowledgements
The authors thank the participants and staff of the Sun Yat-sen University Cancer Center for their contributions. The research reported in this manuscript was supported by Guangdong Basic and Applied Basic Research Foundation (No. 2022A1515111160) to W.W., co-funding by the National Natural Science Foundation of China (No. 82203753) and Guangzhou Science and Technology Program (Grant No. 2023A04J1784) to Q.L.
Author information
Authors and Affiliations
Contributions
W.W., Z.M.D., and L.J.D.: conceptualization, methodology, draft polish. W.T. and Q.Q.L.: provided clinical samples and data collection. H.W.: provided data of the FUDAN cohort. Y.X.: analyzed data. M.F.L., B.B.L., and D.D.: writing-original draft. All the authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wan, T., Li, Q., Hao, W. et al. HRD in endometrial cancer: LST loss drives distinct genomic profile and platinum response. npj Precis. Onc. (2026). https://doi.org/10.1038/s41698-026-01347-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-026-01347-8