Abstract
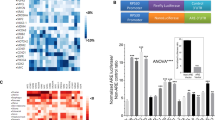
The recent successes of HER2-targeting agents, even in tumors characterized by FDA-approved molecular testing as HER2-negative or non-amplified, have underscored the limitations of current diagnostic approaches for accurately identifying patients with actionable HER2/EGFR activation/phosphorylation. We therefore performed a multi-omic investigation integrating clinical next-generation sequencing with a CLIA-certified reverse-phase protein array (RPPA) assay and laser microdissection-enriched tumor samples to characterize ERBB2/HER2 at the DNA, RNA, protein, and phosphoprotein level in patients with advanced pan-cancer solid tumor malignancies. Functional pathway activation mapping by RPPA revealed several patients with ERBB2 genomic or transcriptomic alterations and/or HER2Total-positivity by immunohistochemistry who exhibited no significant HER2Y1248 activation/phosphorylation. In contrast, other patients lacking ERBB2 genomic/transcriptomic alterations demonstrated significant HER2Y1248 activation/phosphorylation with co-activation of EGFRY1173, a marker associated with prognostic significance. Our results highlight the weak concordance between ERBB2 genomic/transcriptomic alterations and downstream activation of HER family signaling and support the inclusion of functional proteomic/phosphoproteomic analysis as an essential component of precision oncology pipelines to more accurately guide selection of HER2- and EGFR-targeted therapies.
Similar content being viewed by others
Data availability
Sequencing data supporting the findings of this study were generated by Tempus AI, Inc. The deidentified data used in the research were collected in a real-world healthcare setting and are subject to controlled access for privacy and proprietary reasons. When possible, derived data supporting the findings of this study have been made available within the paper and its supplementary figures and tables. The RPPA abundance data is available in Supplementary Data 1.
Code availability
Code sharing is not applicable to this article as no custom codes were generated or analyzed during the current study.
References
Cheng, X. A comprehensive review of HER2 in cancer biology and therapeutics. Genes 15 (2024).
Yarden, Y. & Sliwkowski, M. X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol. 2, 127–137 (2001).
Sergina, N. V. & Moasser, M. M. The HER family and cancer: emerging molecular mechanisms and therapeutic targets. Trends Mol. Med. 13, 527–534 (2007).
Li, B. T. et al. 654O Efficacy and safety of trastuzumab deruxtecan (T-DXd) in patients (pts) with solid tumors harboring specific HER2-activating mutations (HER2m): Primary results from the international phase II DESTINY-PanTumor01 (DPT-01) study. Ann. Oncol. 34, S459–S460 (2023).
Meric-Bernstam, F. et al. Efficacy and safety of trastuzumab deruxtecan (T-DXd) in patients (pts) with HER2-expressing solid tumors: DESTINY-PanTumor02 (DP-02) interim results. J. Clin. Oncol. 41, LBA3000–LBA3000 (2023).
Tarantino, P. et al. HER2-Low Breast Cancer: Pathological and Clinical Landscape. J. Clin. Oncol. 38, 1951–1962 (2020).
Schettini, F. et al. Clinical, pathological, and PAM50 gene expression features of HER2-low breast cancer. NPJ Breast Cancer 7, 1 (2021).
Modi, S. et al. Trastuzumab Deruxtecan in previously treated HER2-low advanced breast cancer. N. Engl. J. Med. 387, 9–20 (2022).
Johnston, L. E. et al. Proteomics based selection achieves complete response to HER2 therapy in HER2 IHC 0 breast cancer. npj Precis. Oncol. 8, 203 (2024).
Randall, J. et al. Quantitative proteomic analysis of HER2 protein expression in PDAC tumors. Clin. Proteom. 21, 24 (2024).
Wulfkuhle, J. D. et al. Molecular analysis of HER2 signaling in human breast cancer by functional protein pathway activation mapping. Clin. Cancer Res 18, 6426–6435 (2012).
Wulfkuhle, J. D. et al. HER family protein expression and activation predicts response to combination T-DM1/pertuzumab in HER2+ patients in the I-SPY 2 TRIAL. J. Clin. Oncol. 37, 3133–3133 (2019).
Wulfkuhle, J. D. et al. Evaluation of the HER/PI3K/AKT family signaling network as a predictive biomarker of pathologic complete response for patients with breast cancer treated with Neratinib in the I-SPY 2 TRIAL. JCO Precis. Oncol. 1–20 (2018).
Hunt, A. L. et al. Real-time functional proteomics enhances therapeutic targeting in precision oncology molecular tumor boards. NPJ Precis. Oncol. 9, 111 (2025).
Mosele, F. et al. Trastuzumab deruxtecan in metastatic breast cancer with variable HER2 expression: the phase 2 DAISY trial. Nat. Med. 29, 2110–2120 (2023).
Pierobon, M. et al. Multi-omic molecular profiling guide’s efficacious treatment selection in refractory metastatic breast cancer: a prospective phase II clinical trial. Mol. Oncol. 16, 104–115 (2022).
Gallagher, R. I. et al. Protein signaling and drug target activation signatures to guide therapy prioritization: Therapeutic resistance and sensitivity in the I-SPY 2 Trial. Cell Rep. Med. 4, 101312 (2023).
Tarantino, P. et al. Quantitative pre-treatment assessment of trastuzumab deruxtecan (T-DXd) antibody target (HER2) and payload target (topoisomerase 1, Topo1) to predict outcomes in metastatic breast cancer (MBC). J. Clin. Oncol. 43, 1032–1032 (2025).
Schlam, I., Tarantino, P. & Tolaney, S. M. Overcoming Resistance to HER2-directed therapies in breast cancer. Cancers 14, https://doi.org/10.3390/cancers14163996 (2022).
Esserman, L. J. et al. Pathologic complete response predicts recurrence-free survival more effectively by cancer subset: results From the I-SPY 1 TRIAL—CALGB 150007/150012, ACRIN 6657. J. Clin. Oncol. 30, 3242–3249 (2012).
Edfors, F. et al. Gene-specific correlation of RNA and protein levels in human cells and tissues. Mol. Syst. Biol. 12, 883 (2016).
Ørntoft, T. F., Thykjaer, T., Waldman, F. M., Wolf, H. & Celis, J. E. Genome-wide study of gene copy numbers, transcripts, and protein levels in pairs of non-invasive and invasive human transitional cell carcinomas. Mol. Cell. Proteom. 1, 37–45 (2002).
Greenbaum, D., Colangelo, C., Williams, K. & Gerstein, M. Comparing protein abundance and mRNA expression levels on a genomic scale. Genome Biol. 4, 117 (2003).
Hunt, A. L. et al. Extensive three-dimensional intratumor proteomic heterogeneity revealed by multiregion sampling in high-grade serous ovarian tumor specimens. iScience 24, 102757 (2021).
Burdett, N. L. et al. Multiomic analysis of homologous recombination-deficient end-stage high-grade serous ovarian cancer. Nat. Genet. https://doi.org/10.1038/s41588-023-01320-2 (2023).
Hunt, A. L. et al. Mapping three-dimensional intratumor proteomic heterogeneity in uterine serous carcinoma by multiregion microsampling. Clin. Proteom. 21, 4 (2024).
Tian, Q. et al. Integrated genomic and proteomic analyses of gene expression in mammalian cells. Mol. Cell. Proteom. 3, 960–969 (2004).
Chen, G. et al. Discordant protein and mRNA expression in lung adenocarcinomas. Mol. Cell Proteom. 1, 304–313 (2002).
Bateman, N. W. et al. Proteogenomic analysis of enriched HGSOC tumor epithelium identifies prognostic signatures and therapeutic vulnerabilities. npj Precis. Oncol. 8, 68 (2024).
Gonçalves, E. et al. Widespread post-transcriptional attenuation of genomic copy-number variation in cancer. Cell Syst. 5, 386–398.e384 (2017).
Hunter, F. W. et al. Mechanisms of resistance to trastuzumab emtansine (T-DM1) in HER2-positive breast cancer. Br. J. Cancer 122, 603–612 (2020).
Saleh, K. et al. Mechanisms of action and resistance to anti-HER2 antibody-drug conjugates in breast cancer. Cancer Drug Resist. 7, 22 (2024).
Luque-Cabal, M., García-Teijido, P., Fernández-Pérez, Y., Sánchez-Lorenzo, L. & Palacio-Vázquez, I. Mechanisms behind the resistance to Trastuzumab in HER2-amplified breast cancer and strategies to overcome it. Clin. Med Insights Oncol. 10, 21–30 (2016).
Unni, A. M., Lockwood, W. W., Zejnullahu, K., Lee-Lin, S. Q. & Varmus, H. Evidence that synthetic lethality underlies the mutual exclusivity of oncogenic KRAS and EGFR mutations in lung adenocarcinoma. Elife 4, e06907 (2015).
Lee, S. M. & Oh, H. RAS/RAF mutations and microsatellite instability status in primary colorectal cancers according to HER2 amplification. Sci. Rep. 14, 11432 (2024).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Suzuki, R. & Shimodaira, H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006).
Clark, A. S. et al. Neoadjuvant T-DM1/pertuzumab and paclitaxel/trastuzumab/pertuzumab for HER2(+) breast cancer in the adaptively randomized I-SPY2 trial. Nat. Commun. 12, 6428 (2021).
Balk, E. M. et al. Effects of Statins on nonlipid serum markers associated with cardiovascular disease. Ann. Intern. Med. 139, 670–682 (2003).
Vogel, C. & Marcotte, E. M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 13, 227–232 (2012).
de Sousa Abreu, R., Penalva, L. O., Marcotte, E. M. & Vogel, C. Global signatures of protein and mRNA expression levels. Mol. Biosyst. 5, 1512–1526 (2009).
Li, X., Zhao, L., Li, W., Gao, P. & Zhang, N. HER2-targeting CAR-T cells show highly efficient anti-tumor activity against glioblastoma both in vitro and in vivo. Genes Immun. 25, 201–208 (2024).
Ahmed, N. et al. HER2-specific T cells target primary glioblastoma stem cells and induce regression of autologous experimental tumors. Clin. Cancer Res 16, 474–485 (2010).
Zhang, C. et al. ErbB2/HER2-Specific NK cells for targeted therapy of Glioblastoma. J. Natl. Cancer Inst. 108, https://doi.org/10.1093/jnci/djv375 (2015).
Ramezani, M., Siami, S., Rezaei, M., Khazaei, S. & Sadeghi, M. An immunohistochemical study of HER2 expression in primary brain tumors. Biomedicine 10, 21–27 (2020).
Xu, B. et al. The expression and prognostic value of the epidermal growth factor receptor family in glioma. BMC Cancer 21, 451 (2021).
Beaubier, N. et al. Clinical validation of the Tempus xT next-generation targeted oncology sequencing assay. Oncotarget 10 (2019).
Beaubier, N. et al. Integrated genomic profiling expands clinical options for patients with cancer. Nat. Biotechnol. 37, 1351–1360 (2019).
Tempus A. I., I. Tempus xT Validation Summary. Tempus AI, Inc. Official Website. https://www.tempus.com/resources/document-library/Tempus-xT_Validation (2024).
Hunt, A. L. et al. Integration of multi-omic data in a molecular tumor board reveals EGFR-associated ALK-inhibitor resistance in a patient with inflammatory myofibroblastic cancer. Oncologist https://doi.org/10.1093/oncolo/oyad129 (2023).
Baldelli, E. et al. in Molecular Profiling: Methods and Protocols Vol. 1606 (ed Virginia Espina) 149–169 (Springer New York, 2017).
Youden, W. J. Index for rating diagnostic tests. Cancer 3, 32–35 (1950).
R Core Team _R: A Language and Environment for Statistical Computing_. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/ (2024).
Wickham, H. Welcome to the Tidyverse. J. Open Source Softw. 4 https://doi.org/10.21105/joss.01686 (2019).
Pantano, L. et al. DEGreport: Report of DEG analysis. https://doi.org/10.18129/B9.bioc.DEGreport, R package version 1.44.0, https://bioconductor.org/packages/DEGreport, https://bioconductor.org/packages/release/bioc/html/DEGreport.html (2025).
Aragon T epitools: Epidemiology Tools. R package version 0.5-10.1, https://CRAN.R-project.org/package=epitools (2020).
Acknowledgements
Acknowledgements: The authors would like to acknowledge Dr. Paulette Mhawech-Fauceglia for histopathology image analysis support.
Author information
Authors and Affiliations
Contributions
Study concept and design: T.L.C., E.F.P., T.P.C. Management of clinical records: J.R., L.J., W.S. Data acquisition, analysis, and interpretation: A.L.H., J.D.O., S.M., M.S., V.C., T.P.C., E.F.P., T.L.C. Writing and/or reviewing: A.L.H., J.R., J.D.O., L.J., W.S., V.C., G.L.M., N.W.B., T.P.C., E.F.P., T.L.C. All authors gave final approval of the completed work and are accountable for accuracy and integrity. The views expressed in this article are those of the author(s) and do not necessarily reflect the official policy or position of the Uniformed Services University of the Health Sciences (USUHS), Department of the Navy, Department of the Air Force, Department of the Army, Department of War, or the United States Government. Mention of trade names, commercial products, or organizations does not imply endorsement by the U.S. Government.
Corresponding authors
Ethics declarations
Competing interests
T.P.C. is a Thermo Fisher Scientific, Inc. SAB member and receives research funding from AbbVie. E.F.P. receives research funding from Genentech, Pfizer, Mirati, Springworks Therapeutics, Deciphera, AbbVie, and is a co-inventor of the RPPA Technology described herein, and related HER2 biomarker patents, and receives royalties on the related license agreements.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hunt, A.L., Randall, J., Ogata, J.D. et al. Pan-cancer multi-omic ERBB2-HER2 characterization using next-generation sequencing and quantitative proteomics. npj Precis. Onc. (2026). https://doi.org/10.1038/s41698-026-01351-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-026-01351-y