Abstract
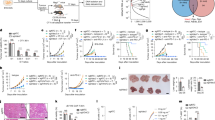
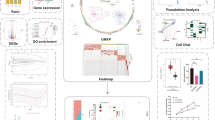
VDAC2’s known role in cancer and immune regulation via enhancing the CD8+ T cell-mediated killing, and it is worth systematically digging out the role of VDAC2 in pan-cancer based on this research. Bulk RNA sequencing, single-cell RNA sequencing, and spatial transcriptomic analyses were utilized to explore the role of VDAC2 from multiple perspectives in pan-cancers. RT-PCR, cell co-culture, CCK-8 assay, Transwell invasion assays, and ELISA were performed to validate the expression level and biological function. VDAC2 was upregulated in the majority of pan-cancers, and functional enrichment analyses displayed that VDAC2 may take part in the biological progress of energy metabolism, mitochondrial damage and cell proliferation. The landscape of VDAC2 expression and immune infiltration was constructed, and the VDAC2-BAK1-IFNγ pathway was identified in digestive cancer. VDAC2 had the potential to serve as a novel prognostic, screening cancer indicator and immune therapeutic target sensitive to various drugs. Overexpression of VDAC2 significantly promoted gastric cancer cell proliferation, invasion and immune invasion, as validated in vitro experiments. In short, our pan-cancer analysis constructed a comprehensive landscape of VDAC2’s oncogenic role, establishing VDAC2 + -BAK1-IFNγ as an important pathway in tumor progression and immune evasion. VDAC2 emerges not only as a valuable prognostic biomarker but also as a promising novel therapeutic target.
Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Chin, H. S. et al. VDAC2 enables BAX to mediate apoptosis and limit tumor development. Nat. Commun. 9, 4976 (2018).
Yuan, Z. et al. Key residues in the VDAC2-BAK complex can be targeted to modulate apoptosis. PLoS Biol. 22, e3002617 (2024).
Yuan, S. et al. VDAC2 loss elicits tumour destruction and inflammation for cancer therapy. Nature 640, 1062–1071 (2025).
Patel, S. J. & Chen, Z. J. VDAC2 brake release: unleashing inflammation via IFNγ. Trends Pharm. Sci. 46, 695–696 (2025).
Li, K. et al. Differential regulation of BAX and BAK apoptotic activity revealed by small molecules. Sci. Adv. 11, eadr8146 (2025).
Wang, S. et al. Identification of a ferroptosis-related gene signature (FRGS) for predicting clinical outcome in lung adenocarcinoma. PeerJ 9, e11233 (2021).
Aono, Y. et al. Sulindac sulfone inhibits the mTORC1 pathway in colon cancer cells by directly targeting voltage-dependent anion channel 1 and 2. Biochem. Biophys. Res. Commun. 505, 1203–1210 (2018).
Shixiang, W. et al. UCSCXenaShiny: an R/CRAN package for interactive analysis of UCSC Xena data. Bioinformatics 38, 527–529 (2021).
Shi, J. et al. The web-based portal spatialTME integrates histological images with single-cell and spatial transcriptomics to explore the tumor microenvironment. Cancer Res. 84, 1210–1220 (2024).
Xun, Z. et al. Reconstruction of the tumor spatial microenvironment along the malignant-boundary-nonmalignant axis. Nat. Commun. 14, 933 (2023).
Arthur, L. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1139–1140 (2011).
Huating, Y. et al. CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res. 47, D900–D908 (2018).
Sonja, H., Robert, C. & Justin, G. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Suoqin, J. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Taiwen, L. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res 48, W509–W514 (2020).
Tu-Xiong, H. & Li, F. The immune landscape of esophageal cancer. Cancer Commun. 39, 79 (2019).
Chenchen, F. et al. KnockTF 2.0: a comprehensive gene expression profile database with knockdown/knockout of transcription (co-)factors in multiple species. Nucleic Acids Res 52, D183–D193 (2023).
Jun-Hao, L. et al. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42, D92–97 (2013).
Michael, R. et al. TCGASpliceSeq: a compendium of alternative mRNA splicing in cancer. Nucleic Acids Res 44, D1018–1022 (2015).
Balázs, G. et al. Transcriptome-level discovery of survival-associated biomarkers and therapy targets in non-small-cell lung cancer. Br. J. Pharmacol. 181, 362–374 (2023).
Geeleher, P., Cox, N. & Huang, R. S. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE 9, e107468 (2014).
Geeleher, P., Cox, N. J. & Huang, R. S. Clinical drug response can be predicted using baseline gene expression levels and in vitro drug sensitivity in cell lines. Genome Biol. 15, R47 (2014).
Danielle, M. & Robert, F. G. oncoPredict: an R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Brief Bioinform. 22, bbab260 (2021).
Tathiane, M. M. et al. Machine learning identifies stemness features associated with oncogenic dedifferentiation. Cell 173, 338–354 (2018).
Chen, Y. et al. A survey of optimal strategy for signature-based drug repositioning and an application to liver cancer. Elife 11, e71880 (2022).
Kai, Z. et al. VDAC2 interacts with PFKP to regulate glucose metabolism and phenotypic reprogramming of glioma stem cells. Cell Death Dis. 9, 988 (2018).
Zhao, W. B. et al. Superantigen-fused T cell engagers for tumor antigen-mediated robust T cell activation and tumor cell killing. Mol. Ther. 32, 490–502 (2024).
Acknowledgements
We thank the Home for Researchers editorial team (www.home-for-researchers.com) that helps as for the language editing service and Figdraw for drawing pictures. We thank Sparkle (https://grswsci.top/analyze/) for multi-omics data analysis. This study was funded by the Ningbo Top Medical and Health Research Program (No. 2023020612) and the Key Scientific and Technological Projects of Ningbo (No. 2021Z133).
Author information
Authors and Affiliations
Contributions
Yan and Shao made outstanding contributions to identify this manuscript. Ye revised it critically for vital intellectual content. Dong and Wang completed PCR and functional experiment in vitro. Yan drew the figure and wrote the draft. All authors contributed to the figures and approved the final submitted manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yan, J., Wang, J., Dong, H. et al. Perspectives from machine learning and multi-omics to decoding the effects of VDAC2 malignant subsets on tumor evolution. npj Precis. Onc. (2026). https://doi.org/10.1038/s41698-026-01394-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-026-01394-1