Abstract
Traditional disease classification models often disregard the clinical significance of misclassifications and lack interpretability. To overcome these challenges, we propose a hierarchical prototypical decision tree (HPDT) for skin lesion classification. HPDT combines prototypical networks and decision trees, leveraging a class hierarchy to guide interpretable predictions from general to specific categories. By incorporating a hierarchy-based distance matrix, the model prioritizes less severe misclassifications while maintaining diagnostic accuracy. Evaluated on a dataset of 235,268 dermoscopic images across 65 conditions, HPDT outperforms flat classifiers and existing hierarchical methods in accuracy, error severity reduction, and interpretability. It also generalizes effectively to unseen classes. These results highlight the value of integrating clinical hierarchies into model design and training to improve diagnostic reliability and decision transparency, demonstrating HPDT’s potential for clinical decision support.
Similar content being viewed by others
Introduction
Recent advances in deep learning and the availability of large medical datasets have greatly improved the accuracy of classification algorithms for medical image diagnosis1. Typically, these algorithms are trained with semantically independent, one-hot encoded labels, to produce flat N-way predictions. The standard paradigm for evaluating a model is to treat all classes other than the ground truth as equally wrong. However, this flat classification approach has two major limitations: it conveys little insight into the algorithmic decision-making process, while clinical diagnosis demands transparency of predictions from algorithms. Moreover, it fails to account for the semantic relationships between disease classes, where certain classes may be more semantically related to each other than to other classes. This implies that some misclassifications may have different clinical implications than others. For example, in dermatology, misclassifying a malignant lesion as benign poses significantly higher risks than confusing two benign conditions. While utility-based frameworks have been proposed to minimize the risk of misclassification by presenting multiple plausible classes for ambiguous predictions2,3, these approaches often lack mechanisms to assess and manage the clinical severity of error predictions–a critical consideration in high-stakes applications like diagnostics. To build more clinically applicable algorithms, it is desirable to enhance transparency in a model’s decision-making process and reduce the risk of making a life-threatening misdiagnosis.
Incorporating disease class hierarchies into algorithm development through hierarchical classification presents a promising solution to addressing these limitations. First, hierarchical models shed light on the decision process by breaking a complicated classification decision into a set of intermediate decisions and then making predictions in a sequential manner4,5. Therefore, humans can easily trace and locate where an error decision occurs by inspecting the sequential prediction path. Second, class relationships can be expressed by a taxonomic hierarchy tree of class labels defined by clinical experts. Training a model with such hierarchical class labels can facilitate the classification and reduce the severity of mistakes by restricting the error predictions to be close to their true labels in the class tree6,7,8, ensuring the predicted class and ground truth belong to the same super class. Despite these benefits, the integration of deep learning models with class hierarchies for medical image classification remains relatively unexplored.
Existing hierarchy-aware models can be grouped into three categories: (1) hierarchical label embeddings9, (2) hierarchical losses10, and (3) hierarchical architectures11,12,13,14,15. Hierarchical label embeddings and hierarchical losses are considered indirect methods for modeling class hierarchies, as they modify the label space or the objective function without explicitly producing tree-structured hierarchical outputs. Hierarchical label embedding methods primarily focus on encoding class labels into continuous vectors that capture semantic correlations from the hierarchy. The classification process, in this context, involves matching image embeddings to these label vectors. In contrast, hierarchical loss methods adapt the loss function based on the class hierarchy, imposing higher penalties for incorrect predictions that are further from the true label within the hierarchy. However, these indirect methods often suffer from accuracy degeneration8,16 and the single-label outputs lack the details about how a decision is formed. Hierarchical architectures, on the other hand, are designed to produce multi-granularity outputs according to class hierarchies by introducing multi-branch structures or sequence modules12. Despite this, these hierarchical models provide limited insight into the decision-making process, as they perform hierarchical classifications independently at each level. This separation can lead to inconsistent predictions across different hierarchical levels14,15.
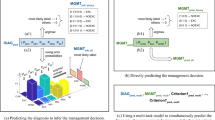
In this study, we propose a hierarchical classification model for skin disease diagnosis to produce sensible sequential decisions while reducing the severity of mistakes and maintaining diagnostic accuracy. Our approach combines prototypical networks with decision trees to form a hierarchical prototype decision tree model (HPDT). First, the prototypical network jointly learns image representations and class prototypes by optimizing the distribution of Euclidean distance between them17. A decision tree is then constructed on top of the prototypical network to make path predictions along the class hierarchy, rather than independent predictions at each hierarchical level. Unlike previous works that learn separate prototypes for class nodes in the taxonomy4,18, we compute prototypes for the internal class nodes by aggregating the prototypical vectors of leaf class nodes. This oblique decision tree enables flexible adaptation of the model to different hierarchical trees without extensive architectural modification. To assess the severity of misclassifications, we introduce a metric based on the height of the lowest common ancestor (LCA) in the class tree19,20 (see Fig. 1). This LCA-based distance quantifies the semantic relationship between classes by measuring their shared ancestry; lower LCA heights indicate greater similarity. By incorporating this metric into both the evaluation and loss functions, we align the decision-making process with the class hierarchy, ensuring that, when misclassifications occur, they remain within taxonomically similar categories, thus reducing diagnostic severity (see Fig. 2). The contributions of this paper can be summarized as follows:
The class distance is the height of the LCA. The height of a node is defined as the length of the longest path from that node to a leaf node, with leaf nodes having a height of zero.
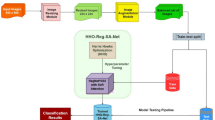
Expert knowledge defines a taxonomy of disease categories, which guides the construction of a hierarchical tree model. The hierarchical tree provides interpretable path predictions, offering insight into the decision-making process. Class relationships are encoded in a distance matrix, which quantifies the severity of misclassifications and informs the learning process of the decision tree. This framework supports error severity assessment and improves the overall transparency and reliability of predictions.
1) We propose a prototypical decision tree model for hierarchical image classification (Fig. 3), which decomposes complex classification tasks into sensible, sequential steps within the hierarchy. Our model not only maintains high accuracy but also provides transparent decision paths for clinician verification. Moreover, it adapts flexibly to different disease taxonomies without architectural changes, making it suitable for evolving medical standards.
The model begins with a prototypical network that jointly learns image features and class prototypes. A prototypical decision tree is then constructed using these prototypes to produce sequential path predictions that align with the hierarchical structure of the classes. During training, a hierarchical class distance matrix is utilized to guide prototype learning, ensuring that the learned representations reflect semantic relationships among classes.
2) We introduce a normalized LCA height metric to quantify the severity of misclassification and further incorporate this metric to direct the learning process of the proposed decision tree model. This approach enables the model to learn semantic-aware prototypes and prioritize less severe mistakes, aligning model behavior more closely with practical, clinically relevant error management.
3) We validate our approach on a dataset of over 230,000 dermoscopic images covering 65 skin diseases across three taxonomic levels. Collected from Molemap clinics in Australia and New Zealand, this dataset represents one of the largest hierarchically structured skin disease datasets evaluated. Our results show that the model performs competitively with flat classifiers and existing hierarchical approaches while offering added interpretability, error severity insights, and improvement in unseen class recognition, demonstrating potential for clinical applicability.
Results
Study design and overview
We systematically evaluate our proposed HPDT framework’s effectiveness in addressing key challenges in medical image diagnosis: the need for transparent decision-making processes and clinically relevant error management. Our evaluation framework encompasses four specific dimensions: performance comparison with existing methods, model decision interpretability, generalization to unseen classes, and component analysis through ablation studies. We evaluate our approach on a large-scale teledermatology dataset from Molemap clinics, comprising 235,268 dermoscopic images spanning 65 skin conditions organized in a three-level hierarchy. The conditions include 53 benign and 12 malignant classes, reflecting the natural distribution and hierarchical relationships of skin lesions in clinical practice. Our evaluation metrics include both traditional classification metrics (accuracy) and hierarchy-aware measures (hierarchical F1 score, severity of mistakes using normalized LCA height, and top-k hierarchical distance), quantifying both diagnostic accuracy and clinical relevance of predictions. We first benchmark HPDT against 15 baseline models representing different paradigms in hierarchical classification, from traditional CNN-based hierarchy-agnostic approaches to state-of-the-art hierarchical architectures. To ensure statistical rigor, we conduct analysis using Cohen’s d effect size across all evaluation metrics. We then analyze the interpretability of our model’s decision process through its hierarchical prediction pathway, demonstrating how it provides transparent, sequential decisions for clinical verification. This is followed by evaluating its capability to handle unseen skin conditions, a critical requirement for real-world clinical applications. Finally, through ablation studies, we examine how each model component contributes to both classification performance and error severity reduction.
Comparison to existing methods
We compare our method to several baseline models, chosen to represent diverse approaches in hierarchical classification. Our selection includes a basic CNN model21 and prototype network17 as hierarchy-agnostic baselines to demonstrate the impact of incorporating hierarchical information. For hierarchy-aware models, we carefully selected methods across three categories representing different paradigms for handling hierarchical relationships: Hierarchical loss: MC22, HXE6, YOLO-v223. For HXE, we report results under different settings of the alpha; b) Hierarchical label embedding: Softlabel6, hyperspherical embeddings24, and hyperbolic embeddings9. Similar to6, we experiment with different settings of beta to regulate the hardness of soft labels; c) Hierarchical architecture: multi-task CNN, chang et al.16, HAF8, DHC25, and a CNN-Transformer modified from26. Different from26, we let each query token to be responsible for a classification task at a hierarchical level. For all models, we use the same ImageNet pre-trained ResNet3427 as the backbone. These methods represent both classical approaches and current state-of-the-art techniques in hierarchical classification. The selection particularly emphasizes models that have shown promise in medical image analysis or have characteristics relevant to fine-grained visual classification tasks.
We conduct both quantitative metric comparisons and statistical significance analysis using Cohen’s d effect size, where ∣d∣ < 0.2 indicates a negligible effect and ∣d∣ > 0.8 indicates a large effect. The comparison results are summarized in Table 1, and Fig. 4. We observe that most hierarchy-aware models, such as MC, HXE, Hyperbolic-proto (fixed), and Yolo-V2, achieve less severe mistake severity in top-1 predictions and smaller hierarchical distance in top-5 predictions compared to the hierarchy-agnostic CNN models. This is supported by moderate negative effect sizes (−0.5 to −0.7) in error severity metrics. However, their overall accuracy is lower than that of the basic CNN and prototypical CNN, with effect sizes ranging from −0.3 to −0.7 for accuracy comparison. This result indicates that these hierarchy-aware models indeed learn class relationships, as indicated by semantically closer top-5 predictions to the true labels, but they also produce more low-severity error predictions, which ultimately degrade the accuracy. While models with hierarchical label embeddings or hierarchical loss functions incorporate class hierarchy during training, they make flat predictions directly at the leaf level. As a result, the hierarchy is not explicitly accessible during inference, which limits interpretability for analyzing decision processes at intermediate levels. In contrast, hierarchical architectural models, such as HAF, DHC, and CNN-Transformer directly leverage class hierarchy to construct hierarchical architecture. This improves the accuracy as well as the hierarchical metrics compared to the basic CNN model, demonstrated by small to moderate positive effect sizes (0.2–0.5) across multiple metrics. However, because these models perform classification independently at each taxonomic level, they may produce inconsistent predictions across levels (see Figs. 6, 7), reducing the interpretation of the decision pathway.
Each heatmap shows pairwise Cohen’s d effect sizes between different models, where color intensity and values indicate the magnitude and direction of differences. Blue indicates positive effect sizes (improvement over comparison model) while red indicates negative effect sizes (decline compared to comparison model). a Effect sizes for accuracy differences. b Effect sizes for hierarchical F1 differences. c Effect sizes for error severity differences, where negative values indicate reduced error severity. d Effect sizes for top-5 hierarchical distance differences.
Our HPDT approach, on the other hand, provides a coherent sequence of decisions by computing path probability rather than giving separated predictions at each hierarchical level. HPDT achieves a competitive accuracy (60.87 for HPDT and 60.91 for HPDT-Guided) and hierarchical F1 score (81.56 for HPDT and 81.62 for HPDT-Guided), while with lower mistake severity (0.616 for HPDT-Guided) and superior hierarchical distance in top-5 predictions (0.437 for HPDT-Guided) compared to all other methods. The effect size analysis strongly supports these improvements: HPDT and HPDT-guided show large positive effect sizes for accuracy (d = 1.9–2.1) and hierarchical F1 score (d = 0.8–2.2), while demonstrating significant reductions in error severity (d = −0.8 to −2.1) and top-5 hierarchical distance (d = −1.0 to −1.5). These consistent large effect sizes across all metrics provide strong statistical evidence for the practical significance of our method’s improvements over existing approaches. Figure 5 illustrates the relationship between accuracy and three hierarchical metrics for all models. It highlights the trade-off between accuracy and the model’s ability to capture hierarchical relationships, showcasing the advantages of our method.
The scatter plots show relationships between classification accuracy and three hierarchical metrics: Hierarchical F1 score, Severity of Mistake, and Top-5 Hierarchical Distance.
Analysis on the interpretability of decision-making
The proposed decision trees provide insight into the decision-making process by decomposing a complex decision into smaller intermediate decisions. Humans can understand the model’s behavior by inspecting the probability when traversing nodes in a class tree. Figures 6, 7 show the probability of path traversals for four samples in three hierarchical levels as well as predictions from the two other best-performing hierarchical models. The DHC makes inconsistent decision results in Figs. 6 a, b and 7b. While the CNN-Transformer has a decision disagreement in Fig. 7b. We quantitatively assess the inconsistent hierarchical predictions for the hierarchical models and illustrate the ratio of inconsistent predictions of all other hierarchical models in Fig. 9a. Among them, the ratio of decision inconsistency is 7.6% and 7.2% for the DHC model and CNN-Transformer respectively. Our HPDT avoids this issue and captures well the model’s decision-making process with the sequential decisions obtained by traversing class nodes from coarse to fine.
a A malignant basal cell carcinoma (bcc) lesion correctly classified by HPDT (green path), while DHC provides an inconsistent prediction at Level 2 compared to its Level 3 prediction and CNN-Transformer fails at all levels. b A benign melanocytic atypical lesion correctly predicted by both HPDT and CNN-Transformer, whereas DHC fails at Level 3 and provides an inconsistent prediction relative to Level 2 (green checkmarks indicate correct predictions, red crosses indicate incorrect predictions, and yellow exclamation marks indicate inconsistent predictions. This comparison highlights the strengths of HPDT in hierarchical classification).
a A benign melanocytic blue lesion is misclassified as benign melanocytic nevus by HPDT (red path), with DHC and CNN-Transformer succeeding at Levels 1–2 but failing at Level 3. b A benign keratinocytic solar lentigo is misclassified as benign keratinocytic nevus by HPDT, while DHC and CNN-Transformer fail at Level 2 and show inconsistent predictions between Level 2 and 3 (green checkmarks indicate correct predictions, red crosses indicate incorrect predictions, and yellow exclamation marks indicate inconsistent predictions).
In some cases, the intermediate decisions may have very high uncertainty. These samples are likely ambiguous cases or hard cases for our model. To find them, we compute the max entropy for probabilities under each node along the prediction path in the class tree and then pick out samples with the largest decision entropy. Figure 7a, b are two top-ranked confusing samples. In Fig. 7a, the HPDT confuses benign melanocytic blue lesion as benign melanocytic nevus because the lesion exhibits a pattern of blue nodular. The model disperses among benign melanocytic blue, benign melanocytic compound, and benign melanocytic nevus. For Fig. 7b, our model has a highly diverse prediction under the benign melanocytic class. In Fig. 8, we calculate the accuracy and severity of mistakes for high-uncertainty samples. We rank the samples according to the entropy of the probability prediction and select the top 30% as high-uncertainty samples. Our model achieves an accuracy of 41.29% which is ~4% higher than the two flat models. This improvement demonstrates the model’s effectiveness against error propagation, even with highly uncertain samples. In addition, the mistake severity of HPDT is lower than the flat models by ~0.1.
From left to right: basic CNN, prototypical CNN, and HPDT. The comparison highlights differences in accuracy and severity of mistakes across models, with HPDT demonstrating improved performance in handling uncertain cases.
Result on the unseen class classification
An appealing aspect of hierarchical models is their capability of classifying unseen classes. Following the methodology4, we evaluate this capability through a systematic assessment of coarse-grained classification accuracy for unseen classes. In our experimental setup, we define unseen classes as those excluded from training but whose superclass categories exist within the known class hierarchy. For example, if “superficial spreading melanoma” is designated as an unseen class, while other melanoma subtypes are included in training, the model should still recognize it as belonging to the melanoma superclass. We evaluate prediction correctness at the parent node level: a prediction is considered correct if the predicted class and the true class share the same immediate parent node in the hierarchy.
We compare our decision tree model to flat models, i.e. basic CNN and prototypical CNN. For the tree model, the coarse-grained class predictions are directly computed from the outputs of inner nodes. While for flat models, we obtain the predictions as the superclass that the fine-level prediction belongs to. We experiment on three different number settings of seen class and unseen class: 50 seen classes vs. 15 unseen classes (50/15), 40 seen classes vs. 25 unseen classes (40/25) and 30 seen classes vs. 35 unseen classes (30/35). Results are listed in Table 2. As can be seen, the proposed HPDT consistently outperforms the basic CNN and prototypical CNN under all the class settings by a substantial margin. This performance advantage is particularly notable as the number of unseen classes increases, highlighting our model’s robust generalization capability.
It is important to note that this evaluation focuses on unseen classes within known superclass categories, distinct from out-of-distribution (OOD) detection where test samples may belong to entirely novel disease categories. For true OOD cases that fall outside our known hierarchy, the model defaults to classification at the coarsest level (malignant vs. benign), reflecting the fundamental diagnostic categories in clinical practice. Future extensions of our work could incorporate explicit OOD detection mechanisms28, such as prototype distance thresholds or uncertainty quantification, to identify samples that may require special consideration in clinical settings.
Result of ablation study
Here, we present ablation results to demonstrate how different configurations affect our model’s performance. Our model is optimized with two cross-entropy losses (\({{\mathcal{L}}}_{DCE}\) and \({{\mathcal{L}}}_{TCE}\)) to separate leaf and internal class nodes. We report two distinct accuracy metrics: flat accuracy, based on single-level class predictions, and tree accuracy, which evaluates the entire classification path using the oblique decision tree. By default, our model reports tree accuracy.
First, to assess the impact of distance metrics and the \({{\mathcal{L}}}_{TCE}\) loss for hierarchical representation learning, we compare the decision tree built on top of the prototypical CNN using cosine distance and Euclidean distance respectively, shown in Fig. 9b and c. Our results indicate that using Euclidean distance yields a higher tree accuracy than Cosine distance by ~16%, even without explicitly training the inner class nodes with surrogate tree loss \({{\mathcal{L}}}_{TCE}\). When applying the \({{\mathcal{L}}}_{TCE}\), the HPDT model using Euclidean distance produces better tree accuracy while maintaining its original flat accuracy. in contrast, with Cosine distance, applying the \({{\mathcal{L}}}_{TCE}\) improves the tree accuracy but slightly reduces flat accuracy.
a The ratio of inconsistent hierarchical predictions across compared hierarchical architecture models. b, c Performance comparisons of the HPDT trained with Cosine distance and Euclidean distance, respectively, highlighting the impact of distance metrics on both flat accuracy and tree accuracy.
Next, to examine the benefits of hierarchical modeling via the oblique decision tree, we compare three configurations: a flat model trained solely with leaf-level cross-entropy loss (Fig. 9b, cosine distance, without TCE), a regular top-down classifier that incorporates path probability through global optimization on the oblique decision tree (Fig. 9c, cosine distance, with TCE), and our full model. The flat model achieves a flat accuracy of 59.80%, while the regular top-down classifier achieves 57.62% in flat accuracy but improves to 60.62% in tree accuracy by leveraging the hierarchical structure. These results underscore the value of incorporating hierarchical relationships to enhance performance, particularly when evaluating the entire classification path. The superior performance of Euclidean distance further highlights its effectiveness in aligning feature representations with class prototypes and in accurately modeling hierarchical relationships within the oblique decision tree.
Then, we report the effect of ICD loss. Figure 10b shows that the accuracy improved from 60.73% to 60.91% when increasing λ from 0 to 0.4. While the accuracy drops when using a larger λ. Figure 11 shows HPDT trained with ICD loss has a more compact feature distribution. In Fig. 10, we calculate the average intra-class distance during model training. Both of these results suggest that ICD loss is capable of ensuring a compact feature space and an appropriate setting of λ can benefit the performance. Then, we experiment with different normalization settings and report results in Table 3. It can be seen that both batch normalization and layer normalization significantly improve the performance with a margin of 2.3% and 2.4% in accuracy, respectively. However, the L2 normalization degrades the performance with a decrease of ~6% in accuracy and an increase of 0.04 in the severity of mistakes.
a Average intra-class distance changes during training, demonstrating the impact of normalization and ICD loss on feature compactness. b Model accuracy under different weight settings for ICD loss, highlighting the trade-off between compactness and accuracy. c Discrepancy between the class distance matrix derived from learned prototypes and the hierarchical class tree, comparing models trained with and without hierarchical guidance.
The top and bottom rows represent Level 2 and Level 3 classes in the hierarchy, respectively. Comparisons include HPDT without ICD loss, HPDT with ICD loss, and HPDT with ICD loss and hierarchical guidance, illustrating the progressive improvement in feature separation and alignment with hierarchical class relationships.
Finally, we illustrate the effect of guided prototype learning with class distance. Figure 12 shows that the distance between prototypes learned with the explicit guidance of the hierarchical distance matrix preserves semantic relations. For the model learned without guidance, there are no clear relations in their class distance matrix. We calculate the discrepancy between the class distance matrix of learned prototypes and the hierarchical distance matrix in Fig. 10c using Eq. (11). The discrepancy of HPDT w/o guided is slightly lower than prototypical CNN which indicates the decision tree to some extent helps the prototypical network capture the semantic relations. The class distance guidance contributes most to learning the semantic-aware prototypes as the guided HPDT achieves the smallest discrepancy which is significantly lower than the baseline.
The comparison highlights the improved alignment of prototype distances with hierarchical relationships when hierarchical guidance is applied during training.
Discussion
Existing deep classification algorithms for medical diagnosis, which produce flat N-way predictions, lack transparency in the decision process and treat all misclassifications equally-overlooking the fact that some diagnostic errors carry greater consequences than others. For successful deployment in real clinical settings, diagnostic models must provide insight into the model’s decision-making and prioritize avoiding high-stakes errors. In this study, we address these needs by combining a decision tree model with class hierarchy regularized prototype learning. Decision trees are easy to understand and interpret because they transparently arrange decision rules in a hierarchical structure5,29. While the hierarchy-aware class prototypes enable a reduction in the model’s mistake severity by restricting error predictions into sibling classes of the ground truth.
We evaluate and compare our method to existing hierarchy-aware models on a large-scale skin lesion dataset. Our results reveal a trade-off between accuracy and mistake severity for models using hierarchical loss and hierarchical embeddings (shown in Fig. 5), consistent with the findings from nature image classification studies6,16. When inspecting prediction errors, we empirically observed that improvements in mistake severity metrics often come at the cost of increased low severity errors. The accuracy trade-off occurs primarily due to the inherent tension between capturing fine-grained class distinctions and respecting hierarchical structure. Hierarchical loss methods often struggle to optimize performance across all levels simultaneously, as emphasizing coarse-level accuracy can come at the expense of fine-grained discrimination8. Hierarchical label embedding methods face challenges in aligning image features with label embeddings in continuous space, which is complex and error-prone, requiring effective regularization or embedding strategies9. Furthermore, using pre-computed or fixed label embeddings can result in significant accuracy drops, as shown in the comparison between the hyperbolic-proto (fixed) and hyperbolic-proto (guided) models in Table 1.
Hierarchical models with architecture modification bypass this dilemma with improved accuracy and mistake measurement than the hierarchy-agnostic model. However, these hierarchical models provide little insight into the model’s decision process as they perform classifications at each hierarchical level independently, potentially leading to inconsistent prediction results. This issue, as noted in a recent study8, may be due to interference between coarse- and fine-level classifications within the feature space. In this study, we avoid explicitly performing classification at every taxonomical level but generate sequential decisions by computing path probability. Our method hence captures the decision process well and achieves superior performance.
Our model also mitigates error propagation, a common issue in hierarchical classification where early errors cascade through the hierarchy due to independent, top-down decisions and hard thresholds30,31. We achieve this by integrating two core mechanisms: soft decision-making and global optimization, with class hierarchy-guided prototype learning. Our soft oblique decision tree provides probabilistic outputs at each node, allowing the model to retain uncertainty without committing prematurely. Additionally, we apply loss across the entire path probability, jointly optimizing the decision tree and image encoding network. This design helps the model find the most probable path with a global perspective, tolerating intermediate errors and reducing cascading misclassifications.
Compared to traditional hierarchical classification approaches–often referred to as top-down classifiers30 and primarily developed before the deep learning era–such as local classifier per parent node (LCPN)30,32, nested dichotomies33,34, conditional probability trees35, and probabilistic classifier trees31, our model is specifically designed for transparency and severity-sensitive decision-making in medical applications. Traditional methods focus primarily on computational efficiency and scalability by breaking down multi-class tasks into binary decisions along a hierarchy, optimizing for speed in large datasets. However, these approaches often rely on randomly generated or computationally optimized hierarchies, which may not address the needs of clinical contexts, where decision transparency and sensitivity to error severity are critical. By incorporating clinically informed hierarchies and path-based optimization, our model enhances interpretability and aligns decision-making with clinical relevance.
However, our study has several limitations: First, we only explored learning the proposed model on a pre-defined class hierarchy which was built based on semantic similarity, whereas deep learning models prefer a class tree constructed based on visual similarity. This explains why performance improvement is limited. Combining semantic and visual class hierarchies could enhance classification performance. Second, our model was developed on the Euclidean space, however, hyperbolic space has been demonstrated to be more suitable for hierarchical modeling36,37. Hence, one more step forward will be to enhance our model with hyperbolic geometry. Lastly, it remains uncertain to what extent the sequential decisions would aid clinicians in practice. A reader study is needed to evaluate the real-world impact of our model on clinical diagnostic workflows.
Methods
Dataset
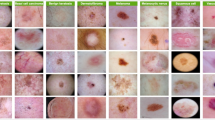
The Molemap dataset used in this study comprises 235,268 dermoscopic images, collected and verified through teledermatology practices in clinical settings. Each image is labeled by professionals following teledermatology standards and systematically organized into a three-level hierarchical taxonomy. This taxonomy includes 65 distinct skin conditions, divided into 53 benign and 12 malignant categories, based on their degree of malignancy. The hierarchical structure mirrors clinical diagnostic pathways, facilitating a systematic analysis of disease relationships and misclassification severity. Figure 13 illustrates the complete hierarchical taxonomy, detailing the organization of conditions from broad categories to specific diagnoses. Additionally, the class distribution and representative image samples are shown in Fig. 14, providing insights into the dataset’s diversity and complexity.
The structure illustrates the relationships between broad disease classes (e.g., malignant and benign) and their finer subcategories, highlighting the hierarchical organization used for classification tasks.
For each class, a representative dermoscopic image is provided to illustrate the variability within the dataset. The hierarchy spans benign keratinocytes, melanocytic, and vascular conditions, as well as malignant basal cell carcinoma, squamous cell carcinoma, and melanoma.
Model training
We implement a structured training protocol to ensure consistent evaluation across all experiments. The dataset is divided into training (70%), validation (10%), and testing (20%) sets. To enhance model generalization and robustness, we apply standard image augmentation techniques including random resized cropping, color transformation, and horizontal flipping uniformly across all experiments. All dermoscopic images are standardized to 320 × 320 pixels for consistent processing. The training process employs the ADAM optimizer with a batch size of 72 over 36 epochs. We implement a two-tier learning rate strategy: 3 × 10−5 for the pre-trained backbone network and 3 × 10−4 for newly added layers. The learning rate follows a step decay schedule with a decay factor of 0.1 applied every 10 epochs.
Hierarchical prototype decision tree
The overview of the proposed method is shown in Fig. 3. Our model consists of a feature encoding network and a set of leaf prototypes which constitute the prototypical decision tree for producing hierarchical predictions. In addition, the class distance regularization is introduced in prototype learning to incorporate semantic relations from the hierarchy such that the learned prototypes preserve the inter-class correlation in the embedding space.
A prototype network is often characterized by an image embedding network \({{f}}_{e}\left(\cdot \right)\) and a set of prototypical vectors \({\bf{A}}=\{{{\bf{a}}}_{j}| j\in \{1,2,...,K\}\}\) for K classes. Each prototype corresponds to a class and can be treated as the agent for that class. The classification process is performed by matching image representations to the nearest class prototype:
where \(d\left(\cdot \right)\) is the distance measurement.
Existing methods for prototype network learning17,38 directly compute posterior distribution for each image embedding over all classes by applying a softmax on Euclidean distances to the prototypes:
and then optimize the model using cross-entropy loss on the distance-based probability distribution:
where y is the true class and b denotes the batch size. The idea is such that the image representations are pushed close to the true class prototype, whereas it is repelled by the other prototypes.
However, the DCE loss does not necessarily ensure a compact intra-class distribution. This is because the output probability of softmax function in Eq. (2) depends on the relative scale between values in the input vector of distances. Moreover, when the dimension of image embeddings is very large, the magnitude of distances may explode and fall in the saturation area of the softmax39 during the optimization. The distance explosion can lead to the sparse gradient problem which will slow down the training process.
To address above problems, we first introduce a regularization term in the objective function for controlling the intra-class compactness. In each batch, the loss of intra-class distance (ICD) considers the relations between feature embeddings from same classes and regulates the distance to the true prototypes:
where A+ is a set of prototypical vectors that the corresponding classes are included in the current training batch. Second, the exploded distance in prototype learning is mainly caused by the discrepancy between the scale of image embeddings and class prototypes. Hence, we use layer normalization in the encoding network to restrain the mean and the variance of image embeddings during the model training.
To produce hierarchical prediction with a predefined class hierarchy, we construct a hierarchical prototype decision tree (HPDT) on top of the prototype network. Each node in the decision tree is associated with a prototypical vector and corresponds to a class in the hierarchy. The proposed decision tree classifies an image by matching the image feature with class nodes at every taxonomic level and then gives a sequence of discrete predictions along the class tree.
Formally, given a class hierarchy \({\mathcal{H}}=\left(V,E\right)\), where V and E represent class nodes and edges in the directed acyclic graph of \({\mathcal{H}}\), respectively, we formulate the implementation of the decision tree as follows:
1) Hierarchical class prototype assignment: Let’s represent K leaf nodes as \({V}_{leaf}=\left\{{{\bf{a}}}_{1},...,{{\bf{a}}}_{K}\right\}\) and denote N inner nodes as \({V}_{inner}=\left\{{{\bf{a}}}_{K+1},...,{{\bf{a}}}_{K+N}\right\}\). We assign a raw vector for each leaf node while computing the representations for internal nodes by averaging prototypical vectors of its child nodes. Take k-th inner class node as an example, we find all the leaves in its subtree and compute the mean prototypical vectors:
2) Computing class node probability: We then compute classification probability for class nodes by applying a softmax on the Euclidean distance between an image embedding to class prototypes of child nodes under each inner node. For j-th child node of ai ∈ Vinner, the probability is:
where \(j\in child\left(i\right)\).
3) Leaf class prediction with path probability: As each leaf node in Vleaf has a unique path connecting it to the root, we compute the product of node probabilities along the path and pick the leaf class with the largest probability as the final prediction.
where \(path\left(k\right)\) indicates the path traversing \({\mathcal{H}}\) from the root node to the leaf node k.
For training of the decision tree, we consider both the separation of class prototypes for Vleaf and Vinner. The loss Eq. (3) and Eq. (4) are responsible for separating leaf classes whereas they cannot guarantee the separation of inner class nodes. We thus add a surrogate loss defined on the class distribution of path probability in the hierarchical tree to learn representations for inner nodes (Eq. (7)):
Class distance guided prototype learning
Ideally, the learned class prototypes in the HPDT should follow a semantic distribution corresponding to their relations in the class hierarchy. That is, prototypes of sibling classes should have larger similarities than that of sub-classes from different super-classes. The semantically distributed prototypes are beneficial to align the prototypical tree with the class hierarchy and therefore enable the model to make semantically meaningful mistakes. However, though we build the decision tree by aggregating the leaf prototypes according to the class hierarchy, the learned prototypical vectors do not warrant such semantic distribution. We illustrate the arrangement of prototypes in the embedding space in Fig. 15. Building upon the metric-guided prototype learning framework introduced by Landrieu et al.38, we adapt their approach for hierarchical skin lesion classification. We propose to regularize the arrangement of prototypes in the embedding space with the hierarchical distance derived from the class tree during training.
The left panel shows prototypes learned without semantic guidance, while the right panel shows prototypes learned with semantic guidance. The guided approach demonstrates better alignment with hierarchical relationships, enhancing the semantic coherence of the embedding space across Levels 1, 2, and 3.
In6 and19, the authors measure dissimilarity between two classes in a hierarchy with the height of LCA (shown in Fig. 1). For each leaf class, we calculate a distance vector using the height of the LCA for measuring the semantic relations of the class node to all other class nodes. Sibling classes tend to have smaller LCA heights than that classes from different parent classes. Based on this concept, we can easily represent relationships between all leaf classes with a matrix of class distance \({\bf{D}}\in {\left[0,h\right]}_{+}^{K\times K}\).
As prototypes of inner nodes are calculated from leaf class prototypes, We explicitly introduce such class relations into class prototypes of leaf nodes and propose to guide the prototype learning by constraining the distance between leaf prototypes to be consistent with the class distance in the D. According to40 and following the formulation in38, the distortion of a mapping between the finite metric space of hierarchical class distance \({\bf{D}}\left[i,j\right]\in {\left[0,h\right]}_{+}\) and the continuous metric space of prototype distance \(d\left({{\bf{a}}}_{i},{{\bf{a}}}_{j}\right)\in {{\mathbb{R}}}_{+}\) can be defined as:
A low-distortion mapping means that the learned prototypes preserve well the relations between classes defined by the D. However, achieving low-distortion mapping requires the prototypes arranged in the embedding space with a specific distance constraint, and this may conflict with the DCE loss that encourages the distance between an embedding to negative class prototypes to be as large as possible. Hence, following38, we adopt a scale in the formulation of the distortion for removing the discrepancy between the scale of the prototype distance and the hierarchical class distance:
Then, we define the following smooth surrogate of the distos for optimization:
The final objection function for optimizing the proposed hierarchical model is:
where the λ is a constant coefficient and α is a factor gradually decreasing from 1 to 0 over training epochs. The combination of the \({{\mathcal{L}}}_{DCE}\) and \({{\mathcal{L}}}_{ICD}\) enables to jointly learning image embeddings and class prototypes, while the \({{\mathcal{L}}}_{disto}\) enforces a semantic-aware arrangement of leaf prototypes in the embedding space. The \({{\mathcal{L}}}_{TCE}\) is used to learn prototypes of internal class nodes in the decision tree.
Evaluation metrics
To evaluate the model’s performance, we used standard accuracy, the Hierarchical F1 Score, and two hierarchical metrics based on the LCA height. Detailed definitions of these hierarchical measurements are provided below.
Hierarchical F1 Score (H-F1): This metric extends the traditional F1 score by considering the hierarchical relationships between classes. For each prediction, it computes precision and recall based on the overlap between the ancestor sets of the predicted and true classes. Formally, for each prediction pair \((y,\hat{y})\), where y is the true class and \(\hat{y}\) is the predicted class, the hierarchical Precision (HP), hierarchical Recall (HR) are defined as follows:
where Anc(y) represents the set of ancestors of class y in the hierarchy (including y itself). The overall hierarchical F1 score for n samples is then calculated as:
Severity of Mistake: Following the approach in ref.6, we measure the severity of misclassification based on the height of the LCA between the true class and the incorrectly predicted class, indicating how closely related the misclassified class is to the true label. We extend this measure by using the normalized height of the LCA, which accounts for the relative depth of the nodes:
where R is the root node. This normalization allows us to handle incomplete hierarchies more effectively and facilitates comparisons across trees of varying sizes by reflecting hierarchical costs consistently across different tree structures.
Top-k Hierarchical Distance (HD): This metric calculates the mean normalized LCA height between the true class and each of the top-k predicted classes. It is particularly valuable in cases where multiple high-probability predictions are considered, as it reflects the semantic proximity of the model’s top predictions to the correct class, offering insight into the reliability of alternative suggestions within a hierarchical context.
Statistical analysis
To rigorously evaluate and compare model performance, we conducted paired statistical analyses across our test set of 47,082 samples. Rather than relying solely on traditional p-value testing, which often shows statistical significance with large datasets, we employed Cohen’s d-effect size analysis to quantify the practical significance and magnitude of performance differences between models. For each pair of models, we calculated the effect size as the standardized mean difference of paired observations, where d = (mean of differences)/(standard deviation of differences). The effect size analysis was performed across four key metrics: accuracy, hierarchical F1 score, error severity, and top-5 hierarchical distance. Following standard interpretations, we considered ∣d∣ < 0.2 as negligible effect, 0.2 < ∣d∣ < 0.5 as small effect, 0.5 < ∣d∣ < 0.8 as medium effect, and ∣d∣ > 0.8 as large effect.
Data availability
The complete Molemap dataset used in this study was collected in a clinical setting under strict privacy and ethical guidelines. Due to patient privacy concerns and data protection regulations, the full dataset cannot be made publicly available. However, to support reproducibility and facilitate future research, we provide a curated subset of the dataset consisting of de-identified dermoscopic images that can be used for testing and validation purposes. This subset maintains the original three-level hierarchical taxonomy structure spanning 65 skin conditions (53 benign and 12 malignant classes) and includes representative samples from each class. Researchers interested in accessing the test subset or additional data for academic purposes may contact Dr. Zhen Yu (zhen.yu1@monash.edu) at AIM for Health Lab, Faculty of Information Technology, Monash University. Any inquiries about collaboration or the full dataset should be directed to the corresponding author, Prof. Zongyuan Ge (zongyuan.ge@monash.edu), subject to ethical approval and data usage agreement.
Code availability
To ensure reproducibility and facilitate future research, we provide a comprehensive implementation of our method. The complete codebase, including model architecture, training scripts, and evaluation code, is available at our GitHub repository (https://github.com/Zakiyi/Hierarchical-skin-lesion-image-classification-with-prototypeical-decision-tree.git). We also provide trained model weights and a detailed Jupyter notebook demonstrating the testing pipeline with example usage. The implementation is in PyTorch and includes documentation for environment setup, data preprocessing, model training, and evaluation procedures.
Change history
21 February 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41746-025-01494-5
References
Rajpurkar, P., Chen, E., Banerjee, O. & Topol, E. J. Ai in health and medicine. Nat. Med. 28, 31–38 (2022).
Mortier, T., Wydmuch, M., Dembczyński, K., Hüllermeier, E. & Waegeman, W. Efficient set-valued prediction in multi-class classification. Data Min. Knowl. Discov. 35, 1435–1469 (2021).
Zaffalon, M., Corani, G. & Mauá, D. Evaluating credal classifiers by utility-discounted predictive accuracy. Int. J. Approx. Reason. 53, 1282–1301 (2012).
Hase, P., Chen, C., Li, O. & Rudin, C. Interpretable image recognition with hierarchical prototypes. In Proceedings of the AAAI Conference on Human Computation and Crowdsourcing, vol. 7, 32–40 (2019).
Wan, A. et al. Nbdt: Neural-backed decision tree. International Conference on Learning Representations (2021).
Bertinetto, L., Mueller, R., Tertikas, K., Samangooei, S. & Lord, N. A. Making better mistakes: Leveraging class hierarchies with deep networks. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 12506–12515 (2020).
Karthik, S., Prabhu, A., Dokania, P. K. & Gandhi, V. No cost likelihood manipulation at test time for making better mistakes in deep networks. International Conference on Learning Representations (2020).
Garg, A., Sani, D. & Anand, S. Learning hierarchy aware features for reducing mistake severity. European Conference on Computer Vision, 252–267 (Springer, 2022).
Yu, Z. et al. Skin lesion recognition with class-hierarchy regularized hyperbolic embeddings. International Conference on Medical Image Computing and Computer-Assisted Intervention,594–603 (Springer, 2022).
Ju, L., Yu, Z., Wang, L., Zhao, X., Wang, X., Bonnington, P. & Ge, Z. Hierarchical Knowledge Guided Learning for Real-World Retinal Disease Recognition. IEEE Trans Med Imaging. 43, 335–350 (2024).
Yang, J. et al. Hierarchical classification of pulmonary lesions: A large-scale radio-pathomics study. International Conference on Medical Image Computing and Computer-Assisted Intervention, 497–507 (Springer, 2020).
Barata, C., Marques, J. S. & Celebi, M. E. Deep attention model for the hierarchical diagnosis of skin lesions. 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), 2757–2765 (IEEE, 2019).
Barata, C. & Marques, J. S. Deep learning for skin cancer diagnosis with hierarchical architectures. 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), 841–845 (IEEE, 2019).
Liu, Y. et al. A deep learning system for differential diagnosis of skin diseases. Nat. Med. 26, 1–9 (2020).
Kowsari, K. et al. Hmic: Hierarchical medical image classification, a deep learning approach. Information 11, 318 (2020).
Chang, D. et al. Your" flamingo" is my" bird": Fine-grained, or not. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 11476–11485 (2021).
Snell, J., Swersky, K. & Zemel, R. Prototypical networks for few-shot learning. Proceedings of the 31st International Conference on Neural Information Processing Systems, 4080–4090 (2017).
Yang, Z., Bastan, M., Zhu, X., Gray, D. & Samaras, D. Hierarchical proxy-based loss for deep metric learning. Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, 1859–1868 (2022).
Barz, B. & Denzler, J. Hierarchy-based image embeddings for semantic image retrieval. 2019 IEEE Winter Conference on Applications of Computer Vision (WACV), 638–647 (IEEE, 2019).
Russakovsky, O. et al. Imagenet large scale visual recognition challenge. Int. J. Comput. Vis. 115, 211–252 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. 770–778, https://doi.org/10.1109/CVPR.2016.90 (2016).
Dhall, A. et al. Hierarchical image classification using entailment cone embeddings. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, 836–837 (2020).
Redmon, J. & Farhadi, A. Yolo9000: better, faster, stronger. Proceedings of the IEEE conference on computer vision and pattern recognition, 7263–7271 (2017).
Mettes, P., van der Pol, E. & Snoek, C. Hyperspherical prototype networks. Adv. Neural Inf. Process. Syst. 32, 1487–1497 (2019).
Gao, D. Deep hierarchical classification for category prediction in e-commerce system. Proceedings of The 3rd Workshop on e-Commerce and NLP, 64–68 (2020).
Carion, N. et al. End-to-end object detection with transformers. European Conference on Computer Vision, 213–229 (Springer, 2020).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE conference on computer vision and pattern recognition, 770–778 (2016).
Mehta, D. et al. Revamping AI models in dermatology: Overcoming critical challenges for enhanced skin lesion diagnosis. arXiv preprint arXiv:2311.01009 (2023).
Costa, V. G. & Pedreira, C. E. Recent advances in decision trees: an updated survey. Artif. Intell. Rev. 56, 4765–4800 (2023).
Silla, C. N. & Freitas, A. A. A survey of hierarchical classification across different application domains. Data Min. Knowl. Discov. 22, 31–72 (2011).
Dembczyński, K., Kotłowski, W., Waegeman, W., Busa-Fekete, R. & Hüllermeier, E. Consistency of probabilistic classifier trees. Machine Learning and Knowledge Discovery in Databases: European Conference, ECML PKDD 2016, Riva del Garda, Italy, September 19–23, 2016, Proceedings, Part II 16, 511–526 (Springer, 2016).
Secker, A. D. et al. An experimental comparison of classification algorithms for hierarchical prediction of protein function. Expert Update (Magazine of the British Computer Society's Specialist Group on AI) Vol. 9, 17–22 (University of Kent, 2007).
Frank, E. & Kramer, S. Ensembles of nested dichotomies for multi-class problems. Proceedings of the twenty-first international conference on Machine learning, 39 (2004).
Melnikov, V. & Hüllermeier, E. On the effectiveness of heuristics for learning nested dichotomies: an empirical analysis. Mach. Learn. 107, 1537–1560 (2018).
Beygelzimer, A., Langford, J., Lifshits, Y., Sorkin, G. & Strehl, A. Conditional probability tree estimation analysis and algorithms. In Proceedings of the Twenty-Fifth Conference on Uncertainty in Artificial Intelligence, 51–58 (2009).
Atigh, M. G., Keller-Ressel, M. & Mettes, P. Hyperbolic Busemann learning with ideal prototypes. Thirty-Fifth Conference on Neural Information Processing Systems (2021).
Ghadimi Atigh, M., Keller-Ressel, M. & Mettes, P. Hyperbolic Busemann learning with ideal prototypes. Advances in Neural Information Processing Systems Vol. 34 (2021).
Landrieu, L. & Garnot, V. S. F. Leveraging class hierarchies with metric-guided prototype learning. British Machine Vision Conference (BMVC) (2021).
Chen, B., Deng, W. & Du, J. Noisy softmax: Improving the generalization ability of dcnn via postponing the early softmax saturation. In Proceedings of the IEEE conference on computer vision and pattern recognition, 5372–5381 (2017).
Sala, F., De Sa, C., Gu, A. & Ré, C. Representation tradeoffs for hyperbolic embeddings. International conference on machine learning, 4460–4469 (PMLR, 2018).
Acknowledgements
This study was supported by the NHMRC Centre of Research Excellence in Melanoma (Grant No. 1135285). Z. Ge is funded by the Airdoc Research Australia Centre and the Monash-NVIDIA Joint Research Centre. Victoria Mar is supported by a National Health and Medical Research Council Early Career Fellowship and an Australian Cancer Research Foundation infrastructure grant, which facilitated the establishment of the Australian Centre of Excellence for Melanoma Imaging and Diagnosis, including a 3D total body imaging network (Vectra, Canfield Scientific). Both V. Mar and Z. Ge have received a Victorian Medical Research Acceleration Fund grant, with matching industry funding provided by MoleMap.
Author information
Authors and Affiliations
Contributions
Z.Y. conceived the methodology, designed the framework, and implemented the algorithm. T.N. prepared the data and supported algorithm development by contributing to portions of the code. Z.G. and V.M. supervised the project, offering strategic guidance and overseeing the organization and refinement of the manuscript. L.J. and Y.G. assisted in the experimental design and critically reviewed the results. M.S., P.B., and L.Z. contributed to writing and carefully refining the discussion section. All authors participated in discussions and reviewed the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yu, Z., Nguyen, T.D., Ju, L. et al. Hierarchical skin lesion image classification with prototypical decision tree. npj Digit. Med. 8, 26 (2025). https://doi.org/10.1038/s41746-024-01395-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-024-01395-z