Abstract
Determining tumor microsatellite status has significant clinical value because tumors that are microsatellite instability-high (MSI-H) or mismatch repair deficient (dMMR) respond well to immune checkpoint inhibitors (ICIs) and oftentimes not to chemotherapeutics. We propose MSI-SEER, a deep Gaussian process-based Bayesian model that analyzes H&E whole-slide images in weakly-supervised-learning to predict microsatellite status in gastric and colorectal cancers. We performed extensive validation using multiple large datasets comprised of patients from diverse racial backgrounds. MSI-SEER achieved state-of-the-art performance with MSI prediction by integrating uncertainty prediction. We achieved high accuracy for predicting ICI responsiveness by combining tumor MSI status with stroma-to-tumor ratio. Finally, MSI-SEER’s tile-level predictions revealed novel insights into the role of spatial distribution of MSI-H regions in the tumor microenvironment and ICI response.
Similar content being viewed by others
Introduction
Patients whose cancers are microsatellite instability-high (MSI-H)/deficient in mismatch repair proteins (dMMR) have better outcomes than patients with microsatellite stable (MSS) tumors1,2. Additionally, MSI-H/dMMR tumors are highly sensitive to immune checkpoint inhibitors (ICIs) and may not respond to traditional chemotherapy3,4,5.
MSI testing is recommended for all newly diagnosed gastric6 and colorectal cancers7,8. Current methods of testing include immunohistochemistry (IHC) for detecting MMR status and polymerase chain reaction (PCR) for determining MSI status. However, these assays are time-intensive and costly, and many patients do not undergo the recommended molecular profiling9. Numerous studies have demonstrated the feasibility of using deep learning algorithms to analyze hematoxylin and eosin (H&E)-stained whole-slide images (WSIs) to predict MSI status10,11,12,13,14,15,16,17,18,19,20,21,22. Thus, incorporating artificial intelligence (AI) into the clinical workflow may provide cost-efficient and widely accessible MSI testing.
The adoption of AI-based MSI-status prediction into routine clinical practice requires extensive validation in large, diverse patient cohorts. The inclusion of heterogeneous patient cohorts is particularly important as there may be biological differences associated with race and ethnicity23,24,25. A recent study showed that a model trained on a predominantly non-Hispanic White patient cohort with gastric cancer, performed poorly when it was tested on samples from Asian patients10. These data highlight the fundamental need to validate novel clinical tools across diverse populations.
The ability to quantify uncertainty in predictions is not only crucial to enhance a model’s predictive accuracy, but it also may guide physicians to make more informed decisions. Cases with high predictive uncertainty will require nuanced decision-making by human experts. While numerous deep learning methods, including convolutional neural networks (CNNs) and vision-transformer-based methods, have been applied to MSI status prediction problems, most do not capture the uncertainty in the prediction as point estimation methods. A prediction model must not only deliver accurate predictions, but also quantify the uncertainty of the predictions. Finally, previously reported algorithms also focus solely on MSI prediction without providing insights into ICI responsiveness, which limits their clinical utility.
To address these challenges, we propose a novel MSI prediction algorithm that we named MSI-SEER, which analyzes H&E-stained WSIs by utilizing deep Gaussian processes (DGPs)26 in weakly supervised learning, which is a form of inexact supervision tasks27. MSI-SEER predicts MSI-H status by first calculating the probability of being MSI-H for each tile within a WSI, then aggregating these tile-level probabilities to assess the overall MSI-H status of the slide. This approach provides a predictive distribution that quantifies the uncertainty of predictions, thereby enhancing the precision of MSI-H assessments and informing clinicians about the need for additional confirmatory lab testing. Additionally, by calculating the MSI-H status at both the tile and slide levels, our model provides new insights into the tumor microenvironment, as related to ICI responsiveness in gastric cancer.
Results
Datasets
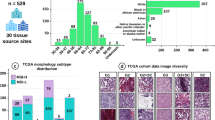
In this study, we analyzed H&E-stained WSIs from 12 distinct datasets comprised of colorectal and gastric cancers, with 2091 and 1101 slides, respectively. These datasets included a diverse patient population comprised of Asian, Black or African American, and White patients treated at multiple international sites, including Yonsei University and, Seoul St. Mary’s Hospital in Korea, Mayo Clinic in the USA, and various international sites (Table 1). The MSI testing methods used for these datasets are summarized in Supplementary Table 1.
The colorectal cancer datasets were TCGA-CRC, Yonsei-1, Yonsei-1-remade, Yonsei-2, STMary-Colon, CPATC-COAD, and Mayo Clinic. We used the TCGA-CRC and Yonsei-1 for training and the rest for validation. Of note, the Yonsei-1-remade dataset was generated by re-cutting slides and performing H&E staining from existing blocks of the Yonsei-1 dataset to explore how staining variability affected MSI-H prediction.
For gastric cancer, we analyzed datasets named TCGA-STAD, Yonsei-Classic, STMary-GC, GC-ICI, and Molecular subtypes. TCGA-STAD or Yonsei-Classic were used to train the model for gastric cancer sample analyses, and the remaining datasets were used for validation. The MSI status of the samples in the Molecular subtypes dataset was determined by both PCR and IHC. Thus, this dataset represents the gold standard for our validation efforts. Finally, the GC-ICI dataset consisted of gastric cancer patients treated with ICIs and allowed us to test the clinical utility of MSI-SEER to predict ICI response.
Developing and training the MSI-SEER model
The workflow of this study is summarized in Fig. 1. Our tumor MSI status prediction model MSI-SEER consists of two main components: a feature extractor and a prediction model (Fig. 1a illustrates the MSI prediction pipeline, while Supplementary Fig. 1 details the DGP-based prediction model within the weakly supervised learning framework). The feature extractor utilizes pre-trained deep learning model within the transfer learning framework to compute feature vectors from image tiles in a WSI. Tumor tiles in a slide were transformed into image features using either CTransPath28 or one of the nine MSIDETECT CNNs12. The prediction model is based on DGP in a weakly supervised learning framework. The DGP model first estimated the probability of each individual tile being MSI-high, using the extracted tile features. The final slide-level MSI-high prediction was then obtained by aggregating these tile-level probabilities. This aggregation was performed using the extremized geometric mean of the probabilities, a method shown to achieve superior calibration performance compared to other pooling techniques, as evaluated by the Brier score29. Additionally, a weight was assigned to each tile within the geometric mean pooling operator, minimizing attention to irrelevant tiles for MSI prediction.
a MSI-SEER The method consists of two core components, an image feature extractor and a DGP-based MSI prediction model. Tumor tiles in a whole slide image (WSI) were color normalized using Macenko method and transformed into feature vectors using a pre-trained CNN model. The slide-level MSI-H predictions were made by aggregating the tile-level MSI-H predictions using the weighted version of the extremized geometric mean of the tile-level MSI-H probabilities (σ represents the sigmoid function, ϕ is the output of the DGP model for each tile, and a is the extremized parameter). b Output of MSI-SEER provided MSI-H prediction probability at both the tile-level and slide-level and quantified predictive uncertainty at the slide-level. c Uncertainty quantification identified cases that should be referred to human experts for further investigation. Selective exclusion of highly uncertain predictions improved the model’s prediction performance. d Information provided by MSI-SEER predicted immune check point inhibitor (ICI) treatment response. We can predict ICI-treatment response using predicted MSI-H tumor region by MSI-SEER and stromal composition obtained by an image-based cell-type classification method CellViT. The figure was created in BioRender. Park, S. (2025) https://BioRender.com/p31j116.
We trained the DGP-based MSI prediction models using dropout variational inference30, leveraging the fact that, with random feature expansion31, a DGP can be reduced to a specific structure of a Bayesian Neural Network (BNN)32. Detailed information on the model formulation in weakly supervised learning, along with the training and inference processes, is provided in the Methods section.
We first explored the DGP models based on the CTransPath feature extractor to determine the optimal number of GP layers. We trained the model with different numbers of layers, from 1 to 7, and selected the best one based on the 3-fold cross validation (CV) performance (Supplementary Fig. 2). While we did not see a wide variation in performance with different numbers of GP layers, the model performance increased up to six GP layers, which was then used for all the training datasets for both DGP models integrated with CTransPath and with MSIDETECT CNN models.
We implemented our model in ensemble learning, where the model was trained using bootstrapped samples of training data 10 times, and the final dropout samples from all ensemble models were aggregated to make the final prediction. Of note, the DGP integrated with MSIDETECT CNN models included nine different ensemble models depending on which CNN model was used for feature extraction. Since selecting the best performer from the nine ensemble models for a test slide is challenging, we combined these nine ensemble models by using the same aggregation method used to combine 10 models trained using bootstrapped samples. Finally, we observed that the DGP integrated with MSIDETECT CNN models generally outperformed the DGP integrated with CTransPath (Supplementary Table 2 and 3). Unlike the MSIDETECT CNNs, the CTransPath model was trained without MSI status labels in self-supervised learning. Therefore, we will utilize the DGP models integrated with MSIDETECT CNNs, which we will refer to as MSI-SEER.
MSI-SEER predicted MSI status with accuracy similar to previously published models
To compare the predictive capability of MSI-SEER to previously published models, we adapted the experimental designs from Laleh et al.33. We compared MSI-SEER to previously reported CNN-based deep learning models, ResNet34, ShuffleNet35, and EfficientNet36. We also compared these models to the MSIDETECT CNN models. We evaluated the colorectal and gastric cancer samples separately so that each model was tested on the datasets only in the same cancer type to which the training dataset belonged. We did not observe a significant performance improvement when the models were trained using the datasets from both cancer types (Supplementary Table 4).
We trained ResNet, ShuffleNet, and EfficientNet using the same training steps as described by Laleh et al.33, where the CNNs were trained in supervised learning under the assumption that all image tiles in a WSI shared the same label assigned to the WSI. We also attempted to retrain MSIDETECT, but observed severe performance degradation due to catastrophic forgetting, the phenomenon in which neural networks lose knowledge gained from previous tasks. We therefore used the pre-trained MSIDETECT models for the comparisons in the rest of the experiments. We used 3-fold CV to evaluate the performance of the training datasets (TCGA-CRC and Yonsei-1 for colorectal cancer and TCGA-STAD and Yonsei-Classic for gastric cancer). To evaluate the inter-cohort prediction performance, we trained each model using all data points in each training dataset, tested the models on the validation datasets in the same cancer type. Figures 2 and 3 show the prediction performance of the methods in terms of the area under the ROC curve (AUC) as a heatmap. We also used other metrics, recall, precision and F1 measure, to evaluate the performance of the models in Supplementary Tables 5 and 6.
The 3-fold cross-validation (CV) performance is evaluated for the training data, while the inter-cohort performance is evaluated for the validation datasets. Area under the ROC curve (AUC) values are shown. MSI-SEER, ResNet, EfficientNet, and ShuffleNet were trained using TCGA-CRC, Yonsei-1, and the combined data of TCGA-CRC and Yonsei-1. The 3-fold CV performance of MSIDETECT and MSI-SEER on TCGA-CRC was not evaluated because this dataset was already included in the training data for MSIDETECT. (*) denotes the dataset used for training and the remaining datasets were used for validation.
The threefold cross-validation (CV) performance is evaluated for the training data, while the inter-cohort performance is evaluated for the validation datasets. Area under the ROC curve (AUC) values is shown. MSI-SEER, ResNet, EfficientNet, and ShuffleNet were trained using TCGA-STAD, Yonsei Classic and the combined data of TCGA-STAD and Yonsei Classic. (*) denotes the dataset used for training and the remaining datasets were used for validation.
In the model performance heatmaps (Figs. 2 and 3), best and worst for MSI-SEER represent the best and worst performing models among the nine DGP ensemble models. The aggregate MSI-SEER had performances that were comparable to the best performing individual MSI-SEER models in most cases. Similarly, best and worst for MSIDETECT represent the best and worst performing models among nine MSIDETECT CNNs. We observed that no single model among the nine MSIDETECT CNNs performed best for all datasets, and there was a wide variation between the best and worst performance (Supplementary Fig. 3). Similar to MSI-SEER, we defined the aggregated model of the MSIDETECT CNNs by averaging the output MSI-H probabilities of the CNNs on a test WSI, where a slide-level prediction is also made by averaging the MSI-H probabilities over the tiles in the slide.
For the colorectal datasets (Fig. 2), we found that the MSI-SEER had better predictive performance for more cases when trained using Yonsei-1 than when trained using TCGA-CRC. However, we also found that the model trained on the combined data from both datasets did not perform better than the models trained on either dataset. This may be because the feature extractors were already trained on the datasets that included TCGA-CRC, and thus the combined data did not provide enough new information to MSI prediction. Therefore, for unseen colorectal cancer slides, we recommend using the aggregated MSI-SEER model trained on Yonsei-1. For the gastric cancer datasets (Fig. 3), we found that the performance of the aggregated MSI-SEER was best when trained on a combined data from TCGA-STAD and Yonsei-Classic datasets, as compared to the models trained on each dataset alone. Thus, for unseen slides in gastric cancer, we recommend using the aggregated model trained on the combined TCGA-STAD and Yonsei-Classic datasets. In the rest of the paper, MSI-SEER refers to the aggregated MSI-SEER model trained on Yonsei-1 for colorectal cancer and on the combined dataset for gastric cancer.
For colorectal cancer datasets (Fig. 2), MSI-SEER had AUC ranging from 0.815 to 0.953, demonstrating that MSI-SEER worked well for colorectal cancer WSIs obtained from a diverse patient cohort. We next compared the MSI status prediction performance of the MSI-SEER model to the other models using DeLong’s method37, which tests whether the AUC of one model is significantly different from that of another model. We found that MSI-SEER generally had comparable performance as the other methods, and in some cases, such as the CPATC-COAD and Mayo Clinic dataets, MSI-SEER showed better performance (Supplementary Fig. 4). For the gastric cancer datasets (Fig. 3), we first found that ResNet, ShuffleNet, and EfficientNet trained on TCGA-STAD, which consisted of a diverse patient cohort, performed worse on the validation datasets generated from Korean patients. However, their predictive performance did not improve much on these validation datasets even when the models were trained on the combined data from TCGA-STAD and Yonsei-Classic. In contrast, MSI-SEER performed well on these validation datasets generated from Korean patients, with AUC ranging from 0.761 to 0.937 (Fig. 3c). Using DeLong’s method, we also showed that MSI-SEER significantly outperformed the other methods (Supplementary Fig. 5). We also compared our DGP-based MSI prediction models, including MSI-SEER and the DGP models using either CTransPath or MSIDETECT, with representative multiple instance learning (MIL) models, such as attention-based MIL38, CLAM39, TransMIL40, and RRT-MIL41, as detailed in Supplementary material. The experimental results, presented in Supplementary Figs. 6–11, confirm that our models achieved comparable prediction performance. Notably, our method demonstrates stable prediction performance across various validation datasets, even when trained on different training data.
Incorporating predictive uncertainty improved MSI-SEER performance
For MSI-SEER, we quantified predictive uncertainty through a Bayesian confidence score (BCS)42, where, \({bcs}\,({y}_{* })=1-2\sqrt{{\text{var}}({y}_{*})}\), with \({\text{var}}({y}_{*})\) representing the variance of the slide-level predictive distribution for a testing point (slide). High BCS values indicated higher model confidence in the prediction. Our model’s inference, implemented using Monte Carlo dropout, estimated the likelihood of slide-level MSI-H status using extremized geometric means of tile-level MSI prediction probabilities. Higher computed variance occurred when the weighted sum of log-odds is near zero, indicating ambiguity in prediction (see Methods for more details). Unlike attention-based weakly supervised learning methods that aggregate tile-level features28,39, our method computed MSI-H probabilities for each tile and aggregated these to slide-level predictions. This tile-level analysis allowed for exploration of spatial MSI-H patterns within tumors, providing unique insights compared to other methods that may not offer detailed local predictions.
Figure 4 shows representative tile-level prediction results from our model, displaying the mean probability estimate of each tile being MSI-H on a heatmap on an WSI. Tiles with a dominant MSI-H or MSS morphological pattern generally yielded higher BCS, indicating more confident predictions (Fig. 4a, b). Conversely, tiles with heterogeneous MSI-H probabilities lead to lower BCS, reflecting higher uncertainty due to the complex patterns (Fig. 4c, d). In this case, despite the complex pattern, the weighted sum of the log-odds was approximately zero, and thus the slide-level MSI-H prediction probability was marginal (≈0.5) with the high uncertainty. WSIs with high predictive uncertainty (Fig. 4c) showed a spatially random distribution of MSI-H-like and MSS-like tiles. This correlation between spatial randomness and slide-level uncertainty was seen in each of our datasets, and there was a significant and strong negative correlation between the BCS and spatial (Altieri’s) entropy43 (Supplementary Figs. 12 and 13).
a, c WSI images with tile-level MSI-H prediction probability heat maps. b, d Histograms quantifying tiles based on predicted probability of MSI-H status. While both slides were considered MSI-H, the sum predictive probability was 0.996 (BCS = 0.871) for the highly confident sample, and the sum predictive probability was 0.503 (BCS << 0.001) for the highly uncertain slide.
We next tested the effects of excluding the most uncertain predictions as based on Deodato et al.42. We found that removing the most uncertain predictions enhanced the overall model performance by 1.1% and 2.6% in AUC in the colorectal (Fig. 5a–b) and gastric cancer datasets (Fig. 5c–d), respectively (Supplementary Table 7 for the other evaluation metrics). This selective approach of discarding the most uncertain predictions, consistently improved performance metrics, in contrast to random exclusions that showed no beneficial effect (Fig. 5). Details about how to discard the most uncertain predictions are provided in the Method section. These findings demonstrate the potential of leveraging predictive uncertainty to refine diagnostic accuracy, with further details on test data sets and various training scenarios show in Supplementary Figs. 14 and 15.
In this model, the aggregated version of MSI-SEER was trained using Yonsei-1 in colorectal cancer or the combined data from TCGA-STAD and Yonsei-Classic in gastric cancer. a All test datasets in colorectal cancer, except the training data Yonsei-1, were combined and tested. The numbers of WSIs classified correctly are green, and those classified incorrectly are orange. The predictive uncertainty as measured by the Bayesian confidence scores are shown. b The changes in the prediction performance (in terms of area under the curve (AUC)) when the predictions are discarded at increasing rates, i.e. #discarded WSIs/#total WSIs, in each data cohort. The red line represents the change in the performance when the most uncertain predictions (as measured by the BCSs obtained by our model) are discarded while the black line is the average change in the performance where the predictions are randomly discarded 1000 times at each rate. c All the test datasets in gastric cancer, except the training data TCGA-STAD and Yonsei-1, were combined and tested. The correctness of classification for gastric cancer datasets is shown. d The change in prediction performance is shown for the gastric cancer test datasets.
Predicted MSI-H regions and stromal composition correlated to immunotherapy response in gastric cancer
To further assess the clinical utility of MSI-SEER, we performed a targeted analysis within the GC-ICI cohort, which included 21 slides from patients who responded to ICIs and 75 slides from patients who did not. We tested the association between the proportion of predicted tumor MSI-H regions and ICI response. To determine the MSI status for each tile, we used 0.5 as the cutoff for the mean predicted probability of the tile being MSI-H. We found that responders had an average of 62% of tumor MSI-H predicted regions, while non-responders had only an average of 30% (P < 0.001, Fig. 6a).
a The MSI-H fraction, defined as the proportion of MSI-H tiles within a whole slide image (WSI). Responders demonstrated a significantly higher fraction of MSI-H tiles compared to non-responders (Wilcoxon test, p << 0.001). To determine MSI-H and MSS tiles in a slide, we used 0.5 as the cutoff for the mean predictive probability of each tile being MSI-H over the dropout samples. MSI status is provided for reference and was not used in the comparison. Unknown denotes slides without available MSI status information. b Analysis of the stromal-to-tumor ratio within MSI-H predicted tiles, comparing responders and non-responders (Wilcoxon test, p = 0.0016). This metric assesses the microenvironment’s cellular composition, providing insights into the tumor-stroma dynamics that may influence ICI responsiveness.
Notably, there were 5 responders who were categorized as having MSS tumors by traditional testing methods. Three of the samples had more than 85.3% of tumor MSI-H predicted regions, and the other 2 had a similar amount of tumor MSI-H predicted regions as other confirmed MSI-H samples. Conversely, there was one clinically determined MSI-H tumor that did not respond to ICI, and it had only 7.7% of tumor MSI-H predicted regions. On review by board-certified pathologists (JHP and JYS), we found that these patients displayed histopathological features consistent with borderline state between MSS and MSI-H, suggesting that the MSI-SEER algorithm may uncover MSI-H patterns not detected by standard laboratory tests. Thus, MSI-SEER may help refine patient selection for ICI therapy by identifying patients with MSS tumors who may benefit and MSI-H tumors who may not benefit.
Based on our previous work showing that stromal content is associated with ICI response44, we next compared the stromal fraction within the predicted MSI-H tiles in responders and non-responders. Within each MSI-H tumor tile, we used CellViT to classify each cell as tumor or stromal45. Since the average number of tumor cells per tile was significantly higher in the responder group than in the non-responder group (Supplementary Fig. 16), the stromal cell count was normalized by the tumor cell count. The majority of predicted MSI-H tiles in non-responders contained a high number of stromal cells compared to those in responders. For example, more than 56% of the predicted MSI-H tiles in non-responders contain more than 50 stromal cells in the MSI-H tiles, while only 38.2% of the predicted MSI-H tiles in responders do. Figure 6b shows that a high abundance of stromal cells in the MSI-H tiles is significantly associated with ICI non-responsiveness.
Finally, we developed a rule-based classifier incorporating predicted MSI-H fraction and stroma-to-tumor ratio within predicted MSI-H regions to predict ICI response. We treated 17 samples that do not have MSI status information in the cohort as test data and used the remaining samples to define the classification rule. Based on the data ranges of the predicted MSI-H fraction and the stroma-to-tumor ratio of responders obtained from samples that have MSI status information (Supplementary Fig. 17), we defined a patient with an MSI-H fraction greater than 0.5 and a stroma-to-tumor ratio of less than 3.6 in a predicted MSI-H tile as a responder. Using this simple rule, we were able to stratify the test samples into responders and non-responders with an accuracy of 94.1% (Supplementary Table 8). These results demonstrate the potential of MSI-SEER to predict ICI responsiveness.
Discussion
Determining tumor MSI status provides clinically actionable information for both colorectal and gastric cancers. Deep learning models that analyze WSI may provide cost-efficient and widely accessible MSI testing. Indeed, previous work has shown that AI-based models have promising utility to predict MSI status for both colorectal and gastric cancers10,14,46,47,48,49,50. However, most of these models were not validated in diverse patient cohorts or did not report the racial makeup of their patient samples, thus raising the question of the generalizability of these models. The necessity of models externally validated in large and diverse patient cohorts have been demonstrated by several recent studies. Wagner et al. analyzed colorectal samples from multiple countries and found decreased generalizability of their model on samples from different races47. Similarly, Kather et al. found that their model for gastrointestinal cancers performed poorly on an Asian cohort when trained on data from predominantly non-Asian patients10. We were able to overcome these limitations with MSI-SEER, which is a prediction model based on DGP in weakly supervised learning, through extensive experiments on large datasets comprised of patients from diverse racial backgrounds and collected from multiple international sites. By training MSI-SEER on samples collected from diverse patient cohorts, we were able to improve our model’s performance.
Our Bayesian approach also provided significant advantages over traditional neural networks by quantifying uncertainty. Previous models are point estimation methods, which provide only binary (MSS or MSI-H) results. These outputs do not effectively capture or interpret uncertainty51,52, often leading to overconfident53 and potentially misleading results especially when used in complex decision-making processes such as clinical care. With MSI-SEER, we are able to provide uncertainty quantification while maintaining comparable prediction performance as previously developed deep learning MSI prediction tools, such as MSIDETECT. However, the ability of MSI-SEER to quantify the uncertainty in the prediction makes MSI-SEER a more clinically useful tool as uncertain results can be augmented with expert review for more nuanced decision making.
Finally, MSI-SEER may predict ICI response in gastric cancer. MSI-SEER was able to identify a subset of patients who were classified as MSS by traditional testing methodologies yet showed a clinical response to ICI treatment. This predictive capability extended to analyzing the stromal composition within MSI-H predicted regions. We previously found that high tumor ACTA2 expression, a marker of cancer-associated fibroblasts, is associated with ICI non-response44. With MSI-SEER, we similarly observed that increased stromal cells within tumors correlated with ICI non-responsiveness. By combining MSI-H fraction with stroma-to-tumor ratio we were able to achieve high accuracy in predicting ICI responsiveness in patients with gastric cancer. These findings highlight the significant influence of tumor microenvironment on therapy effectiveness.
Our model does have potential limitations that warrant mention. In order to capture real-world diversity and increase generalizability, we collected data from multiple institutions across different geographical regions. This approach naturally introduced variability in sample handling and clinical practices. To mitigate this variability, we generated new H&E slides from paraffin blocks under standardized staining protocols for certain cohorts and applied Macenko color normalization to reduce color discrepancies. Because our analysis was retrospective in nature, variability in treatment protocols–including in the GC-ICI cohort–was unavoidable. Treatment decisions were made according to the discretion of the treating physician and prevailing clinical practices, which evolved over time. However, we believe this diversity underscores our model’s robustness, as it was able to predict ICI responsiveness in a range of real-world treatment settings.
To our knowledge MSI-SEER is the first computational model that predicts ICI response from WSIs in gastric cancers. Given that MSI-H gastric cancer may not be responsive to cytotoxic chemotherapies3, using MSI-SEER to predict ICI response may spare select patients the toxicities associated with chemotherapy and allow them to receive more optimal treatment in the form of ICIs.
Methods
Datasets
The datasets used in this study are H&E-stained colorectal and gastric cancer slides collected from multiple institutions containing multiple racial groups. For the colorectal cancer data, we first used publicly available multi-center data from The Cancer Genome Atlas (TCGA) project: TCGA-CRC. Yonsei-1 and Yonsei-2 were collected from Gangnam Severance Hospital, Yonsei University College of Medicine, Seoul, Republic of Korea. Yonsei-1-remade was reprocessed from Yonsei-1: the remaining tumor tissues from a subset of the patients in Yonsei-1 were scanned to produce whole slide images. STMary-Colon was collected from Seoul St. Mary’s Hospital, Seoul, Republic of Korea. The CPTAC-Colon dataset is from The Clinical Proteomic Tumor Analysis Consortium Colon Adenocarcinoma Collection (CPTAC-COAD). Mayo Clinic slides are from Colon Cancer Family Registry (CCFR).
For gastric cancer data, we also used publicly available data from TCGA, (TCGA-STAD). We then included Yonsei-Classic data which was collected from patients who received D1 gastrectomy plus capecitabine and oxaliplatin chemotherapy or surgery alone. Molecular subtypes dataset is from Seoul St. Mary hospital in Korea, and the GC-ICI dataset was obtained from Yonsei University College of Medicine and Seoul St. Mary hospital.
TCGA (TCGA-CRC and TCGA-STAD), CPATC-colon and Mayo clinic datasets data contained primarily white patients, whereas all other datasets were collected from Asian (Korean) patients. Table 1 contains the summary statistics of each dataset in both colorectal- and stomach cancers. WSIs without clinical MSI status were excluded from experiments where MSI-H prediction performance was evaluated.
As outlined in Supplementary Table 1, the samples in our datasets were primarily tested for MSI using PCR, which is widely recognized as the gold standard for MSI detection54. For the datasets collected in Korea across both cancer types, with the exception of Yonsei-Classic, the PCR tests analyzed the markers BAT-25, BAT-26, D2S123, D5S346, and D17S250. Tumors were classified as MSI-H (≥2 unstable markers), MSI-L (1 unstable marker), or MSS (no instability). We classified MSI-L and MSS as MSS due to their similar clinical characteristics. For Yonsei-Classic, the PCR tests analyzed the markers BAT-25, BAT-26, NR-21, NR-24, and MONO-27 (MSI Analysis System, Version 1.2, Promega, Madison, WI), with the classification criteria: MSI-H for ≥2 unstable markers and MSS otherwise. For Mayo Clinic colon data cohort, testing for MMR status was performed via PCR and/or MMR protein immunohistochemistry. In this dataset, for tumors evaluated by immunohistochemistry, MMR deficient (dMMR) was defined by loss of at least one MMR protein among MLH1, MSH2, PMS2, and MSH6. For tumors evaluated by PCR, tumors were classified as dMMR if >30% of the markers showed instability, and MMR proficient (pMMR) if 0% to 29% of the markers showed instability. For TCGA-CRC, TCGA-STAD, and CPATC-COAD the MSI status was determined via sequencing10,55,56,57. Based on multiple prior reports of concordance rates among MSI testing methods, PCR, IHC, and sequencing-based approaches approaching 100%58,59,60,61,62,63,64,65, variability in MSI testing methods across data cohorts should not significantly impact the training or inference results.
WSIs from the Yonsei-1, Yonsei-1-remade, and Yonsei-2 datasets (in.mrxs format) were generated using the PannoramicⓇ 250 Flash III scanner (3DHISTECH, Budapest, Hungary) at a pixel resolution of 0.2428 m/pixel. All WSIs from the Mayo Clinic colon cohort were scanned at 40X magnification using a Leica GT450 scanner. For the Yonsei-Classic dataset, slides were scanned at 40x magnification using the Leica Aperio AT scanner (University of Leeds, UK). Slides from the ST Mary-Colon, ST Mary-GC, and Molecular Subtypes cohorts were scanned using either Aperio or Hamamatsu at 40x equivalent magnification (0.25 m/pixel). For TCGA-CRC and TCGA-STAD, we obtained the processed tumor tiles from ref. 10. To address potential batch effects across datasets, Macenko color normalization66, a widely used method for processing H&E-stained WSIs67, was applied to all the tumor tiles.
All patient data used in this study were obtained either with informed consent or under an exemption for retrospective studies, as determined by the Institutional Review Board (IRB). The relevant IRBs include Gangnam Severance Hospital IRB (Approval No: 3-2020-0035 and 3-2021-0367), the IRB of Severance Hospital, Yonsei University (Approval No: 4-2020-0724), the IRB of the College of Medicine at The Catholic University of Korea (Approval No: KC20RISI0329 and KC19SESI0518), and the Mayo Clinic IRB (Approval No: 806-96). The study was conducted in full compliance with IRB-approved protocols, ensuring adherence to ethical guidelines for human research. Where applicable, the IRB granted a waiver of informed consent in accordance with institutional and regulatory policies, due to the retrospective nature of the study and minimal risk to participants.
Weakly supervised learning and prediction uncertainty analysis
Prediction tasks using WSIs can naturally be formulated as a weakly supervised learning problem or a set problem68. Typical WSIs can exceed gigapixels in size, so one of the most common approaches is to divide a WSI into multiple, non-overlapping small image tiles. In contrast to standard supervised learning, where an input point and corresponding label are paired, in the weakly supervised learning that we utilized for the current study, a set of input points (image tiles) are given a slide-level single label, which in our study is the PCR-determined MSI status or IHC-detection of mismatch repair protein presence. Many early MSI prediction methods10,12,14,16,17 do not fully solve the prediction problem in weakly supervised learning, due to simplicity in implementation and computational limitations. These models were trained in standard supervised learning, which trains models to associate each image tile with its slide-level label. Within the scope of weakly supervised learning, the prediction process was based on aggregating individual tile predictions within the slice by mean or max pooling. However, due to intratumor heterogeneity, some image tiles, e.g., MSI-H-like tiles in an MSS slide or MSS-like tiles in an MSI-H slide, may introduce noise into the models, and all tiles in a slide may not be relevant for prediction.
Multiple instance learning (MIL)69 was applied to address prediction tasks involving inputs of variable size. MIL is a specialized form of weakly supervised learning, where each data point consists of a bag of multiple instances paired with a bag-level label. Most MIL methods derive a bag-level representation by aggregating instance features, a strategy known as the embedding-level MIL approach38. The simplest approaches are pooling methods, such as mean or max pooling, but these operations are non-trainable. The attention-based methods aggregate instance-level features through a weighted sum, where the weights are trainable and learned from the data38. Variants of transformers (multi-head self-attention)70 have been integrated into MIL methods to model the correlation among instances in a bag40. Recently, self-supervised learning methods have been integrated into the MIL framework to learn improved representations in an unsupervised manner71,72. Previous studies on MSI prediction models have utilized one or a combination of these MIL learning frameworks, including attention mechanisms, transformers, and self-supervised learning47,73,74,75. As previously mentioned, our DGP models aggregate scores (log odds) from individual tiles within a slide to generate slide-level predictions. While our approach may have less learning capacity compared to the embedding-level MIL approach, it offers increased interpretability, such as providing spatial MSI distribution across a slide. In addition, the experimental results, when compared with multiple MIL methods, demonstrate that our approach achieves comparable prediction performance while exhibiting consistent and stable results across diverse validation and training datasets.
It is important to note that caution must be exercised when applying MIL methods to the MSI prediction task. The assumption on the label generation process in standard MIL may be overly restrictive and fail to accurately reflect how slide-level MSI labels are assigned in current MSI testing methods, such as PCR or IHC. In standard MIL, the bag-level (slide-level) label is determined by the presence of positive instances (tiles) within the bag69. For instance, a slide is labeled as positive if it contains even a single positively labeled tile among numerous negative tiles. While this assumption is well-suited for tasks like tumor detection, where a slide is classified as normal only if no tumor is present, it is less appropriate for MSI prediction. For example, in IHC-based MSI testing, tumors are classified as dMMR (MSI-H) if there is a complete absence of nuclear staining for any MMR protein in tumor cells, while normal cells retain nuclear expression. Tumors are labeled as pMMR (MSS) otherwise. Due to tumor heterogeneity, MSS slides can still contain MSI-H regions within the tumor. This distinction highlights a fundamental difference between MSI prediction and tumor detection tasks. This restrictive assumption about the label generation process in MIL may not significantly affect most MIL methods when predicting unseen slides. However, certain MIL methods rely heavily on this assumption during training. For instance,76 employed self-supervised contrastive learning based on the assumption that all instances from negative bags inherently belong to the same set (class). This approach may introduce noise into the model due to the tumor heterogeneity, as discussed earlier. We therefore use the term “weakly supervised learning” rather than “multiple instance learning” to describe the MSI prediction using whole slide images.
Quantifying the uncertainty in the prediction allows us to understand what a prediction model does or does not know. A high degree of uncertainty at a specific test point may indicate that the model’s lack of confidence in its prediction, possibly because the test point is out of the training data distribution, or because there are unknown variables or noise within the observations. These underlying causes of high prediction uncertainty align closely with two different types of uncertainty: epistemic and aleatoric uncertainty77,78. Epistemic uncertainty is related to the randomness of the model parameters due to the insufficient number of training data and can be reduced if we collect more data. Conversely, aleatoric uncertainty refers to intrinsic noise in the observations and is irreducible. Once the prediction uncertainty is computed, we can refer the test points, or WSIs in this context, with high uncertainty to human experts for in-depth evaluation. This step can potentially reduce prediction errors on challenging test points, thereby enhancing the overall performance of the prediction model. Standard deep learning-based methods, limited to providing point estimates, are unable to capture prediction uncertainty51. The final outputs of these methods (e.g., softmax probabilities in neural networks) are frequently misinterpreted as uncertainty, but unfortunately, they are known to be overconfident for test points far from the training data53 and miscalibrated52. On the other hand, Bayesian approaches can intrinsically generate uncertainty in the prediction, providing a distribution over a prediction in Bayesian learning (by averaging the likelihood over the posterior distribution of the model parameters). Gaussian Process (GP)79, a nonparametric Bayesian method for nonlinear function estimation, allows us to compute a prediction distribution in the form of a Gaussian distribution, where the variance captures the uncertainty in the prediction. DGP26,80, a multi-layer hierarchical extension of GPs, inherits the attractive properties of GPs, including nonparametric prior modeling and well-calibrated uncertainty estimation, while providing a more flexible and generalizable prior distribution than GPs. It is noteworthy that a GP is nothing more than a special case of a DGP (i.e., a single-layer DGP).
Image preprocessing
Each pathology image was divided into multiple non-overlapping patches (the size of each image tile is set to 256 × 256 μm). Only image tiles containing mainly tumor were used in the experiments: an image tile containing non-tissue regions or consisting of ≥50% of white background was discarded. To detect tumor from image patches, we trained a ResNet-18 model on TCGA-CRC data. For the GC-ICI cohort, a U-Net-based tumor detection model was employed to predict tumor regions. This approach was selected because the cohort includes biopsy slides, which are smaller in size compared to resection slides and are believed to require higher-resolution, pixel-level tumor predictions rather than tile-level predictions. All remaining image tiles were color normalized using Macenko normalization method66 to suppress possible variations across samples or different data cohorts. Then, each image patch was fed to the trained tumor detection model.
Feature transfer learning
For our DGP model, we used transfer learning to extract features from image patches: a feature vector for an image tile was calculated using a pre-trained model (feature extraction in transfer learning81). Tumor tiles were transformed into image features using either CTransPath28 or one of the nine MSIDETECT CNNs12. These features were subsequently utilized as inputs for our DGP models described below.
Problem definition
The task of predicting the MSI status from a whole slide image can be defined in weakly supervised learning. We are given pairs of an input image and an output label, i.e., \({\{({{\mathcal{I}}}_{i},{y}_{i})\}}_{i = 1}^{N}\), where \({{\mathcal{I}}}_{i}\) represents the ith image, yi is a binary label, i.e., yi ∈ {0, 1}, where yi = 1 for MSI-H and yi = 0 for MSS, and N is the total number of the training images. Each image is divided into multiple non-overlapping small image patches each of which can be processed separately. Using transfer learning to deal with a small number of training labels, each image \({{\mathcal{I}}}_{i}\) can be represented by a set of image feature vectors, i.e., \({{\boldsymbol{X}}}_{i}={[{{\boldsymbol{x}}}_{i1},{{\boldsymbol{x}}}_{i2},...{{\boldsymbol{x}}}_{{N}_{i}}]}^{\top }\in {{\mathbb{R}}}^{{N}_{i}\times D}\), where \({{\boldsymbol{x}}}_{ij}\in {{\mathbb{R}}}^{D}\) and Ni is the number of image patches in the ith image. All training input data can be denoted by \({\boldsymbol{X}}={[{{\boldsymbol{X}}}_{1}^{\top },...,{{\boldsymbol{X}}}_{N}^{\top }]}^{\top }\in {{\mathbb{R}}}^{{N}_{T}\times D}\), where \({N}_{T}=\mathop{\sum }\nolimits_{i = 1}^{N}{N}_{i}\), and all training output labels by \({\boldsymbol{y}}={[{y}_{1},...,{y}_{N}]}^{\top }\). The objective is to learn a classifier that takes a set of N* image tiles of a new WSI, i.e., \({{\boldsymbol{X}}}_{* }={[{{\boldsymbol{x}}}_{* 1},...,{{\boldsymbol{x}}}_{* {N}_{* }}]}^{\top }\in {{\mathbb{R}}}^{{N}_{* }\times D}\) as input and estimates its prediction probability given the training data i.e., \({p}({{y}_{*}}=1|{{\boldsymbol{X}}_{*}},\,{\boldsymbol{X}},\,{{\boldsymbol{y}}})\).
In order to deal with the weakly supervised learning problem described above, where there is only a single slide-level label available for a set of multiple feature vectors computed from image tiles in a WSI, we propose to use the aggregation of image tile-level probability estimates of MSI-H based on the geometric mean of the odds operator29. We first assume that we can access the probability of each images tile (xij) being MSI-H in the ith WSI, i.e., pij = P(yij = 1∣xij) and that the log-odds (of the tile-level probability estimates) are sampled from the Normal distribution centered at the true log-odds as in29. Then, the maximum likelihood estimator of the true log-odds in this model setting is given by the geometric mean of the odds:
where σ is the sigmoid function and a > 0 is the extremization parameter. Note that a larger value of a makes \({\overline{p}}_{i}\) more extreme (\({\overline{p}}_{i}\) becomes closer to either 1 or 0). We then introduce a simple extension of (1) to the uneven tile weights by replacing the uniform weights (1/Ni) with weight terms \({\tilde{\xi }}_{ij}\ge 0\) (with \(\mathop{\sum }\nolimits_{j = 1}^{{N}_{i}}{\tilde{\xi }}_{ij}=1\)). We define a nonlinear function mapping from image tile vectors, i.e., \({{\mathbb{R}}}^{D}\), to \({{\mathbb{R}}}^{2}\) each output dimension of which are denoted by [ϕ]1 or [ϕ]2: [ϕ]1 directly models the log-odds of each image tile, i.e., \({[\phi ({{\boldsymbol{x}}}_{ij})]}_{1}=\log (\frac{{p}_{ij}}{{1-p}_{ij}})\) and [ϕ]2 models the weights with additional transformations, i.e., \({\tilde{\xi }}_{ij}={\xi }_{ij}/\mathop{\sum }\nolimits_{j = 1}^{{N}_{i}}{\xi }_{ij}\), where \({\xi }_{ij}=0.1+0.9* \sigma ({[\phi ({{\boldsymbol{x}}}_{ij})]}_{2})\). To ensure notational consistency throughout the paper, we define the (normalized) likelihood of the mapping functions ϕ with the introduction of a loss function \(\mathcal{L}\) as follows82:
where \({\overline{p}}_{i}^{\tilde{\xi }}=\sigma \left(a\mathop{\sum }\nolimits_{j = 1}^{{N}_{i}}{\tilde{\xi} }_{ij}\log \left(\frac{{p}_{ij}}{1-{p}_{ij}}\right)\right)\) is the uneven weighted extension of (1). Note that, when the cross entropy is used as the loss function, i.e., \({\mathcal{L}}({y}_{i},{\overline{p}}_{i}^{\xi })={y}_{i}\log {\overline{p}}_{i}^{\xi }+(1-{y}_{i})\log (1-{\overline{p}}_{i}^{\xi })\), where the right side on (2) becomes nothing but \(\sigma \left({\tilde{y}}_{i}a\mathop{\sum }\nolimits_{j = 1}^{{N}_{i}}{\tilde{\xi} }_{ij}\log \left(\frac{{p}_{ij}}{1-{p}_{ij}}\right)\right)\), where \({\tilde{y}}_{i}\) is the signed binary label, i.e., \(\left({\tilde{y}}_{i}=2({y}_{i}-0.5)\right)\).
DGPs with random feature expansion
This subsection shows that the mapping function ϕ can be modeled using a DPG with random feature (RF) expansions31,32. More formally, we assume that ϕ is modeled by L layers of GPs (i.e., a DGP with L layers):
where the superscript (l), where 1 ≤ l ≤ L, denotes the lth layer and f(l) in each layer is a multivariate function whose the output dimensionality is D(l), i.e., \({f}^{(l)}\in {{\mathbb{R}}}^{{D}^{(l)}}\). Each output dimension is assumed to be modeled by an individual GP, and thus there are D(l) number of GPs in each layer.
To understand this modeling more clearly, let us consider the latent function values of the all the training data points X up to the lth layer: \({{\boldsymbol{F}}}^{(l)}\in {{\mathbb{R}}}^{{N}_{T}\times {D}^{(l)}}={\phi }^{(l)}({\boldsymbol{X}})\triangleq \left({f}^{(l)}\,{\circ}\,{\ldots}\,{\circ}\,{f}^{(1)}\right)({\boldsymbol{X}})\). Assuming that all the GPs in each layer share the same covariance matrix, we have \(p({{\boldsymbol{F}}}^{(l)}| {{\boldsymbol{F}}}^{(l-1)})=\mathop{\prod }\nolimits_{k = 1}^{{D}^{(l)}}{\mathcal{N}}({{\boldsymbol{F}}}_{:,k}^{(l)}| 0,{{\boldsymbol{K}}}^{(l)})\), i.e., each column \({{\boldsymbol{F}}}_{:,k}^{(l)}\) is modeled by a GP with the covaraince matrix \({{\boldsymbol{K}}}^{(l)}\in {{\mathbb{R}}}^{{N}_{T}\times {N}_{T}}\) whose \((i,{i}^{{\prime} })\) element is defined over the output function values of the previous layer i.e., \({\kappa }^{(l)}({{\boldsymbol{f}}}_{i}^{(l-1)},{{\boldsymbol{f}}}_{{i}^{{\prime} }}^{(l-1)})\), where κ(l) is the covariance function the lth layer and fi is the ith row vector of \({{\boldsymbol{F}}}^{(l-1)}\in {{\mathbb{R}}}^{{N}_{T}\times {D}^{(l-1)}}\), i.e., \({{\boldsymbol{f}}}_{i}^{(l-1)}={({{\boldsymbol{F}}}_{i,:}^{(l-1)})}^{\top }\). Note that, the layer depth at zero is defined to be the input layer, i.e., \({{\boldsymbol{F}}}^{(0)}={\boldsymbol{X}}\in {{\mathbb{R}}}^{{N}_{T}\times D}\).
Note that, the total number of instances, NT, can be massive even if the number of images N is small (e.g., the number of whole slide images is a few hundreds, but each image can have thousand image tiles). In addition, the memory space of and the time complexity of algebraic operations on each covariance matrix are \({N}_{T}^{2}\) and \({N}_{T}^{3}\), respectively, which makes a GP prohibitive even for a dataset of hundreds of images. To alleviate this computational issue, we consider the low-rank approximation of the covariance matrices {K(l)}:
where \({{\mathbf{\Phi }}}^{(l)}\in {{\mathbb{R}}}^{{N}_{T}\times m}\) and m ⩽ NT. This approximation leads a Bayesian linear model that can approximate the GP latent function values83. Using the notational abuse, let us define F(l) ≜ Φ(l)W(l), where the priors over the linear weight matrix \({{\boldsymbol{W}}}^{(l)}\in {{\mathbb{R}}}^{m\times {D}^{(l)}}\) assumed to be i.i.d. Gaussians, i.e., \({{\boldsymbol{W}}}^{(l)}={\prod }_{n,k}{\mathcal{N}}({{\boldsymbol{W}}}_{n,k}^{(l)}| 0,1)\). One can easily see the validity of this linear model approximation by checking \({\rm{cov}}\,({{\boldsymbol{F}}}_{:,k}^{(l)}| {{\boldsymbol{F}}}^{(l-1)})={{\mathbf{\Phi }}}^{(l)}{\mathbb{E}}[{{\boldsymbol{W}}}_{:,k}^{(l)}{({{\boldsymbol{W}}}_{:,k}^{(l)})}^{\top }]{{\mathbf{\Phi }}}^{(l)}\approx {{\boldsymbol{K}}}^{(l)}\).
To implement the low-rank approximation in (4), we employ random feature expansions31,32. First consider the arc-cosine kernel function as the covariance between two input points h and \({{\boldsymbol{h}}}^{{\prime} }\) 84 (in our case the input h is assumed to be the output of the previous layer of the DGP model, i.e., \({\boldsymbol{h}}={{\boldsymbol{f}}}_{i}^{(l-1)}\) and \({\boldsymbol{h}}={{\boldsymbol{f}}}_{j}^{(l-1)}\) with arbitrary indices i and j):
where H is the Heaviside step function. Note that we can approximate the integration with finite samples drawn from the Gaussian (\({\omega }_{1},...,{\omega }_{m} \sim {\mathcal{N}}({\boldsymbol{\omega }}| 0,{\boldsymbol{I}})\)). With the fact that when p = 1, H(⋅)(⋅)p becomes ReLU( ⋅ ), the low rank matrix Φ(l) in the approximation (5) can be written by
where \({{\mathbf{\Omega }}}^{(l-1)}=[{{\boldsymbol{\omega }}}_{1},...,{{\boldsymbol{\omega }}}_{m}]\in {{\mathbb{R}}}^{{D}^{(l-1)}\times m}\) and \({{\boldsymbol{F}}}^{(l-1)}\in {{\mathbb{R}}}^{{N}_{T}\times {D}^{(l-1)}}\) are the outputs of the previous layer by the definition. Recalling that F(l) = Φ(l)W(l) and W(l) or Ω(l) can be defined as network weights with a prior distribution (Gaussian), a DGP with random feature expansion can be reduced to a Bayesian neural network.
Model inference
We train the model in Bayesian learning framework with the black-box (BB) α-divergence minimization85,86 which can be, roughly speaking, understood as a stochastic gradient version of power expectation and propagation (EP)87. Power EP, which is based on the local α-divergence minimization88, generalizes EP to include variational inference (VI) (α → 0) or EP (α = 1) as a special case with a general setting of the parameter α. However, it does not scale well because its implementation involves storing a local approximation parameter (also known as a site parameter) of each likelihood factor (each data point) in memory. Furthermore, since Power EP (as well as EP) is based on message passing, its solution is not guaranteed to converge to a stationary point of the energy function. On the other hand, since the BB-α divergence minimization directly optimizes the energy function with respect to the global (single) parameter which is combined from the site parameters without performing message passing85, we can directly apply any gradient descent methods or desirably stochastic gradients methods for the optimization and thus this method is applicable for large-scale problems. In particular, in this work we use the further approximate version of BB-α divergence minimization proposed in86 because it leads a simple and efficient (variational) inference method along with the use of Monte Carlo Dropout30,89.
We first define the approximate posterior distribution over the model parameters \(\left(\psi \triangleq {\{{{\mathbf{\Omega }}}^{(l)},{{\boldsymbol{W}}}^{(l)}\}}_{l = 0}^{L-1}\right)\). We show here only the case of the linear model parameter W(l) (the superscript for the layer number is omitted for rotational brevity), but the posterior distributions over the other variables can be defined in exactly the same way. The posterior distribution over W is defined as a mixture of two Gaussian distributions30:
where
where πw ∈ [0, 1]. Let us then define MW = [mW,1, . . . , mW,m]. Introducing a binary variable vector zw each element of which follows the Bernoulli distribution, i.e., zw,k ~ Bernoulli(πw) for k = 1, . . . , m and letting τ tend to zero, random samples drawn from the posterior (7) can be approximated by
where the operator diag[v] is a diagonal matrix with the vector v and each element in the binary vector zw can switch on or off the corresponding column of \(\widehat{{\boldsymbol{W}}}\) with the probability πw. We apply the same construction to the posterior distributions of the other variables and define the variational parameters \(\theta ={\left\{{{\boldsymbol{M}}}_{\Omega }^{(l)},{{\boldsymbol{M}}}_{W}^{(l)}\right\}}_{l = 1}^{L}\). Now, we can define the BB-α divergence energy function86 of our model for a mini-batch \({{\mathcal{O}}}_{b}\) using Monte Carlo expectation with random samples (\({\{{\widehat{\psi }}_{s}\}}_{s = 1}^{S}\)) drawn from the variational posterior distribution over the model parameters which is parameterized by the variational parameters θ, i.e., qθ(ψ):
where π = πw = πΩ and \({\widehat{\overline{p}}}_{i,s}^{\,\xi }\) is a forward pass of \({\overline{p}}_{i}^{\xi }\) at the parameter \({\widehat{\psi }}_{s}\), p0 is the prior distribution (a Gaussian distribution for one column in each parameter matrix) and the Kullback-Leibler (KL) divergence in the first line was also approximated as in30. To create a mini-batch \({{\mathcal{O}}}_{b}\), a batch-sized number (i.e., \(| {{\mathcal{O}}}_{b}|\)) of slides were randomly selected. Since each whole slide image (WSI) can contain thousands of tumor image tiles, up to 300 tiles per slide were randomly sampled using the tlle-level attention weights for model training. This method was inspired by a recent study demonstrating that random subsampling can enhance prediction performance over no sampling for binary classification in multiple-instance learning with WSIs when a sufficient number of tiles (100 to 1000) are included90.
Prediction for a new test image
The predictive distribution of a test image (feature vectors) \({{\boldsymbol{X}}_{*}}\) can be computed as
Again, the above expectation can be approximated using random samples of the model parameters by resampling the binary Bernoulli variables (i.e., using Monte Carlo dropout). For example, the predictive distribution of a test image being MSI-H is given by \(q({y}_{* }=1| {{\boldsymbol{X}}}_{* })\approx \frac{1}{S}\mathop{\sum }\nolimits_{s = 1}^{S}\exp \{{-\mathcal{L}}(1,{\widehat{\overline{p}}}_{* ,s}^{\,\xi }| {\widehat{\psi }}_{s})\}=\frac{1}{S}\mathop{\sum }\nolimits_{s = 1}^{S}{\widehat{\overline{p}}}_{* ,s}^{\,\xi }\) (recall that σ is the sigmoid function and \({\widehat{\overline{p}}}_{* ,s}^{\,\xi }\) is a forward pass of \({\overline{p}}_{* ,s}^{\xi }\) from the DGP model with the sampled model parameter \({\widehat{\psi }}_{s}\) at the test image X*). To evaluate the uncertainty in prediction, we calculate the variance of the predictive distribution as follows91
where the first term in the last equation is aleatoric uncertainty and the second term epistemic uncertainty78,91. Again, the variance can be approximated with random samples drawn from qθ(ψ):
where \({\widehat{\overline{p}}}_{* }^{\xi }=\frac{1}{S}\mathop{\sum }\nolimits_{s = 1}^{S}{\widehat{\overline{p}}}_{* ,s}^{\xi }\).
Prediction improvement using prediction uncertainty
As MSI-SEER typically made misclassifications at low prediction uncertainty, prediction performance can be improved by discarding the most uncertain predictions. First, Bayesian confidence scores (BCSs) are computed from the predictive variances defined in Eq. (13). Predictions are then sorted based on their BCSs, and the most uncertain predictions (those with the lowest BCSs) are discarded at a specified discard rate.
Performance evaluation
For the evaluation metrics, we used the area under the ROC curve (AUC), whose value range is from 0 to 1. An AUC close to 1 indicates that a model has good predictive power. To compare the performance of two classification methods, we used DeLong’s method37, which tests whether the AUC of one model is significantly different from that of another model.
Implementation
In addition to the number of layers in the DGP model (L), our DGP model incorporates several user-defined hyperparameters. These include the parameter α in the black-box α divergence formulation (Eq. (10)) in Methods), and the rank of the approximate covariance matrix in each GP layer (m). The black-box α divergence formulation simplifies to expectation and propagation (EP) when α = 1 or to variational inference (VI) when α converges to 0. For all experiments, we set α = 0.5, as prior studies have shown that using a non-standard setting of α, e.g., α = 0.5, outperformed EP (α = 1) or VI (α → 0)85. The rank of the approximate covariance matrix (m) was fixed at 100 across all GP layers and experiments.
To train our DGP models and optimize the objective function, Eq. (10) in Methods, we employed stochastic gradient methods. Specifically, we used the Adam optimizer with the learning rate of 0.001, β1 = 0.9, β2 = 0.999 and ϵ = 1e−7. The maximum number of training epochs was set to 100.
We trained CNN-based deep learning models according to the same training steps described by Laleh et al.33 for comparison. We used ResNet, ShuffleNet, and EfficientNet as the backbone CNN Models pretrained on ImageNet. The models were finetuned end-to-end for the MSI prediction task with a training epoch of 8 and a patience of 5. Training was stopped if the validation loss did not decrease. For all CNNs, we trained models with learning rates set to 1e−4, weight decay at 1e−5, batch size of 512; using Adam optimizer and freeze ratio of layers at 0.5. Both DGP models and CNNs were implemented in PyTorch using Python 3.7.
Data availability
The Macenko color-normalized and downsampled tile images of TCGA-CRC and TCGA-STAD, utilized in the experiments described in the main article, are available at https://zenodo.org/records/2530835. Whole slide images of the CPATC-COAD cohort are available at https://www.cancerimagingarchive.net/collection/cptac-coad/. Whole slide images from the remaining data cohorts are available from the corresponding authors upon reasonable request. For inquiries related to the Mayo Clinic colon cohort, please contact Rish K. Pai at pai.rish@mayo.edu.
Code availability
The DGP models were implemented in Python, with accompanying Jupyter notebooks illustrating the training and inference processes. The source codes are available at https://github.com/hwanglab/MSI-SEER.
References
Cristescu, R. et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat. Med. 21, 449–456 (2015).
Miceli, R. et al. Prognostic impact of microsatellite instability in Asian Gastric Cancer Patients Enrolled in the ARTIST trial. Oncology 97, 38–43 (2019).
Smyth, E. C. et al. Mismatch repair deficiency, microsatellite instability, and survival: an exploratory analysis of the medical research council adjuvant gastric infusional chemotherapy (MAGIC) trial. JAMA Oncol. 3, 1197–1203 (2017).
Ribic, C. M. et al. Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N. Engl. J. Med. 349, 247–257 (2003).
Sargent, D. J. et al. Defective mismatch repair as a predictive marker for lack of efficacy of fluorouracil-based adjuvant therapy in colon cancer. J. Clin. Oncol. 28, 3219–3226 (2010).
Ajani, J. A. et al. Gastric Cancer, Version 2.2022, NCCN clinical practice guidelines in oncology. J. Natl Compr. Cancer Netw. 20, 167–192 (2022).
Cervantes, A. et al. Metastatic colorectal cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann. Oncol. 34, 10–32 (2023).
Benson, A. B. et al. Colon cancer, version 2.2021, NCCN Clinical Practice Guidelines in Oncology. J. Natl Compr. Cancer Netw. : JNCCN 19, 329–359 (2021).
Eriksson, J. et al. Mismatch repair/microsatellite instability testing practices among US physicians treating patients with advanced/metastatic colorectal cancer. J.Clin. Med. 8, 558 (2019).
Kather, J. N. et al. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat. Med. 25, 1054–1056 (2019).
Cao, R. et al. Development and interpretation of a pathomics-based model for the prediction of microsatellite instability in colorectal cancer. Theranostics 10, 11080–11091 (2020).
Echle, A. et al. Clinical-grade detection of microsatellite instability in colorectal tumors by deep learning. Gastroenterology 159, 1406–1416.e11 (2020).
Krause, J. et al. Deep learning detects genetic alterations in cancer histology generated by adversarial networks. J. Pathol. 254, 70–79 (2021).
Yamashita, R. et al. Deep learning model for the prediction of microsatellite instability in colorectal cancer: a diagnostic study. Lancet Oncol. 22, 132–141 (2021).
Wang, T. et al. Microsatellite instability prediction of uterine corpus endometrial carcinoma based on H&E histology whole-slide imaging. In: 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI), 1289–1292 (2020).
Lee, S. H., Song, I. H. & Jang, H.-J. Feasibility of deep learning-based fully automated classification of microsatellite instability in tissue slides of colorectal cancer. Int. J. Cancer 149, 728–740 (2021).
Echle, A. et al. Artificial intelligence for detection of microsatellite instability in colorectal cancer-a multicentric analysis of a pre-screening tool for clinical application. ESMO Open 7, 100400 (2022).
Leiby, J.S., Hao, J., Kang, G.H., Park, J. W. & Kim, D. Attention-based multiple instance learning with self-supervision to predict microsatellite instability in colorectal cancer from histology whole-slide images. In: 2022 44th Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), 3068–3071 (2022).
Schirris, Y., Gavves, E., Nederlof, I., Horlings, H. M. & Teuwen, J. DeepSMILE: contrastive self-supervised pre-training benefits MSI and HRD classification directly from H&E whole-slide images in colorectal and breast cancer. Med. Image Anal. 79, 102464 (2022).
Guo, B., Li, X., Jonnagaddala, J., Zhang, H. & Xu, X. S. Predicting microsatellite instability and key biomarkers in colorectal cancer from H&E-stained images: achieving SOTA predictive performance with fewer data using Swin Transformer. J. Pathol. Clin. Res. 9, 223–235 (2022).
Lou, J. et al. PPsNet: an improved deep learning model for microsatellite instability high prediction in colorectal cancer from whole slide images. Comput. Methods Prog. Biomed. 225, 107095 (2022).
Zhu, J. et al. Computational analysis of pathological image enables interpretable prediction for microsatellite instability. Front. Oncol. 12, 825353 (2022).
Dietze, E. C., Sistrunk, C., Miranda-Carboni, G., O’Regan, R. & Seewaldt, V. L. Triple-negative breast cancer in African-American women: disparities versus biology. Nat. Rev. Cancer 15, 248–254 (2015).
Yuan, J. et al. Integrated analysis of genetic ancestry and genomic alterations across cancers. Cancer Cell 34, 549–560.e9 (2018).
Wang, S. C. et al. Hispanic/Latino patients with gastric adenocarcinoma have distinct molecular profiles including a high rate of germline CDH1 variants. Cancer Res. 80, 2114–2124 (2020).
Damianou, A. & Lawrence, N. Deep Gaussian processes. In: Carvalho, C. M. & Ravikumar, P. (eds.) Proceedings of the International Conference on Artificial Intelligence and Statistics (AISTATS), vol. 31, 207–215 (Scottsdale, Arizona, USA, 2013).
Zhou, Z.-H. A brief introduction to weakly supervised learning. Natl. Sci. Rev. 5, 44–53 (2018).
Wang, X. et al. Transformer-based unsupervised contrastive learning for histopathological image classification. Med. Image Anal. 81, 102559 (2022).
Satopää, V. A. et al. Combining multiple probability predictions using a simple logit model. Int. J. Forecast. 30, 344–356 (2014).
Gal, Y. & Ghahramani, Z. Dropout as a Bayesian approximation: representing model uncertainty in deep learning. In: Balcan, M. F. & Weinberger, K. Q. (eds.) Proceedings of The 33rd International Conference on Machine Learning, vol. 48, 1050–1059 (PMLR, New York, New York, USA, 2016).
Rahimi, A. & Recht, B. Random features for large-scale kernel machines. In: Platt, J., Koller, D., Singer, Y. & Roweis, S. (eds.) Advances in Neural Information Processing Systems, vol. 20 (Curran Associates, Inc., 2008).
Cutajar, K., Bonilla, E. V., Michiardi, P. & Filippone, M. Random feature expansions for deep gaussian processes. In: Precup, D. & Teh, Y. W. (eds.) Proceedings of the 34th International Conference on Machine Learning, vol. 70, 884–893 (PMLR, International Convention Centre, Sydney, Australia, 2017).
Laleh, N. G. et al. Benchmarking weakly-supervised deep learning pipelines for whole slide classification in computational pathology. Med. Image Anal. 79, 102474 (2022).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (2015).
Zhang, X., Zhou, X., Lin, M. & Sun, J. ShuffleNet: an extremely efficient convolutional neural network for mobile devices https://arxiv.org/abs/1707.01083 (2018).
Tan, M. & Le, Q. Efficientnet: rethinking model scaling for convolutional neural networks. In: International conference on machine learning, 6105–6114 (PMLR, 2019).
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44 3, 837–45 (1988).
Ilse, M., Tomczak, J. M. & Welling, M. Attention-based deep multiple instance learning. In: Proceedings of the International Conference on Machine Learning (ICML) 80, 2127–2136 (2018).
Lu, M. Y. et al. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat. Biomed. Eng. 5, 550–570 (2021).
Shao, Z. et al. Transmil: transformer based correlated multiple instance learning for whole slide image classification. Adv. Neural Inf. Process. Syst. 34, 2136–2147 (2021).
Tang, W. et al. Feature re-embedding: towards foundation model-level performance in computational pathology. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 11343–11352 (2024).
Deodato, G., Ball, C. & Zhang, X. Bayesian neural networks for cellular image classification and uncertainty analysis. https://www.biorxiv.org/content/10.1101/824862v1.full (2019).
Altieri, L., Cocchi, D. & Roli, G. A new approach to spatial entropy measures. Environ. Ecol. Statist. 25, 95–110 (2018).
Park, S. et al. ACTA2 expression predicts survival and is associated with response to immune checkpoint inhibitors in gastric cancer. Clin. Cancer Res. 29, 1077–1085 (2023).
Hörst, F. et al. CellViT: Vision Transformers for precise cell segmentation and classification. Med. Image Anal. 94, 103143 (2024).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Wagner, S. J. et al. Transformer-based biomarker prediction from colorectal cancer histology: a large-scale multicentric study. Cancer cell 41, 1650–1661.e4 (2023).
Rubinstein, J. C. et al. Deep learning image analysis quantifies tumor heterogeneity and identifies microsatellite instability in colon cancer. J. Surg. Oncol. 127, 426–433 (2023).
Saldanha, O. L. et al. Direct prediction of genetic aberrations from pathology images in gastric cancer with swarm learning. Gastric Cancer 26, 264–274 (2023).
Hinata, M. & Ushiku, T. Detecting immunotherapy-sensitive subtype in gastric cancer using histologic image-based deep learning. Sci. Rep. 11, 22636 (2021).
Gal, Y. Uncertainty in deep learning. PhD Thesis, University of Cambridge (2016).
Guo, C., Pleiss, G., Sun, Y. & Weinberger, K. Q. On calibration of modern neural networks. In: Precup, D. & Teh, Y. W. (eds.) Proceedings of the 34th International Conference on Machine Learning, vol. 70, 1321–1330 (PMLR, 2017).
Hein, M., Andriushchenko, M. & Bitterwolf, J. Why ReLU networks yield high-confidence predictions far away from the training data and how to mitigate the problem. https://arxiv.org/abs/1812.05720 (2019).
Li, K., Luo, H., Huang, L., Luo, H. & Zhu, X. Microsatellite instability: a review of what the oncologist should know. Cancer Cell Int. 20, 16 (2020).
Liu, Y. et al. Comparative molecular analysis of gastrointestinal adenocarcinomas. Cancer Cell 33, 721–735.e8 (2018).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385.e18 (2018).
Vasaikar, S. et al. Proteogenomic analysis of human colon cancer reveals new therapeutic opportunities. Cell 177, 1035–1049.e19 (2019).
Shimozaki, K. et al. Concordance analysis of microsatellite instability status between polymerase chain reaction based testing and next generation sequencing for solid tumors. Sci. Rep. 11, 20003 (2021).
Bartels, S. et al. Concordance in detection of microsatellite instability by PCR and NGS in routinely processed tumor specimens of several cancer types. Cancer Med. 12, 16707–16715 (2023).
Salipante, S. J., Scroggins, S. M., Hampel, H. L., Turner, E. H. & Pritchard, C. C. Microsatellite instability detection by next generation sequencing. Clin. Chem. 60, 1192–1199 (2014).
Chen, J. et al. Microsatellite status detection of colorectal cancer: evaluation of inconsistency between PCR and IHC. J. Cancer 14, 1132–1140 (2023).
Chen, M.-L. et al. Comparison of microsatellite status detection methods in colorectal carcinoma. Int. J. Clin. Exp. Pathol. 11, 1431–1438 (2018).
Yamamoto, G. et al. Concordance between microsatellite instability testing and immunohistochemistry for mismatch repair proteins and efficient screening of mismatch repair deficient gastric cancer. Oncol. Lett. 26, 494 (2023).
Salins, A. G. D. D. et al. Discordance between immunochemistry of mismatch repair proteins and molecular testing of microsatellite instability in colorectal cancer. ESMO Open 6, 100120 (2021).
Ali-Fehmi, R. et al. Analysis of concordance between next-generation sequencing assessment of microsatellite instability and immunohistochemistry-mismatch repair from solid tumors. JCO Precis. Oncol. 8, e2300648 (2024).
Macenko, M. et al. A method for normalizing histology slides for quantitative analysis. In: 2009 IEEE International Symposium on Biomedical Imaging: From Nano to Macro, 1107–1110 (2009).
Smith, B., Hermsen, M., Lesser, E., Ravichandar, D. & Kremers, W. Developing image analysis pipelines of whole-slide images: Pre- and post-processing. J. Clin. Transl. Sci. 5, e38 (2020).
Zaheer, M. et al. Deep sets. In: Proceedings of the 31st International Conference on Neural Information Processing Systems, NIPS’17, Guyon, I. et al. (eds.) 3391–3401 (Curran Associates, Inc., Red Hook, NY, USA, 2017).
Dietterich, T. G., Lathrop, R. H. & Lozano-Pérez, T. Solving the multiple instance problem with axis-parallel rectangles. Artif. Intell. 89, 31–71 (1997).
Vaswani, A. et al. Attention is all you need. In: Guyon, I. et al. (eds.) Adv. Neural Information Processing Systems, 30, 5998–6008 (Curran Associates, Inc., 2017).
Li, B., Li, Y. & Eliceiri, K. W. Dual-stream multiple instance learning network for whole slide image classification with self-supervised contrastive learning. In: 2021 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 14313–14323 (2021).
Wang, Y., Zhao, Y., Wang, Z. & Wang, M. Robust self-supervised multi-instance learning with structure awareness. In: Proceedings of the thirty-seventh AAAI conference on artificial intelligence and thirty-fifth conference on innovative applications of artificial intelligence and thirteenth symposium on educational advances in artificial intelligence, AAAI’23/IAAI’23/EAAI’23 (AAAI Press, 2023).
Leiby, J. S., Hao, J., Kang, G. H., Park, J. W. & Kim, D. Attention-based multiple instance learning with self-supervision to predict microsatellite instability in colorectal cancer from histology whole-slide images. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2022, 3068–3071 (2022).
Niehues, J. M. et al. Generalizable biomarker prediction from cancer pathology slides with self-supervised deep learning: a retrospective multi-centric study. Cell Rep. Med. 4, 100980 (2023).
Gustav, M. et al. Deep learning for dual detection of microsatellite instability and POLE mutations in colorectal cancer histopathology. npj Precis. Oncol. 8, 115 (2024).
Qu, L. et al. Rethinking multiple instance learning for whole slide image classification: a good instance classifier is all you need. IEEE Trans. Cir. Sys. Video Technol. 34, 9732–9744 (2024).
Kiureghian, A. D. & Ditlevsen, O. Aleatory or epistemic? Does it matter? Structure Saf. 31, 105–112 (2009).
Kendall, A. & Gal, Y. What uncertainties do we need in bayesian deep learning for computer vision? In: Guyon, I. et al. (eds.) Advances in neural information processing systems, vol. 30 (Curran Associates, Inc., 2017).
Rasmussen, C. E. & Williams, C. K. I. Gaussian processes for machine learning https://gaussianprocess.org/gpml/chapters/RW.pdf (MIT Press, 2006).
Bui, T. D., Hernández-Lobato, J. M., Hernández-Lobato, D., Li, Y. & Turner, R. E. Deep Gaussian processes for regression using approximate expectation propagation. In: Proceedings of the International Conference on Machine Learning (ICML) https://arxiv.org/abs/1602.04133 (2016).
Li, Z. & Hoiem, D. Learning without forgetting. In: Leibe, B., Matas, J., Sebe, N. & Welling, M. (eds.) Computer vision - ECCV 2016, 614–629 (Springer International Publishing, Cham, 2016).
Lecun, Y., Chopra, S., Hadsell, R., Ranzato, M. A. & Huang, F. J. A tutorial on energy-based learning. In: Bakir, G., Hofman, T., Scholkopt, B., Smola, A. & Taskar, B. (eds.) Predicting structured data (MIT Press, 2006).
Tran, G.-L., Bonilla, E., Cunningham, J. & Filippone, M. Calibrating deep convolutional gaussian processes. In: 22nd International Conference on Artificial Intelligence and Statistics 89, 1554–1563 (2019).
Cho, Y. & Saul, L. Kernel methods for deep learning. In: Bengio, Y., Schuurmans, D., Lafferty, J., Williams, C. & Culotta, A. (eds.) Advances in Neural Information Processing Systems, vol. 22 (Curran Associates, Inc., 2009).
Hernandez-Lobato, J. et al. Black-box alpha divergence minimization. In: Balcan, M. F. & Weinberger, K. Q. (eds.) Proceedings of the International Conference on Machine Learning (ICML), vol. 48, 1511–1520 (New York, New York, USA, 2016).
Li, Y. & Gal, Y. Dropout inference in Bayesian neural networks with alpha-divergences. arXiv https://arxiv.org/abs/1703.02914 (2017).
Minka, T. Power EP. Tech. Rep. MSR-TR-2004-149, Microsoft Research Cambridge (2004).
Amari, S. Differential geometrical methods in statistics. 11–65 (Springer, 1985).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Tarkhan, A., Nguyen, T. K., Simon, N. & Dai, J. Investigation of training multiple instance learning networks with instance sampling. In: Resource-efficient medical image analysis: first MICCAI Workshop, REMIA 2022, Singapore, 95–104 (Springer-Verlag, Berlin, Heidelberg, 2022).
Kwon, Y., Won, J.-H., Kim, B. J. & Paik, M. C. Uncertainty quantification using Bayesian neural networks in classification: Application to biomedical image segmentation. Comput. Stat. Data Anal. 142, 106816 (2020).
Acknowledgements
We acknowledge funding support from the National Cancer Institute (NCI) through grant R01CA276690 awarded to THH and SW, grant R37CA265967 to SW and THH, and grant U01CA294518 to THH and SW. Additional support was provided by the Department of Defense (DOD) under grant CA190578 to THH. THH is also supported by the Eric and Wendy Schmidt Foundation’s AI Innovation Award through the Mayo Clinic Foundation and the AACR Innovation and Discovery Grant. MFP is supported by the American College of Surgeons Resident Research Scholarship and the National Cancer Institute (3R37CA265967-01A1S1). SCW is a UT Southwestern Disease Oriented Clinical Scholar. SHL is supported by a grant from the National Research Foundation of Korea (NRF-2022R1A2C2010644). This study was conducted in accordance with the Declaration of Helsinki. Mayo Clinic H&E images used in this manuscript are part of the Colon Cancer Family Registry (CCFR, www.coloncfr.org), which is supported in part by funding from the National Cancer Institute (NCI), National Institutes of Health (NIH) (award U01 CA167551). This study was approved by the Institutional Review Boards (IRBs) of Gangnam Severance Hospital (Approval Nos: 3-2020-0035 and 3-2021-0367), the College of Medicine at The Catholic University of Korea (Approval Nos: KC20RISI0329 and KC19SESI0518), Severance Hospital, Yonsei University (Approval No: 4-2020-0724), and Mayo Clinic (IRB No: 806-96).
Author information
Authors and Affiliations
Contributions
T.H.H. conceptualized the study. J.H.C., J.K., S.H.L., Y.J.C., I.H.K., R.K.P., I.B. (Banerjee), J.J.S., I.B. (Barnfather), J.R.C,. H.K., and Y.K. curated the datasets. S.P. developed and implemented the methodology and conducted the numerical experiments. M.K. and I.J. also conducted the numerical experiments. M.F.P., S.C.W., S.P., and T.H.H. drafted the manuscript. J.H.P. and J.Y.S. reviewed the whole slide images from the GC-ICI cohort and interpreted the results. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
T.H.H. is a co-founder of kure.ai therapeutics and kure.s. This involvement is independent of the research described in this paper. The companies kure.ai therapeutics and kure.s had no influence on the study design, data collection and analysis, decision to publish, or preparation of the paper. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, S., Pettigrew, M.F., Cha, Y.J. et al. Deep Gaussian process with uncertainty estimation for microsatellite instability and immunotherapy response prediction from histology. npj Digit. Med. 8, 294 (2025). https://doi.org/10.1038/s41746-025-01580-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01580-8
This article is cited by
-
Deep learning-based mismatch repair prediction using colorectal cancer macroscopic images: a diagnostic study
Journal of Gastroenterology (2025)