Abstract
Keratinocyte carcinoma, such as Actinic Keratosis (AK) and Basal Cell Carcinoma (BCC), share similar clinical presentations but differ significantly in prognosis and treatment, highlighting the importance of effective screening. Optical coherence tomography (OCT) shows promise for diagnosing AK and BCC using signal intensity and skin layer thickness, but variability due to skin characteristics and system settings underscores the need for a standardized diagnostic method. Here, we propose an automated diagnostic method using the optical attenuation coefficient (OAC) and a joint learning strategy to classify AK, BCC, and normal skin. OAC images extracted from OCT data revealed notable disparities between normal and cancerous tissues. By incorporating probability distribution function (PDF) information alongside OAC images, the model achieved an accuracy of over 80% and approaching 100% by utilizing 3D OAC data to enhance robustness. This approach highlights the potential of OAC-based analysis for automated, intelligent diagnosis of early-stage non-melanoma skin cancers.
Similar content being viewed by others
Introduction
Keratinocyte carcinoma (KC), which includes basal cell carcinoma (BCC), squamous cell carcinoma (SCC), and actinic keratosis (AK), is the most common form of skin cancer, accounting for over 95% of all reported cases1. Among them, AK is considered to be a precursor of SCC, which may be transformed into SCC. The global incidence of KC is estimated to be in the millions annually, with a noticeable rise in recent decades. One of traditional gold standard for in-clinic diagnosis is pathological examination. While histopathological biopsy provides valuable insights, it poses additional burdens on patients in terms of time, cost, and invasiveness. Patients with early-stage cancer often exhibit limited willingness to undergo biopsy procedures due to concerns about the invasive nature of the procedure or associated costs. Therefore, the development of non-invasive detection methods is essential for early skin cancer screening. Currently, dermoscopy and reflectance confocal microscopy (RCM), which provide superficial imaging at depths of approximately 100–200 micrometers, optical coherence tomography (OCT), which offers medium-depth imaging ranging from 1 to 3 millimeters, and ultrasound imaging, which enables deep-depth imaging at 3 to 15 millimeters, are increasingly being used as supplementary diagnostic tools, allowing for preliminary diagnoses before biopsy2. Among these methods, OCT stands out due to its high speed, high resolution (3–15 μm), high signal sensitivity, and adequate imaging depth (1–3 mm), making it particularly suitable for detecting KC.
BCC and AK may exhibit certain similarities in their clinical manifestation, such as abnormal growths or rough patches on the skin, particularly with prolonged sun exposure. However, due to substantial variations in prognosis and treatment strategies for these conditions, it is imperative to prioritize efficient screening methods for patients3. Moreover, AK is a chronic precancerous lesion that typically does not metastasize and can be treated conservatively. In contrast, BCC is a malignant skin tumor that rarely metastasizes but exhibits locally invasive growth, often requiring surgical resection to prevent tissue destruction. Previous studies have defined some morphological criteria for diagnosing BCCs and AKs using OCT4,5,6. As OCT technology has advanced, it has revealed more structural details of the skin, increasing diagnostic accuracy. These details help identify BCC subtypes and distinguish AKs from SCCs and normal skin7,8. However, reliance on expert experience in these studies can pose diagnostic challenges due to the lack of quantifiable methods.
The study by Schuh et al.9 demonstrates that BCCs and AKs can be objectively diagnosed and differentiated using standardized signal intensity and layer thickness measurements with OCT. This quantitative diagnostic method can help inexperienced clinicians accurately distinguish between BCC and AK. However, signal intensity can be influenced by factors such as the patient’s skin condition and the intensity of incident light. Additionally, accurately measuring layer thickness can be challenging, particularly when the distinction of the epidermis-dermis junction (EDJ) in AK is unclear. These challenges present significant hurdles for clinical OCT diagnosis. Another quantitative diagnostic approach involves using the optical attenuation coefficient (OAC) derived from OCT images, which has been validated in identifying oral cancers10. The OAC offers an improved depiction of biological tissue heterogeneity, enhancing image contrast compared to traditional intensity signals. However, due to the complexity of skin cancers, additional measures are still required to assist in diagnosis.
Recent studies have shown that machine learning methods can classify BCC and AK with ~0.76 sensitivity and 0.695 AUC using dermoscopy11 and ~74% overall accuracy/precision/recall using near-infrared spectroscopic imaging12. However, these accuracies are insufficient for reliable automated diagnostics, largely due to the limited structural information beneath the skin’s surface. With advancements in artificial intelligence, OCT has emerged as a promising tool for automated diagnosis, particularly in ophthalmology13,14,15. Its ability to leverage the layered structure of the fundus has enabled deep learning models to distinguish various ophthalmic diseases with impressive accuracy, including in multi-classification tasks16. In dermatology, Li et al. were the first to use neural networks for feature extraction from BCC images17, applying machine learning techniques for automated detection. Further, the use of full-field optical coherence tomography (FF-OCT) has demonstrated potential for improving diagnostic accuracy by acquiring higher-resolution images for automated classification18. However, FF-OCT’s relatively slow acquisition speed can introduce depth-directional jitter, and its limited penetration depth of 200 μm restricts its ability to detect deeper skin structures, hindering its application in distinguishing BCC from AK, both of which exhibit pathological features in the dermis and share similarities in OCT images19. To date, no studies have fully resolved the automated diagnosis of BCC and AK using OCT.
The aim of this study is to develop an automated diagnostic framework for AK and BCC based on OCT and OAC features. We extracted OAC images from OCT data and processed them through noise reduction and contrast enhancement. Extracting the OAC allows for more precise differentiation of variations along the depth direction of biological tissues, thus enhancing contrast in fine details. By analyzing the correlation between OAC and the degree of malignancy, we observed notable disparities in OAC between normal and cancerous tissues. Additionally, a ResNet34 network was employed to analyze the two-dimensional structural information of the OAC images, while a custom multilayer fully connected network processed the one-dimensional probability density function (PDF) features. Subsequently, OAC images and PDF data were jointly trained using both networks. OAC image can provide higher contrast than OCT image, and PDF can reflect the intrinsic property changes of the essence of the tissue. Combined with the data of these two models for joint training can better improve the accuracy of diagnosis. In this study, a dataset of 14,501 images (6137 AK, 4547 BCC, and 3817 normal skin) was used for training, resulting in a training accuracy of ~95% and a validation accuracy of ~80% for distinguishing AK, BCC, and normal tissues. To further enhance accuracy and robustness for clinical applications, 3D OAC data of lesions were fully incorporated, pushing the automated diagnosis accuracy towards 100%. This approach reduces reliance solely on skin feature structures for differentiation and effectively addresses the challenge posed by the varied structures of diseased tissues, thereby leading to clinically meaningful diagnostic accuracy.
Results
Study design and patient characteristics
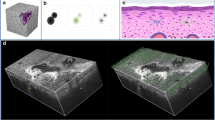
A graphical overview of the proposed framework is depicted in Fig. 1. Figure 1a shows our custom-built OCT skin imaging system (see details in Methods). This system is used to perform OCT imaging and data collection on patients prior to biopsy. As shown in Fig. 1b, we utilize a depth-resolved estimated method to extract the OAC of the skin by B-scan OCT images (see details in Methods). After OCT acquisition, biopsy diagnoses were obtained and used as labels for the data input into the neural network (Fig. 1c). We further isolate a fixed-size region of interest in the OAC images and compute the PDF within this region. 2D OAC images are input into the ResNet34 network, while 1D PDF data are fed into the FC network for joint learning (Fig. 1d and Methods). The outputs from both networks are merged and passed to the classification layer for the final diagnosis of AK, BCC, and normal skin (Fig. 1e).
a The schematic diagram of the OCT system. b Extraction of OAC images from the OCT data of AK, BCC, and normal skin, followed by flattening, cropping, and other post-processing steps. c Biopsy diagnosis of the patients, followed by OCT acquisition. d The network architecture for joint learning involves inputting OAC images into the ResNet34 network and PDF data into the FC network. The outputs from both networks are then merged and passed into the classification layer for final prediction. e Classification results of 2D OAC images for AK, BCC, and normal skin tissue.
The clinical study was conducted at a single center and approved by the ethics committees of Huashan Institutional Review Board (HIRB) (Approval Number: 2021-944, Approval Date: Jan. 14, 2022). All patients provided informed consent before participating in the study. The study subjects were patients diagnosed with AK or BCC by clinical examination between January 2022 and January 2024, excluding patients who are unable to cooperate with image acquisition and skin biopsy surgery. Among the 31 patients, 17 had AK and 14 had BCC. Among the AK patients, there were 2 males (13%) and 15 females (87%), ranging in age from 64 to 92 years old, with a mean age of 76 years old. In some patients, multiple lesions were collected, and a total of 24 cases were collected, which were distributed in the face (13 cases), around the eyes (2 cases), cheeks (5 cases) and nose (4 cases). Among BCC patients, there were 5 men (35%) and 9 women (65%), with ages ranging from 34 to 78 years and a mean age of 58 years. A total of 19 cases were collected, distributed across the head (16 cases), chest (2 cases), and legs (1 case). Subtypes of BCC included superficial type (8 cases), nodular type (10 cases), and invasive type (1 case). OCT data collection occurred before biopsy surgery, and biopsy results were used as the final diagnostic results. The research methodology followed the principles outlined in the Helsinki Declaration.
Comparison of OCT and OAC images in normal skin, AK, and BCC
To demonstrate the enhancement of internal details in biological tissues provided by OAC images, we compared the contrast differences between OCT and OAC images in normal skin, AK, and BCC (Fig. 2). In OCT images (Fig. 2a–c), the dermo-epidermal junction (DEJ) is difficult to delineate, whereas in OAC images (Fig. 2d–f), the boundary is clearly distinguishable. The thickening of the epidermal layer, a key feature for distinguishing AK, relies on accurate DEJ identification. Figure 2g–i further illustrates how OAC images enhance tissue details, with superior contrast and clarity in the dermal tissue compared to OCT images. This enhancement makes OAC images particularly effective for classification tasks, as they better represent the attenuation characteristics of biological tissues, improving the distinction between skin tissues and aiding in disease diagnosis.
a–c Depict OCT images of normal skin, AK, and BCC, respectively. d–f Show their corresponding OAC images. g–i Provide magnified views of the yellow boxed regions from the left images. All scale bars represent 0.5 mm.
Statistical comparison of pathological images with OAC images in AK and BCC
The standard method for diagnosing KC relies on pathology. For AK, in histopathological findings, dysplasia and parakeratosis are typical feature of AK. At the same time, collagen degeneration in the superficial dermis and epidermal thickening are frequently observed. Compared to normal collagen tissue, which appears pale pink in Fig. 3a, the superficial dermis in the AK pathological image exhibits degenerated collagen tissue, marked by a yellow circle in Fig. 3b.
20x pathological images of a normal skin and b AK, respectively. c Segmented heatmap of the OAC image for normal skin tissue collected from the biopsy region, corresponding to the pathological image in (a). d PDF distribution plot for the image in (c), with PPD representing pixel probability density. e Segmented heatmap of the OAC image and f PDF distribution from an AK patient, corresponding to the pathological image in (b). g Segmented heatmap of the OAC image and h PDF distribution from another AK patient with a lesion. Scale bars: a and b represent 0.1 mm; c, e, and g represent 0.5 mm.
As OAC images provide more structural detail than OCT images, they enable a better distinction between normal and cancerous tissue. We further conducted a statistical analysis of OAC intensity (see details in Methods). Figure 3c, d illustrates the segmented heatmap and PDF of the OAC image of normal skin tissue, while Fig. 3e, f shows the corresponding OAC images from adjacent skin sites of the same patient with an AK lesion. Comparing Fig. 3c–e, the OAC in the superficial dermis of AK significantly decreases, indicating collagen degeneration. This observation aligns closely with the histopathological characteristics of AK. In contrast, such a finding is absent in the analyzed basal cell carcinoma cases. This is more clearly reflected in the PDF, as shown in Fig. 3d, f. Although the distribution pattern of the OAC in normal skin still persists, as marked by the black arrow, the OAC of skin affected by AK exhibits an overall decrease, as shown by the noticeable downward shift in the proportion of OAC values greater than 3 mm−1. The emergence of a peak around 1 mm−1 to 1.5 mm−1, marked by the yellow arrow, represents degenerated collagen tissues. Therefore, the overall decrease in OAC in the superficial dermis and the signal of collagen degeneration may serve as a key feature to differentiate AK from BCC, as they correspond to the pathological hallmark of collagen degeneration. To exclude randomness, we analyzed the OAC distribution in skin samples from multiple patients with AK. Figure 3g shows an example of the OAC image from another AK patient, where prominent skin lesions are observed, in contrast to Fig. 3e. Despite variations in epidermal morphology, a significant reduction in OAC extent is observed compared to normal skin tissue (Fig. 3h).
Figure 4 presents pathological images, OAC images, and PDF plots of BCC at various stages of lesion severity. Compared to AK, BCC exhibits distinct structural features in pathology, with cancer cells forming oval-shaped nests, as shown in Fig. 4a. These regions are evident in the extracted OAC images, highlighted by the yellow circles in Fig. 4b, where the OAC values are not significantly reduced compared to the collagen tissue in AK. This is further corroborated by the PDF distribution, which shows a slight elevation near the value of 2 mm−1 (denoted by the red arrow in Fig. 4c). As the severity of BCC lesions increases, these features become more pronounced (Fig. 4d–i). The pathological images reveal progressively larger areas of cancer cell nests. This progression is also reflected in the corresponding PDFs, where the characteristic peaks gradually widen and rise, indicating an expansion of the lesion tissue, as shown in the gray-shaded areas of Fig. 4c, f, i. Notably, the trend of OAC above the value of 4 mm−1 does not show the significant decrease as observed in AK.
a Pathological image of a BCC lesion. b Segmented heatmap of the OAC image corresponding to the same lesion region shown in (a). c PDF distribution derived from the OAC image in (b). d–i Pathological images, segmented OAC heatmaps, and PDF plots of BCC lesions with increasing severity. Yellow circles highlight the same lesion areas across both pathological and OAC images of the same patient. The gray-shaded areas mark the characteristic region associated with BCC lesion tissue in the PDF plots. All scale bars represent 0.5 mm.
The analysis above highlights the similarities and differences in OAC among AK, BCC, and normal skin tissue by comparing their OAC images and PDF distributions. However, it is important to note that not all PDF distributions of AK and BCC strictly follow the above mentioned pattern, as variations depend on lesion severity and skin condition. In manual diagnosis, the structural and PDF similarities can confuse doctors, leading to diagnostic difficulties and potential errors. To address this, we propose a machine learning approach using neural networks and extensive datasets to improve classification and diagnostic accuracy.
The performance of the automated diagnosis using joint learning
To improve accuracy, we developed and tested a joint learning approach that integrates OAC structural features with PDF features, enabling more reliable automated diagnosis (see details in Methods). We evaluated the performance of this joint learning model using the provided dataset and compared it with the 2D-OAC model, which utilizes only 2D OAC images. The PDF, a robust feature reflecting the extent of pathological changes, shows significant potential for aiding diagnosis and assessing cancer progression. The joint learning model demonstrates the enhanced classification performance achieved by incorporating PDF information.
Table 1 presents the datasets and parameters used in training the neural network models. Although the number of normal tissue samples is fewer than that of AK and BCC cases, this has a limited impact on model training. The primary reason is that the optical properties of normal tissue are relatively stable, with fewer variations compared to diseased tissues. The current dataset is sufficient for the model to achieve high accuracy, and further expansion of normal tissue data is unlikely to yield significant improvements in performance. In contrast, AK and BCC exhibit more complex pathological morphology and greater individual variability in optical properties, requiring a larger sample size to capture their diversity.
We employed ResNet34 and a customized joint learning network, conducting 200 epochs to sufficiently capture the characteristics of diseased tissues. Additionally, to ensure the model’s generalizability, the test set consisted entirely of samples from different patients, enhancing the model’s reliability in practical applications. The batch size was set to 25, and the learning rate was maintained at 10–5 to ensure stable and effective training. Although the normal tissue samples were relatively limited, their characteristic patterns were consistent, allowing the model to learn key features even with fewer examples. As a result, this data distribution effectively supports robust model learning while ensuring reliable performance in real-world applications.
Table 2 presents the classification results for normal skin, AK, and BCC, comparing the performance of the 2D-OAC model with the joint learning model. The results highlight the advantages of integrating additional information beyond OAC structural features. For normal tissue classification, both models achieved high accuracy. However, in classifying diseased tissues, particularly AK and BCC, the joint learning model consistently outperformed the 2D-OAC model in accuracy, sensitivity, and precision. This improvement is particularly notable for AK cases, where the joint learning model achieved a 3.9% higher accuracy and a 4.5% improvement in precision compared to 2D-OAC model. Similarly, for BCC, the joint learning model exhibited superior sensitivity and precision, leading to an overall more balanced classification performance. Overall, the joint learning model demonstrated a higher overall accuracy (78.8% vs. 75.1%), sensitivity (83.3% vs. 80.5%), and precision (78.8% vs. 75.1%), confirming its effectiveness in distinguishing between different skin conditions. These findings suggest that incorporating additional features beyond OAC structural information enhances the model’s ability to identify and differentiate skin lesions, making it a more robust approach for automated diagnosis.
Figure 5a–c compare the Area Under the Curve (AUC) between the 2D-OAC model and the joint learning model for different classes. For normal cases, both models achieved an AUC of 1.0, indicating perfect classification performance. However, for AK, the joint learning model achieved an AUC of 0.870, outperforming the 2D-OAC model, which achieved an AUC of 0.805. This difference was statistically significant, with a Z-score of −5.1461 and a p-value of 2.66E−07. Similarly, for BCC, our model achieved an AUC of 0.855, compared to 2D-OAC’s 0.716, with a Z-score of −9.9721 and a p-value of 2.02E−23, further highlighting the significant improvement in classification performance.
ROC curves comparing the performance of the joint learning model (green) and the 2D-OAC model (blue) in classifying a normal skin tissues, b AK, and c BCC. The confusion matrix of the test set was obtained using d 2D-OAC model and e the joint learning model, respectively.
Crucially, by incorporating one-dimensional PDF information, our approach significantly improved classification performance. This validates the effectiveness of PDF information in aiding diagnosis and underscores its crucial role in classification tasks. Compared to the 2D-OAC model, which relies solely on OAC structural feature, the joint learning model further enhances classification accuracy, highlighting its potential value in clinical diagnostics. This also shows that relying solely on image data, even OAC images with high contrast, is not enough to achieve a satisfactory diagnostic accuracy. The possible reason is that due to the variability of the structure, the addition of multi-angle data can achieve a higher diagnostic accuracy.
Six untrained datasets of AK, BCC, and normal skin tissues were used for prediction to assess the network’s overfitting. The confusion matrix of the prediction results is shown in Fig. 5d, e. Compared to the 2D-OAC model, the joint learning method achieved a 5% increase in accuracy on the test set, reaching 80.2% for BCC and 70.6% for AK. In clinical diagnosis, lesion data are typically collected as 3D volumetric images, meaning multiple OCT images correspond to a single lesion. Therefore, we recommend using the majority label from the entire volumetric dataset as the final prediction. Testing this method on the six unseen volumetric datasets resulted in 100% prediction accuracy at the volumetric level.
Discussion
Non-invasive diagnostic methods for distinguishing between benign and malignant skin tumors help reduce unnecessary pain and costs for patients, as well as alleviate the economic burden on the healthcare system. Although advancements in OCT technology and imaging quality have provided more structural details for distinguishing these conditions, the accuracy of these methods still heavily depends on the physician’s experience. There is an urgent need for quantitative, automated diagnostic methods to overcome the limitations of experience-based manual diagnosis.
In clinical practice, a major challenge in using OCT for non-invasive diagnosis of keratinocyte carcinoma skin lesions is distinguishing between AK and BCC4. Although previous studies have demonstrated that OCT can quantitatively differentiate AK and BCC based on intensity signals and thickness variations, manual delineation of the epidermis and dermis is still required to calculate the average intensity or thickness in specific areas9. This manual process is complicated by confusion and damage at the epidermal-dermal junction, which can compromise diagnostic accuracy and increase both complexity and time consumption. A review of relevant studies highlights the difficulty of distinguishing between AK and BCC based solely on structural characteristics, due to the wide range of subtypes and diverse morphologies of these lesions4,20.
In pathological terms, AK is characterized by collagen degeneration, while BCC is characterized by the formation of cancer cell nests. These differences are reflected in OAC images as varying ranges of low OAC values. In OAC images, AK shows widespread reductions in OAC, with more severe cases demonstrating a gradual replacement of high attenuation areas in normal collagen tissue. In contrast, BCC displays localized decreases in OAC due to cancer cell nests, which are distinguishable from degenerated collagen tissue. In severe BCC cases, large areas of cancer cell nests may extend beyond the OCT detection depth, leading to extensive OAC reductions in the dermis, unlike the partial attenuation observed in AK.
Despite the existence of these mappings, confusion can still arise in diagnosing OAC images, as degenerated collagen tissue may resemble regions with similar structure features to cancer cell nests. To address this, our work further extracts PDF features from OAC images and employs deep-learning networks to assist in diagnosis. This approach combines the high contrast of two-dimensional OAC images with the intuitive representation of cancerous tissues provided by one-dimensional PDF information. By jointly training a ResNet34 network and an FC network, we achieved nearly 80% accuracy in three-class classification and an AUC of 0.85. This represents a significant improvement over using OAC images alone for classification. The p-values for the two models were well below 0.05, indicating a statistically significant difference, primarily due to the inclusion of the PDF feature, which enhanced the performance of automate diagnosis. Given the current constraints of our dataset size, our study has primarily focused on distinguishing between AK and BCC. To advance this work, we are actively expanding our dataset to incorporate SCC and Bowen’s disease, while also initiating multicenter validation to address population heterogeneity and enhance the generalizability of our model. These steps are integral to our long-term objective of developing a comprehensive diagnostic framework for common cutaneous malignancies.
While we are actively expanding our dataset to include SCC and Bowen’s disease, with the long-term objective of establishing a comprehensive diagnostic framework for common cutaneous malignancies, the proposed method already demonstrates potential for clinical implementation as a non-invasive imaging tool to assist in differentiating AK from BCC. Within diagnostic workflows, clinicians can utilize OCT imaging to capture high-resolution cross-sectional views of suspicious lesions, enabling real-time algorithmic support for enhanced decision-making. Our system offers several advantages: (1) The contact-free imaging protocol eliminates consumable costs and minimizes patient discomfort; (2) The simplified workflow and compliance with established OCT standards expedite regulatory approval pathways; (3) high imaging speed ensures practical clinical utility, reducing motion artifacts and improving patient throughput. While the upfront cost of custom OCT hardware requires meticulous assessment, the absence of recurring expenses and compatibility with automated diagnostic framework position this approach as a cost-effective solution for dermatological practice.
In this study, we have established an OCT-based non-invasive early skin cancer database to provide sufficient clinical data for developing a deep-learning-based automatic diagnosis. This database includes OCT volumetric data, pathological images, high-resolution camera images, and de-identified personal information such as age, gender, and lesion location for each patient. This long-term initiative is crucial due to the skin’s complexity and regional structural variations. Such a database will enhance research efficiency, standardize data, facilitate data sharing, and enable customized training of deep learning networks for early skin cancer diagnosis. Furthermore, our work demonstrated that high-contrast OAC images extracted from OCT data effectively reflects the differences and severity between AK and BCC. By incorporating PDF information with OAC images, we developed a joint learning approach for intelligent diagnostic tasks. The joint learning model improved classification accuracy by approximately 5% compared to using OAC images alone, with similar improvements observed in both AK and BCC classification. It further achieves over 80% accuracy in the three-class classification task using the 2D dataset, and nearly 100% accuracy when incorporating 3D OAC data to enhance robustness. Therefore, we believe that the development of more efficient classification networks, together with the expansion of skin cancer databases, holds significant potential to enhance diagnostic accuracy and improve the assessment of cancer progression.
Methods
OCT system for skin imaging
The OCT data was acquired by using a custom-made spectral-domain OCT system located within the Dermatology Department of Huashan Hospital, Fudan University (Shanghai, China). As shown in Fig. 1a, the OCT system utilizes a broadband light source with a center wavelength of 830 nm and a bandwidth of 100 nm (BLM2-D-840-B-10, Superlum). Compared to the central wavelength of 1310 nm, the central wavelength of 830 nm can achieve higher spatial resolution, which is comparable to the size of skin cells, enabling a more accurate reflection of pathological changes at the cellular level. The spectrometer consisted of a transmission diffraction grating with 1435 lines/mm (Wasatch Photonics, USA), an imaging lens with a focal length of 100 mm, and a line-scan camera (EV71YO1CCL2210-BB3, Teledyne e2v, 2048 pixels, 10 μm pixel width, USA). The spectral resolution is 0.051 nm/pixel, and the maximum imaging depth can reach 3.4 mm in air. The sensitivity of falloff is 3 dB at 1 mm and 6 dB at 1.6 mm, which ensures an adequate signal-to-noise ratio for signals more than 1 mm under the skin surface. This system achieves a 5-μm lateral resolution and a 5-μm axial resolution. The target population consisted of individuals suspected of having AK and BCC who had not yet undergone pathological surgery. For each lesion site, we collected one or more three-dimensional volumetric datasets, with the number depending on the size of the lesion. Each OCT volumetric dataset has a field of view (FOV) of 5 mm by 5 mm and a pixel resolution of 1000 by 1000 with an acquisition time of 10 s.
Depth-resolved estimation of optical attenuation coefficients
Typically, OCT relies on detecting backscattered light to reflect the optical reflectivity of biological tissues along the depth direction. With the penetration increasing, the intensity of the reflected light correspondingly decreases. Therefore, this approach may not accurately capture the intrinsic optical properties of biological tissues, such as light absorption and backscattering. In contrast, the OAC is an inherent optical property of biological tissues that describes the extent of light attenuation within the tissue, encompassing light absorption and scattering. Thus, we calculate and extract the OAC from OCT images in our work.
The OAC in biological tissue is related to the intensity of light that is back-scattered from the sample and measured by the OCT system. Several methods have been proposed to estimate the OAC from OCT data by fitting curve to a segment of an A-scan over which the OAC is assumed to be constant. However, such curve-fitting algorithms can only handle cases where the confocal function is known and it cannot distinguish small variations in the OAC at the pixel level. To reflect the cellular-level changes in OAC, we employed a proposed depth-resolved OAC calculation method21. This approach allows the calculation of the OAC for each pixel, and given our resolution is similar to that of skin cells, it enables the reflection of OAC changes at the cellular level.
This method introduced the following model:
The OAC corresponding to a single pixel can be defined as follows:
where \(\mu\) represents the optical attenuation coefficient and \(z\) denotes the depth. The variable \(i\) corresponds to the pixel index along the depth direction and \(I\) indicates the light intensity. \(I\left[i\right]\) is derived from the integrated signal over a small spatial interval defined by the pixel size Δ, which is typically associated with the coherence length of the light source. To further specify, let \(u\) and \(v\) represent the lower and upper boundaries of a pixel, respectively, such that \(u\) = \(i\Delta\), \(v\,=u+\Delta\), and \(u\le z\le v\).
By the above formula, we can calculate the OAC from OCT signals. However, in practical applications, OCT signals often have noise floor and depth-dependent signal decay. To obtain accurate OAC, we conducted corresponding calibrations. The resulting noise floor signal N(z) was subtracted from all phantom measurements. And the signal decay S(z) caused by roll-off can be expressed as22:
where \({k}_{s}\) is calculated as \({(k}_{\max }-{k}_{\min })/{N}_{{SAMP}}\), \({N}_{{SAMP}}\) represents the number of pixels collected per spectrum, and \({k}_{\max }\) and \({k}_{\min }\) represent the maximum and minimum wavenumbers of the light source. For a spectrometer-based OCT system, \({k}_{o}\) represents the optical resolution in wavenumbers, which is determined by the dispersion line width of the spectrometer. Another signal decay factor is the confocal point spread function (PSF):
where \({z}_{f}\) and \(z\) are the depth positions of the focus and the tissue concerning zero-delay, respectively, and \({Z}_{R}={\rm{\pi }}{w}_{0}^{2}/{\lambda }_{0}\) is the Rayleigh length of the sample arm optics measured in air. \({w}_{0}\) is the beam waist at the focus and \({\lambda }_{0}\) is the center wavelength of the OCT source. The PSF can be calibrated by knife-edge measurements of \({w}_{0}\). \(n\) is refractive index.
The final intensity \({{\rm{I}}}_{{\rm{final}}}({\rm{z}})\) obtained after removing the interference term is:
Denoising and extraction of statistical features from OAC
Despite our efforts to eliminate baseline noise and correct the signal decay with depth to restore the final OCT intensity image, there still exist certain high-intensity noise points. These noise points can cause fluctuations and distortion in the extracted OAC. Therefore, further denoising operations are still necessary based on the obtained OAC images. Typically, denoising in OCT employs supervised deep-learning algorithms. However, as the OCT system collects clinical data for skin cancer, obtaining high-quality ground truth is challenging. Moreover, we must consider the random and discontinuous nature of noise points that appear between consecutive frames in OAC images. Consequently, we use an unsupervised 3D deep learning method to denoise the three-dimensional OAC data23. This method is based on the 3D-Unet network, which extracts odd and even frames of a 3D dataset to form two separate 3D datasets. By calculating the loss between these two datasets, it learns the noise distribution of each frame, ultimately achieving unsupervised noise removal. This approach is suitable for continuous or time-varying 3D data. With this method, we can effectively remove abnormal noise points and enhance the image quality.
Afterward, the OAC images were flattened and cropped to focus on the specific area of interest and minimize interference from signal-free regions located deeper within the skin. The precision in capturing the lesion area enhances the accuracy of disease diagnosis and eliminates interference from irrelevant signals. The following post-processing steps were applied to convert the 2D OAC input images from Fig. 1b–d: Firstly, the Canny edge detection algorithm was employed to extract the skin edges in the OAC images. Secondly, the coordinates of the edge points closest to the upper surface were extracted, smoothed, and subjected to linear fitting using the obtained contour points of the skin surface. Finally, we flattened the image based on the fitted slope and cropped a fixed-size region. This step ensures that the extracted features contain the majority of the valid information while reducing the number of parameters input into the network.
Additionally, we applied color encoding to the OAC images to generate segmented heatmaps: pixels in the 0.7–1.5 mm–¹ range were shown in purple, those in the 1.5–3 mm–¹ range in red, and those in the 3–6 mm–¹ range in blue.
Probability density function of OAC images
The Probability Density Function (PDF) describes the likelihood of a continuous random variable taking a specific value. Mathematically, a PDF is a non-negative function f(x) such that the probability of the variable falling within an interval [a,b] is given by the integral:
In our study, f(x) represents the PDF describing the likelihood of any pixel in the OAC image taking a specific OAC value.
In each OAC image, we tallied pixel counts within the OAC range of 0.7–6 and calculate the PDF distributions. This ensures that our obtained distribution predominantly reflects the relative distribution among OAC within the designated range. Based on previous studies, we believe that cancerous changes would lead to alterations in the skin’s OAC. Drawing from pathological diagnostic criteria for AK and BCC, we found that the PDF distribution of OAC images could reflect the differences between the two. This feature, distinct from methods based on skin structural characteristics, is particularly well-suited for diagnosing diseases like AK and BCC, which are characterized by diverse and complex lesion structures.
The joint learning network architecture
We propose a joint training strategy by combining robust 1D PDF information with 2D high-order detail structure information. The joint training approach effectively balances the network, reducing the risk of overfitting and enhancing robustness. The specific network structure is illustrated in Fig. 1d.
To accommodate the characteristics of one-dimensional data in the PDF distribution, a one-dimensional fully connected (FC) network with only ten layers was employed with the number of neurons in each layer being 512 × 3, 256 × 3, 128 × 3, and 64 × 1, respectively. The simplicity of the network structure not only offers advantages in practical applications but also helps prevent overfitting, avoiding the extraction of excessive high-dimensional details. The input PDF is divided into 1000 small intervals, distributed in the OAC range from 0.7 to 6. The first nine FC layers use the ReLU activation. The final FC layer employs the softmax activation. The optimizer used is Adam with a learning rate of 0.00001. The loss function used is cross-entropy, which can be represented by the following formula:
where M is the number of categories, \({y}_{{ic}}\) is the sign function (0 or 1), if the true class of sample \(i\) equals c, take 1; otherwise, take 0, \({p}_{{ic}}\) is the predicted probability of sample\(\,i\) belonging to class c. This network is trained to classify AK, BCC, and normal skin. By utilizing one-dimensional data, the network structure is significantly compressed, enabling it to perform classification tasks with just a few FC layers.
We experimented with multiple classical network structures for classification tasks to train OAC image alone, ultimately selecting ResNet34 for its superior performance in 2D image feature extraction. The 2D feature maps extracted by ResNet34 are flattened into a 1D vector and transformed into a 64-channel vector through FC layers. This vector is then concatenated with the 64-channel feature vector obtained from the 1D feature extraction, and the combined vector is fed into a final classification layer.
After training the two networks separately, a joint training phase is conducted to fine-tune the model. The final network parameters are selected based on the best performance on the validation set. To enhance diagnostic accuracy in clinical scenarios, data is typically collected as a volume from the lesion site. This series of B-scan data is input into the network for classification, generating a probability distribution that reflects the likelihood of disease across the entire volume. The final diagnostic result is then determined based on this distribution.
Evaluation metrics
We assessed our method using five commonly used metrics in multi-class classification tasks, namely accuracy (ACC), sensitivity (SE), precision (PR), specificity (SP), and confusion matrix:
where N represents the number of classes, \({{TP}}_{i}\) is the true positives, \({{TN}}_{i}\) is the true negatives, \({{FP}}_{i}\) is the false positives, \({{FN}}_{i}\) is the false negatives for the i-th classification label.
Area under the curve (AUC), as a comprehensive measure of the model’s classification ability, is also used in our study. To avoid decreased reliability of accuracy due to uneven sample distribution, we calculate the AUC separately for AK, BCC, and normal tissue. In addition, we also calculated the confusion matrix as a parameter to provide a clearer representation of error distribution, aiding in better understanding and improving the diagnostic process.
Data availability
Anonymized data are available from the corresponding author upon reasonable request.
Code availability
The underlying code for this study are available from the corresponding author upon reasonable request.
References
Pandey, M. et al. Site-Specific Vesicular Drug Delivery System for Skin Cancer: A Novel Approach for Targeting. Gels 7, 218 (2021).
Bakos, R. M., Blumetti, T. P., Roldan-Marin, R. & Salerni, G. Noninvasive Imaging Tools in the Diagnosis and Treatment of Skin Cancers. Am. J. Clin. Dermatol 19, 3–14 (2018).
Kandolf, L. et al. European consensus‐based interdisciplinary guideline for diagnosis, treatment and prevention of actinic keratoses, epithelial UV‐induced dysplasia and field cancerization on behalf of European Association of Dermato‐Oncology, European Dermatology Forum, European Academy of Dermatology and Venereology and Union of Medical Specialists (Union Européenne des Médecins Spécialistes). J. Eur. Acad. Dermatol. Venereol. 38, 1024–1047 (2024).
Olsen, J. et al. Diagnostic accuracy of optical coherence tomography in actinic keratosis and basal cell carcinoma. Photodiagnosis Photodyn. Ther. 16, 44–49 (2016).
Jørgensen, T. M., Tycho, A., Mogensen, M., Bjerring, P. & Jemec, G. B. E. Machine‐learning classification of non‐melanoma skin cancers from image features obtained by optical coherence tomography. Ski. Res. Technol. 14, 364–369 (2008).
Mogensen, M. et al. Assessment of Optical Coherence Tomography Imaging in the Diagnosis of Non-Melanoma Skin Cancer and Benign Lesions Versus Normal Skin. Dermatologic Surg. 35, 965–972, (2009).
Ulrich, M. et al. The sensitivity and specificity of optical coherence tomography for the assisted diagnosis of nonpigmented basal cell carcinoma: an observational study. Br. J. Dermatol. 173, 428–435 (2015).
Boone, M. A. L. M. et al. A new algorithm for the discrimination of actinic keratosis from normal skin and squamous cell carcinoma based on in vivo analysis of optical properties by high‐definition optical coherence tomography. J. Eur. Acad. Dermatol. Venereol. 30, 1714–1725 (2016).
Schuh, S., Kaestle, R., Sattler, E. C. & Welzel, J. Optical coherence tomography of actinic keratoses and basal cell carcinomas – differentiation by quantification of signal intensity and layer thickness. J. Eur. Acad. Dermatol. Venereol. 30, 1321–1326 (2016).
Yang, Z., Shang, J., Liu, C., Zhang, J. & Liang, Y. Identification of oral cancer in OCT images based on an optical attenuation model. Lasers Med. Sci. 35, 1999–2007 (2020).
Guan, H. et al. Dermoscopy-based Radiomics Help Distinguish Basal Cell Carcinoma and Actinic Keratosis: A Large-scale Real-world Study Based on a 207-combination Machine Learning Computational Framework. J. Cancer 15, 3350–3361 (2024).
Courtenay, L. A. et al. Using computational learning for non-melanoma skin cancer and actinic keratosis near-infrared hyperspectral signature classification. Photodiagnosis Photodyn. Ther. 49, 104269 (2024).
Lee, C. S., Baughman, D. M. & Lee, A. Y. Deep Learning Is Effective for Classifying Normal versus Age-Related Macular Degeneration OCT Images. Ophthalmol. Retin. 1, 322–327 (2017).
Li, F. et al. Deep learning-based automated detection of retinal diseases using optical coherence tomography images. Biomed. Opt. Express 10, 6204–6226 (2019).
Shah, M., Roomans Ledo, A. & Rittscher, J. Automated classification of normal and Stargardt disease optical coherence tomography images using deep learning. Acta Ophthalmol. 98, e715–e721 (2020).
Lu, W. et al. Deep Learning-Based Automated Classification of Multi-Categorical Abnormalities From Optical Coherence Tomography Images. Transl. Vis. Sci. Technol. 7, 41 (2018).
Li, A. et al. Automated basal cell carcinoma detection in high-definition optical coherence tomography. in 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC). 2885–2888 (IEEE, 2016).
Mandache, D. et al. Basal cell carcinoma detection in full field OCT images using convolutional neural networks. in 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018). 784-787 (IEEE, 2018).
Ho, C. J. et al. Detecting mouse squamous cell carcinoma from submicron full‐field optical coherence tomography images by deep learning. J. Biophotonics 14, e202000271 (2020).
Boone, M. A. L. M. et al. High‐definition optical coherence tomography algorithm for the discrimination of actinic keratosis from normal skin and from squamous cell carcinoma. J. Eur. Acad. Dermatol. Venereol. 29, 1606–1615 (2015).
Vermeer, K. A., Mo, J., Weda, J. J., Lemij, H. G. & de Boer, J. F. Depth-resolved model-based reconstruction of attenuation coefficients in optical coherence tomography. Biomed. Opt. Express 5, 322–337 (2013).
Chang, S. & Bowden, A. K. Review of methods and applications of attenuation coefficient measurements with optical coherence tomography. J. Biomed. Opt. 24, 1–17 (2019).
Li, X. et al. Real-time denoising enables high-sensitivity fluorescence time-lapse imaging beyond the shot-noise limit. Nat. Biotechnol. 41, 282–292 (2023).
Acknowledgements
This work was supported in part by the National Key R&D Program of China (2022YFF0708700), Shanghai Basic Research Special Zone Program (22TQ020), Natural Science Foundation of Shanghai (22ZR1404300, 22ZR1409500), Shanghai Science and Technology Innovation Action Plan (22S31905500), Medical Engineering Fund of Fudan University (yg2021-032, yg2022-2), Young Talents of Shanghai Health Commission (2022YQ043), Huashan Hospital Innovation Fund (2024CX06).
Author information
Authors and Affiliations
Contributions
L.Z. and X.L. built the experimental setup and performed data collection and analysis. L.Z., W.C., and Y.G. contributed to coding, comparisons, and visualization. H.W. and Z.L. contributed to data collection, quality checking, and annotation. H.W., Z.L., and B.D. supervised the project, contributed to the conceptualization, and designed the experiment. L.Z., X.L., H.W., Z.L., and B.D. wrote the manuscript. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
B.D. has a financial interest in Lishi Intelligent Science & Technology (Shanghai) Co., Ltd. and MicroLux (Shanghai) Intelligent Science & Technology Co., Ltd., though these companies did not support this work. All other authors declare no competing interests.
Ethics
The clinical study was conducted at a single center and approved by the ethics committees of Huashan Institutional Review Board (HIRB) (Approval Number: 2021-944, Approval Date: Jan. 14, 2022). All patients provided informed consent before participating in the study. The study subjects were patients diagnosed with AK or BCC by clinical examination between January 2022 and January 2024, excluding patients with mental disorders, prisoners, or those in custody. Data collection occurred before biopsy surgery, and biopsy results were used as the final diagnostic results. The research methodology followed the principles outlined in the Helsinki Declaration.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, L., Li, X., Chen, W. et al. A joint learning approach for automated diagnosis of keratinocyte carcinoma using optical attenuation coefficients. npj Digit. Med. 8, 232 (2025). https://doi.org/10.1038/s41746-025-01634-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01634-x