Abstract
The widespread adoption of real-world data has given rise to numerous healthcare-distributed research networks, but multi-site analyses still face administrative burdens and data privacy challenges. In response, we developed a Collaborative One-shot Lossless Algorithm for Generalized Linear Mixed Models (COLA-GLMM), the first-ever algorithm that achieves both lossless and one-shot properties. COLA-GLMM ensures accuracy against the gold standard of pooled data while requiring only summary statistics and completes within a single communication round, eliminating the usual back-and-forth overhead. We further introduced an enhanced version that employs homomorphic encryption to reduce the risks of summary statistics misuse at the coordinating center. The simulation studies showed near-exact agreement with the gold standard in parameter estimation, with relative differences of 7.8 × 10−6%–3.0% under various cell suppression settings. We also validated COLA‑GLMM on eight international decentralized databases to identify risk factors for COVID‑19 mortality. Together, these results show that COLA‑GLMM enables accurate, low‑burden, and privacy-preserving multi‑site research.
Similar content being viewed by others
Introduction
Largely driven by the widespread adoption of EHRs in U.S. healthcare settings1, real-world data (RWD) have yielded more robust and generalizable findings, including those that would be impossible to develop with confidence using smaller datasets. These approaches have been facilitated by the development of distributed research networks (DRNs), where “distributed” refers to settings in which patient-level observational clinical data are collected and maintained across multiple sites. DRNs can be broadly classified into two categories: centralized and decentralized. In centralized DRNs, patient-level data from participating sites can be shared across the network and are typically stored in a central repository accessible to authorized researchers within the network. In contrast, decentralized DRNs operate under stricter data-sharing constraints, where patient-level data remain at the local sites and cannot be shared, even with other participants within the same network.
Notable examples of centralized research networks include ~60% of the networks within the National Patient-Centered Clinical Research Network (PCORnet)2,3, which is a network of networks, such as PEDSnet4, OneFlorida+5, INSIGHT6, etc. Beyond the scope of PCORnet, there are additional centralized networks such as IBM Watson Health7, the All of Us Consortium8, Flatiron Database9, Optum10,11, UK biobank12, and eMERGE13. In recent years, particularly within the context of the COVID-19 pandemic, centralized initiatives like the National COVID Cohort Collaboration (N3C)14,15 and the RECOVER initiative16 have been launched. In contrast, about 40% of PCORnet’s networks adopt a decentralized research infrastructure, featuring networks such as the PaTH17 and STAR18 networks, as well as the Greater Plains Collaborative19. Other decentralized initiatives aimed at facilitating international studies include The Consortium for Clinical Characterization of COVID-19 by Electronic Health Records (4CE)20,21, a key player in COVID-19 research, and the Observational Health Data Sciences and Informatics (OHDSI) community22, a key stakeholder in the European Health Data and Evidence Network23.
Despite the conceptual appeal of these DRNs, their use remains logistically challenging because sharing patient-level data across clinical sites typically involves legal agreements, secure file transfers, and repeated back-and-forth communication, each requiring dedicated advocates at each institution. Centralized DRNs have invested extensive effort to streamline these processes. Nonetheless, the administrative burden remains a large and costly process, adding delay and drag.
To address the challenge in data sharing, the development and adoption of federated learning algorithms enable the conduct of multi-site studies with larger sample sizes without requiring patient-level data sharing. These algorithms also enhance the power and provide more generalizable clinical evidence. The federated learning process is particularly beneficial for decentralized DRNs, especially those involving international collaborators, where sharing patient-level data among hospitals is rarely possible due to privacy concerns. Ideally, federated learning algorithms should be designed to achieve both lossless and one-shot properties. The “lossless” property ensures that results from the federated algorithms align with those obtained in the ideal setting where patient-level data are pooled together for analysis, also known as pooled analysis, which is considered the gold standard. The “one-shot” property refers to achieving results in a single communication round, eliminating the typical back-and-forth data sharing required across collaborating sites and thereby streamlining the process significantly. To date, only a few algorithms have successfully achieved both lossless and one-shot properties simultaneously, notably linear regression24 and linear mixed models (i.e., DLMM25).
Another significant and practical consideration in multi-site analysis is the potential existence of between-site patient heterogeneity. Different sites often attract patients varying considerably in illness severity, comorbidities, social circumstances, and healthcare needs. This between-site heterogeneity creates confounding bias unless recognized and accounted for. The federated learning algorithm named DLMM deploys a linear mixed-effects model to account for the site-level heterogeneity and has been successfully implemented in real-world settings for continuous outcomes within linear models25.
Several federated learning algorithms have been developed for fitting Generalized Linear Mixed Models (GLMMs)26,27,28. For example, Li et al. proposed an iterative algorithm for fitting GLMM in a multi-site setting. While this approach avoids sharing patient-level data, it requires dozens or even hundreds of rounds of parameter updates to achieve model convergence27. To reduce communication overhead, Luo et al. introduced the dPQL method, which completes GLMM fitting in fewer than five rounds of communication by transferring only summary-level statistics. This represents a substantial improvement in communication efficiency and enhances the method’s practicality for real-world applications28. Yan et al. developed Fed-GLMM based on a quadratic surrogate likelihood approach, which approximates the global likelihood function. However, to achieve the accuracy of pooled analysis, which is the gold standard, multiple rounds of communication are still recommended26,29. These methods have successfully extended federated learning to various outcome types, including binary, categorical, count, and time-to-event outcomes, by employing non-linear estimating equations common in regression models. Nonetheless, the need for iterative communication between data partners and the coordinating center can introduce delays and necessitate additional computational infrastructure, which may limit scalability in practical settings. While DLMM and these iterative federated learning algorithms for GLMM possess a lossless nature, no federated algorithm for GLMM has yet been developed that combines both lossless and one-shot properties.
To address the methodological gap in existing federated learning algorithms for GLMM that lack the possession of both lossless and one-shot properties, we introduce a novel federated learning algorithm designed to meet the following criteria: (1) requires only summary statistics instead of patient-level data; (2) accounts for between-site heterogeneity at both patient-level and site-level; (3) maintains the lossless property; and (4) achieves results in a single round of communication. Specifically, we propose the Collaborative One-shot Lossless Algorithm for GLMM (COLA-GLMM), the first-ever algorithm that achieves both lossless and one-shot properties, developed to meet the practical demands of data privacy and efficiency. To accommodate various operational environments, we offer two versions of COLA-GLMM. The first is tailored for a “trusted environment,” where both the coordinating center and data contributors rigorously follow the specified analysis protocol. In this setting, the coordinating center will never attempt to infer sensitive information from the summary statistics collected from the data partners. The second version, known as HE-enhanced COLA-GLMM, incorporates homomorphic encryption (HE)30 within a “semi-trusted environment”30. In this setting, while both data contributors and the coordinating center adhere to the protocol without engaging in malicious actions, the coordinating center may still attempt to derive insights from passively obtained data, indicating a curiosity in extracting information. To further minimize the safety concern of summary statistics at the coordinating center, we adopted full HE to protect the summary statistics contributed from the data partners. The adoption of fully HE to our COLA-GLMM framework preserves the privacy of summary statistics while still maintaining the accuracy of the model fitting and estimation. Although HE introduces an additional round of communication and extends computational time, the total process is confined to just two rounds. The encryption and decryption tasks are performed locally within each data partner’s system, adding only a few hours to the overall computation time in generating the summary statistics. This is both reasonable and manageable in real-world settings, balancing the need for data privacy with the practicality of model implementation.
The key contributions of the proposed COLA-GLMM include: (1) Patient-level information protection: COLA-GLMM enables the model fitting across sites without sharing patient-level data; users may optionally adopt HE under either a fully trusted or semi-trusted environment to meet practical security requirements. (2) Communication efficiency: our proposed framework requires only a single round of summary-level statistic exchange, largely reducing the between-site synchronization and human-in-the-loop overhead and making the method scalable and timesaving in real-world implementation. (3) Statistical accuracy: COLA-GLMM delivers a lossless estimator that is mathematically equivalent to the centralized pooled analysis, ensuring no loss in estimation precision or inference validity. (4) Heterogeneity accommodation: our method incorporates site-specific random effects to account for between-site heterogeneity.
To assess the performance and applicability of our proposed COLA-GLMM algorithm, we conducted extensive simulation studies and a truly decentralized real-world use case involving eight data contributors from three countries within the OHDSI network. The simulation results validate the lossless property of the proposed COLA-GLMM algorithm by comparing it with the benchmark method and demonstrate its robust accuracy across different thresholds suppressing data fields where their small size could inadvertently lead to uniquely identifying people, and thus a loss of privacy. We also conducted a real-world study by collaborating with eight data partners from the OHDSI network, focusing on identifying risk factors for COVID-19 mortality among hospitalized patients. In addition, we also examined the temporal consistency of statistically significant risk factors throughout different phases of the COVID-19 pandemic. This study was conducted in a truly decentralized setting, where patient-level data could not be pooled due to international data-sharing restrictions across participating countries. Our proposed algorithm demonstrated strong scalability, effectiveness, efficiency, and accuracy under these real-world constraints, highlighting its practical utility for global, multi-site research.
Results
Proposed COLA-GLMM in a trusted environment
The conventional approach to fitting the GLMM method via the Penalized Quasi-Likelihood (PQL) technique in a federated manner with decentralized data (as shown in the right panel of Fig. 1a), introduced by Luo et al.28, involves an iterative procedure with fewer than five rounds of communication. In this method, the coordinating center communicates interactively with data contributors by collecting summary statistics and updating the intermediate parameters to send back to the data contributors. However, this iterative procedure incurs relatively high communication costs, requires intensive human labor, and extends the time required for practical collaborative research. To address this limitation and facilitate the scalability of implementation of the federated learning algorithms in practice, the R package “pda” and an “over-the-air” online portal called Privacy-preserving Distributed Algorithm Over the Air (PDA-OTA) (http://pda-ota.pdamethods.org/) have been developed31,32,33. The pseudocode of the proposed COLA-GLMM is summarized in the following Table 1. More details on the notations of GLMM, PQL, and PDA-OTA are available in the “Methods” section.
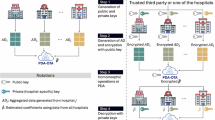
a Multi-site data in centralized distributed research networks and decentralized distributed research networks. b Pipeline of the proposed Collaborative One-shot Lossless Algorithm for GLMM (COLA-GLMM) under a trusted environment. In this scenario, only a single round of communication is needed to transfer summary statistics from the data contributors to the coordination center (i.e., PDA-OTA). c Homomorphic Encryption (HE)-enhanced COLA-GLMM under semi-trusted environment. In this scenario, the nature of the Homomorphic Encryption (HE) encryption-decryption process necessitates two rounds of communication. It is important to note that the addition of HE does not impact the numerical accuracy of the COLA-GLMM method.
However, even with PDA-OTA, the ideal goal for a federated learning algorithm is to achieve the “one-shot” feature, which implies that only a single round of communication is required. In this context, a single round is defined as the coordinating center collecting information from the data contributors just once. The coordinating center is defined as the central server or site responsible for coordinating the study, and it does not contribute any data. Additionally, we aim for the “lossless” feature in our federated learning algorithm, which ensures that the accuracy of parameter estimation remains intact despite data-sharing constraints. This means that the results obtained using the federated learning algorithm with the lossless feature match exactly those of the pooled analysis, where the patient-level data are pooled together.
Here, we propose the COLA-GLMM to fit PQL in a federated manner on the data from a decentralized DRN where the patient-level data are not allowed to be shared across sites, which we refer to this proposed novel method as COLA-GLMM. Initially, we assume a trusted environment where both the coordinating center and data contributors strictly adhere to the predefined study protocol and are sufficiently motivated to follow it faithfully. Such an assumption on the environment of implementing COLA-GLMM could be reasonable because, in practice, the activities of all participating parties would typically be governed by contractual agreements that forbid unauthorized data sharing and tampering. Notably, established collaborative networks like OHDSI34, PCORnet35, and Sentinel36 adhere to similar frameworks, demonstrating the viability of such a cooperative environment. In Fig. 1b, we present the pipeline of the proposed COLA-GLMM operating in a trusted environment.
We assume that all available covariates are categorical and include a standard set of demographic and clinical variables typically found in EHR and claims data. Furthermore, we adhere to the closed world assumption as outlined in the OMOP Common Data Model37,38. Under this assumption, continuous covariates such as age and BMI are categorized or dichotomized based on domain expert guidance, tailored to the specific scientific questions of each project.
In Fig. 2, we present a synthetic summary statistics matrix to be shared among three sites, with two covariates and a binary outcome of interest. The equations and details calculation procedure of the summary statistics are provided in the “Methods” section.
Consider three sites, each with 1000 patients whose records contain two binary covariates, including sex (female) and age >65, and a binary outcome. Because patient‑level data cannot be shared, every site independently and simultaneously generates summary-level statistics based on all four possible combinations of the two binary covariates. Only these summary statistics are transmitted to the coordinating center.
Once the coordinating center collects the summary statistics matrices from all data contributors, it can construct the likelihood function as shown in Eq. (4) in the “Methods” section. This reconstruction enables the estimation of the parameters of interest to obtain the estimated fixed effect and represent the association between the outcome of interest and the covariates, thereby helping to identify the risk factors. Additionally, we can also estimate the random effects, which capture the heterogeneous site-specific effects, allowing for the quantification of between-site heterogeneity and site-specific predictions.
-
Remark 1 (Eliminates need for initial values): An important feature of our proposed COLA-GLMM is that they do not require any initializations conducted within each data contributors. To date, few federated learning algorithms achieve both lossless and one-shot properties without needing initial values, except for linear regression24 and linear mixed models25.
-
Remark 2 (Accounts for heterogeneous patient populations): Healthcare data from different health systems often present inherent heterogeneity due to variations in patient populations and hospital practices arising from various factors such as geographical and operational differences. The proposed COLA-GLMM yields results equivalent to pooled analysis by directly fitting PQL on pooled data and inherently addresses the heterogeneity in covariate distributions by considering a combined overall population.
-
Remark 3 (Adheres to cell size suppression policy): Since the proposed algorithm collects counts for all possible combinations of the covariate vector, differences in population sizes and data availability across sites could result in small cell counts, which pose a risk of re-identification36. To address this, we follow the established policies on the minimum cell size to be reported36,39,40,41, as outlined in data use agreements and enforced by various regulatory agencies. These policies require that cells with non-zero counts below a specified threshold must be masked, collapsed, coarsened, or suppressed to ensure confidentiality and compliance. For example, a threshold of 11 means that counts from 1 to 10 cannot be reported directly. In the COLA-GLMM framework, to comply with these policies, cells with counts smaller than the threshold are replaced with a standardized value of 6 given a threshold of 11. It is important to note that we highly recommend setting a consistent threshold across all data partners to ensure uniformity based on the pre-specified study protocol.
Homomorphic encryption (HE)-enhanced COLA-GLMM in a semi-trusted environment
Though there are research networks that are implementing a trusted environment, in practice, data contributors adhere to the protocol but may not fully trust the coordinating center. While both data contributors and the coordinating center are expected to avoid malicious actions that deviate from the protocol (i.e., they are trusted in operations), concerns remain among data partners regarding the trustworthiness and safety of the coordinating center—the coordinating center might attempt to infer sensitive information from the summary statistics. This scenario is known as a semi-trusted environment, also commonly referred to as honest-but-curious, and has been widely studied and used in many privacy-preserving deep learning techniques42,43,44.
To prevent the coordinating center from deriving information on patient-level data based on the summary statistics, we employ the fully HE algorithm to our COLA-GLMM algorithm, specifically, we deployed the CKKS45 method and propose the HE-enhanced COLA-GLMM. In this HE-enhanced framework, only encrypted summary statistics are transmitted to the coordinating center, PDA-OTA. The semi-trusted PDA-OTA then processes these encrypted statistics to derive results. It’s crucial to note that the use of HE does not compromise the accuracy of the results, ensuring that outcomes using the HE-enhanced COLA-GLMM are identical to those obtained with the proposed method under an honest environment. This approach prevents the coordinating center from ever accessing unencrypted summary statistics, further enhancing the protection of sensitive information within the proposed framework. We refer to Supplementary Table 1 for detailed components of the CKKS algorithm45.
In Fig. 1c, we present the pipeline of the proposed HE-enhanced COLA-GLMM framework. The framework involves four major stages, which are summarized as follows:
Preparation stage
A trusted third party, which could be one of the data contributors or an authorized entity, is selected to generate key pairs. Each key pair consists of a unique private key for each data contributor and a shared public key. These key pairs are distributed to all participating sites through a secure information channel, such as TLS protocols. The public key, generated by the trusted third party, is accessible to all data contributors and the coordinating center. Each contributor’s private key is accessible only to that site and the trusted third party.
Encryption stage
After receiving the key pairs, each participating site encrypts the generated summary statistics using the public key and then sends the encrypted data to the coordinating center.
Aggregation stage
The coordinating center also has access to the public key and uses homomorphic operations on the encrypted summary statistics to generate encrypted results (i.e., estimated parameters). The use of a shared public key across all data contributors and the coordinating center ensures that homomorphic operations on encrypted summary statistics produce results identical to those from operations on unencrypted summary statistics once decrypted.
Decryption stage
The encrypted estimation results are shared with all data contributors. Each site uses its private key to decrypt the results, obtaining the unencrypted outcomes. As each site receives the same encrypted result from the coordinating center, the decrypted results should be consistent across all sites. These results are then returned to the coordinating center for a consistency check.
In a semi-trusted environment, the HE-enhanced version ensures privacy by keeping decryption keys away from the coordinating center, which only accesses the public key for homomorphic operations. This setup requires two communication rounds involving the coordinating center: (1) receiving and processing encrypted summary statistics, and (2) receiving decrypted results for consistency checks. In contrast, the COLA-GLMM under an honest environment involves just one communication round where the coordinating center collects and aggregates summary statistics to obtain results. Despite necessitating an additional round of communication, the HE-enhanced COLA-GLMM framework effectively mitigates risks of inadvertent or intentional data misuse at the level of the coordinating center. Moreover, the one-shot nature of the COLA-GLMM algorithm ensures that the extra communication burden is minimal, enhancing its scalability across numerous data partners.
Simulation studies and data evaluation
To evaluate the performance and accuracy of the proposed algorithm, we conducted simulation studies to compare it with the benchmark pooled analysis, which involves aggregating patient-level data from all contributors. Given that the HE-enhanced version achieves identical accuracy to the version under an honest environment, our simulations exclusively compared the COLA-GLMM in an honest environment to this benchmark.
The synthetic data were simulated based on summary statistics shared by data contributors from the real-world application. Further details on the data and study cohort are available in the subsequent section. We utilized a logistic regression model, distributing data across eight sites with random sample sizes ranging from 500 to 50,000. We generated nine binary risk factors with prevalence rates varying from 10% to 60% and modeled the binary response variable by adjusting coefficients from −0.4 to 0.5. The results from the proposed algorithm were benchmarked against the gold standard estimates. Additionally, to assess the impact of different cell suppression policies on data sharing, we reported results from the proposed algorithm using cell sizes adjusted to 3 for groups ranging from 1 to 5, and to 6 for groups ranging from 1 to 11, following the CMS cell suppression policy46.
Figure 3 displays a Bland-Altman plot comparing the COLA-GLMM method to the pooled analysis. The y-axis shows the differences in estimated fixed effects on a log (odds ratio, OR) scale, while the x-axis presents the average estimated effect sizes for both methods. Points closer to the horizontal line at zero indicate greater accuracy of the COLA-GLMM relative to the pooled analysis. Two covariates are circled to emphasize their proximity in values, showcasing the precision of the proposed method. We have observed a few key findings from the simulation results:
The subfigures are organized as follows: no cell suppression (left), threshold = 5 (middle), and threshold = 11 (right). Each subfigure includes nine solid dots representing nine simulated covariates. The y-axis shows the differences in the estimated fixed effects (on a log(odds ratio) scale) between the pooled analysis and the COLA-GLMM method, while the x-axis displays the average estimated effect sizes for both methods. Two covariates are highlighted with circles to demonstrate the proximity of the values produced by the two methods.
Lossless property
In the absence of cell suppression requirements, the proposed method replicates the point estimates and variances from the pooled analysis, affirming its lossless nature in comparison to the benchmark method.
Robustness to small cell suppression threshold
Adjusting the cell suppression threshold has minimal impact on estimation accuracy, underscoring the robustness of our method under various data privacy settings.
More detailed simulation results and a comprehensive comparison between the pooled analysis and the proposed COLA-GLMM can be found in Supplementary Note 3.
To demonstrate the applicability of the proposed algorithm in a decentralized DRN (i.e., where the patient-level data are not allowed to be shared across sites), we collaborated with eight data contributors from the OHDSI network. We are interested in identifying the risk factors of COVID-19 mortality among hospitalized patients and examining the temporal consistency of these risk factors across three time periods (i.e., pre-Delta, Delta, and Omicron periods or waves) during the COVID-19 pandemic. The data contributors include:
-
Optum® de-identified Electronic Health Record Dataset (Optum® EHR);
-
Optum’s de-identified Clinformatics® Data Mart Database (Clinformatics®);
-
IQVIA Hospital CDM;
-
University of Florida Health;
-
Department of Veterans Affairs;
-
Integrated Primary Care Information (IPCI), The Netherlands;
-
Columbia University Irving Medical Center (CUIMC);
-
Parc Salut Mar Barcelona (PSMAR), Spain.
The pre-Delta period spans from November 2020 to February 2021. The Delta period, when the Delta variant was predominant, ranges from July 2021 to October 2021. Subsequently, the Omicron period, characterized by the predominance of the Omicron variant, extends from November 2021 to March 2022. Our study cohort included patients aged 18 years and older who had an inpatient visit with either a diagnosis of COVID-19 or a positive test for COVID-19 between 21 days prior to the inpatient visit and the end of the inpatient visit, with the visit starting as the index date. Patients were excluded if they had been observed in the database for fewer than 180 days prior to the index date (i.e., date of hospitalization). The primary clinical outcome was patient death during or up to 7 days after the inpatient visit. The covariates included age (categorized as <65, 65–80, and ≥80), sex, and medical characteristics, including a history of chronic obstructive pulmonary disease (COPD), diabetes, hypertension, kidney disease, and obesity. The Charlson Comorbidity Index (CCI) score was also included as a measure of the patient’s overall health state, with higher scores indicating worse health conditions. We dichotomized the CCI score at a threshold of 5. The summary table of the cohort’s characteristics is available in Supplementary Note 2.
To implement the framework with all collaborators, we utilized the PDA-OTA website, which enables synchronization of project information and status. This platform served as the coordinating center for the collaborating sites to upload and manage the summary statistics. In this study, we adopted a streamlined communication process requiring only a single round of communication. Each participating site had two main responsibilities: first, downloading the study protocol file from the PDA-OTA platform, where the details of the study—such as covariates, outcomes, and the statistical model—were specified; and second, executing an R study package to generate the summary statistics using their locally stored patient-level data, followed by uploading the summary statistics to the PDA-OTA platform. To facilitate transparency and reproducibility, we have made the code for cohort definition, data extraction, and aggregated data generation openly accessible on the GitHub repository through the link https://github.com/ohdsi-studies/OLGLMM-COVID (Note: the initial name of COLA-GLMM was OLGLMM).
Figure 4 reports estimated ORs and 95% confidence intervals for identifying COVID-19 mortality risk factors among hospitalized patients using eight decentralized databases using our proposed algorithm. Based on the figure presented, several risk factors have been consistently identified as significant across three study time periods, including:
-
Age: Being aged 80 and above consistently showed the highest ORs among all risk factors across the three periods, indicating a significantly increased risk of mortality. This risk is also notable in the age group of 65–80, though less pronounced than in the older age group.
-
CCI: Higher CCI scores are statistically associated with an increased risk of mortality. This association remains consistent across all periods, with a notably stronger correlation during the Delta period.
-
Sex (female): There is evidence showing that female patients consistently exhibit a lower risk of mortality compared to males across all periods, though the difference is relatively modest.
In accordance with the OHDSI cell suppression policy, all cells containing values less than 11 were imputed as 6 when sharing aggregated data across databases.
These findings align with several published studies on risk factor identification47,48,49,50. Additionally, we observed that a history of obesity, diabetes, and COPD emerged as statistically significant factors in one or two study periods for the cohort. The data suggest that while certain conditions such as age and sex consistently represent significant risk factors for COVID-19 mortality, the impact of other factors like obesity varies depending on the COVID-19 variant. This variability indicates that the interactions between COVID-19 and patient comorbidities may evolve over time as the virus mutates, necessitating further investigation into the temporal changes of these risk factors over an extended period.
Discussion
In this paper, we introduced the COLA-GLMM algorithm, a novel approach designed for multi-site studies utilizing decentralized RWD. Specifically, the proposed algorithm has a few advantages. First, it possesses both lossless and one-shot properties, ensuring that the aggregated results are equivalent to those obtained from a pooled analysis within a single round of communication. These two properties fulfill the ideal requirements for federated learning algorithms. Second, the one-shot property enhances the scalability of the proposed method. By requiring minimal communication rounds, our algorithm is well-suited for sensitive and large-scale studies with numerous data partners. Its efficiency and minimal overhead encourage adoption in diverse research settings, expanding the possibilities of multi-site collaborative research. Third, the proposed algorithm also accounts for the between-site heterogeneity by modeling the site-specific random effects through GLMM. Additionally, regarding privacy considerations, the proposed algorithm ensures a privacy-preserving implementation by only requiring summary statistics, ensuring that patient-level data never leaves its original site. Last but not least, the additional layer of security introduced by the HE-enhanced version addresses potential vulnerabilities in aggregated data safety at the coordinating center, particularly in semi-trusted environments. HE introduces additional computational and communication overhead, which is an expected trade-off for enhanced data protection. However, the extra computations involved, including key generation, encryption, and decryption, typically require only a few seconds on a modern CPU. The size of the keys and encrypted data to be transferred is on the order of several hundred megabytes, which remains well within the capabilities of standard computing systems. For example, encrypting the summary-level statistics for a model with eight binary variables takes only a few seconds on a system equipped with an AMD Ryzen 9 7900X (4.70 GHz, 12-core processor) and 64 GB of RAM.
The proposed method is not only applicable to decentralized settings but also enhances centralized networks where more comprehensive data integration is necessary. For centralized networks, this federated tool would be particularly useful in studying rare diseases and other significant topics where extensive data from external collaborators outside the network are required to enhance research outcomes. It is important to note that the proposed COLA-GLMM framework is highly flexible and broadly applicable beyond the context of COVID-19. Its design makes it well-suited for multi-site studies across a wide range of disease areas and research domains where individual-level data cannot be shared. For example, the proposed method can be applied to conduct comparative effectiveness research in cardiovascular disease, diabetes, cancer, mental health, Alzheimer’s disease and related dementias, opioid use disorder, and other infectious diseases.
Although the proposed algorithm has achieved two optimal properties within federated learning, there is still room for further enhancement in the methods used for fitting GLMM. This area remains a key focus for future work. First, the use of PQL in our study presents certain limitations, particularly in underestimating the variance components, which could impact the reliability of the results in complex models. Additionally, the PQL method relies on parametric assumptions about the distribution of random effects, typically assuming a Gaussian distribution. This assumption may not always hold true in practical scenarios, potentially leading to inaccuracies. Future work could focus on developing more robust methods that relax these Gaussian assumptions and improve the accuracy of the variance estimation. Exploring alternative approximations or leveraging non-parametric techniques might provide more flexibility and accuracy, enhancing the applicability of the model across diverse datasets where the underlying distributions of random effects are unknown or highly irregular.
We also acknowledge that other forms of heterogeneity often arise in real-world observational clinical data, particularly in multi-site analyses. While the proposed COLA-GLMM addresses heterogeneity related to covariate distributions and outcome–covariate relationships, other complexities, such as differences in measurement practices, patterns of missing data, variability in data quality, and site-specific coding systems, may still pose significant challenges for federated learning analyses. In future work, we plan to extend COLA-GLMM to manage structured missingness and time-varying data quality. Addressing these additional dimensions of heterogeneity will be crucial for further improving the robustness and generalizability of multi-site clinical research.
In summary, we propose COLA-GLMM, a COLA-GLMM. To our knowledge, this is the first federated learning algorithm for GLMMs that is both lossless and one-shot, offering a unique combination of statistical rigor and communication efficiency. COLA-GLMM achieves estimation accuracy equivalent to that of a pooled patient-level analysis while delivering two key advantages: (1) it requires only summary-level statistics, and (2) it needs only a single round of communication across sites for transferring statistics, significantly reducing synchronization and administrative burden across sites. To further enhance data protection, we introduce a HE-augmented version of COLA-GLMM, which mitigates the risk of misuse of shared statistics at the coordinating center. We believe this framework provides a scalable, secure, and practical solution for multi-site analysis of complex hierarchical data and has broad applicability across clinical and public health research domains.
Methods
Notations of Generalized Linear Mixed Model (GLMM) and the Penalized Quasi-likelihood (PQL) method
GLMM is an extension of the generalized linear model with additional random effects, which model the between-site heterogeneity. Assume there are \(K\) hospitals in total within a network, the kth site has number of patients \({n}_{k}\) and the total number of patients within such a network is \(N={\sum }_{k}{n}_{k}\). For subject \(i\) at hospital \(k\), we denote \({y}_{{ki}}\) the outcome (e.g., COVID-19 mortality status which is a binary outcome), \({{\boldsymbol{x}}}_{{ki}}\) the p-dimensional covariates (e.g., age, sex, race, etc) with fixed effects β, and \({b}_{k}\) the random effect, \(k=1,\ldots ,K,i=1,\ldots ,{n}_{k}\). Conditional on the covariates \({{\boldsymbol{X}}}_{k}={\left({{\boldsymbol{x}}}_{k1},\ldots ,{{\boldsymbol{x}}}_{k{n}_{k}}\right)}^{T}\) and random effects \({b}_{k}\), the outcome \({y}_{k}={\left({y}_{k1},\ldots ,{y}_{k{n}_{k}}\right)}^{T}\) are assumed to be independent observations with means and variances specified by a GLMM as follows:
where \(g(\cdot)\) is the link function that connects the conditional means \({\mu }_{{ki}}\) to the linear predictor \({\eta }_{{ki}}\), and \(v(\cdot )\) is the variance function. The random effects \({b}_{k}\) are assumed to follow a normal distribution with mean 0 and variance θ \(({\rm{i}}.{\rm{e}}.,{b}_{k}\sim N(0,\theta ))\). With the aim of risk factor identification, the parameter of interest in this GLMM model is β, which is the fixed effect associated with the covariates.
Consider a centralized network (as shown in Fig. 1a, left panel) where patient-level data from various contributors are aggregated into a central database or data warehouse. To fit GLMM on this multi-site data, the standard procedure for estimating the GLMM parameters (β, θ) is through maximizing the integrated quasi-likelihood function, which is written as
where \({d}_{{ki}}(y,\mu )=-2{\int }_{y}^{\mu }(y-u)/\nu (u){du}\). However, the numerical integration techniques required for calculating the likelihood function become exceedingly complex when these are irreducibly high-dimensional integrals. Therefore, the PQL method was proposed as an approximate approach for estimating parameters in GLMM51. The PQL method has proven its suitability for practical applications across various fields52,53,54.
Specifically, by applying Laplace’s method for integral approximation, the PQL method leads to iteratively fitting a linear mixed model (LMM). The log-likelihood of LMM with all patient-level data from \(K\) sites can be written as:
where \({X}_{k}\) is the covariate matrix, \({Y}_{k}^{* }\) is the working outcome vector, |·| is the matrix determinant, and \({\Sigma }_{k}={\Sigma }_{k}(\theta )=\theta {{\bf{1}}}_{{n}_{k}}{{\bf{1}}}_{{n}_{k}}^{T}+{W}_{k}^{-1},{W}_{k}={\rm{diag}}\left\{v\left({\hat{{\boldsymbol{\mu }}}}_{{\boldsymbol{k}}}\right)\right\}\). In the scenario where we are interested in analyzing the data from a centralized DRN, the above Eq. (4) is then fitted on the pooled data to obtain \((\hat{{\boldsymbol{\beta }}},\hat{\theta })\). We also refer to such analysis directly using the PQL method on the pooled dataset as the “pooled analysis.” This analysis serves as the benchmark (i.e., gold standard) against which we compare the subsequent implementation and empirical evaluation of the proposed federated learning algorithm.
Summary statistics required by COLA-GLMM
Let \(\chi ={\left\{\mathrm{0,1}\right\}}^{p}\) denote the set of all possible combinations of a binary covariate vector x, where x is a p-dimensional vector to represent the patient-level covariates. The size of this set is denoted as \(q={2}^{p}\)(\({\rm{i}}.{\rm{e}}.,|\chi |=q={2}^{p}.)\), which means that the total number of possible unique combinations of the elements in this vector x is q. For example, if the covariate vector x = (female (0 or 1), age > 65 (0 or 1)), then \(p=2\) given there are two covariates and \(q={2}^{p=2}=4\) as the possible combinations of x include \(\left\{\mathrm{0,0}\right\},\left\{\mathrm{0,1}\right\},\{\mathrm{1,0}\}\) and \(\{\mathrm{1,1}\}\). It is important to note that both values (q and p) are consistent across all data contributors, as we assume that the covariates are available for each participating entity.
Let \({{\boldsymbol{x}}}^{(j)}\in {\left\{\mathrm{0,1}\right\}}^{p}\) denote the jth unique individual combination \({\boldsymbol{x}}\), where \(j=\{1,\ldots ,q\}\). All possible combinations can be represented as the \({{\boldsymbol{x}}}^{(1)},{{\boldsymbol{x}}}^{\left(2\right)},\ldots ,{{\boldsymbol{x}}}^{(q)}\). When implementing COLA-GLMM, within a single round of communication, the summary statistics collected from each data contributor consist of a matrix with dimension \(q\times (p+2+p)\). The first p columns are all the possible combinations, i.e., \({{\boldsymbol{x}}}^{(1)},{{\boldsymbol{x}}}^{\left(2\right)},\ldots ,{{\boldsymbol{x}}}^{(q)}\). The rest additional columns for the kth site include:
-
1.
A q-dimensional vector \({{\boldsymbol{C}}}_{k}=\{{c}_{k1},\ldots ,{c}_{{kq}}\}\), where \({c}_{{kj}}=\mathop{\sum }\limits_{i=1}^{{n}_{k}}I\left({{\boldsymbol{x}}}_{{ki}}={{\boldsymbol{x}}}^{({\rm{j}})}\right)\), counting the number of patients for each combination of x.
-
2.
A q-dimensional vector \({{\boldsymbol{S}}}_{k}=\{{s}_{k1},\ldots ,{s}_{{kq}}\}\), where \({s}_{{kj}}=\mathop{\sum }\limits_{i=1}^{{n}_{k}}{y}_{{ki}}I\left({{\boldsymbol{x}}}_{{ki}}={{\boldsymbol{x}}}^{({\rm{j}})}\right)\), representing the sum of observed outcome values \({y}_{{ki}}\) for patients who have a corresponding combination of x.
-
3.
A q×\(p\) dimensional matrix \({{\boldsymbol{U}}}_{k}=\{{{\boldsymbol{u}}}_{k1}^{T},\ldots ,{{\boldsymbol{u}}}_{{kq}}^{T}\}\), where \({{\boldsymbol{u}}}_{{kj}}=\mathop{\sum }\limits_{i=1}^{{n}_{k}}{{\boldsymbol{x}}}_{{\boldsymbol{ki}}}^{T}{y}_{{ki}}I\left({{\boldsymbol{x}}}_{{ki}}={{\boldsymbol{x}}}^{({\rm{j}})}\right)\), representing the sum of \({{\boldsymbol{x}}}_{{\boldsymbol{ki}}}^{T}{y}_{{ki}}\) for patients who have the corresponding combination of x.
The summary-level statistics \({C}_{k},{S}_{k}\), and \({U}_{k}\) are transferred to the coordinating center from the collaborating sites. These statistics are then used to reconstruct and fit the LMM specified in Eq. (4), thereby enabling the estimation of fixed effects and the identification of risk factors associated with the outcome of interest.
PDA-OTA
The PDA-OTA is an advanced web-based software program designed to facilitate collaborative studies while adhering to the principles of implementing algorithms within the PDA framework31,32,33. It ensures secure, private sharing of summary statistics for multi-site studies, facilitating both national and international collaborations. The platform’s user-centric design supports project leads and data partners by syncing project statuses, providing cloud-based SFTP for data transfer, and tailoring tasks for each model. Users can invite sites, upload data, monitor progress, receive updates, and produce project summaries efficiently. PDA-OTA acts as a central hub, coordinating and disseminating information across contributors, as illustrated in Fig. 1b, c.
Data availability
No datasets were generated or analysed during the current study.
Code availability
The R code is available at https://github.com/Penncil/pda and through our R package: “pda” at https://github.com/Penncil/pda. The code of COVID-19 study within the OHDSI network is available at https://github.com/ohdsi-studies/OLGLMM-COVID.
References
Friedman, C. P., Wong, A. K. & Blumenthal, D. Achieving a nationwide learning health system. Sci. Transl. Med. 2, 57cm29 (2010).
Collins, F. S., Hudson, K. L., Briggs, J. P. & Lauer, M. S. PCORnet: turning a dream into reality. J. Am. Med. Inform. Assoc. 21, 576–577 (2014).
Forrest, C. B. et al. PCORnet® 2020: current state, accomplishments, and future directions. J. Clin. Epidemiol. 129, 60–67 (2021).
Forrest, C. B. et al. PEDSnet: a national pediatric learning health system. J. Am. Med. Inform. Assoc. 21, 602–606 (2014).
Shenkman, E. et al. OneFlorida Clinical Research Consortium: Linking a clinical and translational science institute with a community-based distributive medical education model. Acad. Med. 93, 451–455 (2018).
INSIGHT Clinical Research Network. https://insightcrn.org/.
Healthcare. IBM. https://www.ibm.com/industries/healthcare.
All of Us Research Program Investigators. The “All of Us” research program. N. Engl. J. Med. 381, 668–876 (2019).
Flatiron Health. Reimagining the infrastructure of cancer care. https://flatiron.com/.
Claims Data. Optum. https://www.optum.com/en/business/life-sciences/real-world-data/claims-data.html.
Electronic Health Records (EHR) Data. Optum. https://www.optum.com/en/business/life-sciences/real-world-data/ehr-data.html.
UK Biobank—UK Biobank. https://www.ukbiobank.ac.uk/.
Electronic Medical Records and Genomics (eMERGE) Network. https://www.genome.gov/Funded-Programs-Projects/Electronic-Medical-Records-and-Genomics-Network-eMERGE.
Haendel, M. A. et al. The National COVID Cohort Collaborative (N3C): rationale, design, infrastructure, and deployment. J. Am. Med. Inform. Assoc. 28, 427–443 (2021).
National COVID Cohort Collaborative. National Center for Advancing Translational Sciences. https://ncats.nih.gov/research/research-activities/n3c.
About the Initiative. RECOVER COVID. https://recovercovid.org/.
PaTH Network. https://pathnetwork.org/.
STAR Clinical Research Network. https://starcrn.org/.
GPC—Greater Plains Collaborative. https://gpcnetwork.org/.
Brat, G. A. et al. International electronic health record-derived COVID-19 clinical course profiles: the 4CE consortium. npj Digit. Med. 3, 1–9 (2020).
Hripcsak, G. et al. Characterizing treatment pathways at scale using the OHDSI network. Proc. Natl Acad. Sci. USA 113, 7329–7336 (2016).
Voss, E. A. et al. European Health Data & Evidence Network—learnings from building out a standardized international health data network. J. Am. Med. Inform. Assoc. 31, 209–219 (2023).
Chen, Yixin et al. Regression cubes with lossless compression and aggregation. IEEE Trans. Knowl. Data Eng. 18, 1585–1599 (2006).
Luo, C. et al. DLMM as a lossless one-shot algorithm for collaborative multi-site distributed linear mixed models. Nat. Commun. 13, 1–10 (2022).
Yan Z, Zachrison KS, Schwamm LH, Estrada JJ, Duan R. A privacy-preserving and computation-efficient federated algorithm for generalized linear mixed models to analyze correlated electronic health records data. PloS one. 18, e0280192 (2023).
Li, W. et al. Federated learning algorithms for generalized mixed-effects model (GLMM) on horizontally partitioned data from distributed sources. BMC Med. Inform. Decis. Mak. 22, 1–12 (2022).
Luo, C. et al. dPQL: a lossless distributed algorithm for generalized linear mixed model with application to privacy-preserving hospital profiling. J. Am. Med. Inform. Assoc. 29, 1366–1371 (2022).
Jordan MI, Lee JD, Yang Y. Communication-efficient distributed statistical inference. J. Am. Stat. Assoc. (2019).
Hazay, C. & Lindell, Y. Semi-honest adversaries. Inf. Secur. Cryptogr. 15, 53–80 (2010).
PDA-OTA. https://pda-ota.pdamethods.org/login.
PDA website. https://pdamethods.org/.
CRAN—Package pda. https://cran.r-project.org/web/packages/pda/index.html.
Hripcsak, G. et al. Observational Health Data Sciences and Informatics (OHDSI): opportunities for observational researchers. Stud. Health Technol. Inf. 216, 574–578 (2015).
Fleurence, R. L. et al. Launching PCORnet, a national patient-centered clinical research network. J. Am. Med. Inf. Assoc. 21, 578–582 (2014).
Xia, W. et al. Managing re-identification risks while providing access to the All of Us research program. J. Am. Med. Inf. Assoc. 30, 907–914 (2023).
Reiter, R. On closed world data bases. in Readings in Artificial Intelligence 119–140. https://doi.org/10.1016/B978-0-934613-03-3.50014-3 (1981).
Reich, C. et al. OHDSI Standardized Vocabularies—a large-scale centralized reference ontology for international data harmonization. J. Am. Med. Inform. Assoc. 31, 583–590 (2024).
CMS Cell Size Suppression Policy. ResDAC. https://resdac.org/articles/cms-cell-size-suppression-policy.
Wasserman C, Ossiander E. Department of Health Agency Standards for Reporting Data with Small Numbers. Accessed July 08, 2025. https://www.doh.wa.gov/Portals/1/Documents/1500/SmallNumbers.pdf.
Utah Department of Health Data Suppression/Data Aggregation Guidelines Summary. Accessed July 08, 2025. https://uofuhealth.utah.edu/documents/data-suppression-summary.
Shokri, R. & Shmatikov, V. Privacy-preserving deep learning. In Proc. 2015 53rd Annual Allerton Conference on Communication, Control, and Computing (Allerton) 909–910. https://doi.org/10.1109/ALLERTON.2015.7447103 (IEEE, 2015)
Phong, L. T., Aono, Y., Hayashi, T., Wang, L. & Moriai, S. Privacy-preserving deep learning via additively homomorphic encryption. IEEE Trans. Inf. Forensics Secur. 13, 1333–1345 (2018).
Zhang, C. et al. BatchCrypt: efficient homomorphic encryption for cross-silo federated learning. In Proc. 2020 USENIX Conference on Usenix Annual Technical Conference (USENIX Association, 2020).
Cheon, J. H., Kim, A., Kim, M. & Song, Y. Homomorphic encryption for arithmetic of approximate numbers. In Proc. Advances in Cryptology–ASIACRYPT 2017: 23rd International Conference on the Theory and Applications of Cryptology and Information Security, Hong Kong, China, December 3-7, 2017, Part I 409–437 (Springer, 2017).
CMS Cell Size Suppression Policy. https://resdac.org/articles/cms-cell-size-suppression-policy.
Noor, F. M. & Islam, M. M. Prevalence and associated risk factors of mortality among COVID-19 patients: a meta-analysis. J. Community Health 45, 1270–1282 (2020).
Jawad Hashim, M., Alsuwaidi, A. R. & Khan, G. Population risk factors for COVID-19 mortality in 93 countries. J. Epidemiol. Glob. Health 10, 204–208 (2020).
Albitar, O., Ballouze, R., Ooi, J. P. & Ghadzi, S. M. S. Risk factors for mortality among COVID-19 patients. Diab. Res. Clin. Pract. 166, 108293 (2020).
Comoglu, S. & Kant, A. Does the Charlson Comorbidity Index help predict the risk of death in COVID-19 patients? North Clin. Istanb. 9, 117 (2022).
Breslow, N. E. & Clayton, D. G. Approximate inference in generalized linear mixed models. J. Am. Stat. Assoc. 88, 9–25 (1993).
Wilmut, I., Schnieke, A. E., McWhir, J., Kind, A. J. & Campbell, K. H. S. Viable offspring derived from fetal and adult mammalian cells. Nature 385, 810–813 (1997). 1997 385:6619.
Sampson, R. J., Raudenbush, S. W. & Earls, F. Neighborhoods and violent crime: a multilevel study of collective efficacy. Science 277, 918–924 (1997).
Zhou, W. et al. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341 (2018).
Acknowledgements
This work was supported in part by National Institutes of Health (U01TR003709, U24MH136069, RF1AG077820, 1R01LM014344, 1R01AG077820, R01LM012607, R01AI130460, R01AG073435, R56AG074604, R01LM013519, R01DK128237, R56AG069880, R21AI167418, R21EY034179). This work was supported partially through the Patient-Centered Outcomes Research Institute (PCORI) Project Program Awards (ME-2019C3-18315 and ME-2018C3-14899). All statements in this report, including its findings and conclusions, are solely those of the authors and do not necessarily represent the views of the Patient-Centered Outcomes Research Institute (PCORI), its Board of Governors, or the Methodology Committee. All statements in this report, including its findings and conclusions, are solely those of the authors and do not necessarily represent the views of the Patient-Centered Outcomes Research Institute (PCORI), its Board of Governors, or the Methodology Committee. This study was also partially funded through the US Department of Veterans Affairs under the research priority to “Put VA Data to Work for Veterans” (VA ORD 22-D4V). The funders had no role in the design and conduct of the protocol; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
J.T. and Y.C. designed methods and experiments; G.T. and J.Z. contributed to the HE-enhanced version development. J.R., J.M.R., M.B., S.D., T.F., A.M.F., X.H., M.E.M., M.A.M., B.K.P., K.R.S., M.A.S., B.V., R.D.W., M.Z., and J.B. had the datasets for data analysis and generated the aggregated data; J.T. and Y.C. served as the coordinating center; all authors mentioned above, C.L., Y.L., L.L., F.W., and D.A.A. interpreted the results and provided instructive comments; J.T. and Y.C. drafted the main manuscript. All coauthors provided critical edits of the early draft and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tong, J., Reps, J.M., Luo, C. et al. Unlocking efficiency in real-world collaborative studies: a multi-site international study with one-shot lossless GLMM algorithm. npj Digit. Med. 8, 457 (2025). https://doi.org/10.1038/s41746-025-01846-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01846-1