Abstract
Accurate ovarian cancer screening and diagnosis are critical for patient survival. We present UMORSS, an AI-assisted diagnostic system integrating ultrasound (US) imaging and clinical data with uncertainty quantification for precise ovarian cancer risk assessment. Developed and evaluated using a multicentre dataset (7352 patients, 7594 lesions, 9281 US images), UMORSS employs a two-phase approach: Phase I rapidly triages low-risk lesions via initial US analysis, and Phase II provides uncertainty-aware multimodal analysis for complex cases. Phase I accurately identified 68.7% of physiological cysts and 13.8% of benign tumours as low-risk, with zero false negatives, and Phase II achieved an AUC of 0.955 (internal testing) and 0.926 (external validation). Furthermore, a prospective reader study (n = 284 cases, six radiologists) demonstrated that UMORSS as a human-AI collaborative tool increased radiologists’ average AUC by 10.58% and sensitivity by 22.48%. UMORSS shows strong potential to streamline clinical workflow, optimize resource allocation, and standardize ovarian cancer diagnosis.
Similar content being viewed by others
Introduction
Ovarian cancer is associated with high mortality, largely attributable to delayed diagnosis1, underscoring the urgent need for accurate early risk assessment. Optimal diagnosis requires both high sensitivity—critical for timely specialist referral and thus improved patient survival2—and high specificity, essential to avoid unnecessary and potentially harmful interventions for prevalent benign lesions3. Current clinical practice aims to strike this delicate balance by integrating multimodal patient data including clinical findings, laboratory results, and ultrasound (US) imaging.
However, reliably achieving optimal risk stratification is challenging due to the complexity of evaluation procedures and their dependence on clinicians’ varied expertise4,5,6. Although efforts at standardization, such as the Ovarian-Adnexal Reporting and Data System (O-RADS)7,8 for US interpretation, and decision-support tools such as logistic regression models9 and ADNEX10 which integrate multiple clinical parameters, represent valuable advances, they remain limited by reliance on manual assessments that introduce experience-based variations, highlighting an unmet need for tools capable of robustly integrating multimodal data and performing objective, consistent risk stratification.
Artificial intelligence (AI), particularly deep learning, offers a promising approach to these challenges. Its key advantage is its ability to learn complex feature patterns from massive datasets and consistently apply those standards, thereby mitigating the inter-observer variability inherent in manual assessment11. This potential has been demonstrated across medical domains like breast cancer screening and digital pathology, where AI performance can match or exceed that of human experts4,5,12,13,14,15,16,17,18. A primary barrier to clinical translation, however, is the “black box” nature of these models, which introduces potential clinical risks and limits clinical trust19. Initial attempts to address this have focused on visualization techniques, such as heatmaps, to offer qualitative insights into a model’s decision-making5,20,21. While useful, these methods are not quantifiable and thus provide limited utility for objective risk assessment in high-stakes scenarios. This has led to a growing interest in Uncertainty Quantification (UQ) as a means to provide a quantitative measure of model confidence22,23. While UQ has shown promise in unimodal applications—for instance, flagging challenging pathology slides for expert review or identifying novel retinal anomalies in ophthalmology24,25,26,27—its application in a multimodal context, however, remains a practical challenge.
The challenge of estimating prediction uncertainty when fusing disparate data types, such as imaging, clinical history, and lab tests, is magnified in the diagnosis of ovarian cancer. This clinical problem is defined by a dual layer of uncertainty: the inherent operator dependency of US imaging introduces technical variability28, while the large number of diagnostically ambiguous borderline tumors creates profound clinical uncertainty. The inadequacy of existing qualitative methods, coupled with this practical gap in multimodal UQ, therefore presents a critical bottleneck.
To address this critical gap, this study introduces and validates the Uncertainty-aware Multimodal Ovarian Risk Scoring System (UMORSS)(Fig. 1a, Supplementary Fig. 1). Our work makes four core contributions:
-
A novel two-phase diagnostic framework: by simulating and optimizing the clinical pathway, our framework enhances workflow efficiency while ensuring diagnostic safety, demonstrated by zero false negatives (FNs) in the initial triage phase.
-
A generalizable UQ module: we developed a streamlined bypass method that can be conveniently applied to any pre-trained model, providing a practical means for assessing the credibility of complex multimodal AI systems.
-
An uncertainty-weighted human-AI collaboration scheme: our prospectively validated scheme dynamically integrates AI predictions with radiologists’ judgments, significantly enhancing diagnostic performance and achieving a notable 22.48% average gain in sensitivity.
-
Rigorous validation on large-scale, multicentre cohorts (Fig. 1b): the entire system was comprehensively developed and validated on datasets comprising 7352 patients, ensuring the robustness and generalizability of our findings.
Herein, we demonstrate that by integrating a novel diagnostic pathway with a practical approach to uncertainty, it is possible to significantly improve the accuracy, reliability, and efficiency of ovarian cancer risk assessment. Ultimately, UMORSS provides a clinically-tested, generalizable blueprint for how advanced AI can be safely and effectively embedded into complex, high-stakes clinical decision-making.
Results
Overview of datasets
To develop and validate the UMORSS (Fig. 1a, Supplementary Fig. 1), we retrospectively analyzed data from 97,013 patients who underwent ovarian US examinations at Xijing Hospital between January 2012 and August 2021. After applying strict inclusion criteria, particularly regarding follow-up part, retrospective data from Xijing Hospital ultimately comprised 6931 patients with 8800 US images (Supplementary Figs. 2, 3, detailed in the Methods section). We collected the available clinical records for each patient, including their clinical history and laboratory tests. The study population included 4269 surgical cases (61.6%), with all nonsurgical patients having minimum one-year follow-up data. This high proportion of surgical cases, which is consistent with the characteristics of our tertiary care center, enabled the broad representation of various lesions. We divided the cohort by using May 2018 as the cut-off date: among the 5365 pre-May 2018 cases, 80% were randomly allocated to the training set, and the remaining 20% were randomly allocated to the validation set. The 1566 post-May 2018 cases constituted the independent internal testing cohort.
Moreover, after applying the same inclusion criteria, we evaluated data from 1316 patients from three other centers between October 2020 and October 2021. After screening, 421 patients with 481 US images and corresponding clinical data were included as a multicentre validation cohort (Supplementary Figs. 2, 4). To further validate our model in a prospective setting, we continuously enrolled 284 nonpregnant patients with newly diagnosed ovarian masses from Xijing Hospital between January 2022 and March 2022 (Supplementary Fig. 4). All of the patients in this prospective cohort had either pathologically confirmed ovarian lesions based on surgery or a minimum one-year follow-up. We used the data from this prospective cohort to conduct a reader study involving six radiologists with varying experience levels to evaluate the performance of radiologists alone, AI alone, and AI-assisted radiologists.
Primary risk screening to minimize unnecessary examinations
Phase I of UMORSS involved a primary risk screening before a detailed malignancy assessment. This triage approach aimed to identify low-risk ovarian lesions (predominantly physiologic cysts and some benign tumors) that require only routine follow-up, thus avoiding unnecessary interventions and optimizing clinical resources, and promoting clinical equity18. Identifying physiological cysts within this low-risk group is particularly important, as they are the only type of ovarian mass definitively not requiring surgery (Fig. 1a).
a System development workflow showing the construction, validation and clinical deployment of UMORSS using multimodal data from retrospective and prospective cohorts. b Distribution of pathological categories (physiological cysts, benign, borderline, and malignant tumors) across the four study cohorts.
At this phase, we jointly constrained the outputs of risk prediction and uncertainty scores (the prediction and uncertainty thresholds were set to 0.65 and 0.62, respectively, as detailed in the Methods section), and the model successfully identified an average of 68.7% of physiologic cysts and 13.8% of benign tumors as low-risk cases in the three testing cohorts. Furthermore, there were no FN cases (Fig. 2a, b). These findings suggest that the model’s primary screening maintains safety while effectively identifying cases that are suitable for routine surveillance.
a Proportion of cases identified as low-risk across different pathological categories in three cohorts. b Distribution of true negatives (TNs, lighter shades) and false negatives (FNs, darker shades) across different uncertainty thresholds in all cohorts. Numbers above bars indicate total low-risk classifications (TN + FN). The selected threshold (0.62) is marked in red.
Uncertainty-aware multimodal integration for enhanced performance
Uncertainty analysis revealed distinctive distribution patterns across ovarian lesion categories (Fig. 3a). Borderline tumours presented the highest uncertainty scores, reflecting their inherent diagnostic complexity. Malignant and benign tumours presented intermediate uncertainty levels, whereas physiologic cysts presented consistently low uncertainty levels. Moreover, misclassified cases overall presented higher uncertainty levels, although this difference was not significant for borderline tumours. This distribution aligns with clinical diagnostic challenges, thus enabling us to establish a quantitative threshold (0.52) for identifying cases requiring expert review (detailed in the Methods section and Supplementary Fig. 5).
a Distribution of uncertainty scores across correct (blue) and incorrect (pink) model predictions for different pathological outcomes in the internal testing cohort. b,c Comparison of AUC (b) and F1 scores (c) among the image-only model, UMORSS, and high-reliability predictions in the internal testing cohort. d Distribution of uncertainty scores across pathological outcomes in the multicentre cohort. e, f Comparison of AUC (e) and F1 scores (f) among the three models in the multicentre cohort. Box plots (a,d) show the interquartile range (IQR, 25–75th percentiles) with whiskers extending to 1.5×IQR; outliers are shown as individual points. Bar plots (b, c, e, f) display actual values with 95% confidence intervals (error bars) derived from bootstrap analysis. The P-values were tested with two-sided bootstrap tests (1000 resamples). All analyses incorporate combined results from both phases, including low-risk patients identified in Phase I.
In the internal testing cohort, performance increased progressively following the integration of multimodal data and uncertainty filtering (Fig. 3b, c; Supplementary Fig. 6a,b; Supplementary Table 1). Adding clinical parameters (UMORSS) significantly improved all evaluated metrics compared to the image-only baseline: notably, Area Under the Curve (AUC) increased to 0.947 (95% CI: 0.928–0.964, P < 0.001, Cohen’s d = 1.38), along with significant gains in F1 (to 0.737 [95% CI: 0.696–0.776]), F2 (to 0.819 [95% CI: 0.782–0.852]), and Matthews correlation coefficient (MCC) (to 0.694 [95% CI: 0.648–0.736]) (all P < 0.001). Applying uncertainty filtering (High Reliability UMORSS) further enhanced balanced metrics, significantly improving F1 score (to 0.849 [95% CI: 0.810–0.885], P < 0.001 vs UMORSS) and MCC (to 0.825 [95% CI: 0.784–0.867], P < 0.001 vs UMORSS). Although increases in AUC (to 0.955, P = 0.606) and F2 score (to 0.866, P = 0.083) at this stage were observed, these did not reach statistical significance.
Consistent findings were observed in the external multicentre cohort, confirming generalizability and reproducibility of uncertainty patterns and incremental improvements (Fig. 3d–f; Supplementary Fig. 6c,d; Supplementary Table 2). Specifically, compared to the image-only baseline, UMORSS significantly improved F1 score (P < 0.001), F2 score (P < 0.001), and MCC (P = 0.011), along with an increase in AUC (nominal P = 0.016, Cohen’s d = 0.189 [95% CI: 0.101–0.277]). Subsequent uncertainty filtering (High Reliability UMORSS) again significantly enhanced balanced classification metrics relative to standard UMORSS, indicated by increases in F1 score (to 0.616, P = 0.047) and MCC (to 0.617, P = 0.049). Although positive trends were also noted for AUC (to 0.926, P = 0.940) and F2 score (to 0.757, P = 0.146), these changes did not achieve significance after adjusting for multiple comparisons.
After correcting for multiple comparisons (Holm-Bonferroni), stepwise improvements in balanced metrics (F1 score and MCC) remained statistically robust across both internal and external cohorts, whereas the observed increases in AUC and sensitivity-focused F2 score—although nominally significant in certain cases (e.g., external cohort AUC nominal P = 0.016)—did not consistently survive multiple testing correction. This pattern likely reflects high baseline performance levels (AUC) as well as inherent sensitivity trade-offs from uncertainty-based sample filtering. Furthermore, robust performance stability was demonstrated across diverse imaging equipment (Supplementary Fig. 7; Supplementary Table 3). These findings reinforce the primary utility of our integrated multimodal approach with uncertainty awareness for reliably enhancing diagnostic classification.
Comparing the performance of AI and radiologists
We conducted a comparative evaluation of UMORSS against the performance of radiologists in both the internal testing cohort and the prospective cohort. On the basis of the validation set, we established optimal threshold values to convert the model’s continuous risk predictions (0–1) into O-RADS grades (ranging from 2–5). For the internal testing cohort with large-scale samples, we compared our model with a senior radiologist who performed O-RADS annotation. The UMORSS (AUC: 0.947, 95% CI: 0.928, 0.964) significantly outperformed senior radiologists (P = 0.006, DeLong test, Cohen’s d = 3.07); the UMORSS was further improved after filtering out high-uncertainty predictions (AUC: 0.955, 95% CI: 0.933, 0.974).
To obtain a more thorough assessment of the AI system against clinical practice, we conducted an independent reader study using the prospective cohort (n = 284). Six radiologists with varying experience (stratified into senior [>10 years, n = 2] and junior [≤10 years, n = 4] groups) independently assessed the cases following standardized image presentation. We compared their independent performance (‘Only Doctor’) against our AI models (Supplementary Table. 4).
Our UMORSS demonstrated strong standalone performance across a comprehensive suite of metrics. As visualized by its ROC curve (Fig. 4), it achieved an AUC of 0.984 (95% CI: 0.970–0.994). Other key metrics included Accuracy of 0.916 (95% CI: 0.880–0.947), Sensitivity of 0.954(95% CI: 0.881–1.000), Specificity of 0.909 (95% CI: 0.869–0.944), F1 score of 0.774 (95% CI: 0.674–0.857), F2 score of 0.872 (95% CI: 0.797–0.928), and MCC of 0.744 (95% CI: 0.649–0.834) (Supplementary Table 4). Compared to the individual radiologists’ performance points plotted in ROC space (Fig. 4), UMORSS generally operated at a higher sensitivity level than most readers for a given specificity, positioning its overall performance at the upper end of the observed clinical range.
ROC curves show the performance of the image-only model (dashed line), UMORSS (solid black), and High Reliability UMORSS (solid red). Individual radiologist performances are plotted as diamond markers, with the cross marker indicating the mean and standard deviation of radiologists’ assessments.
Furthermore, the High Reliability UMORSS attained exceptional performance levels, reflected in an AUC of 0.991 (95% CI: 0.980–0.999) and its superior position in the ROC analysis (Fig. 4). Its metrics included Accuracy 0.969 (95% CI: 0.945–0.988), Sensitivity 0.971 (95% CI: 0.903–1.000), Specificity 0.968 (95% CI: 0.943–0.991), F1 score 0.892 (95% CI: 0.807–0.960), F2 score 0.938 (95% CI: 0.875–0.982), and MCC 0.878 (95% CI: 0.789–0.954) (Supplementary Table 4). Notably, this high-reliability performance surpassed the highest score achieved by any individual participating radiologist across several key classification metrics, including F1, F2, and MCC, and its operating point on the ROC curve (Fig. 4) demonstrated a better sensitivity-specificity balance than any single reader. These comprehensive evaluations demonstrate that our AI system achieves expert-level performance in the risk assessment of ovarian tumours and has the potential to assist in clinical decision-making.
AI-assisted diagnosis optimization
To optimize diagnostic accuracy by leveraging complementary AI and radiologist strengths29,30, we developed and evaluated three AI-assistance schemes (average, uncertainty-weighted, prediction-weighted) on the internal testing cohort, combining model outputs with a senior radiologist’s O-RADS grades. The uncertainty-weighted scheme was selected as optimal. While demonstrating the most favorable AUC trend (0.972, 95% CI: 0.960-0.982, Supplementary Fig. 8a) and maintaining comparably high Negative Predictive Value (NPV > 0.98) for low-risk cases (O-RADS ≤ 3), its key advantage was a substantially higher Positive Predictive Value (PPV) (85.4% vs 76.2%/79.2%) for high-risk cases (O-RADS ≥ 4) (Supplementary Fig. 8b, c). This superior positive predictive power in the high-risk group suggests better potential for mitigating clinical overtreatment.
We further validated the optimal uncertainty-weighted scheme in the prospective cohort reader study (Fig. 5, Supplementary Fig. 9, Supplementary Table. 4). The AI assistance substantially improved overall diagnostic performance for all six radiologists (AUC P < 0.05 for all versus ‘Only Doctor’, DeLong test). Crucially, these significant gains across multiple metrics were achieved while maintaining specificity at high levels, generally comparable to the radiologists’ strong baseline performance. Assisted specificities remained high across all readers (ranging from 89.6% to 94.2% for Readers 1–5, and 93.8% for Reader 6). With specificity thus maintained, dramatic improvements were observed, most notably in sensitivity: average increases for the five radiologists with typical baseline performance included +10.58% in AUC, +3.98% in accuracy, +22.48% in sensitivity, and +15.96% in F1 score. The benefit was particularly striking for the radiologist starting with the lowest metrics (Reader 6), who experienced a 38.54% AUC increase, an 8.16% accuracy increase, and remarkable 4.57-fold and 2.04-fold increases in sensitivity and F1 score, respectively. These results demonstrate that our uncertainty-weighted AI assistance provides robust diagnostic enhancement by significantly boosting sensitivity and overall accuracy while preserving high specificity, effectively supporting less experienced radiologists and suggesting potential value for standardizing care, especially in resource-limited settings.
Performance metrics shown for the independent model (dotted line), radiologists’ independent assessments (pink bars), and AI-assisted readings (blue bars). Left panels show two senior radiologists ( > 10 years’ experience); right panels show four junior radiologists ( ≤ 10 years). Metrics include AUC (a), accuracy (b), sensitivity (c), specificity (d), and F1 score (e). For metrics b-e, O-RADS > 3 served as the classification threshold. Bars indicate actual values with 95% confidence intervals (error bars) derived from bootstrap analysis.
Interpreting the basis of model predictions
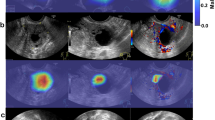
To elucidate the model’s decision-making process, we employed two complementary interpretability methods. SHapley Additive exPlanations (SHAP) analysis of the final multimodal model provided a transparent, quantitative attribution for individual predictions by determining the contribution of each feature (Fig. 6). In parallel, visual interpretation using SHAP-weighted Class Activation Maps (CAM) confirmed that the model’s focus aligns with clinical practice. The resulting heatmaps (Supplementary Fig. 10) demonstrated that high-activation regions consistently localized to sonographic features suggestive of malignancy, such as solid components and papillary projections, while areas corresponding to low-risk features showed minimal activation. These combined analyses suggest the model has learned to base its predictions on clinically relevant indicators.
For three representative cases, (a) Borderline Tumor, (b) Physiologic Cyst, and (c) Malignant Tumor, this figure provides both visual and quantitative explanations.On the left, SHAP-weighted CAM highlight the sonographic regions driving the model’s prediction. On the right, corresponding SHAP waterfall plots quantitatively attribute the final risk score to the contribution of each multimodal feature.
Discussion
The increasing complexity of ovarian cancer diagnosis, coupled with rising healthcare costs, demands more efficient and accurate diagnostic approaches. Current clinical practice faces significant challenges in reliably integrating diverse data sources (clinical findings, labs, US imaging) and managing resource utilization effectively. While recent AI applications have shown promise in analyzing specific data types4,18,31,32, their predominant focus on single modalities often fails to address the inherently multimodal nature of comprehensive ovarian cancer risk assessment. This highlights a critical gap in developing integrated solutions that mirror clinical reality and address workflow bottlenecks.
To bridge this gap, we developed UMORSS, a framework designed to enhance both diagnostic accuracy and workflow efficiency. This study validates the success of this integrated strategy: Phase I acts as a highly sensitive filter, safely excluding a substantial proportion of low-risk lesions (avg. 68.7% cysts, 13.8% benigns in three test cohorts) with zero FNs observed, while Phase II provides robust multimodal assessment. Critically, UMORSS successfully operationalized a ‘sensitivity-first, optimize specificity’ principle, demonstrating exceptionally high sensitivity overall combined with expert-level specificity, as confirmed across multiple cohorts, including a prospective reader study (Supplementary Table. 4). This achievement highlights the potential of such integrated AI systems to significantly improve the consistency and effectiveness of ovarian cancer risk assessment. Furthermore, in a comparative analysis against other state-of-the-art models for ovarian cancer diagnosis, UMORSS demonstrated significantly superior performance, as detailed in Supplementary Table 5.
To further enhance clinical trust and utility, we focused on making the model’s decision-making process more transparent. The challenge of UQ in medical AI extends beyond technical implementation, and while existing approaches have shown success in single-modality applications24,25,26,27, ovarian cancer diagnosis presents unique multimodal challenges. We addressed this through a practical, bypass-based UQ module17. A key advantage of this design is that the uncertainty estimator is trained separately, allowing it to be retrofitted to any pre-trained diagnostic model without architectural changes, a design choice that facilitates its broad application across various clinical diagnostic tasks. Importantly, the system’s behavior is itself interpretable: it assigns the highest uncertainty to borderline tumors and misclassified cases, a rational process that aligns with clinical logic. To provide more granular insight, we also performed a dedicated interpretability analysis. This analysis provides crucial evidence for the model’s clinical plausibility; rather than operating as an unexplainable ‘black box’, the model’s reasoning is shown to be grounded in clinically relevant indicators. Both the quantitative feature attributions from SHAP and the visual evidence from CAM confirm that the model’s focus aligns with established radiological practice (e.g., attending to solid components and papillary projections). This alignment is fundamental for building clinical trust and provides a transparent foundation for effective human-AI collaboration. Together, these features provide a multi-layered explanation for the model’s outputs.
The standardization of ovarian tumor assessment remains a critical challenge in clinical practice. While O-RADS has improved consistency in US interpretation6,8, and previous machine learning approaches have shown promise5, the interaction between AI systems and varying levels of clinical expertise requires careful consideration (Supplementary Fig. 8e). Indeed, our study confirmed variability in diagnostic performance among radiologists with different experience levels (Figs. 4, 5, Supplementary Fig. 9, Supplementary Table. 4), underscoring the inherent subjectivity in clinical practice. Our uncertainty-weighted integration scheme represents a novel approach to this challenge, demonstrating how AI can adapt to different levels of clinical expertise while preserving the standardization benefits of O-RADS. This adaptive capability is particularly important given the variation in radiologist experience and resource availability.
The clinical implications of UMORSS stem from its integrated design. The two-phase architecture directly refines the clinical workflow, acting as an early warning system for high-risk patients to facilitate timely intervention, while also serving as an efficient triage tool for low-risk cases to help conserve expert resources. Beyond workflow optimization, the system’s use of UQ and a human-AI collaboration scheme offers a practical approach to reducing inter-observer variability. This suggests UMORSS could serve as a valuable tool for standardization, helping to enhance diagnostic consistency among clinicians with different levels of experience. Collectively, these features highlight the system’s potential to contribute to a more efficient and reliable standard of care in ovarian cancer diagnosis.
While UMORSS shows promise, its limitations also highlight important avenues for future research. First, the prospective validation was conducted solely at a single center (Xijing Hospital). Although this center serves a broad population, large-scale, multicentre prospective trials are necessary to rigorously evaluate the model’s real-world workflow integration and clinical value. Second, the study’s reliance on static US images omits spatio-temporal information from dynamic video streams; exploring the integration of such data is an important future avenue. Third, further work is needed to evaluate and optimize the human-AI collaboration strategy with larger and more diverse groups of clinicians. Notably, our reader study demonstrated (Supplementary Fig. 11) that providing more static images significantly reduced indeterminate classifications, highlighting the need to optimize information presentation in human-AI interaction. Fourth, our model, utilizing ImageNet pre-training, showed performance comparable (Supplementary Fig. 12) to a large biomedical foundation model (BiomedCLIP33). This might be attributable to our large dataset size diminishing the benefits of domain-specific pre-training. Therefore, developing foundation models specifically pre-trained on large-scale, diverse US datasets remains an important future direction. Fifth, while our model’s performance in classifying borderline tumors reflects the current diagnostic difficulties, this subtype remains challenging to characterize non-invasively. It is important to note that the lack of specific imaging biomarkers for borderline tumors is a broader clinical challenge, not unique to our AI approach. Our uncertainty module is designed to mitigate this risk by flagging these ambiguous cases for expert review. Nevertheless, we believe that with the continuous accumulation of data, future work involving targeted data mining on this specific subtype may yield novel insights and further improve clinical decision-making. Furthermore, to enhance system reliability and clinical applicability, future research should also focus on more advanced methodologies, such as integrating sophisticated uncertainty modeling techniques34,35,36,37,38, and exploring how rapidly evolving AI architectures can more effectively fuse multimodal data39,40.
The core principles of the UMORSS framework—two-phase triage, multimodal fusion, and integrated uncertainty—are inherently generalizable. This framework could be readily adapted to other medical prediction tasks that face similar challenges, such as the risk stratification of breast lesions or lung nodules, thereby extending its potential impact to other clinical domains.
Methods
Dataset description
Data preparation: We retrospectively collected data from patients at Xijing Hospital from January 2012 to April 2018. These data composed the development cohort, which was randomly divided into a training set and a validation set at a ratio of 8:2. Data from the same hospital were collected from May 2018 to August 2021; these data composed the internal testing cohort. Importantly, data splitting was performed on a per-patient basis, such that all US images and corresponding regions of interest (ROIs) from a single patient were assigned entirely to one data subset (e.g., training, validation, or testing). This ensures that the model’s generalization performance was evaluated on data from entirely unseen patients. Moreover, we collected retrospective data between October 2020 and October 2021 from three other hospitals (The Second Affiliated Hospital of Xi’an Jiaotong University, Xi’an Fourth Hospital, Xi’an Aerospace Hospital); these data composed the multicentre validation cohort. Finally, the data for the prospective cohort were collected from 284 eligible patients at Xijing Hospital between January 2022 and March 2022. These data were acquired by three specially trained US doctors who followed unified standards defined according to the O-RADS guidelines7 and used three designated US systems (one GE Voluson E6 US System, and two Mindray M8 Diagnostic US System). As a tertiary hospital, Xijing Hospital has a benign-to-malignant ratio that is distinct from that of other general hospitals. Supplementary Fig. 2,3 outlines the exclusion and inclusion criteria for patients, ovarian lesions, and images. Specifically, to ensure the study sample reflected typical clinical data for ovarian cancer screening and diagnosis, we first excluded patients undergoing examinations during pregnancy (18,955 cases across development and internal testing cohorts; 277 in multicentre cohort). Next, we excluded patients who underwent surgery without clear pathological results or those undergoing follow-up lacking multiple examination records (64,895 cases across development and internal testing cohorts; 419 in multicentre cohort), and patients after surgery or undergoing chemotherapy (3128 cases across development and internal testing cohorts; 149 in multicentre cohort). Finally, we screened for image quality, excluding cases with poor quality images, such as those with corrupted DICOM files, severe blurring, or incomplete imaging of the ovarian mass (3104 cases across development and internal testing cohorts; 100 in multicentre cohort). Then, if patients met the inclusion criteria, we collected their clinical histories and laboratory test results.
During the annotation process, ROIs on US images were annotated by two senior radiologists (L.Zhao and M.Z.). Given that these datasets were large and the annotation task was relatively straightforward, each radiologist annotated half of the samples. Furthermore, all patients in the internal testing cohort and the prospective cohort underwent O-RADS grading on the basis of the US images and the radiologists’ descriptions during image acquisition. In the internal testing cohort, grading was independently conducted by a senior radiologist, whereas in the prospective cohort, six radiologists with varying degrees of experience independently evaluated all patients. The detailed annotation process is presented in Supplementary Fig. 13. By integrating surgical pathology reports or follow-up records of at least one year, we assigned labels representing the risk of malignancy to each lesion. Supplementary Tables 3, 6 and 7 display the number of data points in each cohort, the distribution of patients with different malignancy risk levels in each cohort, and the proportions of data from various US equipment manufacturers, respectively.
This study was approved by the Medical Ethics Committee of Xijing Hospital (KY20212182-F-1). The requirement for informed consent was waived for retrospective cohorts. For the prospective cohort, whilst not an interventional trial, all participants provided written informed consent after receiving comprehensive information about the study.
Data Preprocessing: Data preprocessing comprised the preparation of both US images and structured data. For US images, ROIs were cropped according to radiologist annotations. The training dataset then underwent extensive data augmentation to enhance model robustness. First, to account for potential inter-observer annotation variability, the annotated bounding boxes were randomly scaled by ±10%. The images were then cropped according to these adjusted boxes. Subsequently, a pipeline of further augmentations was applied to these cropped images, including rotation, horizontal flipping, random cropping, affine transformation, scaling, and adjustments to contrast and brightness. All images were subsequently standardized to 352 × 352 pixels. The structured data consisted of 12 features across four categories: (1) clinical history (menopausal status, family history of cancer in immediate family members, history of gynecological surgeries and cancer, hormone therapy history, and tenderness on palpation); (2) laboratory tests (CA125, HE4, CA19-9, AFP, and CEA); (3) maximum diameter of mass; and (4) blood flow characteristics (detailed in Supplementary Table 8). For continuous variables, we performed normalization using the 5 and 95th percentiles as boundaries, with missing values denoted as ‘NaN’. For discrete data, 0 was used to denote missing data (0 also represents “no” or “not”).
Development of UMORSS
We developed UMORSS, a two-phase system for ovarian lesion risk assessment (Supplementary Fig. 1). Phase I (primary risk screening) employed an image classification model to identify low-risk lesions (physiological cysts or benign findings with <1% malignancy risk; O-RADS 1 or 2) using US images. Phase II (multimodal classification) integrated image features with structured clinical data for comprehensive risk assessment. Both phases incorporated uncertainty assessment modules to evaluate prediction reliability.
Image Classification and Feature Extraction: We employed the vision attention network (VAN)41 (VAN-tiny) as the base architecture for both Phase I classification and Phase II feature extraction. VAN utilizes a large kernel attention (LKA) mechanism, combining the advantages of convolution and self-attention operations. The network’s fully connected layer implements a Softmax activation function, with Focal Loss42 addressing data imbalance during training.
The classification strategies differed between phases. In Phase I, physiological cysts were designated as negative cases, while tumor lesions (borderline and malignant) were labeled as positive cases. This classification was designed to distinguish between cases clearly not requiring surgery and those potentially requiring surgery. Benign tumors were excluded during model training but considered in statistical analyses, as their classification as either low-risk or high-risk was acceptable at this screening phase. For Phase II, physiological cysts and benign tumors were categorized as negative cases, while borderline and malignant tumors were designated as positive cases. Both VAN models were initialized with ImageNet pretrained weights and employed balanced sampling in each iteration to avoid performance bias, while also being trained using a progressive learning rate strategy: starting at 1 × 10⁻⁵, increasing to 1 × 10⁻⁴ after a 10-epoch warm-up period, followed by cosine annealing to 1 × 10⁻⁶. Training continued for 200 epochs with a dropout rate of 0.1. All training and testing were performed on six NVIDIA TITAN V GPUs.
Multimodal Classification Model: For multimodal classification, the primary challenge lies in effectively integrating high-dimensional image features with structured clinical data, which often contain mixed variable types and missing values. To address this, we selected XGBoost, a gradient boosting algorithm renowned for its high performance and robustness in handling such heterogeneous tabular data. The initial feature set extracted from the VAN model was high-dimensional (256 features). To create a more streamlined and computationally efficient model by reducing potential feature redundancy, we implemented a feature selection step using the Lasso algorithm. Driven by its intrinsic L1 regularization, which is well-suited for creating parsimonious models, Lasso selected the 155 most informative image features (Supplementary Fig. 14). These selected features were then concatenated with the 12 structured clinical features to train the final XGBoost model. This optimization resulted in a more streamlined model and increased processing speed by 16%, without compromising predictive performance.
To enhance model robustness, we introduced ±0.03 random fluctuations to non-empty components (constrained within 0–1). The XGBoost classifier, using Cross-entropy Loss, was optimized through grid search with the following hyperparameters: learning rate (0.018), estimators (700), maximum tree depth (7), column sampling rate (0.6), minimum child weight (2), subsampling rate (0.9).
Uncertainty Module: We developed an uncertainty module based on the trained classification models using a simple yet generalizable bypass approach43,44. The modules were trained through a regression task, where the classification loss between model predictions and true labels served as the training target.
Uncertainty modules were incorporated into both the Phase I and the Phase II. For Phase I, the cross-entropy loss from the classification task was adopted as the uncertainty prediction target and was calculated as follows:
In contrast, for Phase II, we proposed a comprehensive uncertainty target that was computed as the normalized combination of classification Cross-entropy Loss and the absolute difference between the predicted values and ground truth labels. This target was expressed as follows:
The training of both uncertainty modules was conducted via the mean squared error (MSE) loss function.
For Phase I, we implemented a bypass mechanism that preserved the feature extraction parameters while training three fully connected layers (hidden dimension: 512) with LeakyReLU activation and Softplus normalization. The learning rate was initialized at 5 × 10⁻⁵, with bounds of 1 × 10⁻⁵ and 3 × 10⁻⁶. Phase II’s UQ employed XGBoost regression with identical input features as its classification counterpart. Grid search identified optimal hyperparameters: learning rate (0.0015), estimators (700), maximum depth (6), column sampling rate (0.6), minimum child weight (2), and subsampling rate (0.85).
Following model development, the system was deployed on a standard workstation (single NVIDIA RTX 3070 GPU) using TorchServe. Inference efficiency testing showed that the time to process a single case ranged from 0.124 s to 0.744 s, depending on the number of ROIs (1 to 5).
Selection of operating points
Given that the model outputs continuous values for both classification predictions (risk scores) and uncertainty scores, we established operating points using thresholds to facilitate a two-phase clinical decision-making process.
Phase I thresholds: the primary goal of Phase I was to confidently identify a subset of low-risk lesions, particularly physiologic cysts, which require no intervention, while ensuring very high sensitivity for detecting positive cases (borderline and malignant tumors). Analysis of the validation set (Supplementary Fig. 16a) showed that a classification threshold of 0.65 provided a suitable balance. At this threshold, the sensitivity for positive cases was 95.2%, and the exclusion rate specifically for physiologic cysts was 81.6%. Sensitivity decreased notably if the threshold was set higher, whereas lowering it to further increase sensitivity would significantly compromise the physiologic cyst exclusion rate. Thus, 0.65 was selected as the initial classification threshold. To further minimize potential FNs, an uncertainty threshold was subsequently applied, based on analysis shown in Supplementary Fig. 16b. With the classification threshold fixed at 0.65, various uncertainty thresholds were evaluated. Setting the uncertainty threshold at 0.62 was found to effectively identify low-risk-scoring cases that nonetheless had high uncertainty, capturing additional positive cases. This implementation increased the overall system sensitivity from 97.3% (observed at a higher uncertainty threshold of 0.64) to 99.3% (146/147). Therefore, cases were directed to Phase II for comprehensive evaluation if they met either criterion: risk score ≥0.65 or (risk score <0.65 and uncertainty score ≥0.62).
Phase II Thresholds: For cases proceeding to Phase II, classification thresholds were established to further stratify risk, aligning with O-RADS categories. This was achieved through analysis of sensitivity, specificity, and F2 score curves on the validation set (Supplementary Fig. 17a). Intersection points of these curves determined the boundaries: 0.04 (separating O-RADS 2 vs 3), 0.09 (separating O-RADS 3 vs 4), and 0.43 (separating O-RADS 4 vs 5). The score 0.09 also served as the binary classification threshold for Phase II analyses comparing model performance against radiologists. An uncertainty threshold for Phase II was set at 0.52, based on evaluating trends in patient proportion directed to further assessment, AUC, and F2 scores at different uncertainty levels (Supplementary Fig. 17b). Accordingly, cases with an uncertainty score exceeding the 0.52 threshold were considered to have low prediction reliability, warranting review or examination by experts with more seniority (Supplementary Fig. 5). Consistent with O-RADS guidelines, cases ultimately assigned O-RADS > 3 were classified as positive for final performance evaluations.
Image-only Threshold: For the comparative analysis using only images, the classification threshold was set to 0.33. This value was determined at the intersection point of the sensitivity and specificity curves derived from this specific model’s outputs on the validation set.
Reader study
To evaluate the diagnostic performance of our AI model compared with that of radiologists, we conducted a reader study incorporating radiologists with clinical experience spanning 8–20 years. Readers independently performed O-RADS grading on both the internal testing cohort and the prospective cohort in accordance with standard O-RADS grading guidelines. To ensure unbiased assessment, the cases were presented in randomized order, and the readers were blinded to the AI model predictions, peer assessments, and pathological findings. To compensate for potential information loss in static images, readers were provided with both US images and US examination reports.
In accordance with O-RADS grading guidelines, which define cases above O-RADS 3 as positive, we calculated the accuracy, sensitivity, specificity, and F2 score for each reader and compared these metrics with the AI model’s performance (Fig. 3 and Supplementary Table. 4). Notably, varying numbers of O-RADS 0 (indeterminate) assessments were recorded by readers because of limited image information or case complexity.
Integration of O-RADS and UMORSS
We developed a collaborative approach that integrates radiologists’ expertise with AI capabilities by integrating O-RADS grades labeled by readers (Pdoc), model predictions (Pai), and uncertainty scores (unc). Using the internal testing cohort with O-RADS grades assigned by an experienced radiologist, we evaluated three integration schemes. Using predetermined thresholds (0.04, 0.09, 0.43), we mapped the model’s predictions Pai to corresponding O-RADS grades (2, 3, 4, 5), denoted as Pai’, and designed three AI-assisted radiologist schemes:
Average: We computed the arithmetic mean of Pai’ and Pdoc as the final result.
Uncertainty-weighted: We normalized uncertainty scores (unc) to unc’ using the 5th and 95th percentiles as boundaries, scaling them to the range [0.25, 0.75]. This approach minimized the risk of bias from outliers while ensuring that both the model and the doctor were assigned a fair initial weight of 0.25. The adjusted weights unc’ and (1 - unc’) were then applied to Pdoc and Pai’, respectively, as the final result. In particular, when readers labeled O-RADS 0 (indeterminate), Pai’ was directly used as the final result.
Prediction-weighted: This method followed a similar approach to the uncertainty-weighted scheme but utilized the model’s prediction confidence score Pmax (the predicted probability corresponding to the predicted class) as the weighting basis. Considering that Pmax in binary classification ranged between 0.5 and 1, the score was calibrated by subtracting 0.25 (denoted as Pmax’) to ensure a fair weighting distribution. The adjusted weights Pmax’ and (1-Pmax’) were then applied to Pai’ and Pdoc, respectively, to calculate the final result. Importantly, for patients categorized as low-risk during Phase I, we calculated the weighted confidence using the actual predicted value from that phase rather than the artificially set value of 0.01, and Pai’ corresponds to O-RADS 2. Similarly, Pai’ is directly used as the final result when the O-RADS grade is 0.
To select the optimal AI-assistance scheme, we evaluated these three approaches assessing performance trends (Supplementary Fig. 8a) and clinical utility across O-RADS risk strata. A key finding was that the uncertainty-weighted scheme maintained the essential safety feature of a high NPV ( > 0.98), comparable across all schemes, for ruling out malignancy in the low-risk category (O-RADS ≤ 3). Performance characteristics in the inherently ambiguous intermediate category (3 < O-RADS < 4) were also similar among the schemes. The distinct advantage of the uncertainty-weighted approach emerged in the high-risk group (O-RADS ≥ 4), where it achieved substantially higher PPV, showing absolute improvements of 12.1% and 7.9% compared to the average and prediction-weighted schemes, respectively (Supplementary Fig. 8b, c, d). This demonstrates a markedly superior ability to correctly identify high-risk patients without sacrificing performance in lower-risk categories, thus offering greater potential to mitigate overtreatment and unnecessary referrals more effectively. Based on this favorable balance of maintaining safety while significantly improving positive prediction in the high-risk cohort, the uncertainty-weighted scheme was selected as the final AI-assisted radiologist approach.
Interpretability analysis
To provide both quantitative and visual explanations for the model’s predictions, we employed the SHAP framework. The primary objective was to move beyond “black-box” predictions by attributing outcomes to specific input features. For visual interpretability of the multimodal model, we developed a SHAP-weighted CAM method. For any given prediction, we first calculated the SHAP values for all features input to the XGBoost model. The SHAP values corresponding to the image-derived features were then used as weights for their respective source feature maps from the final convolutional layer of the VAN. These weighted maps were aggregated via a weighted sum and passed through a ReLU activation function to isolate positive contributions. The resulting map was normalized to create a final saliency heatmap, which was then overlaid on the original US image to highlight the specific regions influencing the model’s decision.
Statistical analysis
To comprehensively evaluate the model’s performance from multiple perspectives, we employed a suite of standard evaluation metrics. The definitions of these metrics and their clinical significance within the context of this study are outlined below:
-
AUC: A threshold-independent measure of the model’s overall ability to discriminate between malignant/borderline and benign/physiological cases.
-
Accuracy: The proportion of all cases that were correctly classified.
-
Sensitivity (Recall): The proportion of true positive cases correctly identified. High sensitivity was considered critical to minimize the risk of missed diagnoses.
-
Specificity: The proportion of true negative (TN) cases correctly identified, which is important for reducing unnecessary interventions.
-
F1-Score: The harmonic mean of precision and sensitivity, providing a balanced measure of performance.
-
F2-Score: A variant of the F1-score that places greater emphasis on sensitivity, reflecting the clinical priority of identifying all potential malignancies.
-
MCC: A robust metric for evaluating binary classification, particularly in the presence of class imbalance.
To evaluate the clinical utility of the AI-assisted radiologist scheme, we gave special consideration to predictive values:
-
PPV: The proportion of cases predicted as positive that are true positives. A high PPV indicates the reliability of a high-risk flag from the system.
-
NPV: The proportion of cases predicted as negative that are TNs. A high NPV is a key indicator of a diagnostic tool’s safety, providing confidence in a low-risk classification.
Patient-level assessments were derived from multiple US images per lesion. In Phase I, we selected the highest risk prediction and its associated uncertainty score as the representative result. Patients were classified as low-risk and exempted from further testing when their prediction and uncertainty values were below 0.65 and 0.62, respectively.
For Phase II evaluation, we integrated multiple US images with clinical and laboratory data. Following Phase I’s principle, the highest risk prediction and its corresponding uncertainty score were selected for patient-level assessment. In evaluating UMORSS’s overall performance, low-risk patients were assigned a prediction value of 0.01 (according to the description in the O-RADS guidelines: O-RADS 2, the almost certainly benign category, 1% risk of malignancy8), with 0.09 as the classification threshold for calculating accuracy, sensitivity, specificity, F1 score, F2 score and MCC.
To ensure consistent evaluation metrics across phases, we normalized the uncertainty scores of Phase I low-risk patients by scaling them according to the threshold ratio (0.62:0.52). In our study, Cohen’s d was used to calculate effect sizes (R version 4.4.3, effsize package version 0.8.1). For the comparison of AUC for paired data, specifically when comparing the performance of the AI-assisted doctor versus the doctor alone for each reader individually in the prospective cohort, and to compare the performance of AI alone versus the senior radiologists in the internal testing cohort, the DeLong test was employed (R version 4.4.3, pROC library version 1.18.5). Other statistical comparisons were performed using two-sided bootstrap tests (with 1000 resamples). Performance metrics’ 95% confidence intervals were estimated using 5000 bootstrap resampling iterations (Python scikit-learn version 1.0.2), maintaining original cohort sizes. To further investigate the specific statistical differences among the three groups and control the risk of Type I errors arising from multiple comparisons, we performed post-hoc tests. Specifically, the Holm-Bonferroni method was used to adjust the P-values for pairwise comparisons.
Data availability
Due to patient privacy and institutional review board requirements, the original US images and clinical data are not publicly available. All data requests supporting the results of this study should be directed to the corresponding authors, H.Y.
Code availability
The US image feature extraction and classification were performed using the VAN-tiny model (https://github.com/Visual-Attention-Network/VAN-Classification). The remaining code for this study has been stored on GitHub (https://github.com/MHMAILab/UMORSS) and is intended for non-commercial purposes only.
References
Siegel, R. L., Miller, K. D., Wagle, N. S. & Jemal, A. Cancer statistics, 2023. Ca. Cancer J. Clin. 73, 17–48 (2023).
Bristow, R. E., Tomacruz, R. S., Armstrong, D. K., Trimble, E. L. & Montz, F. J. Survival effect of maximal cytoreductive surgery for advanced ovarian carcinoma during the platinum era: a meta-analysis. J. Clin. Oncol. 20, 1248–1259 (2002).
Yazbek, J. et al. Effect of quality of gynaecological ultrasonography on management of patients with suspected ovarian cancer: a randomised controlled trial. Lancet Oncol. 9, 124–131 (2008).
Gao, Y. et al. Deep learning-enabled pelvic ultrasound images for accurate diagnosis of ovarian cancer in China: a retrospective, multicentre, diagnostic study. Lancet Digit. Health 4, e179–e187 (2022).
Xiang, H. et al. Development and validation of an interpretable model integrating multimodal information for improving ovarian cancer diagnosis. Nat. Commun. 15, 2681 (2024).
Chen, H. et al. Performance of IOTA ADNEX model in evaluating adnexal masses in a gynecological oncology center in China. Ultrasound Obstet. Gynecol. 54, 815–822 (2019).
Andreotti, R. F. et al. Ovarian-adnexal reporting lexicon for ultrasound: a white paper of the ACR ovarian-adnexal reporting and data system committee. J. Am. Coll. Radiol. 15, 1415–1429 (2018).
Andreotti, R. F. et al. O-RADS US risk stratification and management system: a consensus guideline from the ACR ovarian-adnexal reporting and data system committee. Radiology 294, 168–185 (2020).
Timmerman, D. et al. Logistic regression model to distinguish between the benign and malignant adnexal mass before surgery: a multicenter study by the international ovarian tumor analysis group. J. Clin. Oncol. 23, 8794–8801 (2005).
Froyman, W. & Timmerman, D. Methods of assessing ovarian masses. Obstet. Gynecol. Clin. North Am. 46, 625–641 (2019).
Khalifa, M. & Albadawy, M. AI in diagnostic imaging: revolutionising accuracy and efficiency. Comput. Methods Programs Biomed. Update 5, 100146 (2024).
Akazawa, M. & Hashimoto, K. Artificial intelligence in ovarian cancer diagnosis. Anticancer Res. 40, 4795–4800 (2020).
Lu, C., De Brabanter, J., Van Huffel, S., Vergote, I. & Timmerman, D. Using artificial neural networks to predict malignancy of ovarian tumors. in 2001 Conference Proceedings of the 23rd Annual International Conference of the IEEE Engineering in Medicine and Biology Society 1637–1640 (IEEE, 2001). https://doi.org/10.1109/IEMBS.2001.1020528.
Van Calster, B. et al. Evaluating the risk of ovarian cancer before surgery using the ADNEX model to differentiate between benign, borderline, early and advanced stage invasive, and secondary metastatic tumours: prospective multicentre diagnostic study. BMJ 349, g5920–g5920 (2014).
Lu, M. et al. Using machine learning to predict ovarian cancer. Int. J. Med. Inf. 141, 104195 (2020).
Van Calster, B. et al. Validation of models to diagnose ovarian cancer in patients managed surgically or conservatively: multicentre cohort study. BMJ m2614 https://doi.org/10.1136/bmj.m2614 (2020).
Cai, G. et al. Artificial intelligence-based models enabling accurate diagnosis of ovarian cancer using laboratory tests in China: a multicentre, retrospective cohort study. Lancet Digit. Health 6, e176–e186 (2024).
Christiansen, F. et al. International multicenter validation of AI-driven ultrasound detection of ovarian cancer. Nat. Med. 31, 189–196 (2025).
Challen, R. et al. Artificial intelligence, bias and clinical safety. BMJ Qual. Saf. 28, 231–237 (2019).
Qian, X. et al. Prospective assessment of breast cancer risk from multimodal multiview ultrasound images via clinically applicable deep learning. Nat. Biomed. Eng. 5, 522–532 (2021).
Zhou, W. et al. Ensembled deep learning model outperforms human experts in diagnosing biliary atresia from sonographic gallbladder images. Nat. Commun. 12, 1259 (2021).
Kompa, B., Snoek, J. & Beam, A. L. Second opinion needed: communicating uncertainty in medical machine learning. npj Digit. Med. 4, 1–6 (2021).
Chua, M. et al. Tackling prediction uncertainty in machine learning for healthcare. Nat. Biomed. Eng. 7, 711–718 https://doi.org/10.1038/s41551-022-00988-x (2022).
Leibig, C. et al. Combining the strengths of radiologists and AI for breast cancer screening: a retrospective analysis. Lancet Digit. Health 4, e507–e519 (2022).
Wang, M. et al. Uncertainty-inspired open set learning for retinal anomaly identification. Nat. Commun. 14, 6757 (2023).
Linmans, J., Elfwing, S., Van Der Laak, J. & Litjens, G. Predictive uncertainty estimation for out-of-distribution detection in digital pathology. Med. Image Anal. 83, 102655 (2023).
Dolezal, J. M. et al. Uncertainty-informed deep learning models enable high-confidence predictions for digital histopathology. Nat. Commun. 13, 6572 (2022).
Timmerman, D. et al. Subjective assessment of adnexal masses with the use of ultrasonography: an analysis of interobserver variability and experience. Ultrasound Obstet. Gynecol. 13, 11–16 (1999).
Wang, X. et al. Predicting gastric cancer outcome from resected lymph node histopathology images using deep learning. Nat. Commun. 12, 1637 (2021).
Topol, E. J. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 25, 44–56 (2019).
Tang, Z.-P. et al. Ultrasound-based radiomics for predicting different pathological subtypes of epithelial ovarian cancer before surgery. BMC Med. Imaging 22, 147 (2022).
Kawakami, E. et al. Application of artificial intelligence for preoperative diagnostic and prognostic prediction in epithelial ovarian cancer based on blood biomarkers. Clin. Cancer Res. 25, 3006–3015 (2019).
Zhang, S. et al. A multimodal biomedical foundation model trained from fifteen million image–text pairs. NEJM AI 2, 2400640 (2025).
Laves, M.-H., Ihler, S., Fast, J. F., Kahrs, L. A. & Ortmaier, T. Well-calibrated regression uncertainty in medical imaging with deep learning. In Proc. Third Conference on Medical Imaging with Deep Learning 393–412 (PMLR, 2020).
Deng, D., Chen, G., Yu, Y., Liu, F. & Heng, P.-A. Uncertainty estimation by fisher information-based evidential deep learning. In Proc. 40th International Conference on Machine Learning vol. 202 7596–7616 (JMLR, 2023).
Sensoy, M., Kaplan, L. & Kandemir, M. Evidential deep learning to quantify classification uncertainty. In Proc. 32nd International Conference on Neural Information Processing Systems 3183–3193 (Curran Associates Inc., 2018).
Hsu, Y.-C., Shen, Y., Jin, H. & Kira, Z. Generalized ODIN: detecting out-of-distribution image without learning from out-of-distribution data. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 10948–10957 (IEEE, 2020). https://doi.org/10.1109/CVPR42600.2020.01096.
Lakshminarayanan, B., Pritzel, A. & Blundell, C. Simple and scalable predictive uncertainty estimation using deep ensembles. In Proc. 31st International Conference on Neural Information Processing Systems 6405–6416 (Curran Associates Inc., 2017).
Zhang, K. et al. A generalist vision–language foundation model for diverse biomedical tasks. Nat. Med. 30, 3129–3141 (2024).
Hollmann, N. et al. Accurate predictions on small data with a tabular foundation model. Nature 637, 319–326 (2025).
Guo, M.-H., Lu, C.-Z., Liu, Z.-N. et al. Visual attention network. Comput. Vis. Media 9, 733–752 (2023).
Lin, T.-Y., Goyal, P., Girshick, R., He, K. & Dollár, P. Focal loss for dense object detection. In Proc. IEEE/CVF International Conference on Computer Vision 2999–3007 (IEEE, 2017). https://doi.org/10.1109/ICCV.2017.324.
Kirchhof, M., Collier, M., Oh, S. J. & Kasneci, E. Pretrained visual uncertainties. Preprint at https://doi.org/10.48550/arXiv.2402.16569 (2024).
Yoo, D. & Kweon, I. S. Learning loss for active learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 93–102 (IEEE, 2019). https://doi.org/10.1109/CVPR.2019.00018.
Acknowledgements
This work was supported by Natural Science Foundation of China (No. 82172993, 82172860, 82103037), Natural Science Foundation of Shaanxi Province, China (No. 2024JC-ZDXM-52), Shaanxi Provincial Key Industry Innovation Chain (No. 2024SF-ZDCYL-01-17), Shaanxi Provincial Innovation (No. 2023PT-07), Key Research and Development Project of Shaanxi Province (No. 2022ZDLSF04-04) and High-Performance Computing Platform of Xidian University. We are grateful to Xing Han, Rui Li, X. Shuai, Yujiao Luo, Xin Jin, M.Z. and L.Zhao for participation in the reader studies. We also thank Bin Wang and Lei Shang for their support and guidance in statistical analysis.
Author information
Authors and Affiliations
Contributions
H.Y., X.L., and L.W. conceived and guided the project. L.Zou, Z.C., X.S., Y.G., and M.L. collected the original US images and clinical data. L.Zhao and M.Z. annotated data. X.W., J.W., R.Z., B.Z., X.Z., J.Z., and Y.Z. wrote code for different tasks. X.L. and L.Zhao provided clinical guidance. J.W., X.L., and X.W. participated in data analysis. L.W. and L.A. provided guidance for algorithm improvement. X.W., J.W., and X.L. wrote the manuscript with help and feedback from other co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, X., Lv, X., Wang, J. et al. A multimodal uncertainty-aware AI system optimizes ovarian cancer risk assessment workflow. npj Digit. Med. 8, 614 (2025). https://doi.org/10.1038/s41746-025-01986-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-01986-4
This article is cited by
-
Artificial intelligence (AI) and machine learning (ML) in ovarian cancer: transforming detection, treatment, and prevention
Journal of Ovarian Research (2026)
-
Development and validation of an artificial intelligence-based model for diagnosing benign, borderline, and malignant adnexal masses
npj Precision Oncology (2026)