Abstract
Artificial intelligence analysis methods for chest X-ray images are limited by insufficient annotation data and varying levels of annotation, resulting in weak generalization ability and difficulty in clinical dissemination. Here, we present EVA-X, an innovative foundational model based on X-ray images with broad applicability. EVA-X uses a self-supervised learning method capable of capturing both semantic and geometric information from unlabeled images for universal X-ray image representation. It has demonstrated exceptional performance in chest disease analysis and localization, becoming a model capable of spanning over 20 different chest pathologies and achieving leading results in over 11 different pathology detection tasks. Additionally, EVA-X significantly reduces the burden of data annotation in the medical AI field, showcasing strong potential in the domain of few-shot learning. The emergence of EVA-X will greatly propel the development and application of foundational medical models, leading to potential improvements in future medical research and clinical practice.
Similar content being viewed by others
Introduction
Chest X-rays constitute 40% of the 3.6 billion imaging procedures performed annually worldwide due to their efficacy in diagnosing cardiopulmonary abnormalities, including COVID-19, pneumonia, pleural effusions, emphysema, and so on1,2. This imaging technology provides several advantages, such as affordability, minimal radiation exposure, and widespread accessibility. The rapid evolution of Artificial Intelligence (AI) has led to the emergence of numerous deep learning models2, expediting the diagnostic process and improving the accuracy of X-ray image interpretation. However, these models encounter significant challenges. Their heavy reliance on extensive labeled data3,4,5 not only consumes crucial medical resources but also limits their effectiveness and scalability in clinical settings. Moreover, the task-specific nature of current deep learning models restricts their ability to address diverse medical challenges, impacting their adaptability and flexibility in various healthcare settings.
AI foundational models have recently achieved outstanding milestones and become promising solutions to these challenges. Cutting-edge studies are rapidly expanding into medical research6,7,8. Trained on extensive datasets, these models provide precise diagnostic support, facilitating quicker and more accurate decisions for healthcare professionals. They are typically robust and versatile, achieving the best performance across a wide range of healthcare scenarios. Their performance scalability is notable, increasing steadily with data and parameters to well adapt to diverse healthcare needs. Additionally, their interpretability enhances healthcare safety. These advantages eliminate the need for researchers to repeatedly and heavily annotate data and design specific deep-learning models for specific medical scenarios. However, the medical domain has not yet seen an effective, flexible, scalable, and interpretable foundation model for chest X-ray images.
In response, we introduce EVA-X, a foundational model for comprehensive chest X-ray analysis using self-supervised learning. We adopt eight widely used public chest X-ray datasets3,4,5,9,10,11,12,13 for both training and testing, with the pre-training data totaling over 520 k. (see Section “Methods” and Supplementary Section B) Leveraging extensive unlabeled data, EVA-X acquires general visual representations, enabling effective deployment across all chest disease detection tasks based on X-rays. EVA-X demonstrates significant technological advancement by not requiring annotated data for training, thus reducing the demand for medical resources compared to traditional contrastive learning methods14,15,16,17,18. Moreover, EVA-X pioneers a strategy in the X-ray domain to simultaneously learn semantic and geometric features, combining the advantages of contrastive learning pre-training14,15,16,17,18 and mask image modeling pre-training19,20. This innovative approach enhances the universality of its visual representations, facilitating broad utilization across diverse chest disease detection tasks and showcasing exceptional generalization capabilities.
Extensive experiments have demonstrated the superiority of EVA-X in the X-ray domain. From the perspective of pre-trained visual representations, EVA-X is capable of learning without using any annotated data. Compared to 16 previous pre-trained models14,15,16,16,19,20,21,22,23,24,25, EVA-X exhibits greater scalability and flexibility. From the standpoint of transfer learning, we tested EVA-X on 11 X-ray physiological and pathological analysis tasks. The results indicate that EVA-X has significant advantages in semantic understanding and geometric analysis. Moreover, EVA-X can significantly reduce the need for annotated data in downstream tasks. For instance, in COVID-19 detection, EVA-X achieves a 95% accuracy with just 1% of the training data. In terms of interpretability, EVA-X can determine lesion locations using only category information. We argue that EVA-X holds the potential to significantly enhance AI’s diagnostic performance in chest diseases, thereby broadening the application scope of AI within healthcare, reducing the strain on medical resources, and ultimately contributing to the promotion of global public health.
Results
EVA-X is a family of medical foundational models pre-trained specifically for analyzing and diagnosing chest diseases. It utilizes the widely adopted vision transformer architecture21 in computer vision and acquires general visual representations through unlabeled X-ray images.
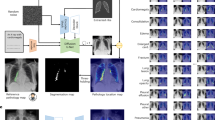
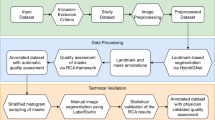
Illustrated in Fig. 1a, our pre-training dataset encompasses more than 20 distinct human chest health conditions, reflecting the diversity and complexity of chest health issues. EVA-X designs a novel self-supervised pre-training approach for X-ray images (Fig. 1b). This approach combines the benefits of contrastive learning and mask image modeling, efficiently capturing semantic and geometric information without requiring manual annotations during training. Due to its diverse training data and superior self-supervised training design, EVA-X can generalize to various X-ray-based chest disease detection scenarios. It is applicable to a wide range of tasks in chest physiology and pathology analysis (Fig. 1c). We evaluate EVA-X’s performance on 11 different X-ray image analysis tasks and compare it with the previous best methods. As depicted in Fig. 1d, EVA-X outperforms all of them, achieving SoTA results across all tasks. To our knowledge, EVA-X represents a comprehensive advancement of the advanced ViT structure over traditional convolutional models in the medical X-ray domain. This innovation heralds a new era in X-ray technology, where robust visual foundational models are likely poised to replace traditional designs.
a Pre-training Dataset: EVA-X pre-training collects and leverages a diverse set of X-ray images encompassing various health conditions.3,4,5b EVA-X Pre-training: EVA-X employs a novel self-supervised pre-training approach that synergistically integrates the strengths of contrastive learning14,15,16,17,18 and mask image modeling19,20. c General Visual Representations: EVA-X exhibits a high degree of transferability, enhancing the comprehensive analysis of X-ray imagery. d Transfer Performance: EVA-X demonstrates state-of-the-art performance across 11 distinct tasks3,4,9,10,11,12,13, outperforming established benchmarks set by previous pre-trained models. (Some icons in the figure sourced from biorender.com).
Below, we analyze the superiority of EVA-X in detail from three major perspectives: pre-training, transfer learning, and interpretability. We discuss the EVA-X self-supervised learning method in Section "Methods", as illustrated in Fig. 2.
EVA-X designs a self-supervised pre-training method combining the advantages of contrastive learning and mask image modeling for Chest X-ray images. Please See Section "Methods" for details.
Pre-training: performance, efficiency, and flexibility
We evaluate the EVA-X pre-training method across three dimensions: the performance of pre-trained visual representations, the number of parameters, and computational FLOPs. Our evaluation employs the CXR14 test set4, which serves as the benchmark dataset in the X-ray domain (see Section "Pre-training Data"). We compare EVA-X with 15 different pre-trained X-ray models, including widely used models such as DenseNet12123, ResNet5022, and ViTs21. Considering the diverse computational demands of medical scenarios, we train three EVA-X models of different scales: EVA-X-Ti, EVA-X-S, and EVA-X-B.
EVA-X demonstrates SOTA performance. As depicted in Fig. 3a left, we categorize these 19 different pre-trained models14,15,16,19,20,21,22,23,24,25 into three comparison groups: tiny models, small models, and base models, based on their parameter counts. Notably, within each group, EVA-X consistently exhibits the lowest parameter count (6M, 22M, 86M). We observe remarkable scalability in EVA-X, with its performance consistently improving as the parameter count increases. Among these models, EVA-X-B stands out as the best pre-trained X-ray model, achieving a visual representation test performance of 83.5 mAUC, surpassing all previous medical self-supervised pre-training methods, such as Medical MAE20, contrastive learning pre-training methods like MGCA14, and well-known pre-training methods for natural images like MAE26 and MoCov227. This achievement sets a new standard for SOTA performance in medical X-ray pre-training.
a Performance and Efficiency of EVA-X Pre-trained Models. Among all pre-trained models14,15,16,19,20,21,22,23,24, EVA-X-B achieves the highest performance. The EVA-X family demonstrates an excellent balance between performance and computational efficiency compared to previous methods. b Performance on Chest Pathologies Classification. EVA-X achieves the best performance in both multi-label and single-label classification tasks for chest pathologies3,4,9. c Performance on Label-efficient Classification. EVA-X shows superior performance across varying amounts of training data, with a particularly notable advantage observed when dealing with extremely limited training data.
EVA-X achieves exceptional efficiency. As depicted in Fig. 3a right, we assess the computational complexity of all methods during testing. To facilitate visualization, we logarithmically scale the FLOPs on the horizontal axis. The purple x marker on the graph signifies the correlation curve between computational complexity and the performance of EVA-X. EVA-X strikes an outstanding balance between performance and computational complexity compared to all other methods.
EVA-X offers a tiny alternative for flexibility. Typically, foundational models aiming for high performance often impose high computational demands, and it could be challenging in resource-constrained medical environments. However, leveraging the impressive capabilities of EVA-X, we not only investigate its performance boundaries but also develop a lightweight variant, EVA-X-Ti. It is worth noting that EVA-X-Ti is the model with the lowest computational complexity (1.26 GFLOPs) among them with incredible performance (82.4 mAUC). We conduct comparative experiments between EVA-X-Ti and 15 previously introduced pre-trained models, most of which have larger parameter counts than EVA-X-Ti. Despite this, EVA-X-Ti, with its streamlined parameters (6M), outperformed 14 of these models in performance metrics. It even outperforms MGCA-B14 (81.8 mAUC) and SelfMedMAE19 (81.5 mAUC), which have 13 times more FLOPs than EVA-X-Ti. This exceptional performance highlights EVA-X-Ti’s potential as a cost-effective alternative to large-scale models, promoting wider adoption and deeper integration of EVA-X technology across various applications.
Transfer learning on chest pathologies classification
X-ray images are one of the important tools for diagnosing chest diseases, with different diseases exhibiting different manifestations on X-ray images. Our experiments demonstrate that the visual representations learned by EVA-X pre-training are universal and can be generalized to diagnostic tasks for all chest pathologies.
Multi-label classification requires the model to make judgments about the presence of multiple different diseases at once. In our work, we evaluate the general pathologies detection capability of EVA-X using two commonly used multi-class chest pathologies diagnosis datasets, Chest X-Ray144, and CheXpert3. We fine-tune the visual representations learned by EVA-X on these two datasets without employing any additional design techniques.
As shown in Fig. 3b CXR14, we compare the results of EVA-X with eight different methods20,28,29,30,31,32,33 on the Chest X-Ray14 dataset. Data are presented as mean ± 95% CI (n = 5). Most of these methods are designed for chest X-ray classification. Among them, our EVA-X-Ti (6M) with 82.4 mAUC exceeds the 82.2 mAUC achieved by Kim et al.33. Their method uses DenseNet121 (8M) as a backbone. Our EVA-X-S (22M) with 83.3 mAUC, exceeds the 82.3 mAUC achieved by Xiao et al.20 with ViT-S 0.823 mAUC. Taken together, EVA-X outperforms the previous best method at two different sizes, reaching new SOTA results. From the perspective of single-pathology diagnosis, EVA-X performs best by achieving the highest accuracy in 12 out of 14 pathology diagnoses (see Supplementary Fig. 1 for more details).
As shown in Fig. 3b CheXpert, we compare EVA-X with 5 previous methods3,20,30,34,35. In terms of individual metrics, EVA-X reaches new SOTA results in 2 categories (see Supplementary Fig. 1 for more details). In terms of mAUC, both EVA-X-Ti, and EVA-X-S outperform all previous methods and reach new SOTA results. Among them, EVA-X-Ti has only 6M parameters, which is smaller than all previous methods, and exceeds the performance of all previous methods, and achieves new SOTA results.
Single-label classification requires the model to make accurate judgments about a specific pathology. In this paper, we test this using COVID-19 as an example. Specifically, we utilize the latest collected and annotated datasets COVID-CXR-3 and COVID-CXR-49 and fine-tune seven different pre-trained models14,15,16,20,22,36, including EVA-X, on each dataset. Data are presented as mean ± 95% CI (n = 5). As shown in Fig. 3b CovidX-CXR-3 and CovidX-CXR-4, EVA-X ranks first among all methods with exceptionally high 99.8 and 99.4 mAUC (benchmark values evaluated on the public dataset). Additionally, EVA-X maintains remarkable stability, demonstrating the most consistent performance across multiple experiments. Specifically, the mean standard deviation of EVA-X on both datasets is 0.03, which is lower than all other methods, including Medical MAE20 (0.045), MGCA14 (0.055), BioViL15 (0.135), etc.
Transfer learning on label efficient classification
The EVA-X model, optimized through large-scale data pre-training, exhibits a high sensitivity to small training data in downstream tasks. It can converge rapidly with minimal data, thereby directly alleviating the pressure of annotation data on the healthcare system. In Fig. 3c, we validate EVA-X’s efficient training capability on COVID-199 and compare it with previous methods14,15,16,20,26,27. To ensure robust results, each model undergoes five independent runs using distinct random seeds (0–4). At the beginning of each training epoch, the training set is shuffled, and a random subset is sampled for model updates. Performance is reported as the mean and standard deviation across the five runs. All models are finally evaluated on the official test set. EVA-X demonstrates the strongest and most stable performance at different data sizes. Especially in the case of very little annotated data, only 1% training data, EVA-X shows a clear advantage over other methods. On the CovidX-CXR-4 dataset, EVA-X achieves 95% diagnostic accuracy with only 1% of training data, highlighting its exceptional learning ability and generalization performance in resource-limited environments.
Transfer learning on chest x-ray segmentation
Medical segmentation demands deep learning models to precisely delineate anatomical structures and identify pathological features in medical images, aiding in diagnosis. We focus on evaluating EVA-X’s performance in both physiological and pathological segmentation tasks. Specifically, we fine-tune seven different medical models14,15,16,20,22,36 across four lung segmentation tasks, encompassing physiological segmentation and pathological segmentation for pneumonia, pneumothorax, and tuberculosis. These tasks demonstrate the model’s robust geometric understanding across diverse health conditions. Quantitative evaluation of segmentation results using Dice and Jaccard metrics, along with visualization of segmentation masks as depicted in Fig. 4, has been conducted through multiple experiments. Data are presented as mean ± 95% CI (n = 5).
a Performance on Chest X-ray Segmentation. EVA-X surpasses six other pre-trained models14,15,16,20,22,36 across all segmentation benchmarks10,11,12,13, exhibiting superior performance on Dice and Jaccard metrics. b Visualization of Segmentation Results. EVA-X demonstrates enhanced accuracy and finer masks across all segmentation tasks.
As shown in Fig. 4a, EVA-X demonstrates outstanding performance across four distinct tasks10,11,12,13. Specifically, in lung segmentation, EVA-X achieves the highest average Dice score of 95.49%. In pneumonia pathology segmentation, EVA-X surpasses both Medical MAE20 (53.16 Dice, 36.20 Jaccard) and BioViL15 (51.96 Dice, 35.10 Jaccard) with Dice and Jaccard scores of 54.51 and 37.47%, respectively. For pneumothorax pathology segmentation, EVA-X outperforms MGCA14 (59.00 Dice, 41.84 Jaccard) and the ImageNet pretrained model22 (57.69 Dice, 40.56 Jaccard) with scores of 60.27 Dice and 43.13% Jaccard. In pulmonary tuberculosis pathology segmentation, EVA-X excels with scores of 60.10 Dice and 42.96% Jaccard, surpassing Medical MAE20 (59.1 Dice, 41.96 Jaccard) and MGCA14 (59.00 Dice, 41.84 Jaccard). Furthermore, as illustrated in Fig. 4b, EVA-X provides more accurate and fine-grained physiological or pathological segmentation, showcasing its exceptional generalization ability in X-ray segmentation tasks.
Interpretability
The interpretability of X-ray deep learning is an essential topic, as highlighted in Baselli et al.37. Utilizing tools like the class activation map (CAM) can help elucidate the rationale behind neural network decisions, as discussed in Grad-CAM38. In the medical domain, pathology diagnosis often hinges on lesion localization. Saporta et al.39 have observed that while deep learning can provide reasonably accurate predictions, there remains a notable gap in its ability to automatically localize compared to human capabilities.
We employ Grad-CAM38 to analyze the gradients of EVA-X in the context of pathology diagnosis. Our analysis involves approximately 1000 images from the Chest X-Ray14 dataset4, as discussed in Sec. “Pre-training Data” each annotated with lesion positions. Subsequently, we select seven different model weights pre-trained as outlined in Sec. “Pre-training: Performance, Efficiency and Flexibility” for comparative evaluation. We get CAMs with each pre-trained model and measure the Intersection over Union (IoU) and Average Precision (AP) between the activation regions and the ground truth (GT) boxes. To determine the optimal performance threshold, we conduct a search within the range of [0.1, 0.6].
We present the corresponding results in Fig. 5a. Furthermore, we visually represent the CAM of EVA-X and the other six models using heatmaps, depicted in Fig. 5b. The results reveal several significant findings. Firstly, EVA-X demonstrates superior performance in terms of quantifiable metrics, such as IoU and AP, compared to the other seven methods. Secondly, consistent with findings in prior research20, ViT pretrained with MAE exhibits notably weaker CAM performance than CNN. However, our experiments indicate a substantial enhancement in ViT’s CAM quality when aided by EVA-X, resulting in a marked increase in mAP from 3.61 to 8.94. Additionally, our visual analysis highlights that EVA-X generates more accurate and distinct activation maps compared to previous methods. While CNN methods14,15,16 exhibit superior map continuity, they may not perform as effectively as EVA-X in localizing smaller lesions.
a Performance on Weakly-Supervised Localization. EVA-X delivers the highest overall performance across all four metrics for weakly supervised localization tasks.4 b Visualization of Grad-CAM. Class Activation Map (CAM)38 is a significant interpretation method for deep learning models. It illustrates that EVA-X can localize pathologies using only classification annotations, showcasing remarkable interpretability.
Real-world data evaluation
To investigate the potential of EVA-X in real-world scenarios, we have conducted an evaluation on an internal, real-world dataset. This dataset includes 10,000 chest X-ray images and reports collected from 14 Chinese hospitals, including Wuhan Tongji Hospital and Wuhan Union Hospital. Following a procedure similar to CheXpert3, we use Deepseek-v340 to analyze the reports and generate annotations for 14 distinct labels. The accuracy of this conversion method is confirmed by validation on a random 1 k subset, which achieves an F1-score of 99% against physician annotations. We then test EVA-X-S, Medical MAE20, MedKLIP16, BioViL15, and MGCA14 on this dataset using a 5-fold cross-validation scheme. The dataset is randomly partitioned into five equal-sized, non-overlapping subsets. In each fold, four subsets (80%) are used for training, and the remaining subset (20%) is used for testing/validation. As shown in Fig. 6, EVA-X achieves the best average AUC of 0.8645 and a maximum AUC of 0.8788, outperforming all other methods and demonstrating its potential superiority in real-world applications.”
In the box plots, the center line is the median, the box spans the 25–75th percentiles (IQR), and whiskers cover 1.5 × the IQR. All data points are shown.
Discussion
We propose EVA-X, medical foundation models tailored for X-ray images. Different from previous work, EVA-X utilizes a self-supervised pre-training strategy, combining previous pre-training methods of contrastive learning14,15,16,17,18 and mask image modeling19,20. It could learn generalizable visual representations for all X-ray tasks without any human-annotated images. This unique advantage makes EVA-X’s pre-training effective, efficient, flexible, and scalable. Compared with over 15 previous pre-trained deep learning models, EVA-X foundation models achieve SOTA performance and computation trade-off. We transfer EVA-X models to 11 downstream tasks3,4,9,10,11,12,13 and compare them with previous SOTA X-ray and natural image pre-trained models. The results show that EVA-X outperforms all previous models in all downstream tasks, demonstrating new SOTA performance on X-ray image classification, segmentation, and interpretation. We argue EVA-X has great potential to become a general foundation model in medical X-ray analysis, facilitating faster and more accurate diagnosis and analysis of chest pathologies.
EVA-X are foundation models designed for medical X-ray images. Its training process is based entirely on data related to chest X-rays. Therefore, its performance on other medical tasks is open to improvement. Due to the unique high-performance self-supervised pre-training strategy of EVA-X and the great potential it shows on X-ray tasks, we believe that EVA-X’s approach is expected to be extended to the entire medical field.
The training data for EVA-X is sourced from the public datasets Chest X-Ray144, CheXpert3, and MIMIC-CXR5. A full analysis of this data’s heterogeneity is provided in Supplementary Table 2. The inherent heterogeneity and potential biases from these sources, such as disparities in disease prevalence and patient demographics (e.g., age distribution), may compromise the model’s performance in specific out-of-domain scenarios41.
Compared to recent publicly available supervised models like Ark+42 and CXR-Foundation43, EVA-X excels with its label-free characteristic. In practice, the self-supervised pre-training of EVA-X can complement supervised methods, enabling the model to leverage more unlabeled clinical data and thereby achieve better performance.
Further clinical validation of EVA-X is an important subsequent step. On one hand, given its high efficiency (the tiny model is only 6M), EVA-X could be rapidly migrated and deployed in most practical scenarios for assistive diagnosis. On the other hand, EVA-X could also serve as a visual encoder combined with large medical language models and agents44,45 to further improve diagnostic performance.
Methods
Pre-training data
EVA-X is trained using exclusively public Chest X-Ray data. Our training set is a combination of three extensive public datasets: Chest X-Ray144, CheXpert3, and MIMIC-CXR5. These datasets are widely recognized for their application in X-ray vision-language pre-training14,16 and image classification31,33. In contrast to previous studies, our approach exclusively leverages pure unlabeled images for pre-training, without the utilization of any annotation or pathology report information.
For these datasets, we specifically process them as follows: (1) Following previous work20, we primarily use frontal view (AP/PA) images and discard lateral view images. (2) We do not use any of the images tested subsequently for training, even though they are unlabeled. (3) To speed up training, similar to CheXpert, we use bilinear interpolation to resize the original images to a resolution of 336 × 336. The combined dataset is called Merged520k (see Fig. 1b). If not otherwise noted, our pre-training experiments will be performed on this dataset. EVA-X utilizes a straightforward data processing method for self-supervised learning. First, we resize the entire Merged-520 k dataset to a uniform size of 336 pixels. During pre-training, we then randomly scale the images to a size between [3/4, 4/3] before taking a random crop of 224 × 224 pixels. Finally, all images are normalized using the mean and standard deviation of the Merged-520 k dataset.
Evaluation data
In the realm of natural images, ImageNet46 typically serves as the primary test dataset for pre-training24,26,47. Similarly, in the domain of X-ray images, it is essential to select a dataset for pre-training evaluation. Among the aforementioned datasets, both Chest X-Ray14 and CheXpert hold prominence as widely utilized categorization datasets31,33,35. They are characterized as multi-label categorical datasets, with labels assigned to 14 distinct pathologies. Notably, these 14 labels are independent of each other.
Here, we have opted for the former dataset, Chest X-Ray14, as our primary test set, which is the most commonly used X-ray classification dataset (as studied by ref. 48). Our decision is based on the following reasons: (1) More rational dataset distribution. The CheXpert dataset comprises a total of 224 k images, but only nearly 200 images are allocated for testing. In Chest X-Ray, the training/validation/test set ratio is 75/11/25 k. (2) Clearer labeling. In the CheXpert dataset, the presence of an “uncertain” annotation indicates that the physician did not identify the condition. Various approaches exist for handling this uncertainty. Some methods uniformly categorize it as “with pathology,” others as “without pathology,” and more complex treatment schemes are also employed. However, the labeling is clearer on the Chest X-Ray14 dataset. The selection of this test dataset is also consistent with the two previous works19,20.
Note that this dataset selection indicates that we perform pre-training studies on this dataset, but does not mean that we only use this dataset to test the final performance of EVA-X. In subsequent sections, we will demonstrate the superior performance of EVA-X on additional datasets.
EVA-X architecture
The pre-training process of EVA-X involves the design of the dual Vision Transformer (ViT)21 (see Fig. 2). The EVA-X transformer is learnable and the tokenizer is frozen. For the convenience of readers, we begin with a brief overview of ViT here.
Assuming the dimensions of the image are H, W, before attention calculation, ViT divides the image into \(n=\frac{H}{P}\times \frac{W}{P}\) different patches, where P represents the patch size. Typically, P can take values like 16, 14, 8, etc. In EVA-X, unless specified otherwise, the patch size for all ViTs is set to 16. For an image patch, ViT uses linear projection to project it into a feature vector of dimension d, which is referred to as image tokens. These vectors form a sequence known as the image token sequence. Additionally, to establish positional relationships between vectors, ViT uses positional encoding added to the image token sequence. After adding the token dedicated to classification, we obtain the final input sequence, as shown in equation (1), denoted as Z.
The transformer block (see Fig. 2b) is a straightforward structure with the same output structure as the input. It mainly consists of two parts: Multi-Head Self-Attention (MHSA) and a Feed Forward Network (FFN). Inspired by Fang et al.24, in EVA-X, we introduce improved structures such as rotational positional encoding, Sub-LN49, and SwiGLU50, which differ slightly from traditional ViT. By stacking any number of transformer blocks, the final ViT is composed. For the input Zi at layer i, the transformer block performs the following calculations to produce the final output Zi+1, where Zi and Zi+1 have the same structure.
EVA-X is a learnable Vision Transformer. Here, we selected three ViTs of different sizes for experimentation: ViT-Ti, ViT-S, and ViT-B, with a patch size of 16 for each structure. Based on the number of parameters, we primarily use EVA-X-Ti (6M) to benchmark against DenseNet12123 (8M), EVA-X-S (22M) against ResNet5022, and EVA-X-B (86M) to explore the effects and influences of scaling up the number of parameters.
To perform mask operations on images in mask image modeling, following previous work24,47, we designed a mask token denoted as m. This token is a learnable d-dimensional vector. Assuming a mask ratio of r, we randomly replace n ⋅ r image tokens with mask tokens. We denote this sequence of masked tokens as mask_list. All mask tokens have the same initialization.
Due to potential dimension differences between EVA-X and Tokenizer, we use a linear projection layer to map the dimension of EVA-X’s image tokens from deva_x to dtgt. We denote the final output sequence of EVA-X as
Self-supervised learning
The role of the Tokenizer is to extract semantically rich features from images, and it is also a ViT structure. Unlike EVA-X, we generally opt for larger-scale ViTs. We primarily investigate two types of structures for Tokenizer’s pre-training performance, namely, natural image CLIP and medical image CLIP. For natural images, we selecte advanced high-performance ViT-B, ViT-L, and ViT-G visual encoders from the EVA-CLIP51 model as our Tokenizer. In the medical field, we chose the ViT-B visual encoder trained with MGCA14 as our Tokenizer. As far as we know, MGCA-ViT-B is currently the best open-source X-ray CLIP model.
Tokenizer takes the sequence Z as shown in the equation below as input and maps it to the target feature sequence Zt, represented by the following equation. During the pre-training process, all parameters of the Tokenizer are kept frozen, and no additional learnable linear mappings are added.
As mentioned earlier, for the token sequences in the equation Z, we randomly select a proportion r of tokens and replace them with randomly initialized mask tokens. Here, we choose a relatively small mask ratio, r = 0.3. We denote the indices of the masked image tokens as mask_list.
For the final output sequences of EVA-X and Tokenizer, we respectively select the image tokens in mask_list to form the sequences \(Z^{\prime}\) and \({Z}_{t}^{{\prime} }\). We aim to maximize the cosine similarity between corresponding tokens in \({Z}_{e}^{{\prime} }\) and \({Z}_{t}^{{\prime} }\), i.e.,
Transfer learning on classification
In the case of classification tasks, we use the simplest decoding strategy uniformly for all models. For CNNs such as ResNet5022, DenseNet12123, etc., we average the features output from their last network layer for pooling, and then input the pooled features into a learnable linear layer to generate predictions. For the ViT21 structure used by methods such as EVA-X, we average all the tokens output from the last block, and then input the corresponding features into a learnable linear layer as well to output the prediction results. This simple structure ensures the ability to directly compare the underlying models, rather than a complex structural design.
We use the mean Area Under the Curve (mAUC) and mean Accuracy (mAcc) as our classification metric, as denoted in Eq (8) and (9), while TPR denotes True Positive Ratio, FPR denotes False Positive Ratio, TP denotes True Positive, TN denotes True Negative, FP denotes False Positive, and FN denotes False Negative.
Transfer Learning on Segmentation
Following the previous methods14,15,16,18, in this paper, we primarily focus on the comparison of pre-trained visual representation performance, without overly emphasizing the potential advantages that structural improvements may bring to the segmentation tasks. Specifically, we build two segmentation models using ResNet5022 and ViT21 backbones, which are the most commonly used structures in X-ray pre-training. For ResNet50, we followed previous work14,16, adopting the structure with a ResNet encoder and a UNet52 decoder. For ViT, we follow common practices in natural images53, initially building a feature pyramid by pooling and deconvolution on the last layer features, and then using UperNet54 as the decoder for segmentation tasks. To ensure the simplicity of the structure as much as possible, we do not employ advanced adaptive structures, to better explore the performance of visual representations, although they may bring improvements in performance.
We use the mean of Dice and the mean of Jaccard as our segmentation metric, as shown in Eq (10) and Eq (11), while S denotes segmentation result and G denotes GT.
Data availability
All datasets used in this study are publicly available. Data preprocessing scripts and model weights are available at our GitHub repository: https://github.com/hustvl/EVA-X. The datasets can be accessed via the following links: Chest X-Ray14: https://www.kaggle.com/datasets/nih-chest-xrays/data; CheXpert: https://stanfordmlgroup.github.io/competitions/chexpert/; MIMIC-CXR: https://physionet.org/content/mimic-cxr/2.1.0/; COVIDx: https://www.kaggle.com/datasets/andyczhao/covidx-cxr2; SIIM-ACR Pneumothorax Segmentation: https://www.kaggle.com/datasets/jesperdramsch/siim-acr-pneumothorax-segmentation-data; RSNA Pneumonia Detection: https://www.kaggle.com/competitions/rsna-pneumonia-detection-challenge; Tuberculosis Chest X-rays: https://www.kaggle.com/datasets/raddar/tuberculosis-chest-xrays-shenzhen; Lung Segmentation: https://github.com/YoushanZhang/Lung_Segmentation.
Code availability
Our codes and models are available at https://github.com/hustvl/EVA-X.
References
Organization, W. H. et al. Communicating radiation risks in paediatric imaging: information to support health care discussions about benefit and risk (WHO, 2016).
Cid, Y. D. et al. Development and validation of open-source deep neural networks for comprehensive chest x-ray reading: a retrospective, multicentre study. Lancet Digital Health 6, e44–e57 (2024).
Irvin, J. et al. Chexpert: A large chest radiograph dataset with uncertainty labels and expert comparison. Proc. Artif. Intell. 33, 590–597 (2019).
Wang, X. et al. Chestx-ray8: Hospital-scale chest x-ray database and benchmarks on weakly-supervised classification and localization of common thorax diseases. In Proc. IEEE Conference on Computer Vision and Pattern Recognition, 2097–2106 (IEE, 2017).
Johnson, A. E. et al. Mimic-cxr-jpg, a large publicly available database of labeled chest radiographs. arXiv preprint arXiv:1901.07042 (2019).
Moor, M. et al. Foundation models for generalist medical artificial intelligence. Nature 616, 259–265 (2023).
Zhou, Y. et al. A foundation model for generalizable disease detection from retinal images. Nature 622, 156–163 (2023).
Ma, J. et al. Segment anything in medical images. Nat. Commun. 15, 654 (2024).
Hemdan, E. E.-D., Shouman, M. A. & Karar, M. E. Covidx-net: A framework of deep learning classifiers to diagnose covid-19 in x-ray images. arXiv preprint arXiv:2003.11055 (2020).
Stein, A. et al. Rsna pneumonia detection challenge https://kaggle.com/competitions/rsna-pneumonia-detection-challenge (2018).
Zawacki, A. et al. Siim-acr pneumothorax segmentation https://kaggle.com/competitions/siim-acr-pneumothorax-segmentation (2019).
Jaeger, S. et al. Two public chest x-ray datasets for computer-aided screening of pulmonary diseases. Quant. Imaging Med. Surg. 4, 475 (2014).
Zhang, Y. Lung segmentation with nasnet-large-decoder net. arXiv preprint arXiv:2303.10315 (2023).
Wang, F., Zhou, Y., Wang, S., Vardhanabhuti, V. & Yu, L. Multi-granularity cross-modal alignment for generalized medical visual representation learning. Adv. Neural Inf. Process Syst. 35, 33536–33549 (2022).
Boecking, B. et al. Making the most of text semantics to improve biomedical vision–language processing. In European Conference on Computer Vision, 1–21 (Springer, 2022).
Wu, C., Zhang, X., Zhang, Y., Wang, Y. & Xie, W. Medklip: Medical knowledge enhanced language-image pre-training. Proceedings of the IEEE/CVF International Conference on Computer Vision (IEEE, 2023).
Zhang, Y., Jiang, H., Miura, Y., Manning, C. D. & Langlotz, C. P. Contrastive learning of medical visual representations from paired images and text. In Machine Learning for Healthcare Conference, 2–25 (PMLR, 2022).
Huang, S.-C., Shen, L., Lungren, M. P. & Yeung, S. Gloria: A multimodal global-local representation learning framework for label-efficient medical image recognition. In Proceedings of the IEEE/CVF International Conference on Computer Vision, 3942–3951 (IEEE, 2021).
Zhou, L., Liu, H., Bae, J., He, J., Samaras, D. & Prasanna, P. Self pre-training with masked autoencoders for medical image classification and segmentation. In IEEE International Symposium on Biomedical Imaging, 1–6 (IEEE, 2023).
Xiao, J., Bai, Y., Yuille, A. & Zhou, Z. Delving into masked autoencoders for multi-label thorax disease classification. In Proc. IEEE/CVF Winter Conference on Applications of Computer Vision, 3588–3600 (IEEE, 2023).
Dosovitskiy, A. et al. An image is worth 16 × 16 words: Transformers for image recognition at scale. In International Conference on Learning Representations (2021).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE conference on computer vision and pattern recognition, 770–778 (IEEE, 2016).
Huang, G., Liu, Z., Van Der Maaten, L. & Weinberger, K. Q. Densely connected convolutional networks. In Proc. IEEE Conference on Computer Vision and Pattern Recognition, 4700–4708 (IEEE, 2017).
Fang, Y. et al. Eva-02: A visual representation for neon genesis. Image Vis. Computing 149, 105171 (2024).
Tayebi Arasteh, S., Misera, L., Kather, J. N., Truhn, D. & Nebelung, S. Enhancing diagnostic deep learning via self-supervised pretraining on large-scale, unlabeled non-medical images. Eur. Radiol. Exp. 8, 10 (2024).
He, K. et al. Masked autoencoders are scalable vision learners. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 16000–16009 (2022).
Chen, X., Fan, H., Girshick, R. & He, K. Improved baselines with momentum contrastive learning. arXiv preprint arXiv:2003.04297 (2020).
Guan, Q. & Huang, Y. Multi-label chest x-ray image classification via category-wise residual attention learning. Pattern Recognit. Lett. 130, 259–266 (2020).
Ma, C., Wang, H. & Hoi, S. C. Multi-label thoracic disease image classification with cross-attention networks. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2019: 22nd International Conference, Shenzhen, China, October 13–17, 2019, Proceedings, Part VI 22, 730–738 (Springer, 2019).
Haghighi, F., Taher, M. R. H., Gotway, M. B. & Liang, J. Dira: Discriminative, restorative, and adversarial learning for self-supervised medical image analysis. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 20824–20834 (IEEE, 2022).
Liu, F. et al. Acpl: Anti-curriculum pseudo-labelling for semi-supervised medical image classification. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 20697–20706 (IEEE, 2022).
Hermoza, R., Maicas, G., Nascimento, J. C. & Carneiro, G. Region proposals for saliency map refinement for weakly-supervised disease localisation and classification. In Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference, Lima, Peru, October 4–8, 2020, Proceedings, Part VI 23, 539–549 (Springer, 2020).
Kim, E., Kim, S., Seo, M. & Yoon, S. Xprotonet: diagnosis in chest radiography with global and local explanations. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 15719–15728 (IEEE, 021).
Seyyed-Kalantari, L., Liu, G., McDermott, M., Chen, I. Y. & Ghassemi, M. Chexclusion: Fairness gaps in deep chest x-ray classifiers. In BIOCOMPUTING 2021: proceedings of the Pacific symposium, 232–243 (World Scientific, 2020).
Pham, H. H., Le, T. T., Tran, D. Q., Ngo, D. T. & Nguyen, H. Q. Interpreting chest x-rays via cnns that exploit hierarchical disease dependencies and uncertainty labels. Neurocomputing 437, 186–194 (2021).
Touvron, H. et al. Training data-efficient image transformers & distillation through attention. In International conference on machine learning, 10347–10357 (PMLR, 2021).
Baselli, G., Codari, M. & Sardanelli, F. Opening the black box of machine learning in radiology: can the proximity of annotated cases be a way? Eur. Radiol. Exp. 4, 1–7 (2020).
Selvaraju, R. R. et al. Grad-cam: Visual explanations from deep networks via gradient-based localization. In Proc. IEEE International Conference on Computer Vision, 618–626 (IEEE, 2017).
Saporta, A. et al. Benchmarking saliency methods for chest x-ray interpretation. Nat. Mach. Intell. 4, 867–878 (2022).
Liu, A. et al. Deepseek-v3 technical report. arXiv preprint arXiv:2412.19437 (2024).
Glocker, B., Jones, C., Roschewitz, M. & Winzeck, S. Risk of bias in chest radiography deep learning foundation models. Radiol. Artif. Intell. 5, e230060 (2023).
Ma, D., Pang, J., Gotway, M. B. & Liang, J. A fully open AI foundation model applied to chest radiography. Nature 643, 488–498 (2025).
Xu, S. et al. Elixr: Towards a general purpose x-ray artificial intelligence system through alignment of large language models and radiology vision encoders. arXiv preprint arXiv:2308.01317 (2023).
Chen, Z. et al. Chexagent: Towards a foundation model for chest x-ray interpretation. arXiv preprint arXiv:2401.12208 (2024).
Thawakar, O. C. et al. Xraygpt: Chest radiographs summarization using large medical vision-language models. In Proc. 23rd Workshop on Biomedical Natural Language Processing, 440–448 (IEEE, 2024).
Deng, J. et al. Imagenet: A large-scale hierarchical image database. In Proc. IEEE Conference on Computer Vision and Pattern Recognition, 248–255 (IEEE, 2009).
Fang, Y. et al. Eva: Exploring the limits of masked visual representation learning at scale. In Proc. of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 19358–19369 (IEEE, 2023).
Çallí, E., Sogancioglu, E., van Ginneken, B., van Leeuwen, K. G. & Murphy, K. Deep learning for chest x-ray analysis: a survey. Med. Image Anal. 72, 102125 (2021).
Wang, H. et al. Foundation transformers. arXiv preprint arXiv:2210.06423 (2022).
Shazeer, N. Glu variants improve transformer. arXiv preprint arXiv:2002.05202 (2020).
Sun, Q., Fang, Y., Wu, L., Wang, X. & Cao, Y. Eva-clip: improved training techniques for clip at scale. arXiv preprint arXiv:2303.15389 (2023).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, October 5-9, 2015, Proceedings, Part III 18, 234–241 (Springer, 2015).
Li, Y., Mao, H., Girshick, R. & He, K. Exploring plain vision transformer backbones for object detection. In European Conference on Computer Vision, 280–296 (Springer, 2022).
Xiao, T., Liu, Y., Zhou, B., Jiang, Y. & Sun, J. Unified perceptual parsing for scene understanding. In Proceedings of the European conference on computer vision (ECCV), 418–434 (ECCV, 2018).
Acknowledgements
This work was supported by the National Key R &D Program of China (2022YFB4500602) and the National Natural Science Foundation of China (NSFC) under Grant No. 62276108.
Author information
Authors and Affiliations
Contributions
J.Y. collected the data, developed the methodology, developed codes, performed data analysis, and wrote the original draft. X.W. and B.W. conceptualized and designed the study, and supervised the research. Y.S. and H.Z. prepared the figures and tables and provided critical revision of the article. J.M. performed the technical validation. Y.C. and W.L. performed statistical analysis. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yao, J., Wang, X., Song, Y. et al. EVA-X: a foundation model for general chest x-ray analysis with self-supervised learning. npj Digit. Med. 8, 678 (2025). https://doi.org/10.1038/s41746-025-02032-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02032-z