Abstract
This study presents an ensemble learning approach for automated screening and severity classification of chronic kidney disease (CKD) using polysomnographic (PSG) phenotypes. We analyzed PSG data from 358 subjects (179 CKD, 179 early-CKD) in the Cleveland Family Study using four ensemble algorithms: Random Forest, XGBoost, LightGBM, and CatBoost. A total of 1210 sleep-related variables were extracted, covering respiration, sleep stages, movement, and cardiovascular features. The models achieved high multiclass classification performance, with AUCs exceeding 89% across CKD stages. Feature importance analysis revealed that disruptions in oxygen saturation, sleep architecture, and heart rate variability were closely associated with CKD severity. These findings highlight the potential of PSG-derived phenotypes combined with ensemble learning for early CKD detection and risk stratification, supporting timely intervention and improved patient management.
Similar content being viewed by others
Introduction
Chronic kidney disease (CKD) is a progressive condition affecting approximately 10% of the global population, with increasing morbidity and mortality rates1. CKD is characterized by a gradual decline in renal function, typically assessed through estimated glomerular filtration rate (GFR) and biomarkers such as serum creatinine and cystatin C levels. As the disease advances, CKD can progress to end-stage kidney disease (ESKD), requiring dialysis or kidney transplantation2. Comorbidities such as anemia, cardiovascular disease, and metabolic dysfunction often accompany CKD, further complicating disease management3. Early detection remains a significant challenge, leading to late-stage diagnosis and limited treatment options. Therefore, there is a critical need for non-invasive screening methods to enable earlier diagnosis and timely interventions.
Emerging research highlights a strong correlation between sleep disorders and CKD4. Conditions such as insomnia, obstructive sleep apnea, and restless leg syndrome are not only risk factors for CKD onset but also indicators of disease severity. Turek et al. 5 reported that sleep disturbances contribute to CKD development and accelerate its progression. Nigam et al. 6 highlighted the high prevalence of sleep disorders among CKD patients, mainly sleep apnea and insomnia, and their association with poorer health outcomes. Similarly, Pierratos et al. 7 observed that sleep disturbances are common in patients with ESKD, and their treatment can improve quality of life and reduce mortality risk. Studies suggest that sleep disorders contribute to CKD pathophysiology through mechanisms such as hypoxia, inflammation, and autonomic dysfunction8. Despite growing evidence, the potential of sleep-derived biomarkers for CKD classification and screening remains underexplored9.
Despite growing interest in the relationship between sleep disorders and CKD outcomes, prior studies have been limited in both scope and methodology. Much of the existing research has relied on self-reported sleep symptoms or single-channel data (e.g., pulse oximetry), which offer limited physiological depth. Furthermore, few studies have utilized full polysomnography (PSG) datasets or employed advanced machine learning techniques capable of capturing complex, high-dimensional phenotypic patterns. Issues related to model interpretability and generalizability across diverse populations have also been largely overlooked. These gaps highlight the need for a comprehensive and explainable AI framework leveraging PSG-derived features for the classification of CKD severity.
Polysomnography (PSG) is a gold-standard diagnostic tool for evaluating sleep disorders and capturing respiratory indices, sleep architecture, and cardiovascular signals10. Although clinicians traditionally use PSG to diagnose sleep-related breathing disorders, its ability to provide comprehensive physiological data offers an opportunity for non-invasive CKD screening. Previous studies have linked PSG-derived features to cardiovascular risk, yet their potential role in CKD detection remains underexplored11. Despite this, researchers have not yet fully investigated the application of PSG for CKD screening. We hypothesize that PSG-derived phenotypic data, when analyzed using machine learning models, can serve as an effective screening tool for classifying CKD severity.
Recent advancements in machine learning (ML) provide powerful tools for analyzing complex physiological datasets. Ensemble learning (EL)12 is a robust ML approach that enhances predictive accuracy by combining multiple base models13. It includes three primary methods: bagging14, boosting15, and stacking16. Bagging techniques, Random forest (RF)17, improve model stability by averaging predictions from multiple decision trees. Boosting algorithms such as Extreme Gradient Boosting (XGBoost)18, Light Gradient Boosting Machine (LightGBM)19, and Categorical Boosting (CatBoost)20 iteratively refine predictions to enhance accuracy. These algorithms have demonstrated strong performance in medical applications, particularly in handling heterogeneous biomedical data21.
Recent advancements in ML, few studies have developed comprehensive and interpretable models that utilize PSG-derived features for CKD severity classification. We hypothesize that PSG-derived phenotypic features encode clinically meaningful signals to address this gap. These signals may support accurate, non-invasive classification of CKD stages and help identify relevant sleep-related features when analyzed through ensemble learning algorithms22. Accordingly, the objective of the present study is to classify CKD severity based on sleep pattern features using an EL-based framework incorporating models such as RF, XGBoost, LightGBM, and CatBoost. This approach aims to enhance screening accuracy, facilitate earlier detection of CKD, and support personalized interventions to slow disease progression.
Results
Our findings revealed a statistically significant difference in gender distribution (p < 0.05), with a higher proportion of females in the CKD group (66.5%) compared to the early-CKD group (51.4%) (Table 1). To minimize potential bias from this imbalance, propensity score matching was performed based on sex, age, and BMI to ensure comparability between the groups. After matching, no statistically significant differences were observed in other demographic and clinical variables, including age, BMI, sleep efficiency, and total sleep time (p > 0.01). While the CKD group exhibited a slightly higher apnea-hypopnea index (AHI) and heart rate than the early-CKD group, these differences were not statistically significant (p > 0.01). Similarly, systolic and diastolic blood pressure values showed no significant variation between the groups. A higher proportion of CKD patients exhibited moderate to severe sleep apnea (AHI ≥ 15 events/hour), although the mean AHI did not differ significantly between groups (5.8 ± 13.7 vs. 4.1 ± 11.8, p = 0.200). Similarly, differences in sleep efficiency and total sleep time between the CKD and early-CKD groups were not statistically significant (p = 0.761 and p = 0.239, respectively). Notably, CPAP therapy was more frequently prescribed among CKD groups, suggesting a greater clinical burden of sleep-disordered breathing. These patterns align with our model’s feature importance analysis, reinforcing the relevance of sleep apnea-related markers in CKD severity classification. The prevalence of diabetes was higher in the CKD group (27.9%) compared to the early-CKD group (18.4%), but this difference did not reach statistical significance (p > 0.01). Among laboratory markers, urinary creatinine levels were significantly elevated in the early-CKD group (125.7 ± 67.4 mg/dL) compared to the CKD group (100.2 ± 63.9 mg/dL, p < 0.001), indicating impaired renal function. However, other metabolic parameters, including total cholesterol, fasting glucose, high-density lipoprotein, and low-density lipoprotein, showed no significant differences between the groups.
We utilized four ensemble learning algorithms to develop a robust predictive model for CKD detection: RF (Supplementary Fig. 1), XGBoost (Supplementary Fig. 2), LightGBM (Supplementary Fig. 3), and CatBoost (Supplementary Fig. 4). The feature importance of the best-performing model, XGBoost, is listed in Table 2, highlighting the most influential factors in CKD classification.
Our analysis revealed that CKD patients exhibited a significantly higher frequency of respiratory-related events, including apneas, hypopneas, and respiratory effort-related arousals (RERAs), often accompanied by oxygen desaturation. A notable proportion of CKD patients experienced severe sleep apnea and frequently required continuous positive airway pressure (CPAP) therapy during sleep. These findings emphasize the strong link between sleep-disordered breathing and CKD progression. Key lifestyle and physiological parameters also played a critical role in model performance. The most influential variables were time since the last cigarette, apnea event durations, and PSG signal quality, all contributing significantly to the model’s predictive accuracy. These insights reinforce the importance of integrating sleep-related biomarkers into CKD risk assessment models (Fig. 1).
a Random Forest (RF), b XGBoost, c LightGBM, and d CatBoost. Each plot highlights the most influential categorical variables, ranked by their contribution to model predictions. These features are critical in CKD classification, providing insights into key phenotypic markers associated with disease severity.
We compared the performance of the four ensemble learning approaches across CKD severity classification with AUC values (Fig. 2). All models demonstrated high AUC values exceeding 85%, with severe CKD stages (stages 4–5) achieving AUC values above 89%, reinforcing strong predictive accuracy. Among the evaluated models, XGBoost outperformed the others, achieving an AUC of 0.95 for stage 2 CKD and near-perfect classification for stages 3–5. In stage 1, XGBoost had a slightly lower AUC of 0.77 but outperformed CatBoost and performed comparably to RF.
The models evaluated include. a Random Forest, b XGBoost, c LightGBM, and d CatBoost. These ROC curves illustrate the classification performance across different CKD stages, comparing original and oversampled PSG-derived data results.
We employed ensemble learning models to examine underlying phenotypic associations within the PSG-derived dataset, followed by a comparative evaluation of algorithmically diverse classifiers— RF (Supplementary Table 1), XGBoost (Supplementary Table 2), LightGBM (Supplementary Table 3), and CatBoost (Supplementary Table 4). This approach aims to enhance predictive performance by leveraging the strengths of each model. Consistent with prior evidence in biomedical classification, our findings (Table 3) demonstrate the effectiveness of ensemble methods in accurately identifying CKD severity based on sleep phenotypes.
We conducted external validation using an independent, large-scale sleep dataset (HCHS/SOL) containing CKD severity stages and PSG-derived phenotypic features to assess the generalizability of the proposed approaches (Fig. 3). The external validation dataset (1,016 participants) demonstrated ROC curve patterns and performance similar to those observed in the original model developed for multiclass classification of CKD stages 1 through 5. Notably, the validation results showed near-perfect discriminative performance for CKD stages 3b, 4, and 5, with AUC values of 0.97, 0.98, and 1.00, respectively. Overall, the proposed model exhibited strong predictive accuracy, particularly in identifying advanced CKD stages, confirming its potential robustness and applicability across diverse clinical settings.
The models evaluated include (A) random Forest, B XGBoost, C LightGBM, and D CatBoost. The area under the curve (AUC) values represent the multiclass classification performance for CKD stages: stage 1 (STG1), stage 2 (STG2), stage 3a (STG3a), stage 3b (STG3b), stage 4 (STG4), and stage 5 (STG5).
Lastly, due to the imbalanced nature of the CKD severity dataset, we have additionally provided the precision–recall curve (Supplementary Figs. 5, 6) and the area under the precision–recall curve (Supplementary Table 5, 6) as supplementary materials.
Discussion
This study presents an AI-driven approach for non-invasive CKD severity classification using PSG-derived phenotypic data. Our findings support the hypothesis that when analyzed using ML models, PSG-derived features can effectively classify CKD severity. Among the ensemble learning models evaluated (RF, XGBoost, LightGBM, and CatBoost), RF demonstrated the highest classification performance, achieving an average accuracy of 80%. In addition, significant 15 features were found for CKD severity classification based on PSG recordings. These results highlight the potential of nocturnal sleep phenotypes as biomarkers for CKD progression, offering a promising avenue for early detection and disease monitoring.
These findings support the initial hypothesis and emphasize the relevance of PSG-derived features in the non-invasive classification of CKD severity. An increasing body of evidence suggests a strong association between sleep-disordered breathing and CKD progression8. Physiological disturbances captured through PSG—such as recurrent nocturnal hypoxia, elevated arousal frequency, and altered autonomic regulation, reflect systemic stress responses that may contribute to renal dysfunction23. These features not only serve as potential indicators of CKD severity but may also play a mechanistic role in disease progression. Thus, leveraging PSG-derived phenotypes presents a biologically grounded and non-invasive approach to improve CKD staging and risk stratification.
Integrating EL-driven CKD screening into clinical workflows has significant potential for improving early detection, risk assessment, and personalized disease management24. Given that PSG is already widely used in sleep medicine, leveraging ML models to analyze PSG-derived features could enable CKD risk identification without requiring additional invasive testing25. Primary care providers and nephrologists could utilize AI-generated PSG risk scores as an early warning system, prompting further renal function assessments, lifestyle modifications, or early interventions. Early-stage CKD detection through nocturnal phenotypic patterns may facilitate timely medical interventions, potentially slowing disease progression and reducing the risk of cardiovascular complications26. ML-based PSG analysis could be incorporated into wearable sleep monitors and telehealth platforms, expanding access to CKD screening, particularly for individuals who do not undergo routine kidney function tests. Moreover, multimodal ML frameworks integrating PSG features with biochemical markers, such as serum creatinine, albuminuria, and cystatin C, could further enhance risk prediction and patient stratification27. These applications underscore the broader impact of PSG-derived EL models in chronic disease management, bridging the gap between sleep medicine and nephrology.
Traditional CKD diagnosis relies on serum creatinine, estimated GFR (eGFR), and albuminuria, which require blood and urine tests that may not be performed regularly, especially in asymptomatic individuals28. Our AI-based approach introduces a potentially more accessible, non-invasive screening method by extracting relevant physiological features from PSG recordings. The advantages of PSG-derived EL models for CKD detection include non-invasiveness, early disease-related physiological changes, integration with existing sleep diagnostics, and potential scalability for home-based screening using ML-powered wearable sleep monitors. While PSG offers rich physiological information29, we acknowledge that it is more time-consuming, costly, and less accessible than standard clinical tests such as eGFR. Therefore, our approach is not intended to replace current clinical practices but to explore the potential of PSG-derived features as a complementary screening tool for CKD—particularly in patients who are already undergoing PSG for sleep-related disorders. In such contexts, repurposing existing PSG data could enable early risk stratification of CKD without the need for additional testing or procedures. We acknowledge that the routine use of PSG for CKD screening is not currently feasible due to cost and accessibility constraints. However, our study aims to assess the feasibility and value of such data in a targeted population, which may inform future work involving more scalable alternatives, such as wearable devices. Importantly, these findings demonstrate that machine learning models can extract clinically relevant signals from PSG data. That said, we agree that further external validation and prospective studies are essential before this approach can be considered for broader clinical application.
Despite its promising findings, this study has several limitations. Firstly, although oversampling techniques were initially applied, residual class imbalance remained, which may have caused model performance to be overestimated. Therefore, future research should recruit larger and more balanced study populations across all CKD severity levels (mild, moderate, and severe) or employ advanced resampling strategies to further mitigate class imbalance. Second, our model relied solely on PSG-derived phenotypic features and did not incorporate key biochemical markers such as serum creatinine, albuminuria, and cystatin C, which are fundamental for CKD diagnosis and monitoring. Including such markers in multimodal models could enhance both prediction accuracy and clinical relevance. Finally, using PSG data may introduce variability due to differences in sleep study protocols, equipment, and scoring standards between sleep centers.
To further refine and expand the applicability of EL-driven CKD screening, future research should focus on developing multimodal AI frameworks that combine PSG-derived features with clinical, biochemical, and genetic data to enhance risk prediction models. Investigating the feasibility of home-based CKD screening using AI-powered sleep-tracking technologies could provide a scalable solution for early detection. Additionally, longitudinal studies should assess how sleep phenotypes evolve and their relationship to CKD progression. Validating EL models in real-world clinical settings will be essential to ensure their integration into routine nephrology practice.
This study introduces a novel EL-driven approach for CKD severity classification using PSG-derived phenotypic data. Our findings confirm that nocturnal sleep phenotypes contain valuable biomarkers for CKD screening, supporting the hypothesis that PSG-derived features, analyzed with machine learning models, can effectively classify CKD severity. By leveraging ensemble learning techniques, we demonstrate the feasibility of non-invasive EL-based CKD detection, potentially revolutionizing early disease identification. Future research should focus on expanding dataset diversity, integrating multimodal biomarkers, and validating EL models in clinical settings to ensure broad applicability and clinical adoption. If successfully implemented, EL-driven CKD screening could improve early diagnosis, personalized treatment strategies, and long-term patient outcomes, bridging the gap between sleep medicine and nephrology for a more holistic approach to chronic disease management. Additionally, prospective studies should explore the integration of EL-driven CKD screening into real-world telehealth platforms and sleep clinics to evaluate its feasibility, usability, and clinical utility.
Methods
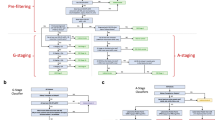
We proposed ensemble learning approaches for automatically screening CKD using PSG-derived phenotypic data. Our study design comprised four key components: study population (Fig. 4a), clinical phenotypic data (Fig. 4b), ensemble learning models (Fig. 4c), and CKD outcomes (Fig. 4d). We implemented four ensemble learning algorithms—RF, XGBoost, LightGBM, and CatBoost—to evaluate the robustness and generalizability of top-ranked nocturnal features associated with CKD severity. These models were chosen for their demonstrated effectiveness in classifying chronic disease severity, particularly in the context of high-dimensional clinical datasets. Each algorithm employs distinct optimization strategies, enabling a comparative analysis of their performance across different ensemble learning paradigms. Detailed descriptions of each model’s architecture and the rationale behind their selection are provided in the Ensemble Learning Models section.
a Study population: This study utilized the Cleveland Family Study data. b Clinical phenotypic data: Nocturnal phenotypic data were extracted and processed for ensemble learning. c Ensemble learning: Four representative algorithms were used to screen CKD stages during sleep. d CKD outcomes: Automatic screening of CKD stages was classified as a multiclass classification into stage 1 (STG1), stage 2 (STG2), stage 3 (STG3a), stage 3 (STG3b), stage 4 (STG4), and stage 5 (STG5).
Study population
To validate our approach, we utilized the Cleveland Family Study (CFS) data, a large, family-based cohort that provides extensive PSG-derived phenotypic and clinical data. The dataset includes 2284 individuals from 361 families, primarily of American and African American descent, recruited initially to investigate the familial aggregation of sleep apnea. Longitudinal data collection occurred between 1990 and 2006 through four follow-up examinations. Additionally, the Hispanic Community Health Study/Study of Latinos (HCHS/SOL) dataset was used for independent external validation. HCHS/SOL is a multi-center epidemiologic study that includes 16,415 Hispanic and Latino participants.
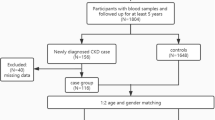
For this analysis, participants were included if they had available PSG recordings, clinical data, and valid eGFR stage classifications. Individuals were excluded if they (1) lacked follow-up data (n = 1549), (2) had missing or insufficient eGFR information (n = 13), or (3) were unmatched in propensity score matching (n = 364). We did not apply any additional exclusion criteria based on specific clinical comorbidities or anatomical abnormalities, as our goal was to capture a broadly representative sample of early CKD and CKD individuals from the dataset (Fig. 5). We included 358 participants after applying exclusion criteria. Initially, 1549 individuals were excluded due to missing follow-up data, and 13 were excluded due to insufficient eGFR stage records. CKD classification was based on eGFR cutoffs, defined as follows:
-
Stage 1 (STG1): eGFR > 90 mL/min/1.73 m²
-
Stage 2 (STG2): eGFR 60–89 mL/min/1.73 m²
-
Stage 3a (STG3a): eGFR 45–59 mL/min/1.73 m²
-
Stage 3b (STG3b): eGFR 30–44 mL/min/1.73 m²
-
Stage 4 (STG4): eGFR 15–29 mL/min/1.73 m²
-
Stage 5 (STG5): eGFR < 15 mL/min/1.73 m²
Detailed flowchart illustrating the subject enrollment process and dataset construction for automated screening of CKD severity.
To minimize confounding effects, we applied propensity score matching using a nearest-neighbor algorithm (1:1 ratio) without replacement based on age, sex, and body mass index (BMI). Standardized mean differences were calculated before and after matching to assess the balance between the CKD and control groups. After matching, we obtained 179 participants with CKD and 179 early-CKD, ensuring well-balanced cohorts and reducing selection bias (Fig. 5).
The final cohort consisted of 58.9% female participants, with an average age of 52.0 ± 18.2 years. BMI levels were similar between groups (CKD: 35.5 ± 11.0 kg/m² vs. Control: 34.8 ± 10.4 kg/m², p = 0.503). Comorbid conditions such as diabetes were more prevalent in the CKD group (27.9%) compared to controls (18.4%), although this difference did not reach statistical significance. Among laboratory markers, creatinine urine levels were significantly elevated in the CKD group (p < 0.001), consistent with impaired renal function. However, other metabolic parameters showed no significant differences between groups, including total cholesterol, fasting glucose, high-density lipoprotein, and low-density lipoprotein (Table 1).
Clinical phenotypic data
We extracted 1210 clinical phenotypic features from PSG recordings of 358 participants to facilitate the automatic screening and classification of CKD severity. These features were categorized into 13 groups, including respiratory indices, apnea-hypopnea parameters, oxygen saturation levels, limb movement patterns, sleep architecture, and signal quality (Table 4).
To optimize model performance and reduce overfitting, we implemented a structured, multi-step feature selection strategy. Initially, features with low variance or high missingness ( > 20%) were removed. Subsequently, univariate statistical tests (chi-square) were conducted within the training set to identify features significantly associated with CKD stages (p < 0.05). Finally, tree-based model importance scores and recursive feature elimination with cross-validation were employed to retain the most informative variables. This approach yielded a refined and discriminative subset of features used as model inputs.
For robust model evaluation, we split the dataset into training (70%), validation (20%), and test (10%) sets. We applied subject-based partitioning to prevent data leakage and maintain independence between the training and test datasets. To address class imbalance, we applied an oversampling strategy to the severe-stage CKD group using the Synthetic Minority Oversampling Technique (SMOTE). Specifically, the number of severe-stage CKD samples was increased from 5 to 28 by generating synthetic instances. This augmentation resulted in a more balanced dataset across CKD severity levels, particularly improving representation for the severe-stage group relative to the early-stage CKD subjects.
Ensemble learning models
In this study, the most influential factors related to CKD were identified, trained, and evaluated using ML techniques to analyze the significance of key parameters. Specifically, we utilized four ensemble learning models30: RF, XGBoost, LightGBM, and CatBoost. These models were applied independently to represent distinct ensemble learning strategies, RF as a bagging-based method, and XGBoost, LightGBM, and CatBoost as gradient boosting techniques. No stacking or meta-ensemble strategy was implemented; rather, model performance was compared to assess the relative effectiveness of these approaches in CKD classification31.
To address the class imbalance across CKD stages, particularly the limited representation in advanced stages, resampling techniques such as SMOTE were avoided to prevent the introduction of synthetic bias. Instead, model evaluation focused on assessing true performance under naturally imbalanced conditions. Classification results for each CKD stage were transparently reported using confusion matrices.
The RF algorithm constructs multiple decision trees and combines their predictions to enhance accuracy. It follows a bootstrap aggregating (bagging) approach, where random subsets of the dataset are sampled with replacement to train individual trees. Approximately 63% of the original observations appear in at least one bootstrap sample, while the remaining data points serve as out-of-bag samples for internal validation. This technique helps assess model performance and reduces overfitting. The final prediction is obtained by averaging or voting across all trees, ensuring a robust classification approach17.
where fi(x) represents individual decision trees in the ensemble.
XGBoost is an optimized gradient-boosting framework designed for computational efficiency and high-performance learning32. It enhances traditional decision tree-based models by employing a sequential learning approach, where weak learners are iteratively trained to minimize residual errors. XGBoost leverages a second-order Taylor expansion for loss function approximation to improve optimization, allowing for more precise gradient calculations and rapid convergence33.
A key advantage of XGBoost is its ability to prevent overfitting through built-in regularization techniques, which help optimize weight distributions across leaf nodes34. Additionally, the model supports parallel processing, enabling fast execution on large datasets while maintaining high predictive accuracy. These characteristics make XGBoost highly effective for classification and regression tasks, particularly in scenarios requiring scalability and computational efficiency35. The fundamental objective function governing XGBoost’s training process is formulated as follows36:
where l(yi,\({\hat{y}}_{i}\)) represents the loss function, ft(xi) denotes the newly added tree, and Ω(ft) is the regularization term that controls model complexity.
LightGBM is a high-performance ensemble learning algorithm for efficient learning on large-scale datasets. Unlike conventional gradient-boosting decision trees (GBDT) that employ a level-wise growth approach, LightGBM adopts a leaf-wise growth strategy to enhance learning efficiency. This method effectively reduces residual errors and minimizes unnecessary node splits, improving learning performance. However, a leaf-wise approach may result in imbalanced tree structures, where specific nodes are split more frequently than others, increasing the risk of overfitting. LightGBM introduces Gradient-based One-Sided Sampling (GOSS) and Exclusive Feature Bundling (EFB) to address this and enhance learning efficiency. GOSS mitigates overfitting by selectively retaining high-gradient data points, contributing significantly to the loss function while sampling only a subset of low-gradient data points. Meanwhile, EFB groups mutually exclusive features into a single unit for processing, improving memory efficiency and computational speed37.
Additionally, LightGBM employs a depth-first growth strategy to generate asymmetric trees, prioritizing the expansion of nodes that offer the greatest improvement in split performance38. As a result, the model delivers faster processing speeds compared to traditional breadth-first growth approaches and ensures high accuracy and efficiency even on large-scale datasets39. The operational mechanism of LightGBM can be further elaborated through Eq. (3) 40.
CatBoost is an advanced gradient-boosting algorithm specifically designed to process categorical data41. Unlike conventional ML models that require extensive preprocessing, such as one-hot encoding or label encoding, CatBoost incorporates automated statistical transformations to convert categorical variables into formats optimized for training42. This built-in feature streamlines data preparation and minimizes the risk of information loss during encoding. CatBoost employs an Ordered Boosting technique to enhance learning efficiency further, ensuring that the model does not leak information from future observations into the training process. This approach mitigates data leakage, improving predictive stability and reducing overfitting, particularly in small datasets. The method maintains strong generalization performance, making it highly effective for structured data applications43.
Additionally, CatBoost supports parallel processing using GPUs, enabling efficient training even on large-scale datasets. It achieves faster training speeds by leveraging GPU-based histogram computation methods than traditional approaches like XGBoost and LightGBM44. This parallel processing capability makes CatBoost well-suited for real-time analytics and large-scale data processing where high performance is essential45.
OTS(xcat,y) represents the ordered target statistics applied to categorical variables.
Model training
PSG data were processed using R (version 4.3.1; R Foundation for Statistical Computing, Vienna, Austria)46 and Python (version 3.11.4; Python Software Foundation, Fredericksburg, VA, USA)47. The proposed EL models were implemented, and the software and hardware configurations were optimized for training. The computing environment consisted of Keras48 running on a TensorFlow backend, with a workstation powered by an Intel i9-13900K CPU (3.5 GHz) and an NVIDIA GeForce RTX 4070 GPU.
The dataset was partitioned into training and testing subsets using the train_test_split function from the sci-kit-learn library, allocating 70% of the data for training and 30% for validation. The data were divided into 64 mini-batches sequentially injected into the model throughout each training epoch to enhance model learning. Receiver Operating Characteristic (ROC) curves were generated to evaluate model performance, while feature importance analysis was conducted to identify key predictors influencing classification accuracy.
Evaluation metrics
We computed accuracy, sensitivity (recall), specificity, and precision to assess model performance. These evaluation measures were defined as follows:
TP represents true positives, TN denotes true negatives, FP refers to false positives, and FN indicates false negatives. These terms correspond to the number of correctly detected normal and abnormal events and the number of misclassified events in the dataset. Additionally, we computed the F1 score, a metric beneficial for handling imbalanced datasets, as it balances precision and recall using the formula:
The recall metric measures how effectively the model detects true positive (TP) cases, whereas precision evaluates how accurately normal cases are classified. Higher values for both metrics, approaching 1, indicate improved reliability49. The ROC curve visualizes the trade-off between False Positive rate (FPR, x-axis) and True Positive rate (TPR, y-axis) across different threshold values. The area under the curve (AUC) quantifies overall model performance, with values closer to 1.0 indicating higher classification accuracy 50.
Data availability
All data were approved by the National Sleep Research Resource (NSRR) for the specific purpose of this study ([https://sleepdata.org/datasets/cfs], [https://sleepdata.org/datasets/hchs]).
References
González, S., García, S., Del Ser, J., Rokach, L. & Herrera, F. A practical tutorial on bagging and boosting based ensembles for machine learning: Algorithms, software tools, performance study, practical perspectives and opportunities. Inf. Fusion 64, 205–237 (2020).
Gupta, R., Woo, K. & Yi, J. A. Epidemiology of end-stage kidney disease. Semin. Vasc. Surg. 34, 71–78 (2021).
Fu, X., Liu, H., Bi, X. & Gong, X. Deep-learning-based CT imaging in the quantitative evaluation of chronic kidney diseases. J. Healthc. Eng. 2021, 3774423 (2021).
Kelly, D. M. & Rothwell, P. M. Disentangling the relationship between chronic kidney disease and cognitive disorders. Front. Neurol. 13, 830064 (2022).
Turek, N. F., Ricardo, A. C. & Lash, J. P. Sleep disturbances as nontraditional risk factors for development and progression of CKD: Review of the evidence. Am. J. Kidney Dis. 60, 823–833 (2012).
Nigam, G., Camacho, M., Chang, E. T. & Riaz, M. Exploring sleep disorders in patients with chronic kidney disease. Nat. Sci. Sleep. 10, 35–43 (2018).
Pierratos, A. & Hanly, P. J. Sleep disorders over the full range of chronic kidney disease. Blood Purif. 31, 146–150 (2011).
Lyons, O. D. Sleep disorders in chronic kidney disease. Nat. Rev. Nephrol. 20, 690–700 (2024).
Gogoi, P. & Valan, J. A. Machine learning approaches for predicting and diagnosing chronic kidney disease: current trends, challenges, solutions, and future directions. Int. Urol. Nephrol. 57, 1245–1268 (2025).
Ogna, A. et al. Sleep characteristics in early stages of chronic kidney disease in the HypnoLaus cohort. Sleep 39, 945–953 (2016).
Lee, P.-L. et al. Recommended assessment and management of sleep disordered breathing in patients with atrial fibrillation, hypertension and heart failure: Taiwan Society of Cardiology/Taiwan Society of Sleep Medicine/Taiwan Society of Pulmonary and Critical Care Medicine joint consensus statement. J. Formos. Med. Assoc. 123, 159–178 (2024).
Sagi, O. & Rokach, L. Ensemble learning: A survey. WIREs Data Min. Knowl. Discov. 8, e1249 (2018).
Dietterich, T. G. Ensemble methods in machine learning. In Multiple Classifier Systems (eds. Kittler, J. & Roli, F.) 1–15 (Springer, https://doi.org/10.1007/3-540-45014-9_1. 2000).
Salman, H. A., Kalakech, A. & Steiti, A. Random forest algorithm overview. Babylon. J. Mach. Learn. 2024, 69–79 (2024).
Mayr, A., Binder, H., Gefeller, O. & Schmid, M. The evolution of boosting algorithms. Methods Inf. Med. 53, 419–427 (2014).
Pavlyshenko, B. Using stacking approaches for machine learning models. In 2018 IEEE Second International Conference on Data Stream Mining & Processing (DSMP) 255–258 (IEEE, https://doi.org/10.1109/DSMP.2018.8478487. 2018).
Rigatti, S. J. Random forest. J. Insur. Med. 47, 31–39 (2017).
Chen, T. et al. Xgboost: extreme gradient boosting. R package version 0.4-2, 1–4 (2015).
Saito, H. et al. Predicting CKD progression using time-series clustering and light gradient boosting machines. Sci. Rep. 14, 1723 (2024).
Ganie, S. M., Dutta Pramanik, P. K., Mallik, S. & Zhao, Z. Chronic kidney disease prediction using boosting techniques based on clinical parameters. PLoS One 18, e0295234 (2023).
Jongbo, O. A., Adetunmbi, A. O., Ogunrinde, R. B. & Badeji-Ajisafe, B. Development of an ensemble approach to chronic kidney disease diagnosis. Sci. Afr. 8, e00456 (2020).
Ganaie, M. A., Hu, M., Malik, A. K., Tanveer, M. & Suganthan, P. N. Ensemble deep learning: A review. Eng. Appl. Artif. Intell. 115, 105151 (2022).
Fink, A. M., Bronas, U. G. & Calik, M. W. Autonomic regulation during sleep and wakefulness: a review with implications for defining the pathophysiology of neurological disorders. Clin. Auton. Res. 28, 509–518 (2018).
Ekundayo, F. Machine learning for chronic kidney disease progression modelling: Leveraging data science to optimize patient management. World J. Adv. Res. Rev. 24, 453–475 (2024).
Strassberger, C. Cardiopulmonary phenotyping during sleep—innovative signal processing for precision medicine in sleep apnea. https://hdl.handle.net/2077/84033 (2024).
Ruilope, L. M. et al. Prevention of cardiorenal damage: Importance of albuminuria. Eur. Heart J. 44, 1112–1123 (2023).
DeGroat, W. et al. Multimodal AI/ML for discovering novel biomarkers and predicting disease using multi-omics profiles of patients with cardiovascular diseases. Sci. Rep. 14, 26503 (2024).
Chen, T. K., Knicely, D. H. & Grams, M. E. Chronic kidney disease diagnosis and management: a review. JAMA 322, 1294–1304 (2019).
Roebuck, A. et al. A review of signals used in sleep analysis. Physiol. Meas. 35, R1–R57 (2013).
Friedman, J. H. Greedy function approximation: a gradient boosting machine. Ann. Stat. 29, 1189–1232 (2001).
Zhou, Z.-H. Ensemble Methods: Foundations and Algorithms. (CRC Press, 2025).
Hakkal, S. & Ait Lahcen, A. XGBoost to enhance learner performance prediction. Comput. Educ. Artif. Intell. 7, 100254 (2024).
Mohril, R. S., Solanki, B. S., Kulkarni, M. S. & Lad, B. K. XGBoost based residual life prediction in the presence of human error in maintenance. Neural Comput. Appl. 35, 3025–3039 (2023).
Dong, Y. et al. A data-driven model for predicting initial productivity of offshore directional well based on the physical constrained eXtreme gradient boosting (XGBoost) trees. J. Pet. Sci. Eng. 211, 110176 (2022).
Wu, J. et al. Prediction and screening model for products based on fusion regression and XGBoost classification. Comput. Intell. Neurosci. 2022, 4987639 (2022).
Ogunleye, A. & Wang, Q.-G. XGBoost model for chronic kidney disease diagnosis. IEEE/ACM Trans. Comput. Biol. Bioinform. 17, 2131–2140 (2019).
Zhang, D. & Gong, Y. The comparison of LightGBM and XGBoost coupling factor analysis and prediagnosis of acute liver failure. IEEE Access 8, 220990–221003 (2020).
Yan, J. et al. LightGBM: accelerated genomically designed crop breeding through ensemble learning. Genome Biol. 22, 1–24 (2021).
Ke, G. et al. LightGBM: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst. 30, 3146–3154 (2017). https://papers.nips.cc/paper/6907-lightgbm-a-highly-efficient-gradient-boosting-decision-tree.pdfNeurIPS.Papers+3BibSonomy+3NeurIPS.Papers+3.
Hancock, J. & Khoshgoftaar, T. M. Leveraging LightGBM for categorical big data. In 2021 IEEE Seventh International Conference on Big Data Computing Service and Applications (BigDataService), 149–154 (IEEE, https://doi.org/10.1109/BigDataService52369.2021.00024ResearchGate. 2021).
Prokhorenkova, L., Gusev, G., Vorobev, A., Dorogush, A. V. & Gulin, A. CatBoost: unbiased boosting with categorical features. Adv. Neural Inf. Process. Syst. 31, 6638–6648 (2018). https://papers.nips.cc/paper/7898-catboost-unbiased-boosting-with-categorical-features.pdfNeurIPS.Papers+1NeurIPS.Papers+1.
Ibrahim, A. A., Ridwan, R. L., Muhammed, M. M., Abdulaziz, R. O. & Saheed, G. A. Comparison of the CatBoost classifier with other machine learning methods. Int. J. Adv. Comput. Sci. Appl 11, 738–748 (2020).
Dorogush, A. V., Ershov, V. & Gulin, A. CatBoost: Gradient boosting with categorical features support. arXiv:1810.11363. https://doi.org/10.48550/arXiv.1810.11363 (2018).
Anghel, A., Papandreou, N., Parnell, T., De Palma, A. & Pozidis, H. Benchmarking and optimization of gradient boosting decision tree algorithms. arXiv:1809.04559. https://doi.org/10.48550/arXiv.1809.04559 (2018).
Li, L. Application of machine learning and data mining in medicine: Opportunities and considerations. In Machine Learning and Data Mining Annual Volume 2023 (IntechOpen, 2023).
Singh, D. & Garg, R. R., language for data analytics. In Proceedings of International Conference on Sustainable Computing in Science, Technology and Management (SUSCOM), Amity University Rajasthan, Jaipur-India (2019).
de Mendonça, F. M. C. Universidade do Algarve (Portugal) (2020).
Gulli, A. & Pal, S. Deep Learning with Keras. (Packt Publishing Ltd, 2017).
Chicco, D., Tötsch, N. & Jurman, G. The Matthews correlation coefficient (MCC) is more reliable than balanced accuracy, bookmaker informedness, and markedness in two-class confusion matrix evaluation. BioData Min. 14, 1–22 (2021).
Davis, J. & Goadrich, M. The relationship between Precision-Recall and ROC curves. In Proceedings of the 23rd International Conference on Machine Learning, 233–240. https://doi.org/10.1145/1143844.1143874 (2006).
Acknowledgements
This research was supported by a grant from the Medical Data-Driven Hospital Support Project through the Korea Health Information Service (KHIS), funded by the Ministry of Health & Welfare, Republic of Korea. The Basic Science Research Program supported this research through the National Research Foundation of Korea(NRF), funded by the Ministry of Education (RS-2023-00249148). This research was supported by the MSIT(Ministry of Science and ICT), Korea, under the National Program in SW (2024-0-00097) supervised by the IITP(Institute of Information Technology Planning & Evaluation) in 2024.
Author information
Authors and Affiliations
Contributions
D.H.S. and D.E. model design, coding, and manuscript writing. S.Y.P., J.T.G., and J.A.L. prepared figures and tables. H.Y.P. and J.T.G. data preprocessing and project administration. J.K. and J.W.Y. are clinical supervisors and proofreaders. J.K. and E.U. funding, experiment design, and result validation. E.U.‘s idea and study design, wrote the main manuscript, and edited and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shin, D.H., Enkhbayar, D., Park, S.Y. et al. Ensemble learning approaches for early prediction of chronic kidney disease based on polysomnographic phenotype analysis. npj Digit. Med. 8, 675 (2025). https://doi.org/10.1038/s41746-025-02039-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41746-025-02039-6