Abstract
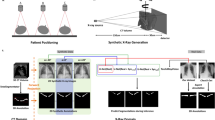
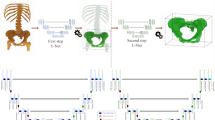
Patient-specific bone models are essential for designing surgical guides and preoperative planning, as they enable the visualization of intricate anatomical structures. However, traditional CT-based approaches for creating bone models are limited to preoperative use due to the low flexibility and high radiation exposure of CT and time-consuming manual delineation. Here, we introduce Semi-Supervised Reconstruction with Knowledge Distillation (SSR-KD), a fast and accurate AI framework to reconstruct high-quality bone models from biplanar X-rays in 30 seconds, with an average error under 1.0 mm, eliminating the dependence on CT and manual work. Additionally, high tibial osteotomy simulation was performed by experts on reconstructed bone models, demonstrating that bone models reconstructed from biplanar X-rays have comparable clinical applicability to those annotated from CT. Overall, our approach accelerates the process, reduces radiation exposure, enables intraoperative guidance, and significantly improves the practicality of bone models, offering transformative applications in orthopedics.
Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are not publicly available because this would compromise the patient confidentiality and privacy agreement with the data-providing hospitals, which prohibits any form of public distribution. The minimal dataset that would be necessary to interpret, replicate, and build upon the findings reported in the article is available from the corresponding authors upon reasonable request.
Code availability
The source code for this study is publicly available on GitHub at https://github.com/xmed-lab/SSR-KD, including the source code for implementing the SSR-KD framework, data generation, and experimental analysis. The code is released under the MIT license. We implemented the network design, model training, and model evaluation using PyTorch56. Key dependencies include TIGRE for X-ray projection simulation and PyTorch3D for calculating the chamfer distance. Other significant libraries used are Open3D, NumPy, Trimesh, skimage, and SimpleITK. The released codebase was tested on a workstation equipped with two NVIDIA GeForce RTX 3090 GPUs (24 GB), Intel Xeon Gold 5218 CPU @ 2.30 GHz, and 128 GB of RAM.
References
Global orthopedic surgery market report. https://www.globenewswire.com/news-release/2019/08/13/1901268/0/en/Global-Orthopedic-Surgery-Market-Report-2017-to-2022-Procedure-Volume-Trends-by-Type-Country-and-Region.html. Accessed 11 May 2023.
Globaldata. https://www.globaldata.com/store/report/usa-orthopedic-procedures-analysis/. Accessed 11 May 2023.
Li, Y.-T. et al. Surgical treatment for posterior dislocation of hip combined with acetabular fractures using preoperative virtual simulation and three-dimensional printing model-assisted precontoured plate fixation techniques. BioMed. Res. Int. 2019, 3971571 (2019).
Woo, S.-H., Sung, M.-J., Park, K.-S. & Yoon, T.-R. Three-dimensional-printing technology in hip and pelvic surgery: current landscape. Hip Pelvis 32, 1–10 (2020).
Portnoy, Y. et al. Three-dimensional technologies in presurgical planning of bone surgeries: current evidence and future perspectives. Int. J. Surg. 109, 3–10 (2023).
Kumar, V., Baburaj, V., Patel, S., Sharma, S. & Vaishya, R. Does the use of intraoperative ct scan improve outcomes in orthopaedic surgery? a systematic review and meta-analysis of 871 cases. J. Clin. Orthop. Trauma 18, 216–223 (2021).
Villarraga-Gómez, H. & Smith, S. T. Effect of the number of projections on dimensional measurements with X-ray computed tomography. Precis. Eng. 66, 445–456 (2020).
Brenner, D. J. & Hall, E. J. Computed tomography—an increasing source of radiation exposure. N. Engl. J. Med. 357, 2277–2284 (2007).
Miglioretti, D. L. et al. The use of computed tomography in pediatrics and the associated radiation exposure and estimated cancer risk. JAMA Pediatr. 167, 700–707 (2013).
Pearce, M. S. et al. Radiation exposure from ct scans in childhood and subsequent risk of leukaemia and brain tumours: a retrospective cohort study. Lancet 380, 499–505 (2012).
Lee, C. I., Haims, A. H., Monico, E. P., Brink, J. A. & Forman, H. P. Diagnostic ct scans: assessment of patient, physician, and radiologist awareness of radiation dose and possible risks. Radiology 231, 393–398 (2004).
Patil, S., Lindley, E. M., Burger, E. L., Yoshihara, H. & Patel, V. V. Pedicle screw placement with O-arm and Stealth navigation. Orthopedics 35, e61–e65 (2012).
Sari, R., Baskan, O., Ozlu, E. B. K. & Elmaci, I. Reduced radiation exposure during o-arm navigation in degenerative lumbar spine surgery. Demiroglu Bilim Univ. Florence Nightingale Tip. Derg. 10, 063–070 (2024).
Sarkalkan, N., Weinans, H. & Zadpoor, A. A. Statistical shape and appearance models of bones. Bone 60, 129–140 (2014).
Baka, N. et al. Statistical shape model-based femur kinematics from biplane fluoroscopy. IEEE Trans. Med. Imaging 31, 1573–1583 (2012).
Zheng, G. et al. A 2d/3d correspondence building method for reconstruction of a patient-specific 3d bone surface model using point distribution models and calibrated X-ray images. Med. Image Anal. 13, 883–899 (2009).
Thusini, X. O. et al. Uncertainty reduction in contour-based 3d/2d registration of bone surfaces. In Proc. Shape in Medical Imaging: International Workshop, ShapeMI 2020, Held in Conjunction with MICCAI 2020, Lima, Peru, October 4, 2020 18–29 (Springer, 2020).
Lamecker, H., Wenckebach, T. H. & Hege, H.-C. Atlas-based 3d-shape reconstruction from X-ray images. In Proc. 18th International Conference on Pattern Recognition (ICPR’06) Vol. 1, 371–374 (IEEE, 2006).
Klima, O., Kleparnik, P., Spanel, M. & Zemcik, P. Intensity-based femoral atlas 2D/3D registration using Levenberg-Marquardt optimisation. In Proc. Medical Imaging 2016: Biomedical Applications in Molecular, Structural, and Functional Imaging Vol. 9788, 113–124 (SPIE, 2016).
Ehlke, M., Ramm, H., Lamecker, H., Hege, H.-C. & Zachow, S. Fast generation of virtual X-ray images for reconstruction of 3D anatomy. IEEE Trans. Vis. Comput. Graph. 19, 2673–2682 (2013).
Zheng, G. Personalized X-ray reconstruction of the proximal femur via intensity-based non-rigid 2D-3D registration. In Proc. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2011: 14th International Conference, Toronto, Canada, September 18-22, 2011, Part II 14 598–606 (Springer, 2011).
Sadowsky, O., Chintalapani, G. & Taylor, R. H. Deformable 2D-3D registration of the pelvis with a limited field of view, using shape statistics. In Proc. Medical Image Computing and Computer-Assisted Intervention–MICCAI 2007: 10th International Conference, Brisbane, Australia, October 29-November 2, 2007, Part II 10 519–526 (Springer, 2007).
Yao, J. et al. Assessing accuracy factors in deformable 2D/3D medical image registration using a statistical pelvis model. In Proc. Ninth IEEE International Conference on Computer Vision 1329–1334 (IEEE, 2003).
Yu, P. et al. Spatial resolution enhancement using deep learning improves chest disease diagnosis based on thick slice ct. npj Digit. Med. 7, 335 (2024).
Bellemo, V. et al. Optical coherence tomography choroidal enhancement using generative deep learning. npj Digit. Med. 7, 115 (2024).
Chen, R. et al. Translating color fundus photography to indocyanine green angiography using deep-learning for age-related macular degeneration screening. npj Digit. Med. 7, 34 (2024).
Čavojská, J. et al. Estimating and abstracting the 3d structure of feline bones using neural networks on X-ray (2D) images. Commun. Biol. 3, 337 (2020).
Jiang, B. et al. Deep learning reconstruction shows better lung nodule detection for ultra-low-dose chest CT. Radiology 303, 202–212 (2022).
Preetha, C. J. et al. Deep-learning-based synthesis of post-contrast T1-weighted MRI for tumour response assessment in neuro-oncology: a multicentre, retrospective cohort study. Lancet Digit. Health 3, e784–e794 (2021).
Magdy, O. et al. Bone scintigraphy based on deep learning model and modified growth optimizer. Sci. Rep. 14, 25627 (2024).
Wu, C. et al. A machine learning-based multiscale model to predict bone formation in scaffolds. Nat. Comput. Sci. 1, 532–541 (2021).
Shiode, R. et al. 2d–3d reconstruction of distal forearm bone from actual x-ray images of the wrist using convolutional neural networks. Sci. Rep. 11, 1–12 (2021).
Kasten, Y., Doktofsky, D. & Kovler, I. End-to-end convolutional neural network for 3D reconstruction of knee bones from bi-planar X-ray images. In Proc. International Workshop on Machine Learning for Medical Image Reconstruction 123–133 (Springer, 2020).
Aubert, B., Vazquez, C., Cresson, T., Parent, S. & de Guise, J. A. Toward automated 3D spine reconstruction from biplanar radiographs using CNN for statistical spine model fitting. IEEE Trans. Med. Imaging 38, 2796–2806 (2019).
Kim, H., Lee, K., Lee, D. & Baek, N. 3D reconstruction of leg bones from X-ray images using CNN-based feature analysis. In Proc. 2019 International Conference on Information and Communication Technology Convergence (ICTC) 669–672 (IEEE, 2019).
Park, J. J., Florence, P., Straub, J., Newcombe, R. & Lovegrove, S. Deepsdf: Learning continuous signed distance functions for shape representation. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 165–174 (IEEE, 2019).
Mescheder, L., Oechsle, M., Niemeyer, M., Nowozin, S. & Geiger, A. Occupancy networks: learning 3D reconstruction in function space. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 4460–4470 (IEEE, 2019).
Lorensen, W. E. & Cline, H. E. Marching cubes: a high resolution 3D surface construction algorithm. In Proc. ACM SIGGRAPH Computer Graphics Vol. 21, 163–169 (ACM, 1987).
Shen, L., Zhao, W. & Xing, L. Patient-specific reconstruction of volumetric computed tomography images from a single projection view via deep learning. Nat. Biomed. Eng. 3, 880–888 (2019).
Reyneke, C. J. F. et al. Review of 2-d/3-d reconstruction using statistical shape and intensity models and x-ray image synthesis: toward a unified framework. IEEE Rev. Biomed. Eng. 12, 269–286 (2018).
Chênes, C. & Schmid, J. Revisiting contour-driven and knowledge-based deformable models: application to 2D-3D proximal femur reconstruction from X-ray images. In Proc. International Conference on Medical Image Computing and Computer-Assisted Intervention 451–460 (Springer, 2021).
Wright, J. M., Crockett, H. C., Slawski, D. P., Madsen, M. W. & Windsor, R. E. High tibial osteotomy. J. Am. Acad. Orthop. Surg. 13, 279–289 (2005).
McNamara, I., Birmingham, T., Fowler, P. & Giffin, J. High tibial osteotomy: evolution of research and clinical applications—a Canadian experience. Knee Surg. Sports Traumatol. Arthrosc. 21, 23–31 (2013).
Fayard, J.-M. et al. Patient-specific cutting guides increase accuracy of medial opening wedge high tibial osteotomy procedure: a retrospective case-control study. J. Exp. Orthop. 11, e12013 (2024).
Yam, M. G. J., Chao, J. Y. Y., Leong, C. & Tan, C. H. 3D printed patient specific customised surgical jig for reverse shoulder arthroplasty, a cost effective and accurate solution. J. Clin. Orthop. Trauma 21, 101503 (2021).
Materialise 3-matic. https://www.materialise.com/en/industrial/software/3-matic. Accessed 10 September 2024.
Khan, F. A., Lipman, J. D., Pearle, A. D., Boland, P. J. & Healey, J. H. Surgical technique: computer-generated custom jigs improve accuracy of wide resection of bone tumors. Clin. Orthop. Relat. Res. 471, 2007–2016 (2013).
Rau, T. S. et al. Concept description and accuracy evaluation of a moldable surgical targeting system. J. Med. Imaging 8, 015003–015003 (2021).
Geiger, L., Zuniga, M. G., Lenarz, T., Majdani, O. & Rau, T. S. Drilling accuracy evaluation of a mouldable surgical targeting system for minimally invasive access to anatomic targets in the temporal bone. Eur. Arch. Oto-Rhino-Laryngol. 280, 4371–4379 (2023).
Yeung, M., Abdulmajeed, A., Carrico, C. K., Deeb, G. R. & Bencharit, S. Accuracy and precision of 3D-printed implant surgical guides with different implant systems: an in vitro study. J. Prosthet. Dent. 123, 821–828 (2020).
Msallem, B. et al. Dimensional accuracy in 3D printed medical models: a follow-up study on SLA and SLS technology. J. Clin. Med. 13, 5848 (2024).
Caiti, G., Dobbe, J. G., Strijkers, G. J., Strackee, S. D. & Streekstra, G. J. Positioning error of custom 3D-printed surgical guides for the radius: influence of fitting location and guide design. Int. J. Comput. Assist. Radiol. Surg. 13, 507–518 (2018).
Milletari, F., Navab, N. & Ahmadi, S.-A. V-net: Fully convolutional neural networks for volumetric medical image segmentation. In Proc. 2016 Fourth International Conference on 3D Vision (3DV) 565–571 (IEEE, 2016).
Newell, A., Yang, K. & Deng, J. Stacked hourglass networks for human pose estimation. In Proc. European Conference on Computer Vision 483–499 (Springer, 2016).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proc. International Conference on Medical Image Computing and Computer-assisted Intervention 234–241 (Springer, 2015).
Paszke, A. et al. PyTorch: an imperative style, high-performance deep learning library. In Proc. Advances in Neural Information Processing Systems Vol. 32, 8024–8035 (Curran Associates, Inc., 2019).
Acknowledgements
This work was supported by a research grant from the Hong Kong Innovation and Technology Fund (Project PRP/041/22FX) and partially supported by the Natural Science Foundation of Zhejiang Province (No. LZ23A050002) and the National Natural Science Foundation of China (No. 12175012 and No. 12575358).
Author information
Authors and Affiliations
Contributions
X. Li, W. Zhao, Y. Lin, and H. Sun conceptualized and designed the study. Y. Lin implemented the method and contributed to the manuscript writing, revision, and analysis of the results. H. Sun, W. Yau, and M. Amangeldy designed and organized the HTO simulation to evaluate its clinical applicability. Y. Li, R. Aslam, T. Cheng, and V. Le Meur coordinated the data annotation. L. Tse, C. Chui, and K. Cho assessed the annotation quality and provided guidance. Y. Ye and J. Zou developed the manuscript outline and guided its preparation. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lin, Y., Sun, H., Li, Y. et al. Real-time reconstruction of 3D bone models via very-low-dose protocols. npj Digit. Med. (2026). https://doi.org/10.1038/s41746-026-02389-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-026-02389-9