Abstract
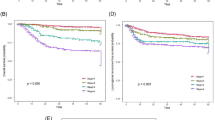
Oral squamous cell carcinoma (OSCC) remains the most common head and neck malignancy, for which early detection is critical yet challenging with current invasive methods. This study aimed to establish a comprehensive diagnostic framework for OSCC by integrating proton transfer reaction-time-of-flight mass spectrometry (PTR-TOF-MS) breath analysis and metagenomic sequencing with artificial intelligence (AI). Exhaled breath and saliva samples were collected from participants in a discovery cohort (n = 222) and an external validation cohort (n = 83). Samples were analyzed using PTR-TOF-MS and metagenomic sequencing, and multimodal diagnostic models were constructed and trained on the discovery cohort data. We identified OSCC-specific biomarkers, including methanethiol and Fusobacterium nucleatum, and developed an interactive online platform (https://bio.futurecnn.com/) enabling real-time predictions and biomarker interpretability. The AI-driven diagnostic model achieved excellent accuracy (ROC-AUC: 0.92) in distinguishing OSCC patients from healthy controls in the external set. This approach offers a practical, noninvasive solution for OSCC screening and establishes an adaptable framework for other breath-based diagnostics.

Similar content being viewed by others
Data availability
The original data generated and analyzed in this study are included in the article and Supplementary Material. Additional information or raw data can be obtained from the corresponding author upon reasonable request. The diagnostic model developed in this study has been fully open-sourced and is publicly accessible at: https://github.com/SunYilan/biomodel.
References
Tan, Y. et al. Oral Squamous Cell Carcinomas: State of the Field and Emerging Directions. Int J. Oral. Sci. 15, 44 (2023).
Bray, F. et al. Global Cancer Statistics 2022: Globocan Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. Ca: A Cancer J. Clinicians 74, 229–263 (2024).
Silveira, F. M., Schuch, L. F. & Bologna-Molina, R. Classificatory Updates in Verrucous and Cuniculatum Carcinomas: Insights From the 5Th Edition of Who-Iarc Head and Neck Tumor Classification. World J. Clin. Oncol. 15, 464–467 (2024).
Ken, K. et al. Doublet Chemotherapy, Triplet Chemotherapy, Or Doublet Chemotherapy Combined with Radiotherapy as Neoadjuvant Treatment for Locally Advanced Oesophageal Cancer (Jcog1109 Next): A Randomised, Controlled, Open-Label, Phase 3 Trial. Lancet 404, 55–66 (2024).
Ahmed, A. et al. Mutation Detection in Saliva From Oral Cancer Patients. Oral. Oncol. 151, 106717 (2024).
Lennon, A. M. et al. Feasibility of Blood Testing Combined with Pet-Ct to Screen for Cancer and Guide Intervention. Science 369, eabb9601 (2020).
Feng, X. et al. Cancer stage compared with mortality as end points in randomized clinical trials of cancer screening: a systematic review and meta-analysis. Jama-J. Am. Med. Assoc. 331, 1910–1917 (2024).
Yan, B. et al. Genai synthesis of histopathological images from raman imaging for intraoperative tongue squamous cell carcinoma assessment. Int. J. Oral. Sci. 17, 12 (2025).
Balakittnen, J. et al. A novel saliva-based mirna profile to diagnose and predict oral cancer. Int. J. Oral. Sci. 16, 14 (2024).
Song, M., Bai, H., Zhang, P., Zhou, X. & Ying, B. Promising applications of human-derived saliva biomarker testing in clinical diagnostics. Int. J. Oral. Sci. 15, 2 (2023).
Zhou, M. et al. Exhaled breath and urinary volatile organic compounds (Vocs) for cancer diagnoses, and microbial-related Voc metabolic pathway analysis: a systematic review and meta-analysis. Int. J. Surg. 110, 1755–1769 (2024).
Li, L. et al. Qualitative and quantitative transformer-cnn algorithm models for the screening of exhale biomarkers of early lung cancer patients. Anal. Chem. 97, 6651–6660 (2025).
Li, X. et al. Volatile organic compounds in exhaled breath: a promising approach for accurate differentiation of lung adenocarcinoma and squamous cell carcinoma. J. Breath Res. 18 (2024).
Roquencourt, C., Grassin-Delyle, S. & Thevenot, E. A. Ptairms: real-time processing and analysis of Ptr-Tof-Ms data for biomarker discovery in exhaled breath. Bioinformatics 38, 1930–1937 (2022).
Roquencourt, C., Lamy, E., Bardin, E., Devillier, P., & Grassin-Delyle, S. A benchmark study of data normalisation methods for Ptr-Tof-Ms exhaled breath metabolomics. J. Breath Res. 18 (2023).
Wang, H. et al. A combined screening study for evaluating the potential of exhaled acetone, isoprene, and nitric oxide as biomarkers of lung cancer. Rsc Adv. 13, 31835–31843 (2023).
Zhang, K. et al. A generalist vision-language foundation model for diverse biomedical tasks. Nat. Med. 30, 3129–3141 (2024).
Beck, A. G. et al. Recent developments in machine learning for mass spectrometry. Acs Meas. Sci. Au. 4, 233–246 (2024).
Brinkman, P. et al. Fulfilling the promise of breathomics: considerations for the discovery and validation of exhaled volatile biomarkers. Am. J. Resp. Crit. Care 210, 1079–1090 (2024).
Deng, F. et al. A novel accurate peak extraction algorithm of mass spectrometry based on iterative adaptive curve fitting. J. Am. Soc. Mass Spectr. 35, 2900–2909 (2024).
Garg, M. et al. Disease prediction with multi-omics and biomarkers empowers case-control genetic discoveries in the UK Biobank. Nat. Genet. 56, 1821–1831 (2024).
Melnikov, A. D., Tsentalovich, Y. P. & Yanshole, V. V. Deep learning for the precise peak detection in high-resolution Lc-Ms Data. Anal. Chem. 92, 588–592 (2020).
Sun, Y. et al. Integrative plasma and fecal metabolomics identify functional metabolites in adenoma-colorectal cancer progression and as early diagnostic biomarkers. Cancer Cell 42, 1386–1400 (2024).
He, X., Zhao, K. & Chu, X. Automl: a survey of the state-of-the-art. Knowl. -Based Syst. 212, 106622 (2020).
Pinichka, C., Chotpantarat, S., Cho, K. H. & Siriwong, W. Comparative analysis of swat and swat coupled with xgboost model using optuna hyperparameter optimization for nutrient simulation: a case study in the upper Nan River Basin, Thailand. J. Environ. Manag. 388, 126053 (2025).
Lu, S., Song, W., Pfob, A. & Gibbons, C. Assessing the representativeness of large medical data using population stability index. Bmc Med. Res. Methodol. 25, 44 (2025).
Gu, Y. et al. Influence of the densities and nutritional components of bacterial colonies on the culture-enriched gut bacterial community structure. Amb. Express 11, 78 (2021).
Hara, T., Sakanaka, A., Lamont, R. J., Amano, A. & Kuboniwa, M. Interspecies metabolite transfer fuels the methionine metabolism of fusobacterium nucleatum to stimulate volatile methyl mercaptan production. Msystems 9, e76423 (2024).
Alon-Maimon, T., Mandelboim, O. & Bachrach, G. Fusobacterium Nucleatum and Cancer. Periodontol 2000 89, 166–180 (2022).
Vassilenko, V., Moura, P. C., & Raposo, M. Diagnosis of carcinogenic pathologies through breath biomarkers: present and future trends. Biomedicines 11, (2023).
Philipp, T. M., Scheller, A. S., Krafczyk, N., Klotz, L., & Steinbrenner, H. Methanethiol: a scent mark of dysregulated sulfur metabolism in cancer. Antioxidants-Basel 12, (2023).
Kwon, I. et al. Detection of Volatile Sulfur Compounds (Vscs) in Exhaled Breath as a Potential Diagnostic Method for Oral Squamous Cell Carcinoma. BMC Oral. Health 22, 268 (2022).
Henderson, B. et al. The Peppermint Breath Test Benchmark for Ptr-Ms and Sift-Ms. J. Breath Res. 15 (2021).
Desai, K. M. et al. Screening of oral potentially malignant disorders and oral cancer using deep learning models. Sci. Rep. -UK 15, 17949 (2025).
Golsanamlu, Z., Jin, H., Soleymani, J. & Jouyban, A. Application of high-resolution analytical techniques in breathomics studies. Microchem. J. 211, 113073 (2025).
Borden, S. A. et al. Characterizing volatile organic compound profiles in oral cancer using multiple sample collection approaches by Gc-Ims and Td-Gc-Ms. Sci. Rep. -Uk 15, 37014 (2025).
Mentel, S. et al. Prediction of oral squamous cell carcinoma based on machine learning of breath samples: a prospective controlled study. Bmc Oral Health 21 (2021).
Bouza, M., Gonzalez-Soto, J., Pereiro, R., de Vicente, J. C. & Sanz-Medel, A. Exhaled breath and oral cavity Vocs as potential biomarkers in oral cancer patients. J. Breath. Res. 11, 16015 (2017).
Jia, Z. et al. Advanced strategy for cancer detection based on volatile organic compounds in breath. J. Nanobiotechnol. 23, 468 (2025).
Le, T. & Priefer, R. Detection technologies of volatile organic compounds in the breath for cancer diagnoses. Talanta 265, 124767 (2023).
Fitzsimonds, Z. R., Rodriguez-Hernandez, C. J., Bagaitkar, J. & Lamont, R. J. From beyond the pale to the pale riders: the emerging association of bacteria with oral cancer. J. Dent. Res. 99, 604–612 (2020).
Xiang, Z. et al. Fusobacterium nucleatum Exacerbates Colitis via Stat3 activation induced by acetyl-Coa accumulation. Gut Microbes 17, 2489070 (2025).
Sun, J. et al. F.nucleatum facilitates oral squamous cell carcinoma progression via Glut1-driven lactate production. Ebiomedicine 88, 104444 (2023).
Cavallucci, V. et al. Proinflammatory and cancer-promoting pathobiont fusobacterium nucleatum directly targets colorectal cancer stem cells. Biomolecules 12 (2022).
Nie, F. et al. The role of Cxcl2-mediated crosstalk between tumor cells and macrophages in fusobacterium nucleatum-promoted oral squamous cell carcinoma progression. Cell Death Dis. 15, 277 (2024).
Wang, Y. et al. Study of the Inflammatory Activating Process in the Early Stage of Fusobacterium Nucleatum Infected Pdlscs. Int. J. Oral. Sci. 15, 8 (2023).
Sun, J., Chen, F. & Wu, G. Potential effects of gut microbiota on host cancers: focus on immunity, DNA damage, cellular pathways, and anticancer therapy. Isme J. 17, 1535–1551 (2023).
Zhu, H. et al. Fusobacterium Nucleatum Promotes Tumor Progression in Kras P.G12D-Mutant Colorectal Cancer by Binding to Dhx15. Nat. Commun. 15, 1688 (2024).
Dan, W., Xiong, C., Zhou, G., Chen, J. & Pan, F. Gut microbiota as a mediator of cancer development and management: from colitis to colitis-associated dysplasia and carcinoma. BBA Rev. Cancer 1880, 189381 (2025).
Association, W. M World medical association declaration of Helsinki: ethical principles for medical research involving human subjects. Jama-J. Am. Med. Assoc. 310, 2191–2194 (2013).
Czippelova, B. et al. Impact of Breath Sample Collection Method and Length of Storage of Breath Samples in Tedlar Bags On the Level of Selected Volatiles Assessed Using Gas Chromatography-Ion Mobility Spectrometry (Gc-Ims). J. Breath Res. 18 (2024).
Yáñez-Serrano, A. M. et al. Glovocs - master compound assignment guide for proton transfer reaction mass spectrometry users. Atmos. Environ. 10, 689–698 (2021).
Kuo, T. et al. Human Breathomics Database. Database-Oxford 2020 (2020).
He, Y. et al. Plasma metabolomics dataset of race-walking athletes illuminating systemic metabolic reaction of exercise. Sci. Data 12, 448 (2025).
Navazesh, M., Mulligan, R. A., Kipnis, V., Denny, P. A. & Denny, P. C. Comparison of whole saliva flow rates and mucin concentrations in healthy caucasian young and aged adults. J. Dent. Res. 71, 1275–1278 (1992).
Acknowledgements
This project was supported by the National Natural Science Foundation of China Outstanding Youth Fund (Grant No. 62322114) and the Medical Engineering Cross Foundation of Shanghai Jiao Tong University (Grant No. YG2023LC06) and the National Natural Science Foundation of China (Grant No. 82272815).
Author information
Authors and Affiliations
Contributions
Y.S.: Conceptualization; methodology; software; data curation; investigation; validation; project administration; resources; visualization; writing—original draft; writing—review and editing; supervision. X.H.: Methodology; data curation; validation; visualization; writing—original draft; writing—review and editing. J.H.: Methodology; data curation; formal analysis. Y.W.: Methodology; visualization. J.L.(Luo): Resources; writing—review and editing. J.Y.: Visualization, writing—review and editing. Y.D.: Supervision; writing—review and editing. X.W.: Supervision; methodology; funding acquisition; writing—review and editing; project administration. J.L.(Liu): Supervision; methodology; funding acquisition; writing—review and editing; project administration.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, Y., Hu, X., Han, J. et al. Rapid and noninvasive artificial intelligence-assisted diagnostic method for oral squamous cell carcinoma. npj Digit. Med. (2026). https://doi.org/10.1038/s41746-026-02527-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-026-02527-3