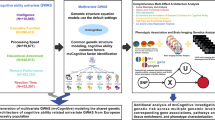
Abstract
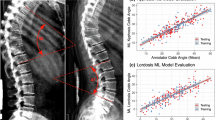
Scoliosis is the most common developmental spinal deformity, but its genetic underpinnings remain only partially understood. To identify scoliosis-related loci, we utilized dual energy X-ray absorptiometry (DXA) scans from 57,588 individuals in the UK Biobank (UKB), and quantified spinal curvature using deep learning-based vertebral segmentation and landmarking to measure cumulative horizontal displacement. On a subset of 150 individuals, our automated image-derived curvature measurements showed a correlation of 0.83 with clinical Cobb angle assessments, supporting their validity as a proxy for scoliosis severity. To connect spinal curvature to genetics, we conducted a genome-wide association study (GWAS). Our quantitative imaging phenotype identified 2 novel loci associated with scoliosis in a European population. These loci are in SEM1/SHFM1 and on an lncRNA on chr 3 located between EDEM1 and GRM7. Genetic correlation analysis revealed significant overlap between our image-based GWAS and ICD-10-based GWAS in the UKB and the Biobank of Japan. We show that our quantitative GWAS identifies more genome-wide significant loci than a case-control scoliosis dataset with ten times the sample size. Our results illustrate the potential of quantitative imaging phenotypes to uncover genetic associations that are challenging to capture with medical records alone and identify new loci for functional follow-up.
Similar content being viewed by others
Data availability
All data used for this study were obtained from the UK Biobank under application number 65439. GWAS summary statistics are currently being uploaded to the GWAS catalog and are available at this Box link: https://utexas.box.com/s/0tyyxhzv2e5x8iucsdcd8dfutobnmcd2.
Code availability
Deep-learning and image processing tools can be found at https://github.com/reb12345/Scoliosis.
References
Janicki, J. A. & Alman, B. Scoliosis: review of diagnosis and treatment. Paediatr. Child Health 12, 771 (2007).
Aebi, M. The adult scoliosis. Eur. Spine J. 14, 925–948 (2005).
Giampietro, P. F. Genetic aspects of congenital and idiopathic scoliosis. Science 2012, 152365 (2012).
Kotwal, S., Pumberger, M., Hughes, A. & Girardi, F. Degenerative scoliosis: a review. HSS J. 7, 257 (2011).
Konieczny, M. R., Senyurt, H. & Krauspe, R. Epidemiology of adolescent idiopathic scoliosis. J. Child. Orthop. 7, 3 (2012).
Takahashi, Y. et al. A genome-wide association study identifies common variants near LBX1 associated with adolescent idiopathic scoliosis. Nat. Genet. 43, 1237–1240 (2011).
Kou, I. et al. Genetic variants in GPR126 are associated with adolescent idiopathic scoliosis. Nat. Genet. 45, 676–679 (2013).
Kou, I. et al. Genome-wide association study identifies 14 previously unreported susceptibility loci for adolescent idiopathic scoliosis in Japanese. Nat. Commun. 10, 1–9 (2019).
Sharma, S. et al. A PAX1 enhancer locus is associated with susceptibility to idiopathic scoliosis in females. Nat. Commun. 6, 6452 (2015).
Khanshour, A. M. et al. Genome-wide meta-analysis and replication studies in multiple ethnicities identify novel adolescent idiopathic scoliosis susceptibility loci. Hum. Mol. Genet. 27, 3986–3998 (2018).
Flynn, B. I. et al. Deep learning based phenotyping of medical images improves power for gene discovery of complex disease. npj Digit. Med. 6, 1–12 (2023).
Faber, B. G. et al. The identification of distinct protective and susceptibility mechanisms for hip osteoarthritis: findings from a genome-wide association study meta-analysis of minimum joint space width and Mendelian randomisation cluster analyses. EBioMedicine 95, 104759 (2023).
Jamaludin, A. et al. Automated measurement of size of spinal curve in population-based cohorts: validation of a method based on total body dual energy X-ray absorptiometry scans. Bone 172, 116775 (2023).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Xu, L. et al. The genetic architecture of and evolutionary constraints on the human pelvic form. Science 388, eadq1521 (1979).
Kun, E. et al. The genetic architecture and evolution of the human skeletal form. Science 381, eadf8009 (1979).
Weng, W. & Zhu, X. U-Net: convolutional networks for biomedical image segmentation. IEEE Access 9, 16591–16603 (2015).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), Vol. 9351, 234–241 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. Proc. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit. https://doi.org/10.48550/arxiv.1512.03385 (2015).
Brink, R. C. et al. Upright, prone, and supine spinal morphology and alignment in adolescent idiopathic scoliosis. Scoliosis Spinal Disord. 12, 6 (2017).
Hasegawa, K., Okamoto, M., Hatsushikano, S., Caseiro, G. & Watanabe, K. Difference in whole spinal alignment between supine and standing positions in patients with adult spinal deformity using a new comparison method with slot-scanning three-dimensional X-ray imager and computed tomography through digital reconstructed radio. BMC Musculoskelet. Disord. 19, 437 (2018).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559 (2007).
Bulik-Sullivan, B. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Ushiki, A. et al. Deletion of Pax1 scoliosis-associated regulatory elements leads to a female-biased tail abnormality. Cell Rep 43, 113907 (2024).
Crackower, M. A. et al. Characterization of the split hand/split foot malformation locus SHFM1 at 7q21.3-q22.1 and analysis of a candidate gene for its expression during limb development. Hum. Mol. Genet. 5, 571–579 (1996).
Mbatchou, J. et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat. Genet. 53, 1097–1103 (2021).
Sollis, E. et al. The NHGRI-EBI GWAS Catalog: knowledgebase and deposition resource. Nucleic Acids Res. 51, D977–D985 (2023).
Watanabe, K. et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 51, 1339–1348 (2019).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Brown, B. C., Ye, C. J., Price, A. L. & Zaitlen, N. Transethnic genetic-correlation estimates from summary statistics. Am. J. Hum. Genet. 99, 76 (2016).
Cheng, J. C. Y. et al. Generalized low areal and volumetric bone mineral density in adolescent idiopathic scoliosis. J. Bone Miner. Res. 15, 1587–1595 (2000).
Epstein, J. A., Epstein, B. S. & Jones, M. D. Symptomatic lumbar scoliosis with degenerative changes in the elderly. Spine (Philos. Pa. 1976) 4, 542–547 (1979).
Gertzbein, S. D. Scoliosis Research Society. Multicenter spine fracture study. Spine (Philos. Pa. 1976) 17, 528–540 (1992).
Wallin, J. et al. The role of Pax-1 in axial skeleton development. Development 120, 1109–1121 (1994).
Olivari, S. et al. EDEM1 regulates ER-associated degradation by accelerating de-mannosylation of folding-defective polypeptides and by inhibiting their covalent aggregation. Biochem. Biophys. Res. Commun. 349, 1278–1284 (2006).
Jacobs, W. B. & Fehlings, M. G. Ankylosing spondylitis and spinal cord injury: origin, incidence, management, and avoidance. Neurosurg. Focus 24, E12 (2008).
Guiliano, D. B. et al. EDEM1 targets misfolded HLA-B27 dimers for endoplasmic reticulum associated degradation. Arthritis Rheumatol. 66, 2976 (2014).
Nicoletti, P. Regulatory elements in SEM1-DLX5-DLX6 (7q21.3) locus contribute to genetic control of coronal nonsyndromic craniosynostosis and bone density-related traits. Genet. Med. Open 2, 101851 (2024).
Otomo, N. et al. Polygenic risk score of adolescent idiopathic scoliosis for potential clinical use. J. Bone Miner. Res. 36, 1481–1491 (2021).
Wright, V. J., Schwartzman, J. D., Itinoche, R. & Wittstein, J. The musculoskeletal syndrome of menopause. Climacteric 27, 466–472 (2024).
Urrutia, J., Diaz-Ledezma, C., Espinosa, J. & Berven, S. H. Lumbar scoliosis in postmenopausal women: prevalence and relationship with bone density, age, and body mass index. Spine 36, 737–740 (2011).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Van Der Walt, S. et al. Scikit-image: image processing in Python. PeerJ https://doi.org/10.7717/PEERJ.453/FIG-5 (2014).
Mason, D. et al. pydicom/pydicom: pydicom 2.3.0. (2022).
Wada, K. Labelme: Image Polygonal Annotation with Python.
Gehan, M. A. et al. PlantCV v2: Image analysis software for high-throughput plant phenotyping. PeerJ 5, e4088 (2017).
Bradski, G. The OpenCV Library. Dr. Dobbas J. Softw. Tools 25, 122–125 (2000).
Blake, J. A. et al. Mouse Genome Database (MGD): knowledgebase for mouse-human comparative biology. Nucleic Acids Res. 49, D981–D987 (2021).
Acknowledgements
V.M.N. was supported on a grant from the Allen Discovery Center program, a Paul G. Allen Frontiers Group-advised program of the Paul G. Allen Family Foundation.
Author information
Authors and Affiliations
Contributions
M.Z. and E.K. wrote the paper with input from all co-authors. M.Z., E.K., S.R., L.X., D.P., J.W., C.L., and N.O. performed the analysis. R.G., C.W. and V.M.N. supervised the analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zeosky, M., Kun, E., Reddy, S. et al. Deep learning-based precision phenotyping of spine curvature identifies novel genetic risk loci for scoliosis in the UK Biobank. npj Digit. Med. (2026). https://doi.org/10.1038/s41746-026-02540-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-026-02540-6