Abstract
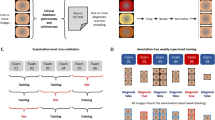
Gastrointestinal (GI) diseases pose a major clinical burden, yet conventional histopathology suffers from subjectivity and limited reproducibility. While existing computational pathology foundation models are often validated across many subspecialties in “broad but shallow” benchmarks, they rarely demonstrate deep clinical utility in real-world scenarios. To address this, we develop Digepath—a disease-specialized foundation model focused exclusively on high-impact GI pathology. Our approach employs a two-stage iterative optimization: first, pretraining on over 353 million multi-scale patches from 210,043 H&E-stained slides; second, fine-tuning on 471,443 expert-annotated regions, balancing tumor and non-tumor samples to enhance lesion perception amid sparse pathology in whole-slide images. Digepath achieves state-of-the-art performance on 32 of 33 systematic downstream tasks in GI pathology—including diagnosis, molecular profiling, and survival prognosis—demonstrating robust generalization. Moreover, we integrate its capabilities into an agent-based clinical reasoning framework that supports end-to-end intelligent diagnostic workflows, paving the way for real-world deployment.
Similar content being viewed by others
Acknowledgements
This work was supported in part by National Natural Science Foundation of China (82430062), the Shenzhen Engineering Research Centre (XMHT20230115004), the Jilin Fuyuan Guan Food Group Co., Ltd., Fujian Provincial Science and Technology Innovation Joint Funds (grant no. 2024Y96010076), and the Fujian Provincial Natural Science Foundation of China (grant no. 2024J011006). We thank Shenzhen Shengqiang Technology Co., Ltd. for providing slide scanners, and H3C Technologies Co., Ltd. for providing the training servers.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, L., Ling, X., Ouyang, M. et al. Subspecialty-specific foundation model for intelligent gastrointestinal pathology. npj Digit. Med. (2026). https://doi.org/10.1038/s41746-026-02684-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-026-02684-5