Abstract
Engineered polyketide synthases (PKSs) have great potential as biocatalysts. These unnatural enzymes are capable of synthesizing molecules that are either not amenable to biosynthesis or are extremely challenging to access chemically. PKSs can thus be a powerful platform to expand the chemical landscape beyond the limits of conventional metabolic engineering. Here we employ a retrobiosynthesis approach to design and construct PKSs to produce δ-valerolactam (VL) and three enantiopure α-substituted VL analogues that have no known biosynthetic route. We introduce the engineered PKSs and pathways for various malonyl-CoA derivatives into Pseudomonas putida and use proteomics, metabolomics and culture condition optimization to improve the production of our target compounds. These α-substituted VLs are polymerized into polyamides (nylon-5) or converted into their N-acryloyl derivatives. RAFT polymerization produces bio-derived polymers with potential biomedical applications. Overall, this interdisciplinary effort highlights the versatility and effectiveness of a PKS-based retrobiosynthesis approach in exploring and developing innovative biomaterials.

Similar content being viewed by others
Main
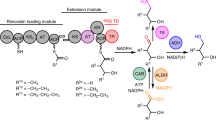
Polyketide synthases (PKSs) are biosynthetic assembly lines responsible for the production of complex and diverse natural products in bacteria and fungi1,2. PKSs consist of catalytic modules, each integrating an acyl coenzyme A (CoA) into the growing polyketide chain3. These modules can be categorized into three types: loading modules, which incorporate a broad range of starter units and usually consist of an acyltransferase (AT)–acyl carrier protein (ACP) didomain; elongation modules, which contain domains such as a ketosynthase (KS), AT and ACP for chain elongation, as well as tailoring domains like ketoreductase (KR), dehydratase (DH) and enoyl reductase (ER) for chain reduction; and termination modules, which offload the fully extended and modified polyketide chain, typically through thioesterase (TE) activity4,5.
Due to their modular nature, with a direct link between PKS gene sequences, protein active sites and the specific chemical building blocks in the final product, these enzymes have garnered substantial attention from metabolic engineers and synthetic biologists hoping to rearrange PKS enzymes into unnatural configurations and produce entirely new compounds5, including commodity and specialty chemicals, polymer precursors6,7 and biofuels8. Moreover, the presence of specific domains enables control over the stereochemistry and types of functional groups within these compounds7. Despite their vast chemical design space, only a handful of PKSs have been harnessed for synthesis of novel compounds due to the limited knowledge and difficulty in PKS engineering7,8. In particular, a largely unexplored field is the PKS-based production of nitrogen-containing monomers, including lactams, due to the relative lack of known PKS pathways.
Lactams are commonly employed directly as monomers for the synthesis of polyamides through ring-opening polymerization (ROP). The most prominent example is the commercial production of nylon-6 from ε-caprolactam (CL), a C6-lactam. More recently, δ-valerolactam (VL), a C5-lactam, has also been polymerized to produce nylon-5 and nylon-6,5, which show potential in thermoplastic and ferroelectric applications9,10. Lactams can also be polymerized via free-radical polymerization of N-vinyl lactams, yielding materials such as N-vinylcaprolactam11, N-vinylpiperidone12 and N-vinylpyrrolidone13, which are useful for water-soluble, temperature-responsive and biocompatible applications.
So far, the biological synthesis of VL has primarily relied on a native pathway from Pseudomonas putida, transforming lysine into 5-aminovaleric acid (5-AVA), followed by chemical or biological lactamization9,14. However, there are currently no known biosynthesis pathways for α-substituted VLs. Industrial applications of α-substituted VLs have remained unexplored, in part due to the absence of convenient chemical or biochemical synthesis pathways, so we addressed this by engineering PKSs to produce enantiomerically pure α-substituted VLs to demonstrate the flexibility and utility of engineering PKS biocatalysts.
In this Article we leverage a PKS-based retrobiosynthesis approach to produce VLs, each featuring functional groups on the α-carbon, in the microbial host P. putida. Specifically, through a combination of rational metabolic engineering in P. putida and PKS reprogramming involving domain and module exchange, we successfully produced VL and three VL analogues that are otherwise unattainable through any other known metabolic pathway. In this study we utilize and showcase the key advantages of the PKS-based biosynthesis platform: the capacity to readily change functional groups appended to the backbone of the target molecule family by exchanging a single PKS domain (AT), the ability to utilize renewable carbon sources, and the inherent stereoselectivity of PKSs for producing enantiopure compounds suitable for synthesizing stereocontrolled polymers without requiring chiral polymerization catalysts. To demonstrate the potential of α-substituted VLs as monomers, we applied step-growth polymerization methods to synthesize the corresponding polyamides and compare their properties. We also employed reversible addition-fragmentation chain transfer (RAFT) polymerization of their N-acryloyl derivatives to create bio-derived polymers with potential applications across diverse industries. Overall, our findings highlight the remarkable versatility and efficient assembly-line features of PKSs, offering a robust platform for producing unnatural compounds and exploring new-to-nature biomaterials.
Results
Host engineering to prevent native lactam metabolism
Pseudomonas putida KT2440 was selected as a host for the PKS-based lactam production due to its diverse carbon metabolism utilizing cost-effective and carbon-neutral feedstocks, its rapid growth rate, its high-GC-content (~62%) genome, which is thought to facilitate the expression of high-GC PKSs, its ease of genetic manipulation, and the promiscuous phosphopantetheinyl transferase (PPTase) for activating PKS systems15,16. To improve the genetic accessibility of this bacterium, we utilized P. putida strain AG5577 with the serine recombinase-assisted genome engineering (SAGE) toolkit, enabling iterative and high-throughput genome engineering (Fig. 1a)17,18.
a, Graphical overview of the genetic modifications. P. putida AG5577 contains a total of nine distinct attB sites in three different locations in its genome. Three genomic regions involved in lactam catabolism were sequentially deleted from AG5577 to generate the LP strain (ΔoplBA, ΔPP_5182 and ΔdavAB). Lys, lysine; 5-AVA, 5-aminovaleric acid; 6-ACA, 6-aminocaproic acid; VL, valerolactam; CL, ε-caprolactam. b, Measurement of 5-AVA and VL levels in AG5577 and comparative analysis of oplBA, PP_5182 and davAB deletion. ND, not detected. c, VL and CL degradation assay with the LP strain using media supplemented with 50 μM VL (4.96 mg l−1) and CL (5.66 mg l−1), respectively. Dashed lines indicate the supplemented levels of VL and CL, respectively. All data are presented as mean and s.d. of three biological replicates.
The versatile metabolic capabilities of P. putida offer substantial advantages over other hosts but can also be challenging if target compounds are being catabolized19. In the case of lactam metabolism, P. putida has been thoroughly studied using random barcode transposon sequencing (RB-TnSeq), revealing OplBA as a crucial enzyme catalysing the ring-opening reaction for C5- and C6-lactams20. Accordingly, we deleted the oplBA genes in P. putida AG5577, which resulted in a minor accumulation of δ-VL (<0.1 mg l−1). In P. putida, VL is a known by-product of l-lysine (l-Lys) degradation20 (Fig. 1a,b). Interestingly, this modification increased production of the VL precursor, 5-AVA. In addition to deleting oplBA, we also deleted PP_5182, encoding an aminotransferase that converts 6-aminocaproic acid (6-ACA) into 6-oxohexanoic acid, assuming that its deletion would prevent the potential loss of non-cyclized product (Fig. 1a)21; however, its deletion had no effect on either endogenous 5-AVA or VL production (Fig. 1b).
5-AVA is known to be activated by endogenous CoA ligases, triggering the spontaneous formation of VL (Fig. 1a)20. To eliminate this natural occurrence and distinguish it from the heterologous production of 5-AVA by our PKSs, we deleted the genes for DavAB, which transform l-Lys into 5-AVA. In spite of abolishing 5-AVA production, we still detected trace amounts of VL, suggesting the potential existence of an alternative VL biosynthesis pathway in P. putida. Given these circumstances, we set the baseline VL production level at 0.1 mg l−1 after 4 days of culture. Culturing the final strain with exogenous VL or ε-caprolactam (CL, C6-lactam) had no impact on their final concentrations, confirming successful genetic modifications (Fig. 1c). In summary, several rounds of genetic modifications generated a P. putida strain, named LP, specifically designed for heterologous lactam production. The strain is deficient in catabolizing C5- and C6-lactams and cannot produce the VL precursor, 5-AVA.
PKS pathway design for C5-lactam production
Most macrolactam polyketides are assembled by hybrids of PKSs and non-ribosomal peptide synthases (NRPSs)22. Another strategy is the formation of an aminoacyl starter unit, which is loaded directly onto the first ACP domain of the PKS. The fluvirucin loading pathway utilizes l-aspartate (l-Asp) as the precursor, and its β-amino-acid derivative, β-alanine (β-Ala), to initiate polyketide synthesis23. A typical two-carbon extension with product release by a lactamizing TE domain would result in the formation of the desired C5-lactams. Based on this design, we reprogrammed the fluvirucin biosynthetic pathway of Actinomadura vulgaris and Actinomadura fulva to produce VL and α-carbon derivatives24. Our PKS-based retrobiosynthesis approach requires three parts: β-amino-acid loading, a chimeric PKS, and the production of extender units (Fig. 2a).
a, Proposed PKS-based VL and α-substituted VL production pathway inspired by the fluvirucin biosynthetic pathway from A. vulgaris and A. fulva. l-Asp, l-aspartate; l-Ala, l-alanine; ACPL, N-terminal loading acyl carrier protein; KS, ketosynthase; AT, acyltransferase; DH, dehydratase; ER, enoyl reductase; KR, ketoreductase; ACP, acyl carrier protein; TE, thioesterase; VL, δ-valerolactam; MVL, α-methylvalerolactam; EVL, α-ethylvalerolactam; IBVL, α-isobutyrylvalerolactam; M-CoA, malonyl-CoA; MM-CoA, (2S)-methylmalonyl-CoA; EM-CoA, ethylmalonyl-CoA; and IBM-CoA, isobutyrylmalonyl-CoA. b, In vitro verification of the β-amino-acid loading pathway with LC-ESI-MS traces of FlvL, FlvK and ACPL modifications. Numbers next to peaks represent their exact m/z value.
To verify the first part of our design, we expressed and purified the enzymes involved in the β-amino-acid loading pathway, including the excised N-terminal loading ACP (ACPL) domain of the fluvirucin PKS (Supplementary Fig. 1). In vitro enzymatic reactions were conducted with purified enzymes and their substrates, followed by monitoring the intact protein mass change using MS. l-Asp loading by the adenylation enzyme FlvN was confirmed through the expected mass shift of the accepting ACP, FlvL. Addition of the pyridoxal-phosphate dependent decarboxylase, FlvO, and the second adenylation enzyme, FlvM, in the presence of l-alanine (l-Ala) enabled the detection of the dipeptidyl-FlvL intermediate. Finally, we confirmed the transfer of this intermediate from FlvL to the ACPL domain by FlvK, a trans-acting AT (Fig. 2b). Overall, these results confirm the functionality of the β-amino-acid loading pathway, successfully loading the C3 intermediate derived from l-Asp onto the ACPL of the fluvirucin PKS.
The chimeric PKS design consists of a single module PKS that extends the loaded intermediate through condensation with malonyl-CoA (M-CoA) or one of its analogues. Depending on which AT domain is used, the recruited extender unit and, consequently, the functional group at the α-carbon of the final lactam can be modified. Following polyketide extension, a full reductive loop, composed of DH, ER and KR domains, fully reduces the β-carbon ketone. After chain extension and modification, FlvJ hydrolyses the l-Ala protection group from the polyketide backbone. Finally, the native TE domain of the fluvirucin PKS releases and cyclizes the amino fatty acyl chain into a lactam.
In vivo production of α-ethylvalerolactam
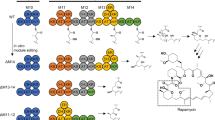
We chose α-ethylvalerolactam (EVL) as the initial target to validate the pathway and assess the feasibility of in vivo production. The PKS-based production pathway was divided into three genetic parts (Fig. 2a), each individually introduced into one of the nine genomic attB sites of the LP strain using SAGE (Fig. 3a).
a, Graphical overview of the three parts integration into the LP strain genome. Co-transformation of plasmids encoding a serine recombinase facilitates the integration of plasmids containing each part into the corresponding genomic attB site. EM, EM-CoA producing operon encoding ccr. b, Protein levels of different codon variants of FluPks1 and FluPks5, along with Ccr for EM-CoA production. Codon optimization methods: GS, GenSmart; UBC, use best codon; MCU, match codon usage. c, EVL production level in strains with combinations of three versions of the β-amino-acid loading part and six PKSs, accompanied by the extender unit production part. ND, not detected. The strain with LM2, EM and FluPks1-MCU is named EVL-S1, and the strain with LM3, EM and FluPks5-MCU is named EVL-S2. d, Enhancing EVL production through the introduction of an additional copy of flvJ and optimizing culture duration. PJ23100-FlvJ plasmid was integrated into the TG1 attB site of EVL-S1 and EVL-S2, resulting in EVL-S1-FlvJ and ELV-S2-FlvJ, respectively. All data are presented as mean and s.d. of three biological replicates.
The first part of our designed pathway is loading of the β-amino acid onto the PKS by expressing genes encoding FlvKLMNO using two operons (FlvNLO and FlvMK). We also designed a third operon (FlvJ and GdmF25) to remove the l-Ala protection group and facilitate the final lactamization step. All three operons were cloned into plasmids with different combinations of promoters, resulting in three different versions of the β-amino-acid loading, product deprotection and cyclization part, named LM1, 2 and 3 (Fig. 3a). The three vectors were integrated into the RV attB site, and proteomic analysis confirmed that all seven enzymes were successfully expressed in M9 minimal medium (Supplementary Fig. 2).
The second genetic part is the chimeric PKS, designed to closely mimic its natural configuration and maintain functionality. We designed two chimeric PKSs, FluPks1 and FluPks5, each incorporating a single unnatural junction between modules within the fluvirucin PKSs. FluPks1 was created by connecting module 1 (KS–AT–DH–ER–KR–ACP), including the ACPL domain, to the TE domain of module 5. Alternatively, for FluPks5, module 5, including the TE domain (AT–DH–ER–KR–ACP–TE), was fused to the C terminus of the ACPL and KS domains of module 1 (Supplementary Fig. 3). Both module 1 and module 5 of the fluvirucin PKS have a full reductive loop and utilize ethylmalonyl-CoA (EM-CoA) as the extender unit24, probably leading to EVL production (Supplementary Fig. 3).
For successful heterologous production, the quantity of correctly folded and soluble PKSs is crucial. The gene sequences of the designed chimeric PKSs are large (~7.5 kbp) and high in GC content (~70%), making their expression particularly sensitive to the effects of codon usage18. We thus recoded the gene sequences for FluPks1 and FluPks5 using three different methods: GenSmart (GS) by GenScript, ‘use best codon’ (UBC) and ‘match codon usage’ (MCU) by BaseBuddy18. A total of six PKS genes with distinct codon usage patterns were integrated into the BxB1 attB site of LP strains containing either LM1, 2 or 3 in their RV attB site, resulting in 18 potential EV-producing strains (Fig. 3a; strains 17–34 in Supplementary Table 4). Proteomic analysis of the strains carrying LM2 along with each of the six PKS genes revealed that the MCU method resulted in the highest PKS protein levels, with at least 2.3-fold higher peptide counts compared to the other methods (Fig. 3b).
The third genetic part for EVL production is responsible for the supply of the extender unit, EM-CoA. Because P. putida lacks an intrinsic pathway for EM-CoA biosynthesis, a heterologous pathway was introduced by expressing crotonyl-CoA carboxylase (Ccr) (that is, the EM part)26, which catalyses the reductive carboxylation of crotonyl-CoA, naturally produced by P. putida, into EM-CoA (Fig. 3a). An expression cassette for the ccr gene was integrated into the R4 attB site of strains containing the LM and PKS parts. The presence of Ccr was confirmed by detection of the corresponding peptides (Fig. 3b).
Liquid chromatography–mass spectrometry (LC–MS) analysis of the final 18 strains containing the complete lactam pathway confirmed that all strains, except for one, successfully produced EVL in the range of 0.02–0.12 mg l−1 (Fig. 3c and Supplementary Fig. 4). The EVL mass peak and fragmentation pattern matched those of an authentic standard (Supplementary Figs. 4 and 5). Importantly, only strains that had all three of the required genetic parts produced EVL, highlighting the indispensable role of each part in the overall design (Fig. 3c and Supplementary Fig. 4a). EVL titres were strongly correlated with PKS protein levels. The MCU codon optimization method resulted in higher protein levels compared to the other optimization methods, ultimately contributing to the increased production of the target compound (Fig. 3b,c). This underlines the importance of codon optimization for the heterologous expression of PKS genes. MCU-optimized FluPks1 and FluPks5 were chosen as the foundational PKSs for further PKS engineering efforts.
To further enhance the EVL titres, we optimized the medium and culture conditions. Initially, a 48-h culture in M9 minimal medium yielded the highest EVL titres with LM2 and FluPks1 (EVL-S1, 0.08 mg l−1) and LM3 and FluPks5 (EVL-S2, 0.12 mg l−1) (Supplementary Fig. 6). Supplementing the media with 5 g l−1 of yeast extract as a protein expression enhancer increased the EVL titres by 1.5- and 1.3-fold, respectively (Supplementary Fig. 6). Despite the negligible improvement in heterologous enzyme expression, a reduction in the expression of the small heatshock protein IbpA was observed (Supplementary Fig. 7a,b). As IbpA serves as an indicator of misfolded proteins, this reduction suggests an increase in properly folded and functional enzymes27. Additionally, supplementing 5 mM l-Asp, a precursor for the EVL loading pathway, increased EVL production by 2.6- (EVL-S1) and 1.8-fold (EVL-S2) compared to the initial medium composition (Supplementary Fig. 6). Proteomics analysis revealed FlvJ as the lowest-expressed heterologous enzyme in the EVL pathway. To address this deficit, we introduced an additional copy of flvJ into the TG1 attB site of the EVL producers, increasing the FlvJ protein levels 2.9-fold (Supplementary Fig. 7c). Consequently, EVL titres for EVL-S1 and EVL-S2 increased by 4.9- and 1.9-fold, respectively. Time-course measurements showed a steady increase in EVL production over 4 days, with the EVL-S1 strain achieving a final titre of 1.82 mg l−1, representing an ~23-fold overall increase (Fig. 3d).
In vivo production of VLs via AT-domain exchange
With the functional PKS system established for EVL production, we could now engineer the PKSs to incorporate extender units with the desired functional groups for producing VL and other α-substituted VLs. Because the AT domains of PKSs determine the incorporated extender unit, we selected seven diverse ATs to replace the AT domains of FluPks1 and FluPks5: three M-CoA-specific ATs, three ATs specific for (2S)-methylmalonyl-CoA (MM-CoA), and one AT that incorporates isobutyrylmalonyl-CoA (IBM-CoA) (Supplementary Table 1).
A common AT-domain exchange strategy includes replacing the entire KS–AT linker (KAL), the AT itself, and the initial segment of the post-AT linker (PAL1)3,28. Following this approach with the updated junction boundaries by Englund et al.3, we exchanged the AT domains in FluPks1 and FluPks5 with seven selected ATs, resulting in a total of 14 AT-exchange mutants (Fig. 4a). Because VL biosynthesis via PKS requires M-CoA, which is naturally available in P. putida, all tested AT domains were expected to exhibit some level of promiscuity towards it. We thus introduced all 14 AT-exchanged PKSs into the BxB1 attB site of the LP strain already containing LM2 or LM3, resulting in 28 combinations (Fig. 4a; strains 35–62 in Supplementary Table 4). The strains were cultured in the optimized condition for EVL production, and VL levels were monitored by LC–MS (Supplementary Figs. 8a and 9a). Notably, all seven AT domains were promiscuous towards M-CoA and enabled VL production. The FluPks1 and LM3 combination generally achieved higher titres than all other combinations, with FluPks1-BorM1AT and LM3 (VL-S1) yielding the highest titre at 0.5 mg l−1 (Fig. 4a).
a, Production of VL in LP strains using 14 AT-exchanged PKSs, combined with LM2 or LM3 as the β-amino-acid loading part. The best producer was named VL-S1. Dashed lines indicate the baseline production level of VL (0.1 mg−1). b, Engineered biosynthetic pathways for extender unit production: MM, MM-CoA producing operon encoding MCM, MeaB and Epi; EM, EM-CoA producing operon encoding Ccr; IBM, IBM-CoA producing operon encoding AnlE, AnlF and AnlG. The MM operon was integrated into the MR11 attB site, and EM and IBM were integrated into the R4 attB site in the genome. Glc, glucose; Pyr, pyruvate; Ac-CoA, acetyl-CoA; IB-CoA, isobutyryl-CoA; Cro-CoA, crotonyl-CoA; Suc-CoA, succinyl-CoA. c, Production of MVL in LP strains using six AT-exchanged PKSs, combined with LM2 or LM3 as the β-amino-acid loading part and MM for extender unit production. The best producer was named MVL-S1. d, Production of IBVL in LP strains using two AT-exchanged PKSs, combined with LM2 or LM3 as the β-amino-acid loading part and IBM for extender unit production. The best producer was named IBVL-S1. e, Enhancing VL and α-substituted VLs production through the introduction of an additional copy of flvJ. The PJ23100-FlvJ cassette was integrated into the TG1 attB site of the best producers, resulting in VL-S1-FlvJ, MVL-S1-FlvJ and IBVL-S1-FlvJ. Significance was assessed by a two-tailed Student’s t-test (***P < 0.001). Exact P values were 3.6 × 10−5, 1.5 × 10−4 and 2.0 × 10−4, respectively. f, VL production from VL-S1-FlvJ in media supplemented with amino-acid mix. ND in all panels indicates not detected. Significance was assessed by a two-tailed Student’s t-test (***P < 0.001). Exact P values were 3.0 × 10−4, 4.0 × 10−4 and 4.0 × 10−6, respectively. All data are presented as mean and s.d. of three biological replicates.
To facilitate α-methylvalerolactam (MVL) and α-isobutyrylvalerolactam (IBVL) production, P. putida requires heterologous pathways for their corresponding extender units: MM-CoA and IBM-CoA (Fig. 4b). For MM-CoA synthesis, we employed an operon from Sorangium cellulosum encompassing MM-CoA epimerase (Epi), MM-CoA mutase (MCM) and an enzyme stabilizer (MeaB) (that is, the MM part)29. For IBM-CoA production, we chose an operon from the Streptomyces sp. CNH18913 ansalactam A biosynthetic gene cluster (that is, the IBM part), which consists of a crotonyl-CoA carboxylase (AnlE), a β-ketoacylsynthase (AnlF) and a dehydrogenase (AnlG)30. Through combinatorial strain construction with AT-exchanged PKSs and corresponding extender unit production parts, we successfully produced MVL and IBVL in the LM2 and LM3 strains, achieving the highest titre of 4.8 mg l−1 (MVL-S1) and 0.4 mg l−1 (IBVL-S1), respectively (Fig. 4c,d, Supplementary Note 1 and Supplementary Figs. 8b,c and 9b,c). To increase the lactam titres, we introduced an additional FlvJ expression cassette into the VL-S1, MVL-S1 and IBVL-S1 strains, resulting in significant improvements (1.44, 10.03 and 0.67 mg l−1, respectively; Fig. 4e). These results are consistent with EVL production, highlighting that low FlvJ levels are a general bottleneck in the designed pathway. Although some VL was produced due to AT promiscuity for M-CoA, the introduction of pathways for the synthesis of the desired extender unit reduced M-CoA incorporation and thus VL production (Supplementary Fig. 10). In some cases, despite containing all necessary components, no production occurs. For example, FluPks5-AnsM8AT with the IBM part in the LM3 strain failed to produce IBM (Fig. 4d). Proteomic analysis showed similar enzyme expression for β-amino-acid loading and PKS, but lower IBM part enzyme levels and higher IbpA levels in LM3 strains (Supplementary Fig. 11).
To validate extender unit production, we assessed the concentration of acids derived from extender unit hydrolysis in strains with the highest titre for VL and α-substituted VLs. P. putida possesses ~13 genes encoding type-II thioesterases, which may convert extender acyl-CoAs into their acid forms31. Thus, measuring the respective acid provides indirect evidence for the production of the extender units. Malonate, derived from M-CoA, was detected in all strains as a consequence of native metabolic pathways. In contrast, methylmalonate, ethylmalonate and isobutyrylmalonate were only observed in strains expressing the corresponding extender unit parts (Supplementary Fig. 12). Unexpectedly, the MVL producer MVL-S1-FlvJ synthesized 0.88 g l−1 and 11.78 g l−1 malonate in a 24-well plate and 1-l fed-batch bioreactor culture, respectively (Supplementary Figs. 12 and 13a). Further validation led us to hypothesize that the MM part increases the metabolic flux towards M-CoA, and the PikM6 AT preferentially hydrolyses M-CoA instead of incorporating it into the polyketide backbone, as previously reported32 (Supplementary Note 2 and Supplementary Fig. 13b).
A 1-l fed-batch cultivation of VL-S1-FlvJ and MVL-S1-FlvJ yielded a similar VL and MVL titre as a 24-well plate culture (Supplementary Fig. 13a,c), suggesting the presence of a production bottleneck that cannot be alleviated through scale-up. To track metabolic changes occurring throughout the strain engineering process and identify potential bottlenecks, we conducted a multi-omics analysis on strains associated with VL production, including LP, LM3, VL-S1 and VL-S1-FlvJ. Global proteomic and metabolomic analysis revealed a severe nitrogen limitation, driven by low availability of amino acids in the media, as a potential bottleneck for the production (Supplementary Note 3 and Supplementary Figs. 14 and 15). Otherwise, β-Ala levels gradually increased with each genetic modification of the strain (Supplementary Fig. 15a), suggesting its release from FlvL or FlvK, potentially with FlvJ-mediated deamination. Introducing an additional flvK expression cassette led to a slight increase in VL titre (Supplementary Fig. 15b). However, the possibility that β-Ala release from FlvL or FlvK is reversible, and that its accumulation reflects an increased metabolic flux toward VL production, cannot be excluded. To validate whether amino-acid starvation limits VL production, we supplemented the medium with a mix of all 20 proteinogenic amino acids and β-Ala. Supplementation at 0.5, 2.5, 5 and 10 mM significantly increased the VL titre of VL-S1-FlvJ by 1.3-, 2.6-, 3.8- and 5.2-fold, respectively (Fig. 4f). It is likely that future optimization of the amino-acid composition or enhancement of endogenous amino-acids biosynthesis could lead to further improvements in the product titres.
In vivo production of VLs via full module exchange
To expand the versatility of PKS engineering beyond AT-domain exchanges, we explored full module (KS–AT–DH–ER–KR–ACP) exchange for VL production. Leveraging the ClusterCAD database33, we identified 20 potential PKS-module candidates for full module exchange for VL production (Supplementary Note 4 and Supplementary Table 2) and selected the five modules (CmiPksM2, VstPksM3, NemPksM3, FluPksM4 and GdnPksM3) with the most distinct protein sequences (Supplementary Fig. 16). Unlike AT-domain exchanges, effective junction sites for full module exchange are not well established. To systematically identify the optimal N-terminal junction site for full module replacement, we performed multiple sequence alignment (MSA) on 300 randomly selected KS–AT amino-acid sequences, revealing 93 conserved positions with over 95% sequence identity (Supplementary Fig. 17). We designed and constructed 140 PKS variants by pairing the five donor modules with FluPks1 as the acceptor PKS, utilizing one of the conserved N-terminal positions and the FD motif at the C-terminal junction (Supplementary Note 4 and Supplementary Fig. 18a). After introducing the PKSs into the LM3 strain, 65 of the 140 variants exhibited VL titres exceeding the baseline production (0.1 mg l−1; Supplementary Fig. 18b). Notably, high titres were achieved for CmiPksM2, NemPksM3 and FluPksM4 when using positions near the end of the KS domain as the N-terminal junction. These findings align with recent proposals to redefine PKS-module boundaries, suggesting that modules terminate with the KS domain rather than initiate with it34. In summary, in addition to AT-domain exchanges, full module replacement proves to be a viable strategy for lactam analogue production. The computational screening from the database enabled us to identify unexplored donor modules, expanding the repertoire for PKS-module engineering.
Sustainable and stereocontrolled PKS-based α-substituted VLs
In addition to the ease of changing the functional group attached to the VL ring through AT-exchange, PKS-based biosynthesis of VLs offers two other key advantages compared to chemical synthesis: renewable carbon substrate utilization and the stereospecific production of enantiopure compounds.
First, P. putida possesses a remarkable ability to exploit renewable carbon sources. To validate this capability, we conducted experiments on the production of VL and α-substituted VLs from a 4× diluted lignocellulosic biomass hydrolysate derived from sorghum35. Notably, all four VL analogues were successfully produced from these plant-derived carbon sources (Fig. 5a). Specifically, the production of MVL exhibited an even higher titre (13.6 mg l−1) compared to M9 medium-based production, suggesting the potential for further engineering to enhance titres by optimizing biomass utilization.
a, Quantification of VL and α-substituted VLs produced from plant biomass. VL-S1-FlvJ, MVL-S1-FlvJ, EVL-S1-FlvJ and IBVL-S1-FlvJ were cultured in 4× diluted sorghum hydrolysate for VL and α-substituted VLs production. All data are presented as mean and s.d. of three biological replicates. b–d, Chiral LC–MS profiles for the separation of enantiomers detected using an MS detector for IBVL (b), MVL (c) and EVL (d). Dashed lines represent S- or R-configured enantiomers, grey lines denote chemically synthesized racemic mixtures, and red lines indicate samples obtained from P. putida strains (IBVL-S1-FlvJ, MVL-S1-FlvJ and EVL-S1-FlvJ).
Second, α-substituted VLs feature a stereocentre on the α-carbon with a substituent, and their chemical synthesis typically results in a racemic mixture of R- and S-configured products (Fig. 5b–d). Producing enantiopure lactam-based monomers enables the synthesis of stereocontrolled polyamides without chiral polymerization catalysts, substantially affecting their thermal, mechanical and optical properties36,37. Varying the enantiomer content within the polymer backbone further allows for tunable thermomechanical properties. Additionally, stereochemistry influences the thermoresponsive properties of water-soluble polymers such as poly(N-isopropylacrylamide) (pNIPAM)38,39. PKS-based production circumvents this issue through its stereoselectivity, which is controlled by reduction-related domains, yielding enantiopure products40,41. The predicted stereochemistry of our system for lactam production is obtained as follows (Supplementary Fig. 19)41: condensation of S-configured M-CoA analogues as extender units by KS and AT domains inverts the α-substituent from the S- to R-configuration; the KR domains of both FluM1 and FluM5 are of the B1-type, maintaining the R-configuration; most DH domains following B-type KR domains form a trans-olefin structure; the ER domains of both FluM1 and FluM5 are of the D-type, generating R-configured intermediates for MVL and EVL, and an S-configured intermediate for IBVL; and cyclization results in (2R)-MVL, (2R)-EVL and (2S)-IBVL, respectively.
Indeed, chiral LC–MS analysis confirmed that chemically synthesized α-substituted VLs are racemic mixtures, whereas those derived from our strains with PKSs are enantiomerically pure. Notably, IBVL presented a particularly clear result, with two distinct peaks for chemical synthesis and a single peak for PKS-based production (Fig. 5b). VL, lacking a stereocentre, consistently exhibited a single peak in both chemical synthesis and PKS-based production (Supplementary Fig. 20). The separation of MVL and EVL enantiomers proved challenging; nevertheless, chemical synthesis resulted in a merged broad peak, whereas PKS-based production yielded a narrower and singular peak (Fig. 5c,d). The stereoselectivity of PKSs thus facilitates enantiopure production, which is often crucial for the functionality of the resulting monomers or polymers. Engineering reduction-related domains in PKSs would allow us to alter the stereochemistry of the substituent on the α-carbon.
Polymerization of α-substituted VLs
To demonstrate the feasibility of polymerizing α-substituted VLs, we performed ROP and copolymerization with CL, butyrolactam and benzocaine to form polyamides (Supplementary Fig. 21). However, steric hindrance from α-carbon substituents impeded the polymerization, in line with the recent finding on α- and β-carbon-methylated CL42. To overcome this issue, we used a ‘Trojan horse’ strategy that has been used recently to polymerize several classes of weakly-strained monomers by ROP43,44. In our system, VL monomers were first activated by converting the lactam into its corresponding tosyl sulfonamide, followed by ring-opening with a diamine and subsequent tosyl group removal to give a new diamine comprising two VL monomer units. This VL-based diamine was reacted with adipoyl chloride via interfacial polymerization to rapidly produce polyamide copolymers at room temperature (Fig. 6a). Unlike random ring-opening copolymerization of VL with other lactam monomers, this step-growth approach ensures a precise sequence-defined microstructure that will be crucial for controlling thermomechanical and electronic properties.
a, Synthesis and interfacial step-growth polymerization of ring-opened VL and MVL. b, Differential scanning calorimetry thermogram of MVL- and VL-derived polyamides synthesized by interfacial step-growth polymerization. c, Synthesis and RAFT polymerization of N-acryloyl VL and the α-substituted VLs monomer. [monomer]0:[CPDT]0:[AIBN]0 = 100.0:1.0:0.2. The conversion rate was determined by 1H NMR analysis of the crude polymerization mixture. The theoretical number-average molecular weight was calculated based on monomer conversion according to Mn,th = (ρ × MWmon × [monomer]0/[CPDT]0) + MWCPDT, where ρ × MWCPDT is the fractional monomer conversion, MWmon is the molecular weight of the monomer, and MWCPDT is the molecular weight of the RAFT agent, CPDT (345.62 g mol−1). Mn,GPC and dispersity Đ were determined by GPC in dimethylformamide, calibrated with polystyrene standards. THF, tetrahydrofuran; conv., conversion.
We applied this method to VL and racemic MVL, both producible on a multigram scale. Unlike the ROP of low-strain monomers, which often yields low molecular weights, our step-growth approach produced polyamides with limited solubility in common organic solvents for polyamides, such as nylon-6 (for example, fluorinated alcohols), suggesting successful synthesis of high-molecular-weight polymers. Although poor solubility precluded molecular weight characterization by gel permeation chromatography (GPC), differential scanning calorimetry thermograms confirmed glass transition temperatures (Tg) of 38 °C and 32 °C for VL- and racemic MVL-derived polyamides, respectively, which are comparable to those of other aliphatic polyamides. Only the VL-derived polymer exhibited a melting temperature (Tm) of 190 °C (Fig. 6b), whereas the racemic MVL-derived polymer—atactic and expected to be amorphous—lacked a Tm. We anticipate that synthesizing polyamides from enantiopure monomers will yield semicrystalline polymers, with melting temperatures dependent on the chiral side-chain substituent (for example, Me, Et and iBu) and stereoregularity.
We next took inspiration from extensive works on the synthesis and application of polymers derived from N-vinyl lactam monomers such as N-vinylcaprolactam, N-vinylpiperidone and N-vinylpyrrolidone11,12,13. Poly(N-vinyl lactam)s comprise all-carbon backbones with pendant lactam side chains, and differ from nylons in that they are water-soluble and biocompatible. These properties have facilitated their widespread application in a range of biomedical fields. The synthesis of N-vinyl lactams typically involves base-promoted conjugate addition of lactam with acetylene. We sought a more accessible and milder route towards synthesizing lactam-based monomers capable of free-radical polymerization and envisioned synthesizing the N-acryloyl analogues of N-vinyl lactams. Substituted N-acryloyl VL monomers were synthesized by sequential reaction of the corresponding α-substituted VLs (MVL, EVL and IBVL) with MeMgBr and acryloyl chloride, followed by flash column chromatography purification (Fig. 6c and Supplementary Figs. 22–29)45. We utilized RAFT polymerization because of its functional group tolerance and molecular weight control. The RAFT polymerization of N-acryloyl VL analogues was conducted at 70 °C in dimethylformamide using 2-cyano-2-propyldodecyl trithiocarbonate (CPDT) as the chain transfer agent (Fig. 6c). Unsubstituted N-acryloyl VL (Fig. 6c, entry 1) resulted in near-quantitative monomer conversion within 22 h, yielding poly(N-acryloyl VL) with sufficient molecular weight (Mn,GPC = 9.6 kDa) and moderately low dispersity (Ð = 1.28). Polymerization of the methyl- and ethyl-substituted N-acryloyl VL derivatives (entries 2 and 3, respectively) resulted in intermediate conversions (46% and 68%, respectively) and proportionally lower molecular weights, probably due to the reduced polymerization rate of these more substituted monomers. The isobutyl-substituted N-acryloyl VL (entry 4) showed even lower conversion (15%) and molecular weight (1.9 kDa), indicating that prolonged reaction times and/or higher reaction temperatures are probably required for higher conversions of the sterically more hindered monomers. Future efforts will explore the solution properties of these lactam-derived polymers and their potential application as biomaterials.
Discussion
PKSs have emerged as pivotal enzymes in the realms of synthetic biology and retrobiosynthetic strategies for synthesizing unconventional compounds. The predominant focus of PKS engineering has revolved around domain or module exchanges, but recent efforts have shifted towards a comprehensive reprogramming of the entire PKS-based pathway. This paradigm shift has enabled the synthesis of a diverse array of commodity chemicals, such as ketones, lactones and fatty-acid derivatives7,8,35. Our research further explores the vast potential of PKSs and has added lactams to this repertoire.
To synthesize C5-lactams, including VL and α-substituted VLs, we reprogrammed the fluvirucin PKS and evaluated 156 chimeric PKS designs through module reconfiguration and domain/module exchanges, enabling the production of VL and three α-substituted VLs in P. putida. This high-throughput approach provided key insights into PKS engineering, highlighting the importance of selecting suitable donor PKSs and defining the boundary of each module as the end of the KS domain for successful domain/module exchanges. The resulting dataset holds the potential to serve as a training set for machine-learning tools, aiding in linking PKS amino-acid sequences and functionality to advance computational PKS design for predicting and optimizing biosynthetic outcomes.
Given the diverse repertoire of over 20 naturally occurring M-CoA/ACP analogues recognized as PKS extender units3, the field of PKS engineering presents a promising opportunity to expand this range and facilitate the synthesis of innovative lactam derivatives. Although concerns about low titres with PKS-based retrobiosynthesis persist, recent studies have demonstrated the feasibility of PKS-based production nearly reaching the g l−1 scale8,46 (Supplementary Note 5). As numerous pharmaceuticals, including antibiotics47 and immunosuppressants48, are industrially produced by native PKSs in their hosts, we anticipate that optimizing engineered PKS expression and host engineering will facilitate industrial-scale production. Our multi-omics-guided approach, with subsequent host engineering and media optimization, resulted in a 23-fold increase for EVL and an eight-fold increase for VL, demonstrating great potential for further optimization.
The PKS-based biosynthesis of lactams has several advantages over chemical synthesis. First, utilizing P. putida as a host enables the use of renewable carbon sources, as demonstrated by the successful production of VL and three α-substituted VLs from plant biomass hydrolysate. Second, the inherent stereoselectivity of PKSs allows precise control over stereochemistry, facilitating the production of enantiopure α-substituted VLs. This is particularly important for the subsequent synthesis of stereocontrolled polymers, eliminating the need for expensive and toxic chiral polymerization catalysts.
Lactams are primarily recognized as monomers for polyamides or temperature-sensitive/water-soluble polymers, showcasing their utility in a diverse range of biomedical applications. To address the challenges of low ring strain and steric hindrance in the ROP of α-substituted VLs, we employed a Trojan horse strategy that has been used recently to polymerize several classes of weakly-strained monomers by ROP43,44. This approach enabled the successful synthesis of polyamides from VL and racemic MVL, allowing property comparison. Although synthesizing stereoregular polyamides remains challenging due to the need for additional efforts to biologically produce larger quantities of each enantiopure VL, we anticipate that the demonstrated polymerization method will also be effective for synthesizing polyamides from other enantiopure α-substituted VLs, crucial for developing nylon-5 derivatives with properties suited for applications such as thermoplastics or ferroelectric materials. Applying PKS-based monomer diversification to C6-lactams, which possess sufficient ring strain for ROP, will enable the feasible synthesis of nylon-6 materials. Additionally, RAFT polymerization of N-acryloyl derivatives of VL and α-substituted VLs has demonstrated their potential as innovative monomers for temperature-sensitive polymers, with applications across various contexts. In summary, the retrobiosynthetic PKS reprogramming demonstrates the versatility and untapped potential of PKSs for broadening the range of compounds that can be biosynthesized from renewable feedstocks.
Methods
Materials
The VL was purchased from MilliporeSigma, and its derivatives were sourced from Enamine. All other unspecified chemicals were purchased from MilliporeSigma.
Culture conditions
Escherichia coli and P. putida were cultured in LB medium for routine strain construction and maintained at 30 °C and 200 r.p.m. with the appropriate selection marker at the following final concentrations: chloramphenicol (20 μg ml−1), kanamycin (50 μg ml−1), gentamicin (30 μg ml−1) and apramycin (50 μg ml−1). For lactam production, overnight cultures grown in LB medium with a selection marker were diluted 100× into 2 ml of M9 medium (1 × M9 salts, 1 mM MgSO4, 0.1 mM CaCl2, 2.0% glucose and 0.5 μg ml−1 thiamine) (Teknova) with 1× trace metals mixture (Teknova) in 24-well plates (VWR) sealed with an AeraSeal film and grown at 30 °C, 200 r.p.m. For media optimization, M9 medium was supplemented with 5 g l−1 of yeast extract (M9YE media) and 5 mM of aspartate (M9YE+Asp media). If not indicated otherwise, all cultures were performed in M9YE+Asp medium. For additional amino-acid supplementation, we added equal concentrations of all 20 proteinogenic amino acids (alanine, arginine, asparagine, aspartate, cysteine, glutamate, glutamine, glycine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, proline, serine, threonine, tryptophan, tyrosine and valine) along with β-alanine into M9YE+Asp. Sorghum hydrolysate was diluted 4× using M9 medium, resulting in a final composition of 11.8 g l−1 glucose, 6.4 g l−1 xylose, 4.2 g l−1 acetic acid, 4.3 g l−1 lactic acid and unquantified amounts of choline and lignin monomers.
Plasmid design and construction
All plasmids and oligonucleotides created in this study are described in Supplementary Tables 3 and 5. They are available through the public version of the JBEI registry (https://public-registry.jbei.org/folders/827). Plasmids and assembly processes were designed using Device Editor, Vector Editor Software and DNAda49,50,51. Primers for plasmid constructions were designed using j5 software52 and purchased from Integrated DNA Technologies (IDT). PKS DNA fragments were codon-optimized using the online tool BaseBuddy18 and synthesized (Genscript). Q5 (NEB) or Phusion (Thermo Fisher Scientific) High-Fidelity DNA Polymerase was used to amplify DNA fragments for plasmid construction. Gibson assembly was used to construct the plasmids with NEBuilder HiFi DNA assembly master mix (NEB), which were then transformed into NEB 5-alpha F’Iq-competent E. coli (NEB), and isolated using a Qiaprep Spin Miniprep kit (Qiagen) following the manufacturer’s instructions.
P. putida strain construction
All strains created in this study are listed in Supplementary Table 4. They are available through the public version of the JBEI registry (https://public-registry.jbei.org/folders/827). The parent strain used in this study is P. putida AG5577, which is derived from strain KT2440 and was provided by A. Guss at Oak Ridge National Laboratory. Deletion mutants of AG5577 were generated using homologous recombination53. Homology fragments of ~1 kbp upstream and downstream of the target gene were assembled and cloned into the pMQ30 plasmid. The plasmids were then introduced into AG5577 through E. coli conjugation. Gene deletions were verified by polymerase chain reaction amplification of genomic regions encompassing the target genes.
After the deletion of oplBA, PP_5182 and davAB from AG5577, the resulting strain was named LP, and served as the recipient strain for subsequent engineering efforts. Plasmids containing β-amino-acid loading, chimeric PKS and extender unit production parts were integrated into the LP genome using the SAGE method. The selection markers and LacI genes present in the SAGE integration plasmids were excised by phiC31 integrase following each integration. Removal of lacI from the genome results in derepression of lacO-containing promoters, such as PLlacO-1, Ptac and PJ5-lacO-RiboJ18. To prepare competent cells for LP strains, the following steps were performed. An overnight culture (1 ml) was washed twice with an equal volume of 300 mM sucrose, and subsequently resuspended in 0.1 ml of 300 mM sucrose. For the co-electroporation of genetic material, 500 ng of a serine integrase attP site plasmid containing the expression cassette for the gene of interest, along with 500 ng of a non-replicating helper integrase expression plasmid, were introduced into the competent cells through electroporation using a 0.2-cm cuvette at 1.8 kV. After electroporation, the cells were resuspended in 0.5 ml of LB medium and incubated at 30 °C for 2 h to allow for recovery. Subsequently, the cells were plated on an LB plate with an appropriate selection marker for the serine integrase attP site plasmid.
Protein expression and purification
All proteins for in vitro analysis were expressed in E. coli and purified54. Plasmids for expressing FlvN, FlvL, FlvO, FlvM, FlvK and ACPL were transformed into E. coli BL21(DE3). The expression strains were cultured in 1 l of Terrific broth medium at 37 °C with appropriate selection markers. Induction was carried out by adding 0.5 mM isopropyl β-d-1-thiogalactopyranoside after 20 min cooling on ice when the optical density at 600 nm (OD600) reached 0.8. After incubating for 16 h at 18 °C, cells were collected (8,000g at 4 °C for 10 min) and resuspended in 40 ml of lysis buffer (50 mM HEPES, pH 8.0, 0.3 M NaCl, 10% glycerol (vol/vol) and 10 mM imidazole). After sonication on ice, cell lysates were collected (8,000g at 4 °C for 10 min) and proteins were purified using HisPur cobalt resin. The proteins were dialysed against 50 mM HEPES, pH 8.0, 1 mM dithiothreitol and 8% glycerol (vol/vol) and concentrated using 3- or 10-kDa Amicon Ultra filters (MilliporeSigma). The final proteins were analysed on sodium dodecyl sulfate–polyacrylamide gel electrophoresis gel for purity, and concentrations were determined using a Nanodrop 1000 spectrophotometer (Thermo Fisher Scientific). The activity of the purified enzymes was assayed in a one-pot enzymatic reaction. When appropriate, the following protein concentrations were used: 50 μM FlvL, 5 μM FlvN, 5 μM FlvO, 5 μM FlvM, 5 μM FlvK and 5 μM ACPL. Protein reactions were incubated with 2 mM ATP, 2 mM amino acids, 1 mM MgCl2, 50 mM HEPES buffer (pH 7.8) and 10% glycerol in a final volume of 50 μl. Reactions were incubated at 30 °C for 60 min. After incubation, the reactions were passed through sterile spin filters with 0.22-µm pores (MilliporeSigma) and transferred to LC–MS vials.
Liquid chromatography-electrospray ionization MS (intact protein MS) analysis
The mobile phase was composed of solvent A (water, purified by a Milli-Q Gradient ultrapure water purification system (MilliporeSigma) to a resistivity of 18.2 MΩ cm, at 25 °C with 1% formic acid) and solvent B (acetonitrile (Optima grade, Thermo Fisher Scientific) with 1% formic acid). For electrospray ionization mass spectrometry (ESI–MS) of the proteins, an Agilent 1260 Infinity II liquid chromatograph equipped with an Agilent 6530 quadrupole time-of-flight (QTOF) LC–MS system (Agilent Technologies) was used. The LC was carried out using a Proswift RP-4H (monolithic phenyl, 1.0 mm × 50 mm, Thermo Fisher Scientific) analytical column. Samples were injected in volumes of 1 μl, and a solvent B elution gradient ranging from 5% to 100% was run at a flow rate of 0.4 ml min−1 over 2 min at 55 °C. The collected mass spectra were subjected to deconvolution analysis using Agilent MassHunter BioConfirm software 10.0. The deconvolution results were plotted using open-source Chartograph software (www.chartograph.com).
Lactam measurements
An aliquot (0.8 ml) of cell culture was mixed with 0.8 ml of methanol and incubated at 4 °C for 24 h; then 1 ml of supernatant was collected (4,000g for 10 min) and filtered with a 0.2-µm regenerated cellulose filter (Macherey-Nagel). Authentic standards of δ-VL and ε-CL were purchased from MilliporeSigma, and α-substituted VL standards were chemically synthesized (Enamine). LC–MS analysis was performed on an Agilent ZORBAX Eclipse XDB-C18 column (5 µm, 4.6 × 150 mm) with an Agilent InfinityLab LC/MSD XT equipped with an Agilent 1260 Infinity II HPLC at 25 °C. The mobile phase was composed of solvent A (water with 1% formic acid) and solvent B (methanol with 1% formic acid). The following gradient was applied: 0–1 min, 5% B; 1–11 min, a linear gradient of B to 40%; 11–14 min, a linear gradient of B to 95%; 14–16 min, 95% B; 16–17 min, a linear gradient of B to 5%; 17–20 min, 5% B. The flow rate was held at 0.6 ml min−1. The eluent was directed to MS using the ESI positive-ion mode with the following conditions: gas temperature, 325 °C; gas flow, 11.0 l min−1; nebulizer pressure, 50 psig; capillary voltage, 3.5 kV. The scan mode was used in the mass range m/z 90–220 kDa.
The stereochemistry of the α-substituted VLs was analysed by LC–MS with an Astec CYCLOBOND I 2000 HP-RSP Chiral HPLC column (5 µm, 25 cm × 4.6 mm; MilliporeSigma). The column temperature was maintained at 40 °C. The mobile phase and MS settings were the same as above. The following gradient was applied: 0–3 min, 2% B; 3–250 min, a linear gradient of B from 2% to 20%; 250–260 min, a linear gradient of B from 20% to 95%; 260–270 min, 95% B; 270–275 min, a linear gradient of B from 95% to 2%; 275–290 min, 2% B. The flow rate was held at 0.2 ml min−1, except for the period 260–270 min, during which it was increased to 0.8 ml min−1.
Verification of EVL production using LC-QTOF-MS/MS
The LC–MS analysis was conducted on a Kinetex XB-C18 column (2.6 μm, 100 mm × 3.0 mm, Phenomenex) using a 1260 Infinity HPLC system (Agilent Technologies). A sample injection volume of 2 μl was used throughout. The sample tray and column compartment were set to 6 and 25 °C, respectively. The mobile phase was composed of solvent A (water with 10 mM ammonium acetate and 0.2% formic acid) and solvent B (90% acetonitrile and 9.8% water with 10 mM ammonium acetate and 0.2% formic acid). The following gradient was applied: 0–4 min, a linear gradient of B from 90% to 70%; 4–5.5 min, 70% B; 5.5–6.0 min, a linear gradient of B to 40%; 6.0–8.5 min, 40% B; 8.5–9.0 min, a linear gradient of B to 90%; 9.0–11.0 min, 90% B. The flow rate was held at 0.6 ml min−1 for 6.5 min, linearly increased from 0.6 ml min−1 to 1 ml min−1 in 0.5 min, and held at 1 ml min−1 for 4 min. The HPLC system was coupled to an Agilent Technologies 6520 QTOF-MS. The QTOF-MS was tuned with Agilent Technologies ESI-L Low concentration tuning mix in the range of 50–1,700 m/z. Drying and nebulizing gases were set to 12 l min−1 and 25 lb in−2, respectively, and a drying-gas temperature of 350 °C was used throughout. ESI was conducted in positive-ion mode, and a capillary voltage of 3,500 V was utilized. The fragmentor, skimmer and OCT 1 RF Vpp voltages were set to 100 V, 50 V and 250 V, respectively. For targeted MS/MS, a precursor ion of 128.10699 m/z was selected for collision-induced dissociation at a collision energy of 20 eV with a narrow isolation width of 1.3 m/z. The acquisition rates for MS and MS/MS were 0.86 spectra s−1 and 1 spectra s−1, respectively. The data acquisition range was from 50 to 1,100 m/z. Data acquisition (Workstation B.08.00) and processing (Qualitative Analysis B.06.00) were conducted using Agilent Technologies MassHunter.
Proteomic analysis to assess heterologous enzyme levels
Protein was extracted from the cell pellets, and tryptic peptides were prepared55. The cell pellets were resuspended in Qiagen P2 lysis buffer (Qiagen) to promote cell lysis. The proteins were precipitated by the addition of 1 mM NaCl and 4× acetone, followed by two additional washes with 80% acetone in water. The recovered protein pellet was homogenized by pipetting with 100 mM ammonium bicarbonate in 20% methanol. Protein concentration was determined by a DC protein assay (BioRad). Protein reduction was accomplished using 5 mM tris 2-(carboxyethyl)phosphine (TCEP) for 30 min at room temperature, and alkylation was performed with 10 mM iodoacetamide (IAM; final concentration) for 30 min at room temperature in the dark. Overnight digestion with trypsin was accomplished with a 1:50 trypsin:total protein ratio. The resulting peptide samples were analysed on an Agilent 1290 UHPLC system coupled to a Thermo Fisher Scientific Orbitrap Exploris 480 mass spectrometer for discovery proteomics56. Peptide samples were loaded onto an Ascentis ES-C18 column (MilliporeSigma) and eluted from the column using a 10-min gradient from 98% solvent A (water with 0.1% formic acid) and 2% solvent B (acetonitrile with 0.1% formic acid) to 65% solvent A and 35% solvent B. Eluting peptides were introduced to the mass spectrometer operating in positive-ion mode and were measured in data-independent acquisition (DIA) mode with a duty cycle of three survey scans from m/z 380 to m/z 985 and 45 MS2 scans with a precursor isolation width of 13.5 m/z to cover the mass range. DIA raw data files were analysed by an integrated software suite DIA-NN57. The databases used in the DIA-NN search (library-free mode) are P. putida KT2440 latest UniProt proteome (UP000000556) FASTA sequences with the protein sequences of the heterologous proteins and common proteomic contaminants. DIA-NN determines mass tolerances automatically, based on first-pass analysis of the samples with automated determination of optimal mass accuracies. The retention time extraction window was determined individually for all MS runs, analysed via the automated optimization procedure implemented in DIA-NN. Protein inference was enabled, and the quantification strategy was set to Robust LC = High Accuracy. The output main DIA-NN reports were filtered with a global false discovery rate (FDR) of 0.01 on both the precursor level and protein group level. The Top3 method, which is the average MS signal response of the three most intense tryptic peptides of each identified protein, was used to plot the quantity of the targeted proteins in the samples58,59. Generated MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD045574 (ref. 60). DIA-NN is freely available for download from https://github.com/vdemichev/DiaNN.
Extender unit acid form measurements
Quantitation was performed on an Agilent 1200 HPLC system (Agilent Technologies) equipped with an Agilent 6120 mass spectrometer. Each sample and standard (2 μl) were injected into a Luna C18(2)-HST column (25 µm, 2.0 × 100 mm, Phenomenex). The column temperature was maintained at 45 °C, and the buffers used to separate the analytes of interest were solvent A (water with 0.16% formic acid) and solvent B (acentonitrile with 0.16% formic acid). The following gradient was applied: 0–1 min, 0% B; 1–7.67 min, a linear gradient of B to 50%; 7.67–10.33 min, a linear gradient of B to 70%; 10.33–11.67 min, 70% B; 11.67–11.68 min, a linear gradient of B to 0%; 11.68–13 min, 0% B. The flow rate was held constant at 0.50 ml min−1. The MS system was set up in negative-ion mode with a gas temperature of 350 °C, drying gas at 12 l min−1, nebulizer pressure set to 35 psig and a Vcap voltage of 3,000 V. A total of four different masses in SIM mode from the MS detector were used to identify and quantitate the analytes of interest. A mass of 103.10 m/z (M-H)- was used for analyte malonic acid, 117.10 m/z (M-H)- for methylmalonic acid, 131.10 m/z (M-H)- for ethylmalonic acid, and 159.10 m/z (M-H)- for isobutylmalonic acid. Each standard was used to construct a calibration curve in the range of 1–100 μg ml−1. A minimum of four calibration levels were used for each analyte with an R2 coefficient of 0.995 or better. A check calibration standard was analysed every ten samples to ensure the integrity of the initial calibration.
Fed-batch cultivation
Bioreactor cultivation was started by inoculating pre-cultures from glycerol stocks into 5 ml of LB medium and incubating overnight at 30 °C and 225 r.p.m. Overnight cultures were then used to inoculate a second seed culture in LB medium starting at an OD600 of 0.2 using a Genesys 140 spectrophotometer (Thermo Fisher Scientific). The second seed cultures were incubated at 30 °C at 225 r.p.m. for 2–3 h until an OD600 of ~1–2 was reached. The seed culture was then washed three times with modified M9 medium (modified Teknova M9 minimal medium with a glucose concentration of 2.8 g l−1 instead of 20 g l−1) and used to inoculate bioreactor vessels containing 0.5 l of modified M9 medium to a starting OD600 of 0.1. Bioreactors were run as dissolved oxygen (DO)-stat batch-fed reactors. In this set-up, bioreactors were monitored with a DO probe which controlled the glucose feed. Cell cultures were sampled from the bioreactors aseptically and centrifuged at 20,000g for 10 min to separate the supernatant. Supernatants were filtered through a 0.22-μm regenerated cellulose syringe filter (Thermo Fisher Scientific) and stored at −20 °C until analysis. Filtered supernatants were evaluated for glucose, secreted acids and lactam concentrations.
Global proteomics and metabolomics
See Supplementary Methods for details of the experimental procedures for global proteomics and metabolomics.
PKS full module exchange
The protein sequences of 20 selected modules, spanning from five residues before the KS domain to five residues after the ACP domain (for the purpose of FluPks1 full module replacement to produce VL) were aligned using the MAFFT software tool61. The resulting alignment data were then transformed into an all-against-all distance matrix, utilizing BioPython and the blosum62 substitution matrix62,63. Employing the MaxMin algorithm, implemented through the RDKit Python Library, we reordered the modules based on maximum diversity. This ensured that each subset, starting from the top of the list, represented the most diverse set of modules of its respective size. Finally, the DNA sequences of the selected five modules were codon-optimized using the MCU method and synthesized.
To explore the ideal N-terminal junction site for full module replacement, we subjected the protein sequences of 300 randomly selected KS–AT di-domains from the initial portion of PKS modules in ClusterCAD to MSA using the MAFFT software tool. We then systematically analysed this 1,413-residue-long MSA for all highly conserved sites with over 95% sequence identity, to use as possible PKS junction sites, considering all possible candidate sites from the beginning of the KS domain to the beginning of the AT domain based on domain boundary coordinates in ClusterCAD, initially computed via AntiSMASH64. This resulted in a total of 93 potential junction sites. Next, we systematically created the combinatorial set of all possible designs replacing these five donor modules into our working FluPks1 for the EVL production design described above. Although we created designs using a diverse array of N-terminal junction sites spanning the entire KS domain and KS–AT linker region, all designs used the same C-terminal junction between the donor ACP and the acceptor TE domain, at the highly conserved ‘FD’ motif. This combination of 93 sites and five donor modules produced a total of 465 PKS designs, which reduced to 200 designs after eliminating designs with duplicate final amino-acid sequences and random downsampling.
Polymerization of VLs
Details of the experimental procedure for step-growth polymerization using the Trojan horse strategy and RAFT polymerization of the N-acryloyl VL analogues monomers are provided in Supplementary Methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this Article.
Data availability
The DNA sequences of all plasmids and strains created in this study are listed in Supplementary Tables 3 and 4 and have been deposited in the public JBEI registry (https://public-registry.jbei.org/folders/827). The latest P. putida KT2440 Uniprot proteome (UP000000556) was used as the database for proteomic analysis, and the proteomics data have been deposited in the PRIDE repository under accession no. PXD045574 and in MassIVE under accession no. MSV000096981. Global metabolomics data are available from the OSF Data depository (https://osf.io/jc7f2/). All plasmids, strains and related data are available from the corresponding authors upon request. Source data are provided with this paper.
References
Risdian, C., Mozef, T. & Wink, J. Biosynthesis of polyketides in Streptomyces. Microorganisms 7, 124 (2019).
Chooi, Y.-H. & Tang, Y. Navigating the fungal polyketide chemical space: from genes to molecules. J. Org. Chem. 77, 9933–9953 (2012).
Englund, E. et al. Expanding extender substrate selection for unnatural polyketide biosynthesis by acyltransferase domain exchange within a modular polyketide synthase. J. Am. Chem. Soc. 145, 8822–8832 (2023).
Weissman, K. J. Genetic engineering of modular PKSs: from combinatorial biosynthesis to synthetic biology. Nat. Prod. Rep. 33, 203–230 (2016).
Hwang, S., Lee, N., Cho, S., Palsson, B. & Cho, B.-K. Repurposing modular polyketide synthases and non-ribosomal peptide synthetases for novel chemical biosynthesis. Front. Mol. Biosci. 7, 87 (2020).
Hagen, A. et al. Engineering a polyketide synthase for in vitro production of adipic acid. ACS Synth. Biol. 5, 21–27 (2016).
Miyazawa, T., Fitzgerald, B. J. & Keatinge-Clay, A. T. Preparative production of an enantiomeric pair by engineered polyketide synthases. Chem. Commun. 57, 8762–8765 (2021).
Yuzawa, S. et al. Short-chain ketone production by engineered polyketide synthases in Streptomyces albus. Nat. Commun. 9, 4569 (2018).
Xu, Y. et al. Metabolic engineering of Escherichia coli for polyamides monomer δ-valerolactam production from feedstock lysine. Appl. Microbiol. Biotechnol. 104, 9965–9977 (2020).
Von Tiedemann, P., Anwar, S., Kemmer-Jonas, U., Asadi, K. & Frey, H. Synthesis and solution processing of nylon‐5 ferroelectric thin films: the renaissance of odd‐nylons? Macromol. Chem. Phys. 221, 1900468 (2020).
Cortez-Lemus, N. A. & Licea-Claverie, A. Poly(N-vinylcaprolactam), a comprehensive review on a thermoresponsive polymer becoming popular. Prog. Polym. Sci. 53, 1–51 (2016).
Ieong, N. S. et al. The missing lactam-thermoresponsive and biocompatible poly(N-vinylpiperidone) polymers by xanthate-mediated RAFT polymerization. Macromolecules 44, 886–893 (2011).
Teodorescu, M. & Bercea, M. Poly(vinylpyrrolidone)—a versatile polymer for biomedical and beyond medical applications. Polym. Plast. Technol. Eng. 54, 923–943 (2015).
Han, T. & Lee, S. Y. Metabolic engineering of Corynebacterium glutamicum for the high-level production of valerolactam, a nylon-5 monomer. Metab. Eng. 79, 78–85 (2023).
Loeschcke, A. & Thies, S. Pseudomonas putida—a versatile host for the production of natural products. Appl. Microbiol. Biotechnol. 99, 6197–6214 (2015).
Nikel, P. I. & de Lorenzo, V. Pseudomonas putida as a functional chassis for industrial biocatalysis: from native biochemistry to trans-metabolism. Metab. Eng. 50, 142–155 (2018).
Elmore, J. R. et al. High-throughput genetic engineering of nonmodel and undomesticated bacteria via iterative site-specific genome integration. Sci. Adv. 9, eade1285 (2023).
Schmidt, M. et al. Maximizing heterologous expression of engineered type I polyketide synthases: investigating codon optimization strategies. ACS Synth. Biol. 12, 3366–3380 (2023).
Nelson, K. E. et al. Complete genome sequence and comparative analysis of the metabolically versatile Pseudomonas putida KT2440. Environ. Microbiol. 4, 799–808 (2002).
Thompson, M. G. et al. Omics-driven identification and elimination of valerolactam catabolism in Pseudomonas putida KT2440 for increased product titer. Metab. Eng. Commun. 9, e00098 (2019).
Schmidt, M. et al. Nitrogen metabolism in Pseudomonas putida: functional analysis using random barcode transposon sequencing. Appl. Environ. Microbiol. 88, e0243021 (2022).
Jørgensen, H. et al. Biosynthesis of macrolactam BE-14106 involves two distinct PKS systems and amino acid processing enzymes for generation of the aminoacyl starter unit. Chem. Biol. 16, 1109–1121 (2009).
Kudo, F., Miyanaga, A. & Eguchi, T. Biosynthesis of natural products containing β-amino acids. Nat. Prod. Rep. 31, 1056–1073 (2014).
Lin, T.-Y., Borketey, L. S., Prasad, G., Waters, S. A. & Schnarr, N. A. Sequence, cloning and analysis of the fluvirucin B1 polyketide synthase from Actinomadura vulgaris. ACS Synth. Biol. 2, 635–642 (2013).
Martín, J. F., Ramos, A. & Liras, P. Regulation of geldanamycin biosynthesis by cluster-situated transcription factors and the master regulator PhoP. Antibiotics (Basel) 8, 87 (2019).
Wilson, M. C. & Moore, B. S. Beyond ethylmalonyl-CoA: the functional role of crotonyl-CoA carboxylase/reductase homologs in expanding polyketide diversity. Nat. Prod. Rep. 29, 72–86 (2012).
Lethanh, H., Neubauer, P. & Hoffmann, F. The small heat-shock proteins IbpA and IbpB reduce the stress load of recombinant Escherichia coli and delay degradation of inclusion bodies. Microb. Cell Fact. 4, 6 (2005).
Yuzawa, S. et al. Comprehensive in vitro analysis of acyltransferase domain exchanges in modular polyketide synthases and its application for short-chain ketone production. ACS Synth. Biol. 6, 139–147 (2017).
Gross, F. et al. Metabolic engineering of Pseudomonas putida for methylmalonyl-CoA biosynthesis to enable complex heterologous secondary metabolite formation. Chem. Biol. 13, 1253–1264 (2006).
Lechner, A. et al. Designed biosynthesis of 36-methyl-FK506 by polyketide precursor pathway engineering. ACS Synth. Biol. 2, 379–383 (2013).
Ma, C. et al. Unravelling the thioesterases responsible for propionate formation in engineered Pseudomonas putida KT2440. Microb. Biotechnol. 14, 1237–1242 (2021).
Bonnett, S. A. et al. Acyl-CoA subunit selectivity in the pikromycin polyketide synthase PikAIV: steady-state kinetics and active-site occupancy analysis by FTICR-MS. Chem. Biol. 18, 1075–1081 (2011).
Tao, X. B. et al. ClusterCAD 2.0: an updated computational platform for chimeric type I polyketide synthase and nonribosomal peptide synthetase design. Nucleic Acids Res. 51, D532–D538 (2023).
Keatinge-Clay, A. T. Polyketide synthase modules redefined. Angew. Chem. Int. Ed. 56, 4658–4660 (2017).
Valencia, L. E. et al. Engineering Pseudomonas putida KT2440 for chain length tailored free fatty acid and oleochemical production. Commun. Biol. 5, 1363 (2022).
Stockmann, P. N. et al. Biobased chiral semi-crystalline or amorphous high-performance polyamides and their scalable stereoselective synthesis. Nat. Commun. 11, 509 (2020).
Worch, J. C. et al. Elastomeric polyamide biomaterials with stereochemically tuneable mechanical properties and shape memory. Nat. Commun. 11, 3250 (2020).
Shi, Y. & He, X. Effect of tacticity sequence of the poly(N-isopropylacrylamide) oligomer on phase transition behavior in aqueous solution. J. Phys. Chem. B 127, 8660–8668 (2023).
García-Peñas, A., Wang, Y., Muñoz-Bonilla, A., Fernández-García, M. & Stadler, F. J. Lower critical solution temperature sensitivity to structural changes in poly(N‐isopropyl acrylamide) homopolymers. J. Polym. Sci. B Polym. Phys. 57, 1386–1393 (2019).
Annaval, T., Paris, C., Leadlay, P. F., Jacob, C. & Weissman, K. J. Evaluating ketoreductase exchanges as a means of rationally altering polyketide stereochemistry. ChemBioChem 16, 1357–1364 (2015).
Barajas, J. F., Blake-Hedges, J. M., Bailey, C. B., Curran, S. & Keasling, J. D. Engineered polyketides: synergy between protein and host level engineering. Synth. Syst. Biotechnol. 2, 147–166 (2017).
Tian, J.-J. et al. Redesigned nylon 6 variants with enhanced recyclability, ductility and transparency. Angew. Chem. Int. Ed. 63, e202320214 (2024).
Parke, S. M., Lopez, J. C., Cui, S., LaPointe, A. M. & Coates, G. W. Polyethylene incorporating Diels-Alder comonomers: a ‘Trojan Horse’ strategy for chemically recyclable polyolefins. Angew. Chem. Int. Ed. 62, e202301927 (2023).
Wu, X.-T. et al. Enabling closed-loop circularity of ‘non-polymerizable’ α,β-conjugated lactone towards high-performance polyester with the assistance of cyclopentadiene. Angew. Chem. Int. Ed. 63, e202404179 (2024).
Palate, K. Y., Yang, Z., Whitwood, A. C. & Unsworth, W. P. Synthesis of medium-ring lactams and macrocyclic peptide mimetics via conjugate addition/ring expansion cascade reactions. RSC Chem. Biol. 3, 334–340 (2022).
Zhang, J., Bista, R., Miyazawa, T. & Keatinge-Clay, A. T. Boosting titers of engineered triketide and tetraketide synthases to record levels through T7 promoter tuning. Metab. Eng. 78, 93–98 (2023).
Goranovič, D. et al. FK506 biosynthesis is regulated by two positive regulatory elements in Streptomyces tsukubaensis. BMC Microbiol. 12, 238 (2012).
Lü, J. et al. Engineering the erythromycin-producing strain Saccharopolyspora erythraea HOE107 for the heterologous production of polyketide antibiotics. Front. Microbiol. 11, 593217 (2020).
Ham, T. S. et al. Design, implementation and practice of JBEI-ICE: an open source biological part registry platform and tools. Nucleic Acids Res. 40, e141 (2012).
Chen, J., Densmore, D., Ham, T. S., Keasling, J. D. & Hillson, N. J. DeviceEditor visual biological CAD canvas. J. Biol. Eng. 6, 1 (2012).
Nava, A. A. et al. Automated platform for the plasmid construction process. ACS Synth. Biol. 12, 3506–3513 (2023).
Hillson, N. J., Rosengarten, R. D. & Keasling, J. D. j5 DNA assembly design automation software. ACS Synth. Biol. 1, 14–21 (2012).
Thompson, M. G. et al. Massively parallel fitness profiling reveals multiple novel enzymes in Pseudomonas putida lysine metabolism. mBio 10, e02577–18 (2019).
Barajas, J. F. et al. Biochemical characterization of β-amino acid incorporation in fluvirucin B2 biosynthesis. ChemBioChem 19, 1391–1395 (2018).
Chen, Y. et al. Alkaline-SDS cell lysis of microbes with acetone protein precipitation for proteomic sample preparation in 96-well plate format. PLoS ONE 18, e0288102 (2023).
Chen, Y., Gin, J. & Petzold, C. J. Discovery proteomic (DIA) LC-MS/MS data acquisition and analysis. protocols.io https://www.protocols.io/view/discovery-proteomic-dda-lc-ms-ms-data-acquisition-kqdg3516qv25/v2 (2021).
Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41–44 (2020).
Ahrné, E., Molzahn, L., Glatter, T. & Schmidt, A. Critical assessment of proteome-wide label-free absolute abundance estimation strategies. Proteomics 13, 2567–2578 (2013).
Silva, J. C., Gorenstein, M. V., Li, G.-Z., Vissers, J. P. C. & Geromanos, S. J. Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol. Cell. Proteom. 5, 144–156 (2006).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Henikoff, S. & Henikoff, J. G. Amino acid substitution matrices from protein blocks. Proc. Natl Acad. Sci. USA 89, 10915–10919 (1992).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Blin, K. et al. antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res. 49, W29–W35 (2021).
Acknowledgements
This work was part of the US Department of Energy (DOE) Joint BioEnergy Institute (https://www.jbei.org), supported by the US DOE, Office of Science, Office of Biological and Environmental Research, through a contract (DE-AC02-05CH11231) between Lawrence Berkeley National Laboratory and the US DOE. This work is also supported by a DOE Office of Science Distinguished Scientist Award to J.D.K. and by grants from the US DOE, Office of Energy Efficiency and Renewable Energy, Bioenergy Technologies Office (EE0008926), the Philomathia Foundation, and The Nancy P. and Richard K. Robbins Foundation. A portion of this research was performed with a project award (https://doi.org/10.46936/reso.proj.2020.51637/60000235) from the Environmental Molecular Sciences Laboratory, a DOE Office of Science User Facility sponsored by the Biological and Environmental Research programme. Pacific Northwest National Laboratory is a multiprogramme national laboratory managed by the Battelle Memorial Institute, operating for the US DOE (DE-AC05-76RL01830). This work was also authored in part by the Alliance for Sustainable Energy, the manager and operator of the National Renewable Energy Laboratory for the US DOE under contract no. DE-AC36-08GO28308. Funding to A.Y., D.S., C.A.S., C.W.J. and W.E.M. was provided by the US DOE Office of Energy Efficiency and Renewable Energy Bioenergy Technologies Office (BETO) for the Agile BioFoundry via contract no. DE-AC36-08GO28308. This work was also supported by the Basque Government through the BERC 2022–2025 programme and by the Spanish Ministry of Science and Innovation MICINN (AEI):BCAM Severo Ochoa excellence accreditation via contract no. CEX2021-001142-S to H.G.M. The US Government retains and the publisher, by accepting the Article for publication, acknowledges that the US Government retains a nonexclusive, paid-up, irrevocable, worldwide licence to publish or reproduce the published form of this work, or allow others to do so, for US Government purposes. We extend our gratitude to A. Guss and his colleagues for generously providing us with the SAGE vector suite and P. putida AG5577 strain. We also acknowledge G. Lan, P. Mellinger and A. Pearson for assistance during the experiment process.
Author information
Authors and Affiliations
Contributions
N.L., R.W.H., T.W.H.B. and J.D.K. conceptualized the study, and T.W.H.B., R.W.H. and J.D.K. supervised the project. All authors designed the experiments. N.L., M.S., C.L., S.K., A.Y., A.L.F., W.E.K. and C.W.J. performed host engineering. C.J.F. and B.A.A. performed chemical polymerization. T.W.H.B., Y.G., M.C.B., G.B., P.D.L. and H.G.M. performed computational analysis. Y.-M.K., Y.C., J.W.G., C.J.P. and E.E.K.B. performed multi-omics analysis. C.A.S., W.E.M. and D.S. performed fed-batch fermentation. N.L., M.S., C.L., C.J.F., B.A.A., T.W.H.B., R.W.H. and J.D.K. wrote the original draft, and all authors reviewed and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
J.D.K. has financial interests in Amyris, Ansa Biotechnologies, Apertor Pharma, Berkeley Yeast, BioMia, Cyklos Materials, Demetrix, Lygos, Napigen, ResVita Bio and Zero Acre Farms. The other authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks Ramya Kumar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Notes 1–5, Figs. 1–29, Tables 1–5, Data 1 and References.
Supplementary Data 1 (download XLSX )
Exact value of each data point.
Source data
Source Data Fig. 1 (download XLSX )
Exact value of each data point.
Source Data Fig. 2 (download XLSX )
Exact value of each data point.
Source Data Fig. 3 (download XLSX )
Exact value of each data point.
Source Data Fig. 4 (download XLSX )
Exact value of each data point.
Source Data Fig. 5 (download XLSX )
Exact value of each data point.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, N., Schmidt, M., Li, C. et al. Retrobiosynthesis of unnatural lactams via reprogrammed polyketide synthase. Nat Catal 8, 389–402 (2025). https://doi.org/10.1038/s41929-025-01325-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41929-025-01325-6