Abstract
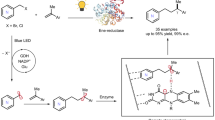
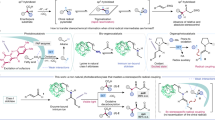
Producing enantioenriched molecules from racemic mixtures is essential for manufacturing. Traditional methods such as resolution, deracemization and enantioconvergent catalysis primarily involve separating or converting enantiomers without altering their structures, or functionalization of stereocentres at or proximal to functional groups. However, there are challenges in enantioselectively forging C–H bonds that are remote from functional groups via hydrogen atom transfer (HAT) with these methods. Here we introduce a strategy for the photoenzymatic stereoablative enantioconvergence of γ-chiral oximes using repurposed flavin-dependent ene-reductases. A photoinduced single-electron reduction of the γ-chiral oxime by an ene-reductase generates an iminyl radical, which then undergoes stereoablative 1,5-HAT at the γ-stereocentre. Subsequent chiral reconstruction through enzymatic HAT and spontaneous imine hydrolysis yields the γ-chiral ketone with high enantioselectivity. This work provides a robust method for remote stereoablative enantioconvergent HAT and broadens the synthetic utility of photobiocatalysis.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are available in the main text or the Supplementary Information. The initial structure for our study was obtained from the crystallographic data of OYE1 (PDB: 3tx9). The atomic coordinates from DFT calculations are provided in the Supplementary Data. Data are available from the corresponding authors upon request. Source data are provided with this paper.
References
Yus, M., Najera, C., Foubelo, F. & Sansano, J. M. Metal-catalyzed enantioconvergent transformations. Chem. Rev. 123, 11817–11893 (2023).
Bhat, V., Welin, E. R., Guo, X. & Stoltz, B. M. Advances in stereoconvergent catalysis from 2005 to 2015: transition-metal-mediated stereoablative reactions, dynamic kinetic resolutions, and dynamic kinetic asymmetric transformations. Chem. Rev. 117, 4528–4561 (2017).
Gao, Y., Hong, G., Yang, B.-M. & Zhao, Y. Enantioconvergent transformations of secondary alcohols through borrowing hydrogen catalysis. Chem. Soc. Rev. 52, 5541–5562 (2023).
Bornscheuer, U. T. et al. Engineering the third wave of biocatalysis. Nature 485, 185–194 (2012).
Arnold, F. H. Directed evolution: bringing new chemistry to life. Angew. Chem. Int. Ed. 57, 4143 (2018).
Kiss, G. et al. Computational enzyme design. Angew. Chem. Int. Ed. 52, 5700–5725 (2013).
Cunha, R. L., Ferreira, E. A., Oliveira, C. S. & Omori, Á. T. Biocatalysis for desymmetrization and resolution of stereocenters beyond the reactive center: how far is far enough? Biotechnol. Adv. 33, 614–623 (2015).
Emmanuel, M. A. et al. Photobiocatalytic strategies for organic synthesis. Chem. Rev. 123, 5459–5520 (2023).
Harrison, W., Huang, X. & Zhao, H. Photobiocatalysis for abiological transformations. Acc. Chem. Res. 55, 1087–1096 (2022).
Miller, D. C., Athavale, S. V. & Arnold, F. H. Combining chemistry and protein engineering for new-to-nature biocatalysis. Nat. Synth. 1, 18–23 (2022).
Peng, Y., Chen, Z., Xu, J. & Wu, Q. Recent advances in photobiocatalysis for selective organic synthesis. Org. Process Res. Dev. 26, 1900–1913 (2022).
Biegasiewicz, K. F. et al. Photoexcitation of flavoenzymes enables a stereoselective radical cyclization. Science 364, 1166–1169 (2019).
Emmanuel, M. A., Greenberg, N. R., Oblinsky, D. G. & Hyster, T. K. Accessing non-natural reactivity by irradiating nicotinamide-dependent enzymes with light. Nature 540, 414–417 (2016).
Ju, S. et al. Stereodivergent photobiocatalytic radical cyclization through the repurposing and directed evolution of fatty acid photodecarboxylases. Nat. Chem. 16, 1339–1347 (2024).
Wang, T.-C. et al. Threonine aldolase-catalyzed enantioselective α-alkylation of amino acids through unconventional photoinduced radical initiation. J. Am. Chem. Soc. 146, 22476–22484 (2024).
Fu, H. & Hyster, T. K. From ground-state to excited-state activation modes: flavin-dependent ‘ene’-reductases catalyzed non-natural radical reactions. Acc. Chem. Res. 57, 1446–1457 (2024).
Cheng, L. et al. Stereoselective amino acid synthesis by synergistic photoredox-pyridoxal radical biocatalysis. Science 381, 444–451 (2023).
Xu, Y. et al. A light-driven enzymatic enantioselective radical acylation. Nature 625, 74–78 (2024).
Huang, X. et al. Photoenzymatic enantioselective intermolecular radical hydroalkylation. Nature 584, 69–74 (2020).
Li, M. et al. Asymmetric photoenzymatic incorporation of fluorinated motifs into olefins. Science 385, 416–421 (2024).
Ye, Y. et al. Using enzymes to tame nitrogen-centred radicals for enantioselective hydroamination. Nat. Chem. 15, 206–212 (2023).
Zhang, Z. et al. Photoenzymatic enantioselective intermolecular radical hydroamination. Nat. Catal. 6, 687–694 (2023).
Harrison, W. et al. Photoenzymatic asymmetric hydroamination for chiral alkyl amine synthesis. J. Am. Chem. Soc. 146, 10716–10722 (2024).
Chen, X. et al. Photoenzymatic hydrosulfonylation for the stereoselective synthesis of chiral sulfones. Angew. Chem. Int. Ed. 62, e202218140 (2023).
Jiang, L. et al. Photoenzymatic redox-neutral radical hydrosulfonylation initiated by fmn. ACS Catal. 14, 6710–6716 (2024).
Shi, Q. et al. Single-electron oxidation-initiated enantioselective hydrosulfonylation of olefins enabled by photoenzymatic catalysis. J. Am. Chem. Soc. 146, 2748–2756 (2024).
Zhao, B. et al. Direct visible-light-excited flavoproteins for redox-neutral asymmetric radical hydroarylation. Nat. Catal. 6, 996–1004 (2023).
Guo, W., Wang, Q. & Zhu, J. Visible light photoredox-catalysed remote C–H functionalisation enabled by 1,5-hydrogen atom transfer (1,5-HAT). Chem. Soc. Rev. 50, 7359–7377 (2021).
Himo, F. Recent trends in quantum chemical modeling of enzymatic reactions. J. Am. Chem. Soc. 139, 6780–6786 (2017).
Sheng, X., Kazemi, M., Planas, F. & Himo, F. Modeling enzymatic enantioselectivity using quantum chemical methodology. ACS Catal. 10, 6430–6449 (2020).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Lefebvre, C. et al. Accurately extracting the signature of intermolecular interactions present in the NCI plot of the reduced density gradient versus electron density. Phys. Chem. Chem. Phys. 19, 17928–17936 (2017).
Trimble, J. S. et al. A designed photoenzyme for enantioselective [2 + 2] cycloadditions. Nature 611, 709–714 (2022).
Sun, N. et al. Enantioselective [2 + 2]-cycloadditions with triplet photoenzymes. Nature 611, 715–720 (2022).
Sandoval, B. A. et al. Photoenzymatic reduction enabled by direct excitation of flavin-dependent ‘ene’-reductases. J. Am. Chem. Soc. 143, 1735–1739 (2021).
Davies, J. et al. Visible-light-mediated generation of nitrogen-centered radicals: metal-free hydroimination and iminohydroxylation cyclization reactions. Angew. Chem. Int. Ed. 54, 14017–14021 (2015).
Li, M. et al. Remote stereocontrol with azaarenes via enzymatic hydrogen atom transfer. Nat. Chem. 16, 277–284 (2024).
Hölzl-Hobmeier, A. et al. Catalytic deracemization of chiral allenes by sensitized excitation with visible light. Nature 564, 240–243 (2018).
Shin, N. Y. et al. Light-driven deracemization enabled by excited-state electron transfer. Science 366, 364–369 (2019).
Huang, M., Zhang, L., Pan, T. & Luo, S. Deracemization through photochemical E/Z isomerization of enamines. Science 375, 869–874 (2022).
Onneken, C. et al. Light-enabled deracemization of cyclopropanes by Al-salen photocatalysis. Nature 621, 753–759 (2023).
Wen, L. et al. Multiplicative enhancement of stereoenrichment by a single catalyst for deracemization of alcohols. Science 382, 458–464 (2023).
Lu, T. & Chen, F. Multiwfn: a multifunctional wavefunction analyzer. J. Comput. Chem. 33, 580–592 (2012).
Acknowledgements
We thank the Core Facilities at the Carl R. Woese Institute for Genomic Biology where most of the experiments in this project were performed. We thank G. Jiang and H. Li for helpful discussions. This work was funded by the DOE Center for Advanced Bioenergy and Bioproducts Innovation (US Department of Energy, Office of Science, Office of Biological and Environmental Research under award number DE-SC0018420 to H.Z.). NMR data were collected at the Carl R. Woese Institute for Genomic Biology Core, on a 600-MHz NMR funded by the National Institute of Health (number S10-RR028833 to H.Z.). We thank the School of Chemical Sciences NMR Laboratory. We acknowledge the Computational Chemistry Commune (http://bbs.keinsci.com/) for their support with the calculations. Our research benefited from the computing resources at Delta, the National Center for Supercomputing Applications, enabled by allocation BIO230215 from the Advanced Cyberinfrastructure Coordination Ecosystem: Services and Support (ACCESS) programme, funded by National Science Foundation grant numbers 2138259, 2138286, 2138307, 2137603 and 2138296. Any opinions, findings, and conclusions or recommendations expressed in this publication are those of the author(s) and do not necessarily reflect the views of the US Department of Energy.
Author information
Authors and Affiliations
Contributions
H.Z. coordinated the project. Z. Zhang and H.Z. conceived the project and designed the experiments. Z. Zhang performed most of the experiments. M.L. performed computational studies. W.H. and Z. Zhao contributed to synthesis. J.L., Y.Y. and W.H. contributed to the site-directed mutagenesis. Z. Zhang, M.L. and H.Z. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic representation of the photoenzymatic enantioconvergent process detailing multiple pathways for the synthesis of chiral ketones with a remote stereocenter.
The diagram illustrates three primary pathways: (1) Direct background hydrolysis of R- and S-2a, leading to non-enzymatic formation of the chiral ketones. (2) Enzyme and light-mediated formation of an iminyl radical from R-2a and S-2a, followed by a HAT to the iminyl radical and subsequent hydrolysis, producing the chiral products. (3) A more complex pathway involving stereoablative 1,5-HAT, chiral HAT, and hydrolysis, further demonstrating the enzymatic control over the stereoselectivity of the product.
Extended Data Fig. 2 Computational and experimental exploration of the generation of the iminyl radical.
a, The photochemical process that results in the formation of an iminyl radical and FMNsq. Calculations were performed at the um062x-D3/def2tzvpp (SMD, ε = 4.0) // um062x-D3/def2svp (gas) level. Quantitative electron transfer dynamics were calculated using the interfragment charge transfer (IFCT) method, facilitated by Multiwfn43 software. Note that the atoms were held constant during geometry optimization, highlighted by green. For the dominant molecular orbital transitions contributing to CT excitation, the colours coding with green and blue represent positive and negative orbital phases, respectively. b, In-depth experimental assessment of hydrogen bond in tyrosine residues. Reactions were performed under the standard conditions.
Extended Data Fig. 3 Optimal structures for transition states and key intermediates.
Calculations were performed at the um062x-D3/def2tzvpp (SMD, ε = 4.0) // um062x-D3/def2svp (gas) level. Note that the atoms were held constant during geometry optimization, highlighted by green.
Extended Data Fig. 4 Comprehensive spin density analysis covering intermediates and HAT transition states.
Utilizing Multiwfn 3.8 software, we conducted an in-depth spin density analysis. Visual representations were constructed with VMD (Version 1.9.3). The green and blue areas depict α and β spin respectively, with all amino acid residues being omitted to ensure clarity in presentation.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–16, Tables 1–21, Methods and References.
Supplementary Data 1 (download PDF )
Atomic coordinates for DFT calculation
Source data
Fig. 2 (download XLSX )
Statistical Source Data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Z., Li, M., Harrison, W. et al. Photoenzymatic stereoablative enantioconvergence of γ-chiral oximes via hydrogen atom transfer. Nat Catal 8, 548–555 (2025). https://doi.org/10.1038/s41929-025-01347-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41929-025-01347-0
This article is cited by
-
Stereoselective cyano translocation reaction enabled by photoenzymatic catalysis
Nature Communications (2026)