Abstract
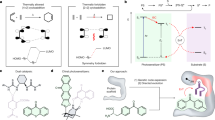
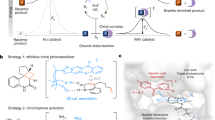
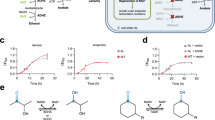
Although artificial enzymes have significantly expanded the scope of enzyme-catalysed transformations, their construction typically relies on the irreversible incorporation of non-natural active sites. Inspired by natural cofactors, here we show a reversible binding strategy for artificial enzyme design using benzophenone adenine dinucleotide (BpAD), a photoactive NAD⁺ analogue that can integrate seamlessly into a broad range of NAD⁺-dependent protein scaffolds. The resulting artificial photoenzymes catalyse both inter- and intramolecular [2 + 2] cycloaddition reactions with excellent enantioselectivity, broad substrate compatibility and notable enantiodivergence. Computational studies confirm the precise binding mode of BpAD and reveal a key exo-attack pathway in the stepwise C–C bond formation mechanism. Notably, BpAD-catalysed reactions are highly orthogonal to those mediated by NAD⁺, allowing simultaneous use without interference. This work introduces a versatile and generalizable approach to artificial enzyme development, leveraging the inherent diversity of NAD⁺-dependent proteins for tailored catalytic applications.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data relating to the materials and methods, experimental procedures, mechanistic studies and computational calculations, HPLC spectra and NMR spectra are available in the Supplementary Information or from the authors on reasonable request. The configurations of MD simulations are available via GitHub at https://github.com/3dwdw58rds/7WBC-2DKN-research.
References
Hanefeld, U., Hollmann, F. & Paul, C. E. Biocatalysis making waves in organic chemistry. Chem. Soc. Rev. 51, 594–627 (2022).
Palmer, T. Enzymes: Biochemistry, Biotechnology, Clinical Chemistry (Woodhead Publishing, 2007).
Kissman, E. N. et al. Expanding chemistry through in vitro and in vivo biocatalysis. Nature 631, 37–48 (2024).
Buller, R. et al. From nature to industry: harnessing enzymes for biocatalysis. Science 382, eadh8615 (2023).
Simić, S. et al. Shortening synthetic routes to small molecule active pharmaceutical ingredients employing biocatalytic methods. Chem. Rev. 122, 1052–1126 (2022).
Vornholt, T. et al. Artificial metalloenzymes. Nat. Rev. Methods Primers 4, 78 (2024).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Drienovská, I. & Roelfes, G. Expanding the enzyme universe with genetically encoded unnatural amino acids. Nat. Catal. 3, 193–202 (2020).
Birch-Price, Z., Hardy, F. J., Lister, T. M., Kohn, A. R. & Green, A. P. Noncanonical amino acids in biocatalysis. Chem. Rev. 124, 8740–8786 (2024).
Lovelock, S. L. et al. The road to fully programmable protein catalysis. Nature 606, 49–58 (2022).
Longwitz, L., Leveson-Gower, R. B., Rozeboom, H. J., Thunnissen, A.-M. W. H. & Roelfes, G. Boron catalysis in a designer enzyme. Nature 629, 824–829 (2024).
Hanreich, S., Bonandi, E. & Drienovská, I. Design of artificial enzymes: insights into protein scaffolds. ChemBioChem 24, e202200566 (2023).
Raines, D. J. et al. Redox-switchable siderophore anchor enables reversible artificial metalloenzyme assembly. Nat. Catal. 1, 680–688 (2018).
Genzink, M. J., Kidd, J. B., Swords, W. B. & Yoon, T. P. Chiral photocatalyst structures in asymmetric photochemical synthesis. Chem. Rev. 122, 1654–1716 (2022).
Chan, A. Y. et al. Metallaphotoredox: the merger of photoredox and transition metal catalysis. Chem. Rev. 122, 1485–1542 (2022).
Romero, N. A. & Nicewicz, D. A. Organic photoredox catalysis. Chem. Rev. 116, 10075–10166 (2016).
Lee, S. H., Choi, D. S., Kuk, S. K. & Park, C. B. Photobiocatalysis: activating redox enzymes by direct or indirect transfer of photoinduced electrons. Angew. Chem. Int. Ed. 57, 7958–7985 (2018).
Candish, L. et al. Photocatalysis in the life science industry. Chem. Rev. 122, 2907–2980 (2022).
Liu, Q. & Wu, L.-Z. Recent advances in visible-light-driven organic reactions. Natl Sci. Rev. 4, 359–380 (2017).
Harrison, W., Huang, X. & Zhao, H. Photobiocatalysis for abiological transformations. Acc. Chem. Res. 55, 1087–1096 (2022).
Fu, H. & Hyster, T. K. From ground-state to excited-state activation modes: flavin-dependent “ene”-reductases catalyzed non-natural radical reactions. Acc. Chem. Res. 57, 1446–1457 (2024).
Emmanuel, M. A. et al. Photobiocatalytic strategies for organic synthesis. Chem. Rev. 123, 5459–5520 (2023).
Emmanuel, M. A., Greenberg, N. R., Oblinsky, D. G. & Hyster, T. K. Accessing non-natural reactivity by irradiating nicotinamide-dependent enzymes with light. Nature 540, 414–417 (2016).
Litman, Z. C., Wang, Y., Zhao, H. & Hartwig, J. F. Cooperative asymmetric reactions combining photocatalysis and enzymatic catalysis. Nature 560, 355–359 (2018).
Biegasiewicz, K. F. et al. Photoexcitation of flavoenzymes enables a stereoselective radical cyclization. Science 364, 1166–1169 (2019).
DeHovitz, J. S. et al. Static to inducibly dynamic stereocontrol: the convergent use of racemic β-substituted ketones. Science 369, 1113–1118 (2020).
Huang, X. et al. Photoenzymatic enantioselective intermolecular radical hydroalkylation. Nature 584, 69–74 (2020).
Fu, H. et al. An asymmetric sp3–sp3 cross-electrophile coupling using ‘ene’-reductases. Nature 610, 302–307 (2022).
Cheng, L. et al. Stereoselective amino acid synthesis by synergistic photoredox-pyridoxal radical biocatalysis. Science 381, 444–451 (2023).
Tseliou, V. et al. Stereospecific radical coupling with a non-natural photodecarboxylase. Nature 634, 848–854 (2024).
Wang, T.-C. et al. Stereoselective amino acid synthesis by photobiocatalytic oxidative coupling. Nature 629, 98–104 (2024).
Xu, Y. et al. A light-driven enzymatic enantioselective radical acylation. Nature 625, 74–78 (2024).
Gu, Y. F., Ellis-Guardiola, K., Srivastava, P. & Lewis, J. C. Preparation, characterization, and oxygenase activity of a photocatalytic artificial enzyme. ChemBioChem 16, 1880–1883 (2015).
Liu, X. H. et al. A genetically encoded photosensitizer protein facilitates the rational design of a miniature photocatalytic CO-reducing enzyme. Nat. Chem. 10, 1201–1206 (2018).
Schwochert, T. D. et al. Design and evaluation of artificial hybrid photoredox biocatalysts. ChemBioChem 21, 3146–3150 (2020).
Fu, Y. et al. Biocatalytic cross-coupling of aryl halides with a genetically engineered photosensitizer artificial dehalogenase. J. Am. Chem. Soc. 143, 617–622 (2021).
Kang, F. Y. et al. Rational design of a miniature photocatalytic CO-reducing enzyme. ACS Catal. 11, 5628–5635 (2021).
Sun, N. et al. Enantioselective [2+2]-cycloadditions with triplet photoenzymes. Nature 611, 715–720 (2022).
Trimble, J. S. et al. A designed photoenzyme for enantioselective [2+2] cycloadditions. Nature 611, 709–714 (2022).
Zubi, Y. S., Liu, B., Gu, Y., Sahoo, D. & Lewis, J. C. Controlling the optical and catalytic properties of artificial metalloenzyme photocatalysts using chemogenetic engineering. Chem. Sci. 13, 1459–1468 (2022).
Fu, Y. et al. Whole-cell-catalyzed hydrogenation/deuteration of aryl halides with a genetically repurposed photodehalogenase. Chem 9, 1897–1909 (2023).
Guo, J. et al. Chemogenetic evolution of diversified photoenzymes for enantioselective [2 + 2] cycloadditions in whole cells. J. Am. Chem. Soc. 146, 19030–19041 (2024).
Chen, K.-Y., Ming, H., Wang, H.-X., Wang, H.-Q. & Xiang, Z. Genetic incorporation of a thioxanthone-containing amino acid for the design of artificial photoenzymes. Angew. Chem. Int. Ed. 64, e202419022 (2025).
Crawshaw, R. et al. Efficient and selective energy transfer photoenzymes powered by visible light. Nat. Chem. https://doi.org/10.1038/s41557-025-01820-0 (2025).
Sun, C. et al. Photosensitizer repositioning affords an enantiocomplementary enzyme for [2 + 2]-cycloadditions. Angew. Chem. Int. Ed. Engl. https://doi.org/10.1002/anie.202503576 (2025).
Yang, X. et al. Bridging chemistry and biology for light-driven new-to-nature enantioselective photoenzymatic catalysis. Chem. Soc. Rev. https://doi.org/10.1039/d4cs00561a (2025).
Poplata, S., Tröster, A., Zou, Y.-Q. & Bach, T. Recent advances in the synthesis of cyclobutanes by olefin [2+2] photocycloaddition reactions. Chem. Rev. 116, 9748–9815 (2016).
Strieth-Kalthoff, F., James, M. J., Teders, M., Pitzer, L. & Glorius, F. Energy transfer catalysis mediated by visible light: principles, applications, directions. Chem. Soc. Rev. 47, 7190–7202 (2018).
Strieth-Kalthoff, F. & Glorius, F. Triplet energy transfer photocatalysis: unlocking the next level. Chem 6, 1888–1903 (2020).
Blum, T. R., Miller, Z. D., Bates, D. M., Guzei, I. A. & Yoon, T. P. Enantioselective photochemistry through Lewis acid-catalyzed triplet energy transfer. Science 354, 1391–1395 (2016).
Li, X., Großkopf, J., Jandl, C. & Bach, T. Enantioselective, visible light mediated aza Paternò–Büchi reactions of quinoxalinones. Angew. Chem. Int. Ed. 60, 2684–2688 (2021).
Yang, P., Wang, R.-X., Huang, X.-L., Cheng, Y.-Z. & You, S.-L. Enantioselective synthesis of cyclobutane derivatives via cascade asymmetric allylic etherification/[2 + 2] photocycloaddition. J. Am. Chem. Soc. 145, 21752–21759 (2023).
Wang, J. et al. Enantioselective [2 + 2] photocycloreversion enables de novo deracemization synthesis of cyclobutanes. J. Am. Chem. Soc. 146, 22840–22849 (2024).
Senn, H. M. & Thiel, W. QM/MM methods for biomolecular systems. Angew. Chem. Int. Ed. 48, 1198–1229 (2009).
Lu, Y. et al. Multiscale QM/MM modelling of catalytic systems with ChemShell. Phys. Chem. Chem. Phys. 25, 21816–21835 (2023).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Grosskopf, J., Kratz, T., Rigotti, T. & Bach, T. Enantioselective photochemical reactions enabled by triplet energy transfer. Chem. Rev. 122, 1626–1653 (2022).
Cabanero, D. C. & Rovis, T. Low-energy photoredox catalysis. Nat. Rev. Chem. 9, 28–45 (2025).
Zhu, M., Zheng, C., Zhang, X. & You, S. L. Synthesis of cyclobutane-fused angular tetracyclic spiroindolines via visible-light-promoted intramolecular dearomatization of indole derivatives. J. Am. Chem. Soc. 141, 2636–2644 (2019).
Zhou, Q. Q., Zou, Y. Q., Lu, L. Q. & Xiao, W. J. Visible-light-induced organic photochemical reactions through energy-transfer pathways. Angew. Chem. Int. Ed. 58, 1586–1604 (2019).
Zhang, Z. et al. Photocatalytic intramolecular [2 + 2] cycloaddition of indole derivatives via energy transfer: a method for late-stage skeletal transformation. ACS Catal. 10, 10149–10156 (2020).
Zhu, M., Zhang, X., Zheng, C. & You, S.-L. Visible-light-induced dearomatization via [2+2] cycloaddition or 1,5-hydrogen atom transfer: divergent reaction pathways of transient diradicals. ACS Catal. 10, 12618–12626 (2020).
Rolka, A. B. & König, B. Bifunctional organic photocatalysts for enantioselective visible-light-mediated photocatalysis. Nat. Synth. 2, 913–925 (2023).
Poplata, S., Troster, A., Zou, Y. Q. & Bach, T. Recent advances in the synthesis of cyclobutanes by olefin [2+2] photocycloaddition reactions. Chem. Rev. 116, 9748–9815 (2016).
Dexter, D. L. A theory of sensitized luminescence in solids. J. Chem. Phys. 21, 836–850 (1953).
Zhou, T.-P., Zhong, F., Wu, Y. & Liao, R.-Z. Regioselectivity and stereoselectivity of intramolecular [2 + 2] photocycloaddition catalyzed by chiral thioxanthone: a quantum chemical study. Org. Biomol. Chem. 19, 1532–1540 (2021).
Fernandez, I. & Bickelhaupt, F. M. The activation strain model and molecular orbital theory: understanding and designing chemical reactions. Chem. Soc. Rev. 43, 4953–4967 (2014).
Wang, S. et al. Copper-catalyzed regiodivergent asymmetric difunctionalization of terminal alkynes. Angew. Chem. Int. Ed. 63, e202410833 (2024).
Huang, J. et al. Accessing ladder-shape azetidine-fused indoline pentacycles through intermolecular regiodivergent aza-Paternò–Büchi reactions. Nat. Commun. 15, 1431 (2024).
Wang, Y.-C., Feng, S., Liang, W. & Zhao, Y. Electronic couplings for photoinduced charge transfer and excitation energy transfer based on fragment particle–hole densities. J. Phys. Chem. Lett. 12, 1032–1039 (2021).
You, Z.-Q. & Hsu, C.-P. Theory and calculation for the electronic coupling in excitation energy transfer. Int. J. Quantum Chem. 114, 102–115 (2014).
Jung, H. W. et al. Distance dependence of electronic coupling in rigid, cofacially compressed, π-stacked organic mixed-valence systems. J. Phys. Chem. B 124, 1033–1048 (2020).
Lai, R. et al. Shallow distance-dependent triplet energy migration mediated by endothermic charge-transfer. Nat. Commun. 12, 1532 (2021).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Maier, J. A. et al. ff14SB: iImproving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Acknowledgements
We thank the National Key Research and Development Program of China (2022YFA1505600 to H.-J.P.), the National Natural Science Foundation of China (22277052 to H.-J.P. and 22121001 to B.W.), the Natural Science Foundation of Jiangsu Province (BK20232016 and BK20220762 to H.-J.P.) and the High-Level Innovation and Entrepreneurial Research Team Program in Jiangsu (JSSCTD202309 to H.-J.P.) for financial support. We also thank X. Lou from Westlake University for her valuable assistance with the transient absorption spectroscopy measurements.
Author information
Authors and Affiliations
Contributions
H.-J.P. conceived and and coordinated the project. P.D. conducted most of the protein scaffold screening, protein evolution and reaction optimization experiments. J.L. designed and synthesized the BpAD cofactor, substrates and standard products. H.L. and J.W. conducted part of the protein evolution work. T.-P.Z. performed the computational studies under the supervision of B.W. W.H. measured the Kd values using ITC. H.-J.P., P.D., J.L., T.-P.Z. and B.W. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks Gui-Juan Cheng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–34, Tables 1–9 and Methods.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Du, P., Li, J., Zhou, TP. et al. An NAD⁺ analogue enables assembly of structurally diverse artificial photoenzymes for enantiodivergent [2 + 2] cycloadditions. Nat Catal 8, 822–832 (2025). https://doi.org/10.1038/s41929-025-01390-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41929-025-01390-x
This article is cited by
-
Unlocking enzymatic deracemization of structurally diverse allenes by designed photoenzymes
Nature Communications (2026)