Abstract
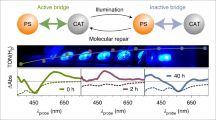
Photobiocatalysis provides a powerful strategy for integrating light and biological catalysts to drive abiological transformations. However, its scalability is hindered by high enzyme loading, reliance on costly cofactors and instability under radical-generating conditions. Here we report the integration of light-driven enzymatic reactions into the cellular metabolism of Escherichia coli, bridging flavin-based photobiocatalysis with biosynthesis. Using synthetic biology strategies, we engineered microbial cells to continuously produce olefin substrates and ene-reductase while regenerating cofactors directly from glucose. By externally supplying radical precursors or introducing synthetic pathways for their in situ production, we enabled fermentation-based microbial photobiosynthesis, achieving high titres and demonstrating feasibility for scale-up in a bioreactor. This approach extends photobiocatalysis from in vitro applications to in vivo semi- and complete biosynthesis, revealing its full potential for integrating light-driven reactions into cellular metabolism.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The DNA sequences of plasmids used in this study are provided in the Supplementary Information. The SnapGene files for the key plasmids can be accessed at https://doi.org/10.5061/dryad.ns1rn8q54 (ref. 50). Source data are provided with this paper. The remaining data are available in the main text or Supplementary Information, or from the corresponding author upon reasonable request.
References
Yang, Y. & Arnold, F. H. Navigating the unnatural reaction space: directed evolution of heme proteins for selective carbene and nitrene transfer. Acc. Chem. Res. 54, 1209–1225 (2021).
Hanefeld, U., Hollmann, F. & Paul, C. E. Biocatalysis making waves in organic chemistry. Chem. Soc. Rev. 51, 594–627 (2022).
Emmanuel, M. A. et al. Photobiocatalytic strategies for organic synthesis. Chem. Rev. 123, 5459–5520 (2023).
Buller, R. et al. From nature to industry: harnessing enzymes for biocatalysis. Science 382, eadh8615 (2023).
Morita, I. & Ward, T. R. Recent advances in the design and optimization of artificial metalloenzymes. Curr. Opin. Chem. Biol. 81, 102508 (2024).
Kissman, E. N. et al. Expanding chemistry through in vitro and in vivo biocatalysis. Nature 631, 37–48 (2024).
Brouwer, B. et al. Noncanonical amino acids: bringing new-to-nature functionalities to biocatalysis. Chem. Rev. 124, 10877–10923 (2024).
Birch-Price, Z., Hardy, F. J., Lister, T. M., Kohn, A. R. & Green, A. P. Noncanonical amino acids in biocatalysis. Chem. Rev. 124, 8740–8786 (2024).
Wu, S. et al. Biocatalysis: enzymatic synthesis for industrial applications. Angew. Chem. Int. Ed. 60, 88–119 (2021).
Peng, Y. et al. Recent advances in photobiocatalysis for selective organic synthesis. Org. Process Res. Dev. 26, 1900–1913 (2022).
Yu, J. et al. Single-electron oxidation triggered by visible-light-excited enzymes for asymmetric biocatalysis. Angew. Chem. Int. Ed. 64, e202419262 (2025).
Harrison, W. et al. Photobiocatalysis for abiological transformations. Acc. Chem. Res. 55, 1087–1096 (2022).
Fu, H. & Hyster, T. K. From ground-state to excited-state activation modes: flavin-dependent “ene”-reductases catalyzed non-natural radical reactions. Acc. Chem. Res. 57, 1446–1457 (2024).
Jain, S., Ospina, F. & Hammer, S. C. A new age of biocatalysis enabled by generic activation modes. JACS Au 4, 2068–2080 (2024).
Cheng, L. et al. Stereoselective amino acid synthesis by synergistic photoredox-pyridoxal radical biocatalysis. Science 381, 444–451 (2023).
Wang, T.-C. et al. Stereoselective amino acid synthesis by photobiocatalytic oxidative coupling. Nature 629, 98–104 (2024).
Xu, Y. et al. A light-driven enzymatic enantioselective radical acylation. Nature 625, 74–78 (2024).
Xing, Z. et al. Synergistic photobiocatalysis for enantioselective triple-radical sorting. Nature 637, 1118–1123 (2025).
Sun, N. et al. Enantioselective [2+2]-cycloadditions with triplet photoenzymes. Nature 611, 715–720 (2022).
Trimble, J. S. et al. A designed photoenzyme for enantioselective [2+2] cycloadditions. Nature 611, 709–714 (2022).
Guo, J. et al. Chemogenetic evolution of diversified photoenzymes for enantioselective [2 + 2] cycloadditions in whole cells. J. Am. Chem. Soc. 146, 19030–19041 (2024).
Tseliou, V. et al. Stereospecific radical coupling with a non-natural photodecarboxylase. Nature 634, 848–854 (2024).
Fu, Y. et al. Whole-cell-catalyzed hydrogenation/deuteration of aryl halides with a genetically repurposed photodehalogenase. Chem 9, 1897–1909 (2023).
Hirschi, S. et al. Synthetic biology: bottom-up assembly of molecular systems. Chem. Rev. 122, 16294–16328 (2022).
Ro, D.-K. et al. Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440, 940–943 (2006).
Luo, X. et al. Complete biosynthesis of cannabinoids and their unnatural analogues in yeast. Nature 567, 123–126 (2019).
Zhang, J. et al. A microbial supply chain for production of the anti-cancer drug vinblastine. Nature 609, 341–347 (2022).
Gao, J. et al. Biosynthesis of catharanthine in engineered Pichia pastoris. Nat. Synth. 2, 231–242 (2023).
Liu, Y. et al. Complete biosynthesis of QS-21 in engineered yeast. Nature 629, 937–944 (2024).
Huang, J. et al. Unnatural biosynthesis by an engineered microorganism with heterologously expressed natural enzymes and an artificial metalloenzyme. Nat. Chem. 13, 1186–1191 (2021).
Huang, J. et al. Complete integration of carbene-transfer chemistry into biosynthesis. Nature 617, 403–408 (2023).
Li, M. et al. Asymmetric photoenzymatic incorporation of fluorinated motifs into olefins. Science 385, 416–421 (2024).
Duan, X. et al. Ground-state flavin-dependent enzymes catalyzed enantioselective radical trifluoromethylation. Nat. Commun. 16, 1225 (2025).
Qi, W. W. et al. Functional expression of prokaryotic and eukaryotic genes in Escherichia coli for conversion of glucose to p-hydroxystyrene. Metab. Eng. 9, 268–276 (2007).
Lee, W.-H., Kim, M.-D., Jin, Y.-S. & Seo, J.-H. Engineering of NADPH regenerators in Escherichia coli for enhanced biotransformation. Appl. Microbiol. Biotechnol. 97, 2761–2772 (2013).
Ajikumar, P. K. et al. Isoprenoid pathway optimization for taxol precursor overproduction in Escherichia coli. Science 330, 70–74 (2010).
McKenna, R. & Nielsen, D. R. Styrene biosynthesis from glucose by engineered Escherichia coli. Metab. Eng. 13, 544–554 (2011).
Huang, X. et al. Photoenzymatic enantioselective intermolecular radical hydroalkylation. Nature 584, 69–74 (2020).
Li, M. et al. Remote stereocontrol with azaarenes via enzymatic hydrogen atom transfer. Nat. Chem. 16, 277–284 (2024).
Liu, Y. et al. Asymmetric synthesis of α-chloroamides via photoenzymatic hydroalkylation of olefins. J. Am. Chem. Soc. 146, 7191–7197 (2024).
Chen, X. et al. Photoenzymatic hydrosulfonylation for the stereoselective synthesis of chiral sulfones. Angew. Chem. Int. Ed. 62, e202218140 (2023).
Harrison, W. et al. Photoenzymatic asymmetric hydroamination for chiral alkyl amine synthesis. J. Am. Chem. Soc. 146, 10716–10722 (2024).
Kumari, A. & Singh, R. K. Medicinal chemistry of indole derivatives: current to future therapeutic prospectives. Bioorg. Chem. 89, 103021 (2019).
Zhao, B. et al. Direct visible-light-excited flavoproteins for redox-neutral asymmetric radical hydroarylation. Nat. Catal. 6, 996–1004 (2023).
Lee, J.-H. & Lee, J. Indole as an intercellular signal in microbial communities. FEMS Microbiol. Rev. 34, 426–444 (2010).
Das, A. et al. Directed evolution of P411 enzymes for amination of inert C–H bonds. Methods Enzymol. 693, 1–30 (2023).
Shimizu, K. & Matsuoka, Y. Feedback regulation and coordination of the main metabolism for bacterial growth and metabolic engineering for amino acid fermentation. Biotechnol. Adv. 55, 107887 (2022).
Yuan, Y. et al. Self-resistance-gene-guided, high-throughput automated genome mining of bioactive natural products from Streptomyces. Cell Syst. 16, 101237 (2025).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Yuan, Y., Li, M., Harrison, W., Zhang, Z. & Zhao, H. Harnessing photoenzymatic reactions for unnatural biosynthesis in microorganisms. Dryad https://doi.org/10.5061/dryad.ns1rn8q54 (2025).
Acknowledgements
We thank L. Burrus and T. Martin for assistance with the fed-batch fermentation experiments. This work was funded by the DOE Center for Advanced Bioenergy and Bioproducts Innovation, under the auspices of the US Department of Energy, Office of Science, Office of Biological and Environmental Research (award DE-SC0018420 to H.Z.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
H.Z. coordinated the project. H.Z., Y.Y. and M.L. jointly conceptualized the project and designed the research experiments. Y.Y., M.L. and H.Z. wrote the paper with input from all authors. Y.Y. executed the strain construction, biosynthesis and fed-batch fermentation experiments. Y.Y. and M.L. constructed the plasmids. M.L. synthesized the product standards. W.H. contributed to construction of the mutant library. W.H. and Z.Z. were responsible for synthesizing radical precursors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–33, Tables 1–7, methods and references.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yuan, Y., Li, M., Harrison, W. et al. Harnessing photoenzymatic reactions for unnatural biosynthesis in microorganisms. Nat Catal 9, 62–72 (2026). https://doi.org/10.1038/s41929-025-01470-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41929-025-01470-y
This article is cited by
-
Light-powered bacteria become living chemical factories
Nature (2026)
-
Light-driven metabolic makeover
Nature Catalysis (2026)