Abstract
CRISPR-based genome editing of pseudogene-associated disorders, such as p47phox-deficient chronic granulomatous disease (p47 CGD), is challenged by chromosomal rearrangements due to presence of multiple targets. We report that interactions between highly homologous sequences that are localized on the same chromosome contribute substantially to post-editing chromosomal rearrangements. We successfully employed editing approaches at the NCF1 gene and its pseudogenes, NCF1B and NCF1C, in a human cell line model of p47 CGD and in patient-derived human hematopoietic stem and progenitor cells. Upon genetic engineering, a droplet digital PCR-based method identified cells with altered copy numbers, spanning megabases from the edited loci. We attributed the high aberration frequency to the interaction between repetitive sequences and their predisposition to recombination events. Our findings emphasize the need for careful evaluation of the target-specific genomic context, such as the presence of homologous regions, whose instability can constitute a risk factor for chromosomal rearrangements upon genome editing.
Similar content being viewed by others
Introduction
Mutations in the neutrophil cytosolic factor 1 (NCF1) gene lead to the phagocyte immunodeficiency p47phox-deficient chronic granulomatous disease (p47 CGD), which may be corrected by viral gene transfer or by genome editing1,2,3,4,5,6. p47 CGD represents a pseudogene-related disorder, in which the mutated gene is flanked by two non-processed pseudogenes, NCF1B and NCF1C, both of which have a high degree of sequence similarity to NCF1.
Most of p47 CGD cases are caused by a frameshift-inducing two-nucleotide GT deletion (ΔGT) at the start of NCF1 exon 2, resulting in a premature termination codon7. This ΔGT sequence is also shared by the NCF1B and NCF1C pseudogenes on the same chromosome. Correction of the ΔGT mutation in NCF1, or either of the pseudogenes, leads to the production of functional p47phox4,8. Although this high number of potentially correctable sites increases gene therapy efficacy, targeting sequences shared by NCF1 and the associated pseudogenes lead to simultaneous cleavage at multiple sites, and may ultimately result in unwanted chromosomal deletions4,9,10.
Here we evaluated diverse corrective CRISPR-based editing approaches that target the ΔGT mutation in NCF1, with the goal of evaluating post-editing genomic outcomes. These approaches include Cas9 ribonucleoprotein (RNP) delivery with single-stranded oligodeoxynucleotide (ssODN) repair templates11,12, dead Cas9 (dCas9) shielding of NCF113, multiplex single-strand nicks using Cas9 nickase (nCas9)14,15, or staggered double-strand breaks (DSBs) using Cas12a16. We aimed at inhibiting the formation of deletions between the targeted NCF1 loci, while maintaining relevant levels of correction. Studies in a cell line model of p47 CGD, PLB-985 NCF1 ΔGT17, and in human hematopoietic stem and progenitor cells (HSPCs) revealed that regardless of the type of genetic manipulation at the NCF1 locus, editing of this region was inevitably associated with induction of chromosomal aberrations.
Results
RNP-mediated correction of NCF1 ΔGT leads to functional correction of the NADPH oxidase but also to deletions between on-target loci
Chromosomal aberrations between the on-target NCF1 loci were previously reported for PLB-985 NCF1 ∆GT cells edited by plasmid-mediated Cas9 delivery4. We, therefore, hypothesized that transient exposure of the genome to CRISPR-Cas9 by RNP delivery11,12 could restrict the frequency of the chromosomal aberrations. To investigate the correction efficacy of the NCF1 GT mutation, we used a myeloid cell line PLB-985 containing either the wild type (WT) GTGT sequence in the NCF1 and ∆GT in both pseudogenes (denoted as PLB-985 WT) or the ∆GT mutation on all three loci (denoted as PLB-985 NCF1 ∆GT). Screening of single-guide RNAs (sgRNAs) identified one selective sgRNA (sgRNA1) able to discriminate between the WT and ΔGT NCF1 WT sequence (Supplementary Fig. S1; Supplementary Table S4).
To achieve high efficiency of correction, we assessed several corrective ssODN repair templates (Supplementary Table S5) and titrated both editing components (RNPs, and corrective ssODN), in the PLB-985 NCF1 ΔGT cell line (Supplementary Fig. S2). Using RNP delivery, we achieved restoration of WT GTGT sequence in 51.9% ± 13.1% of NCF1/NCF1B/NCF1C gene sequences in bulk-treated PLB-985 NCF1 ΔGT cells in the presence of the optimal corrective template (Fig. 1A, B left). Distribution of the number of corrected NCF1 loci per single clone showed considerable variability, as determined by PCR-restriction fragment length polymorphism (RFLP)18 (Fig. 1C).
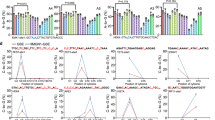
A Schematic representation of CRISPR-Cas9-sgRNA1 cleavage activity in PLB-985 NCF1 ΔGT and PLB-985 WT cell lines. B TIDER analysis of editing frequency and indel formation in PLB-985 NCF1 ΔGT and PLB-985 WT cell lines upon Cas9-sgRNA1 RNP treatment with or without corrective ssODN template (denoted with ‘T’). Reference correction in PLB-985 WT cells refers to the levels of NCF1 GTGT sequence as calculated by TIDER in the non-treated (NT) samples. Statistical analysis with two-tailed unpaired t test (PLB-985 ΔGT samples) or one-way ANOVA followed by Tukey’s multiple comparison post hoc test (PLB-985 WT samples); n = 3, biological replicates, data are expressed as mean ± SD. C PCR-RFLP analysis of single clones generated by FACS from bulk RNP-treated PLB-985 NCF1 ΔGT cell line accompanied by the corrective ssODN. D Flow cytometry analysis of p47phox protein expression in RNP-treated PLB-985 NCF1 ΔGT with corrective ssODN. Representative flow cytometry plots (after gating for live cells and CD11b positive cells). E Percentage of p47phox positive cells within the differentiated population (CD11b positive); n = 3, biological replicates, data are expressed as mean ± SD. p47phox median fluorescent intensity (MFI) ratio of positive population to the MFI of untreated PLB-985 NCF1 ΔGT cells (NT); n = 3, data are presented as median with range. F Assessment of NADPH oxidase activity by DHR assay upon myeloid differentiation. The assay measures the hydrogen peroxide–dependent conversion of DHR 123 to fluorescent rhodamine 123. Percentages (n = 3, biological replicates, data are expressed as mean ± SD) of DHR positive cells within CD11b positive population upon stimulation with PMA, normalized to PLB-985 NCF1 ΔGT untreated (n = 3, data are presented as median with range). Gating strategies are presented in Supplementary Fig. S3. Statistical analysis with one-way ANOVA followed by Tukey’s multiple comparison test.
The p47phox expression was reconstituted in 81.7% ± 5.1% in the bulk PLB-985 NCF1 ∆GT cells treated with Cas9-sgRNA1 alongside ssODN template (sgRNA1+T), with median fluorescence intensity (MFI) comparable to the PLB-985 WT cell line (Fig. 1D, E), and resulted in substantial functional restoration of reactive oxygen species (ROS) production by nicotinamide adenine dinucleotide phosphate (NADPH) oxidase activity in 76.6% ± 20.3% of edited cells (Fig. 1F; Supplementary Fig. S3).
Notably, PLB-985 NCF1 ΔGT cells edited without the ssODN repair templates showed modest restoration of p47phox expression (21.9% ± 5.3%, Fig. 1E) and NADPH oxidase activity (6.4% ± 2.5%, Fig. 1F), despite the lack of detectable sequence correction (Fig. 1B). This was likely due to indel-induced frameshift restoration events, resulting in production of low amounts of functional p47phox protein4.
Selective targeting of NCF1 ΔGT with sgRNA1 in PLB-985 WT cells resulted in cleavage of the pseudogenes and in an increase of the corrected NCF1 sequences (75.9% ± 6.7%), compared to the untreated WT cells (31.2% ± 3.1%), either by homology-directed repair (HDR) with the ssODN, or via intrachromosomal HDR with the healthy copy of NCF1 (Fig. 1B right).
qPCR-based copy number variation (CNV) assessment of genes located between the NCF1 loci (Fig. 2A) identified a high number of PLB-985 clones (19/76) carrying heterozygous deletions. Additionally, one clone exhibited a homozygous intra-chromosomal deletion of the region between NCF1B and NCF1 (Fig. 2B, Supplementary Table S6), an observation that was not previously reported4.
A Schematic representation of the relative positions of NCF1 and its pseudogenes, as well as the regions potentially affected by chromosomal deletions (dotted lines) induced by CRISPR-Cas treatment. The regions upstream of NCF1B (CALN1), between NCF1B and NCF1 (EIF4H), between NCF1 and NCF1C (WBSCR16) and downstream of NCF1C (HIP1) and qPCR target genes (arrows) are indicated. B Relative copy number detected by qPCR in monoclonal lines. Dot colors represent the detected deletions in single clones: clones without deletions (grey), clones with NCF1B:NCF1 (light blue), NCF1:NCF1C (purple) and NCF1B:NCF1C (blue) deletions. The donut chart indicates the number of clones exhibiting deletions between the NCF1 loci.
In sum, CRISPR-Cas9 by RNP delivery improved correction efficiency, but did not prevent nor reduce the incidence of chromosomal aberrations as reported by using plasmid delivery4.
Catalytically inactive Cas9 shields NCF1 loci and hinders simultaneous cleavage of multiple NCF1 ΔGT loci
As an alternative approach to mitigate chromosomal deletions, we tested whether cleavage activity of Cas9 at the NCF1 loci could be modulated by shielding the cleavage sites by catalytically inactive dCas9. The dCas9 RNP complexes were expected to bind the on-target loci with similar affinity as the Cas9 RNP complexes, but without inducing DNA cleavage, thus reducing the probability of multiple simultaneous cleavage events at the NCF1 loci in the same cell. The co-delivered RNP complexes should compete for the same target loci, and therefore result in a dose-dependent reduction in cleavage activity with increasing dose of dCas9, which we confirmed in a cell-free in vitro cleavage assay (Supplementary Fig. S4).
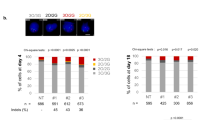
To this end, we co-delivered catalytically inactive dCas9 RNPs along with wild-type Cas9, both complexed with sgRNA1 (Fig. 3A). Treatment of PLB-985 NCF1 ΔGT cells with different dCas9:Cas9 RNPs ratios resulted in a dose-dependent response, with decreasing frequency of editing at increasing dCas9:Cas9 ratios (Fig. 3B). Editing frequency was already reduced at the lowest dCas9:Cas9 tested ratio of 0.5:1. Single clone analysis demonstrated both, a reduced number of corrected clones, and a lower ratio of corrected NCF1 loci per corrected clone at increasing doses of dCas9 (Fig. 3C).
A Representation of the hypothesized mechanism to prevent simultaneous cleavage of multiple on-target NCF1 loci. Catalytically dead Cas9 (dCas9) (grey) complexed with sgRNA1 (blue) is employed as a shield to prevent simultaneous cleavage of multiple on-target NCF1 loci, by competing with active Cas9 (yellow) for on-targets binding. B TIDER analysis of editing efficiency in bulk PLB-985 NCF1 ΔGT cell line upon co-delivery of a constant amount of Cas9-sgRNA1 and increasing ratios of dCas9-sgRNA1 RNP complexes and corrective ssODN template (n = 2 biological replicates, data are expressed as mean ± SD). C PCR-RFLP analysis of 23 monoclonal lines per condition. D Percentage of p47phox positive cells within the differentiated population (CD11b positive gated on live cells) (n = 2 biological replicates, data are expressed as mean ± SD). E Gene copy number analysis in individual clones detected by ddPCR. Each slice of the donut chart represents one clone. 23 clones were analyzed per condition.
Restoration of p47phox expression in bulk culture (Fig. 3D) did not follow the linear reduction of editing efficiency upon increasing the dCas9:Cas9 ratio. Even a low mean correction efficiency in NCF1 sequences, as observed for dCas9:Cas9 ratio 3:1 (4.9% ± 1.3%, Fig. 3B), led to reconstitution of p47phox production in over 20% of edited cells (Fig. 3D). This observation supports the notion that the presence of correctable NCF1 pseudogenes increases the probability of functional correction, despite relatively low editing efficiency.
Next, copy number variation of genes surrounding the NCF1 loci was analyzed by ddPCR in individual clones, obtained from editing experiments in which bulk culture correction levels exceeded 10% (Fig. 3B). A reduction in the number of clones carrying chromosomal aberrations was observed with increasing doses of dCas9 (from 7/23 clones in 1:1 dCas9:Cas9 ratio to 11/23 clones in 0.5:1, Fig. 3E; Supplementary Table S7), which paralleled the reduction of mean editing efficiency.
Interestingly, two of the analyzed clones presented heterozygous deletions, which extended to the NCF1 surrounding regions, as revealed by reduced copy number of HIP1 and CALN1 genes (Fig. 3E). Therefore, although shielding of the NCF1 loci by dCas9 efficiently inhibited on-target editing, it did not prevent induction of deletions between the NCF1 loci, suggesting that chromosomal aberrations are induced already at low levels of editing by Cas9.
Chromosomal aberrations cannot be avoided when editing NCF1 with Cas12a or Cas9 nickases
Structurally distinct DNA strand breaks are processed by structure-specific endonucleases (SSEs)19 and engage different DNA repair pathways20,21,22,23,24. While the Cas9 nuclease generates blunt-ended DSBs25, Cas12a activity yields staggered DSBs with 5’-terminal overhangs16 (Fig. 4A). Paired DNA nicks in close proximity may accomplish precise genome editing through either the induction of two nicks on opposite DNA strands14,23,26, or multiple nicks on the same DNA strand (Fig. 4B)15.
A Schematic representation of the gRNA spacers targeting the NCF1 ΔGT sequence. ssODN template (grey). crRNAs (green) designed for Cas12a. sgRNAs (blue) designed for the double Cas9 nickases approach, targeting adjacent sites on the NCF1 locus with two sgRNAs (sgRNA1 and sgRNA8) sequences. For each gRNA the ΔGT position (red), PAM (black) and cleavage sites (arrowhead in DNA corresponding to gRNAs colors) are indicated. B Different DNA breaks surrounding the ΔGT mutation, induced by complexing gRNAs with different CRISPR-Cas variants. C TIDER analysis of editing frequency in bulk PLB-985 NCF1 ΔGT cells treated with different combination of CRISPR-Cas variants and gRNAs (n = 2 biological replicates, expressed as mean ± SD). D Percentage of p47phox expressing cells within the CD11b+ population in bulk culture upon differentiation (n = 3 biological replicates, data are expressed as mean ± SD). E PCR-RFLP analysis of 23 monoclonal lines per condition. F Copy number analysis in individual clones detected by ddPCR. 23 clones were analyzed per condition.
To induce DNA nicks with different structures, we employed nCas9 variants (D10A and/or H840A)23. The NCF1 ΔGT mutation-specific sgRNA1 was chosen as the targeting guide to prevent re-cleavage upon ΔGT repair (Fig. 4A). sgRNA8 was selected to obtain a sgRNA pair within the requirements for robust gene editing (Fig. 4A)14,23. By complexing the paired sgRNA with combinations of nCas9 variants, we generated staggered DSBs with 45-bp overhangs at 5’ (O1) and 3’ termini (O2), or two adjacent nicks on the same DNA strand (S1 and S2) (Fig. 4B).
The editing activity of the sgRNA8 guide in the NCF1 gene and pseudogenes was confirmed in PLB-985 NCF1 ΔGT cells (Fig. 4C). The relative position of the sgRNA8-guided DSB on the edge of the ssODN homology arm accounts for the absence of knock-in events, while indel-mediated frameshift restoration might explain detectable expression of the p47phox protein (Fig. 4D), as previously observed for treatments with sgRNA1 without corrective template (Fig. 1E). The highest knock-in efficiency (41.0% ± 5.9%, Fig. 4C) and the highest p47phox protein expression (69.7% ± 1.8%, Fig. 4D) could be achieved by treatment with Cas9-sgRNA1.
We achieved NCF1 correction in 24.3% ± 6.0% of targeted sequences in bulk culture with the double-nickase approach, cleaving different DNA strands (O1, O2) with no apparent polarity bias, although with lower efficiency compared to the sample treated with Cas9-sgRNA1 (Fig. 4C–E). Paired cleavage on the same DNA strand (S1, S2) led to markedly lower correction efficiencies (4.9% ± 1.9%). Introduction of a single nick (N1, N2) resulted in low editing frequency (1.4% ± 0.4%) and no detectable correction events, as reflected by the absence of p47phox expression (Fig. 4D).
Compared to cells treated with Cas9-sgRNA1 (11/23, Fig. 3E), treatment with Cas12a-crRNA2 caused a marked increase in the number of individual clones with deletion (14/23) and/or duplication (4/23) events between on-target loci (Fig. 4F). The double-nickase approach, generating double-strand breaks with large sticky overhang ends, resulted in chromosomal aberration frequencies (O1: 10/23, O2: 11/23) comparable to the frequencies observed in cells treated with Cas9-sgRNA1 (11/23, Fig. 3E). The generation of adjacent nicks on the same DNA strand also led to comparable levels of chromosomal aberrations (S1: 11/23, S2: 7/23, Fig. 4F), despite the absence of DSBs.
Additionally, upon induction of a single-strand break at the NCF1 loci, we detected individual clones with chromosomal deletions both within and/or surrounding the NCF1 loci (N1: 1/23, N2: 2/23, Fig. 4F), even in the absence of detectable indels or corrections (Fig. 4C–E).
Cas9 RNP-mediated functional correction of p47phox CGD CD34 + HSCs is accompanied by deletions between on-target loci
To assess the reproducibility of the results obtained in the CGD model cell line in clinically relevant cells, patient-derived p47 CGD CD34+ HSCs were treated with CRISPR-Cas9:sgRNA1 RNPs and a corrective ssODN template (Fig. 5A). Targeting the NCF1 ΔGT in these cells resulted in modest correction efficiency of 5.1% ± 0.9% (Fig. 5B, C). Colony forming unit (CFU) assay of treated cells showed reduced viability upon electroporation, without apparent lineage skewing (Fig. 5D).
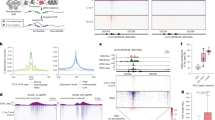
A Experimental workflow of the editing and analysis of p47phox CGD CD34+. B TIDER analysis of editing frequencies at the NCF1 gene and pseudogene loci in HSCs upon RNP treatment with sgRNA1 (n = 2, technical replicates, data are represented as mean ± SD). Statistical analysis with unpaired t test. C Distribution of GTGT-content calculated by PCR-RFLP analysis in CFU derived from Cas9-sgRNA1 RNP-treated CD34+ cells (n = 30). D Colony forming assay of RNP-treated versus untreated CD34+ cells (n = 2, technical replicates, data are represented as mean ± SD). Burst-forming unit-erythroid (BFU-E); CFU-granulocyte, macrophage (CFU-GM); CFU-granulocyte (CFU-G); CFU-macrophage (CFU-M); and CFU-granulocyte, erythrocyte, macrophage, megakaryocyte (CFU-GEMM). E Flow cytometry analysis of p47phox expression and DHR in CD11b positive populations upon myeloid differentiation. Gating strategies are presented in Supplementary Fig. S7. F Copy number detected by ddPCR of regions surrounding the NCF1 loci in CFU of untreated control (NT, n = 33) and sample treated with sgRNA1 and ssODN (sgRNA1+T, n = 40). Dotted lines indicate the upper and lower limit of the confidence interval determined from the NT group, followed by confirmation of the outliers using Z score normalization. G Quantification of chromosomal aberrations in the sample treated with sgRNA1 and ssODN (n = 40), based on ddPCR analysis in F. H–J CAST-Seq analysis of NCF1-edited human CD34+ cells from healthy donor. H Circos plot depicting results of chromosomal rearrangements identified in cells edited with Cas9-sgRNA1 (left) and Cas12a-crRNA2 (right). On-target (ON) genomic aberrations are indicated in green, off-target mediated translocations (OMT) in red, and homology-mediated translocations (HMT) in blue. The arcs highlight identified translocations between ON and other sites; the black outer layer displays chromosomes 1-22, X and Y; the red ring indicates the alignment score against the gRNA sequence with significant score accentuated by red dots; the blue ring indicates length of sequence homology, with significance emphasized by blue dots. I Alignment of the nominated off-target (OT) site on chromosome 3 to the on-target locus (NCF1B/NCF1C) and number of CAST-Seq hits. J CAST-Seq coverage plots show reads aligned to a +/−10 kb region flanking the on-target site NCF1B, as a surrogate for all NCF1 loci (arrow marked surrogate on-target sites, NCF1B/NCF1/NCF1C). Genomic deletions (DEL) are shown in orange and inversions (INV) in purple. The x-axis represents the chromosomal coordinates, the y-axis shows log2 read count per million (CPM), and the dotted line indicates the cleavage site.
RNP-mediated editing of the NCF1 ΔGT mutation in p47 CGD CD34+ HSCs in the presence of the corrective ssODN template restored p47phox protein expression and NADPH oxidase activity upon myeloid differentiation in 20.0% and 8.5% cells, respectively (Fig. 5E). On the other hand, editing without corrective template resulted in a low fraction of p47phox expressing cells (4.9%) and fewer than 1% cells with functional NADPH oxidase (Fig. 5E).
Upon treatment with Cas9:sgRNA1 and a ssODN template, edited patient CD34+ HSCs were differentiated into hematopoietic progenitor cells using a CFU assay, followed by isolation of single CFUs for CNV determination. ddPCR analysis of the regions surrounding and between the cleaved NCF1 gene loci identified a high number of colonies carrying chromosomal aberrations (14/40). While most aberrations were detected in the genomic regions within the NCF1 target loci (12/40), a fraction of colonies carried aberrations extended to the loci upstream of NCF1B (CALN1, 2/40) and downstream of NCF1C (HIP1, 4/40) (Fig. 5F, G). No aberrant gene copy number was detected on the short arm of chromosome 7 (IL6, 0/39), while two colonies exhibited increased copy number (CN) 27 Mb downstream of NCF1C detected (CUX1, 2/39) (Supplementary Fig. S5A). We further investigated the colonies for copy-neutral loss of heterozygosity (LOH), as was recently observed upon CRISPR editing in HSPCs27. No LOH was detected in the colonies at two tested genomic locations, 56 Mb (0/42, rs3735035) and 0.8 Mb (0/19, rs2286822 and rs2286823) downstream of NCF1C (Supplementary Fig. S5B).
Notably, targeting of NCF1 ΔGT in CD34+ cells from healthy donors with sgRNA1, which cleaves only NCF1B and NCF1C, resulted in consistent correction of the NCF1 pseudogenes, with no detectable chromosomal aberration (n = 48 CFU) (Supplementary Fig. S6F, Supplementary Fig. S7).
Lower incidence of chromosomal aberrations upon limiting the cleavage activity to NCF1 pseudogenes was confirmed in the PLB-985 WT cell line (Supplementary Fig. S6G-H). Conversely, targeting of all NCF1 genes with sgRNA8 increased the incidence of chromosomal aberrations in both healthy CD34+ cells (5/23) and in the PLB-985 WT cell line (14/23) (Supplementary Fig. S6E–H). Thus, we conclude that the induction of chromosomal aberrations between Cas9 RNP-targeted NCF1 loci results from the introduction of simultaneous DSBs in the NCF1 (pseudo)genes. These findings are in agreement with other studies indicating that incidence of aberrations is inversely correlated with the distance between the simultaneously targeted loci28,29.
To evaluate potential genotoxic effects on the edited cells, we performed chromosomal aberrations analysis by single targeted linker-mediated PCR sequencing (CAST-Seq), a technology that allows unbiased and sensitive identification of on- and off-target effects by detecting genomic rearrangements30. Due to high sequence similarity between the NCF1 loci and the surrounding regions (Supplementary Fig. S8), a custom reference genome was generated using NCF1B as the surrogate while masking the NCF1 and NCF1C¸ for any chromosomal rearrangements determined by CAST-Seq analyses. Human CD34+ cells from a healthy donor treated with Cas9-sgRNA1 revealed one off-target-mediated translocation (OMT) with a putative off-target site in chromosome 3, as well as three homology-mediated translocations (HMTs) (Fig. 5H, I; Supplementary Fig. S9). In samples treated with Cas12a-crRNA2, two HMTs were identified. Based on the low number of CAST-Seq hits, all translocation events appear to be very rare (Supplementary Tables S8–S10). In addition, CAST-Seq showed that the expected rearrangements at the on-target site were large deletions and inversions (Fig. 5J), although the inversions cannot be assigned to one specific NCF1 gene or its pseudogenes.
Discussion
In this study, we aimed to reduce the incidence of chromosomal aberrations resulting from simultaneous CRISPR-mediated cleavage of on-target sites in the NCF1 gene and its adjacent pseudogenes in p47 CGD4. To enable precise identification of on-target chromosomal aberrations, we have established a ddPCR-based method for fast and reliable quantification of gene CN between and surrounding the targeted NCF1 loci.
None of the RNP-mediated editing strategies tested at the NCF1 loci was able to completely uncouple gene correction from chromosomal rearrangements in the PLB-985 NCF1 ΔGT cell line17. While a modest reduction in the incidence of chromosomal aberrations was observed for several of the editing approaches that we tested, the concomitant reduction of editing efficiency would likely hinder their potential clinical applicability. The observed genomic instability may be attributed to the localization of the NCF1 (pseudo)genes in highly homologous low-copy repeat elements and their predisposition to frequent chromosomal rearrangements, making this locus particularly challenging for genome editing strategies31,32. Importantly, the instability of the chromosomal region containing NCF1 naturally leads to deletions and duplications responsible for two rare genetic disorders, Williams-Beuren syndrome (WBS) with association to blood malignancies33,34,35,36,37, 7q11.23 duplication syndrome38 with implications in myeloid disorders39,40.
In addition to multiple DSBs-mediated loss of intervening genetic sequences, the complex aberration patterns, which include deletions and duplications in different chromosome regions, observed in several clones, are consistent with homology-mediated intra- and inter-chromosomal template-switching during replication41. Of note, a clone carrying a homozygous deletion between NCF1B-NCF1, as described here, can result in an in-frame corrected NCF1 gene. However, such fusion product of NCF1B-NCF1 highlights the danger of detrimental editing outcomes, owing to the large deletion specific to NCF1B-NCF1 that encompasses at least 17 genes in the WBS-associated genomic region33, and the lower promoter activity of the NCF1 pseudogenes reported as compared to the wild-type NCF142.
Unexpectedly, induction of a single-strand nick also led to chromosomal aberrations, likely by triggering the break-induced replication (BIR), and causing replication fork stalling or collapse and conversion of the single-strand break (SSB) to a DSB43,44. Thus, direct CRISPR-mediated generation of DSBs is not required for the induction of chromosomal rearrangements. This observation should warrant caution for the application of nCas9, generally thought to have minimal impact on genome integrity14,15,20, and especially multiplexed nickase-based approaches45, as induction of single nicks can also lead to DSBs and resulting chromosomal rearrangements, albeit with low frequency.
To avoid potential adverse events arising from chromosomal aberrations, alternative CRISPR approaches may be employed. The gene conversion underlying p47 CGD usually results in the transfer of large stretches of pseudogene sequence to the NCF1 gene7. Thus identification and targeting of SNPs in the NCF1 gene is possible5, but requires a patient-specific approach. Alternatively, CRISPR-mediated targeted knock-in of the gene-coding sequence to a specific genomic location46, such as a safe genomic harbor, could permit bypassing the direct targeting of NCF147,48. In addition, the predominant ∆GT mutation could be a target for prime editing49. This approach is currently being explored with reports of high efficacy and long-term correction in immunodeficient mice50. The DSB-independent prime editor platform may potentially improve the safety of direct NCF1 editing as compared to Cas9 DSB-based approaches. However, the genotoxicity risk of prime editors that can arise from the conversion of DNA SSB into DSB during cellular replication is still a concern51.
Upon editing of CGD patient HSPCs, we observed lower colony forming potential as compared to non-treated controls. Although no comparison was made in this context between HSPCs from CGD and from healthy donors, it is known that CGD patients have higher proliferative stress related to persistent chronic inflammatory background52.
The detection of CN aberrations up to 27 Mb away from the targeted NCF1 loci in p47 CGD CD34+ HSCs suggests that multiplexed editing can have a major impact on the integrity of the targeted chromosome, and implicates CRISPR-based editing in the generation of large aberrations and chromosomal loss10,27,53. While we consciously designed this study to mostly evaluate CNV in the proximity of the edited region, additional targets may provide more information on the extent of genomic regions affected by the aberrations. Apart from the described ddPCR method that is intrinsically limited to CNV analysis, we employed CAST-Seq to investigate post-editing genome-wide translocations in healthy donor HSCs. Due to the high sequence similarity between NCF1, NCF1B and NCF1C, analysis of CAST-Seq results had some limitations as all detected chromosomal translocations, including OMTs and HMTs, could not be specifically assigned to any of the three genes. Similarly, the homology of the pseudogenes and the reverse orientation of NCF1 may have led to the misclassification of deletions and inversions at the on-target site. In terms of detecting inversions at the highly conserved NCF1 cluster that spans beyond 1.91 Mb (Supplementary Fig. S11), current long-read sequencing technologies can distinguish the NCF1 cluster with improved alignment and variant calling54,55. Hence, combinations of interphase fluorescence in-situ hybridization, short- and long-read sequencing, and optical mapping will be required to accurately map this region56,57.
In conclusion, we have shown that targeted editing of the NCF1 gene and pseudogenes on the same chromosome can correct the NCF1 ΔGT mutation but is inherently associated with the introduction of chromosomal aberrations in a fraction of the edited cells. Importantly, our presented data provided valuable insights on genome editing involving multiple on-targets on the same chromosome, such as the HBB/HBD and CCR2/CCR5 loci58,59, in which DSB-independent editing systems may be advantageous in terms of safety for future clinical application of pseudogene-associated disorders.
Materials and methods
Cell culture
PLB-985 wild-type (WT) and PLB-985 NCF1 ΔGT cell lines17 were cultured in RPMI 1640 medium (PAN-Biotech, Aidenbach, Germany) supplemented with 10% (v/v) fetal calf serum (FCS) (PAN-Biotech), 100 U/mL penicillin, and 100 mg/mL streptomycin (Thermo Fisher Scientific, Reinach, Switzerland). For granulocyte differentiation, cells were cultured 7–9 days in RPMI 1640 medium supplemented with 5% (v/v) FCS and 0.5% (v/v) N,N-dimethylformamide (Sigma-Aldrich, Buchs, Switzerland). Single clones were generated by FACS with BD FACSAria™ III Cell Sorter (Becton Dickinson, Allschwil, Switzerland).
Human healthy CD34+ cells from mobilized peripheral blood were either purchased (Lonza Group AG, Basel, Switzerland) or acquired from a p47 CGD donor under written informed consent. Sample collection and processing were performed in accordance with ethical principles, applicable local laws and regulations (ethics vote KEK ZH 2015/0135, BASEC-Nr. PB_2016-02202), with the written informed consent from the donor’s parents for these investigations under the local rules concerning general consent for research participation of the University Children’s Hospital Ulm. CD34+ cells were cultured in X-VIVOTM 20 (Lonza), supplemented with 1% (v/v) human albumin (CSL Behring AG, Bern, Switzerland), 300 ng/mL stem cell factor, 300 ng/mL FMS-like tyrosine kinase-3 ligand (Flt3-L) and 100 ng/mL thrombopoietin (CellGenix GmbH, Freiburg, Germany). CD34+ cells were expanded and differentiated in MethoCult™ H4434 Classic (Stemcell Technologies, Cologne, Germany) for 14 days, according to the manufacturer’s instructions.
Guide RNA (gRNA) design
Single guide RNAs (sgRNA) for Cas9, and CRISPR RNAs (crRNA) for Cas12a were designed with CHOPCHOP v3 webtool (https://chopchop.cbu.uib.no/)60 and purchased from Integrated DNA Technologies (IDT, Iowa, USA).
Cas9 and Cas12 nuclease preparation
The Streptococcus pyogenes (Sp)Cas9 expression vector (Addgene 78312) encodes a Cas9 fusion protein with N-terminal hexahistidine-maltose binding protein tag (6xHis-MBP) followed by a tobacco etch virus (TEV) protease cleavage site and a C-terminal hemagglutinin (HA) tag, green fluorescent protein (GFP) tag and three nuclear localization signals (NLS) yielding a 6xHis-MBP-TEV-Cas9-HA-2xNLS-GFP-NLS construct. Point mutations for nickases (dCas9_D10A, dCas9_H840A) and catalytically dead Cas9 (dCas9_D10A, H840A) were introduced by inverse PCR and confirmed by DNA sequencing. The Acidaminococcus sp. (As) Cas12a gene was amplified by PCR (Addgene plasmid 90095) to replace the Cas9 gene in the Cas9 expression vector by Gibson assembly, generating the 6xHis-MBP-TEV-Cas12a-HA-2xNLS-GFP-NLS expression construct. Purification of Cas9 was carried out as described25,61, with minor adjustments. Nickase Cas9 constructs were purified for wild type. Purification of Cas12a was done as described62,63. In brief, Cas9 and Cas12a constructs were expressed in E. coli BL21 Rosetta2 (DE3) cells (Novagen). Cells were lysed in 20 mM Tris pH 8.0, 500 mM NaCl, 5 mM imidazole, 1 µg/mL Pepstatin, 200 µg/mL AEBSF by ultrasonication. Clarified lysate was applied to a 10 ml Ni-NTA (Sigma-Aldrich) affinity column. The column was washed with 20 mM Tris pH 8.0, 500 mM NaCl, 10 mM imidazole, and bound protein was eluted by increasing imidazole concentration to 250 mM. Eluted protein was dialysed against 20 mM HEPES pH 7.5, 250 mM KCl, 10% glycerol, 1 mM dithiothreitol (DTT), 1 mM EDTA (Cas9) or 20 mM HEPES pH 7.5, 250 mM KCl, 1 mM DTT, 1 mM EDTA (Cas12a) overnight at 4 °C in the presence of TEV protease to remove the 6xHis-MBP affinity tag. Cleaved protein was further purified using a HiTrap HP Heparin column (GE Healthcare), eluting with a linear gradient to 1.0 M KCl. Elution fractions were pooled, concentrated, and further purified by size exclusion chromatography using a Superdex 200 (16/600) column (GE Healthcare) in 20 mM HEPES-KOH pH 7.5, 500 mM KCl, 1 mM DTT yielding pure, monodisperse proteins. Aliquots were flash-frozen in liquid nitrogen and stored at − 80 °C.
CRISPR-Cas nuclease RNP electroporation
Ribonucleoprotein (RNP) complexes were formed by complexing Cas protein with gRNAs at 1:1.2 molar ratio and delivered to the cells with the Neon™ Transfection System (Thermo Fisher Scientific). 200,000 PLB-985 cells were electroporated in Buffer R (1350 V, 10 ms, 3 pulses). Unless otherwise stated, 2.7 µM of RNP complexes and 2.5 µM ssODN template (Microsynth AG, Balgach, Switzerland) were delivered per samples. The ssODN template is complementary to the negative strand with two 50 bp homology arms that flanks the ΔGT mutation (Supplementary Table S5). Human CD34+ cells were cultured for 48 h after thawing, and 100,000 cells were electroporated in Buffer T (1400 V, 10 ms, 3 pulses) containing 7.2 µM of RNP complexes and 2.5 µM ssODN template per reaction.
Quantification of editing efficiency
Cleavage and correction efficiency were assessed by the TIDER (https://tide.nki.nl/)64. Correction efficiency in PLB-985 single clones and CFUs were determined by PCR-restriction fragment length polymorphism (PCR-RFLP)18. Briefly, PCR products spanning the target locus were digested with BsrG1 and PstI (New England Biolabs, Ipswich, MA) and developed in a 7.5% polyacrylamide gel. GTGT-content was determined on size-normalized band intensities with ImageJ as previously described18.
CNV analysis with ddPCR
Copy number (CN) variation (CNV) of target genomic regions was assessed in individual PLB-985 clones and in CD34+-derived CFUs by droplet digital PCR (ddPCR) with the QX200 Droplet Digital PCR System (Bio-Rad Laboratories, California, USA). Briefly, genomic DNA was digested with DraI (20 U/µL) (New England BioLabs) at 37 °C for 1 hour and 5 minutes heat inactivation at 65 °C. The PCR reaction consisted of 1x ddPCR Supermix for Probes (No dUTP) (Bio-Rad), 900 nM probes, 250 nM primers (IDT) (Supplementary Table S1) and 100 ng of digested DNA. For droplet generation, 20 µL of PCR reaction mix was transferred to DG8™ Cartridges for QX200™/QX100™ Droplet Generator (Bio-Rad) and processed as per manufacturer’s protocol. Samples were initially denatured for 10 minutes at 95 °C, then amplified with 40 cycles of denaturation (95 °C, 30 sec), annealing (61 °C, 1 min) and extension (72 °C, 1 min), with the final 10 minutes incubation at 95 °C. Droplet reading was carried out with the QX200 Droplet Digital PCR System (Bio-Rad), and data were analyzed with QuantaSoft Analysis Pro Software v1.0 (Bio-Rad).
CNV analysis with qPCR
Quantitative PCR (qPCR) based CNV assessment between the NCF1 loci was performed with the QuantStudio 7 Flex Real-Time PCR System (Thermo Fisher Scientific) and SsoAdvanced Universal SYBR Green Supermix (Bio-Rad), as previously described4.
Heterozygous SNP analysis by Sanger sequencing
Two locations carrying heterozygous single-nucleotide polymorphisms (SNPs) in p47 CGD HSPCs (7q32.3, rs3735035 and 7q11.23, rs2286822 and rs2286823) were identified by Sanger sequencing following targeted PCR amplification of common SNPs on Chr7q. Edited p47 CGD HSPC-derived CFU were analyzed by targeted PCR amplification (Supplementary Table S2) and Sanger sequencing.
Flow cytometry
Cells were stained with a viability dye and CD11b antibody (antibodies listed in Supplementary Table S3) with additional FcR Blocking Reagent (Miltenyi Biotec Swiss AG, Solothurn, Switzerland) for HSCs derived cells. For p47phox staining, cells were permeabilized with a Fixation/Permeabilization Solution Kit (Becton Dickinson). For dihydrorhodamine (DHR) test, surface labeled cells were pre-incubated with 2.9 μM of DHR 123 (Sigma-Aldrich) and 150 U/mL of catalase (Sigma-Aldrich) in PBSgg (0.05% w/v gelatin, 0.09% w/v glucose) for 15 min at 37 °C, and subsequently activated with 1 µg/mL of phorbol 12-myristate 13-acetate (PMA) for 15 min at 37 °C6. Labelled samples were recorded on a BD LSRFortessa™ Flow Cytometer (Becton Dickinson), and the data were analyzed using FlowJo™ Software v10.7 (Becton Dickinson).
CAST-Seq
CAST-Seq analyses were performed on genomic DNA isolated from human CD34+ cells of a healthy donor edited with Cas9-sgRNA1 and Cas12a-crRNA2 as previously described65. The bioinformatics pipeline was modified to accommodate pseudogenes: due to the high similarity of the NCF1 loci (NCF1B:NCF1:NCF1C) and their surrounding regions, the regular CAST-Seq protocol cannot discriminate the three loci. To quantify the homology between NCF1 and its pseudogenes, the surrounding region of NCF1B was screened, extending up to 100 kb downstream of NCF1C, and the percentage of sequence identity was calculated using global-local pairwise alignment. The probability of observing such a high homology by chance was estimated through comparison of the homology score with the empirical cumulative distribution derived from 1000 randomly generated sequences. Because the sequence homology was highly significant (98% vs. 50% in random sequences, p value < 10−4), NCF1B was selected as the surrogate for any chromosomal rearrangements detected in this study. In addition, the sequence chr7:73235945-75272044 includes NCF1 and NCF1C was masked to generate a custom reference genome for the CAST-Seq analyses. Annotation for barcode hopping was included in the CAST-Seq algorithm, and coverage analysis has been revised to reduce the execution time by aligning the gRNAs only to the most covered regions for each site, as previously described65,66. Three technical CAST-Seq replicates were performed on pooled samples from a healthy donor. Only sites identified as significant hits in at least two replicates and showing a read/hit ratio above 10 were considered as putative events.
Statistics and reproducibility
Statistical analysis was performed using GraphPad Prism 8.4.3 (GraphPad Software, La Jolla, CA, USA). All relevant statistical analyses and sample sizes were defined in their respective figure legends. P values were indicated within the figures. For ddPCR data, the upper and lower limit of the confidence interval was determined based on the mean copy number of the NT group, whereby the limit was set to mean ± (3 × SD) as the cutoff criterion for detecting increased/decreased copy numbers. The outliers were confirmed by Z score normalization.
Inclusion and ethics
All authors have fulfilled the criteria for authorship required by Nature Portfolio journals. Participation of all authors was essential for the design and implementation of the study. Our research was determined in collaboration with local partners. Roles and responsibilities were agreed amongst collaborators prior to the research. Our research was not severely restricted in the setting of the researchers and did not result in stigmatization, incrimination, discrimination or personal risk to participants. Primary sample collection and processing were performed in accordance with ethical principles, applicable local laws and regulations (ethics vote KEK ZH 2015/0135, BASEC-Nr. PB_2016-02202), with the written informed consent from the donor’s parents for these investigations under the local rules concerning general consent for research participation of the University Children’s Hospital Ulm. All authors ensured research and literature sources were taken into account in citations.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the study are available within the paper and the Supplementary Information. Numerical source data of all graphs from the main Figs. 1–5 are provided in the Supplementary Data Excel file. Primary data can be made available upon request to the corresponding author. The CAST-Seq data generated in this study can be accessed in the NCBI Gene Expression Omnibus (GEO) repository under accession number GSE263692.
Code availability
All code for CAST-Seq analysis associated with the current submission is available at https://doi.org/10.3389/fgeed.2023.1130736 and https://doi.org/10.1016/j.ymthe.2024.03.00665,66.
References
Bianchi, M. et al. Restoration of NET formation by gene therapy in CGD controls aspergillosis. Blood 114, 2619–2622 (2009).
Brendel, C. et al. Human miR223 promoter as a novel myelo-specific promoter for chronic granulomatous disease gene therapy. Hum. Gene Ther. Methods 24, 151–159 (2013).
Siler, U. et al. Successful combination of sequential gene therapy and rescue Allo-HSCT in two children with X-CGD - importance of timing. Curr. Gene Ther. 15, 416–427 (2015).
Wrona, D. et al. CRISPR-directed therapeutic correction at the NCF1 locus is challenged by frequent incidence of chromosomal deletions. Mol. Ther. Methods Clin. Dev. 17, 936–943 (2020).
Klatt, D. et al. Targeted repair of p47-CGD in iPSCs by CRISPR/Cas9: functional correction without cleavage in the highly homologous pseudogenes. Stem Cell Reports 13, 590–598 (2019).
Schejtman, A. et al. Lentiviral gene therapy rescues p47phox chronic granulomatous disease and the ability to fight Salmonella infection in mice. Gene Ther. https://doi.org/10.1038/s41434-020-0164-6 (2020).
Hayrapetyan, A., Dencher, P. C. D., van Leeuwen, K., de Boer, M. & Roos, D. Different unequal cross-over events between NCF1 and its pseudogenes in autosomal p47(phox)-deficient chronic granulomatous disease. Biochim. Biophys. Acta 1832, 1662–1672 (2013).
Merling, R. K. et al. Gene-edited pseudogene resurrection corrects p47phox-deficient chronic granulomatous disease. Blood Adv. 1, 270–278 (2017).
Kosicki, M., Tomberg, K. & Bradley, A. Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nat. Biotechnol. 36, 765–771 (2018).
Cullot, G. et al. CRISPR-Cas9 genome editing induces megabase-scale chromosomal truncations. Nat. Commun. 10, 1136 (2019).
Kim, S., Kim, D., Cho, S. W., Kim, J. & Kim, J.-S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Liang, X. et al. Rapid and highly efficient mammalian cell engineering via Cas9 protein transfection. J. Biotechnol. https://doi.org/10.1016/j.jbiotec.2015.04.024 (2015).
Sapozhnikov, D. M. & Szyf, M. The PROTECTOR strategy employs dCas orthologs to sterically shield off-target sites from CRISPR/Cas activity. Sci. Rep. 13, 2280 (2023).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR cas9 for enhanced genome editing specificity. Cell. https://doi.org/10.1016/j.cell.2013.08.021 (2013).
Hyodo, T. et al. Tandem paired nicking promotes precise genome editing with scarce interference by p53. Cell Rep. 30, 1195–1207.e7 (2020).
Swarts, D. C. & Jinek, M. Cas9 versus Cas12a/Cpf1: structure–function comparisons and implications for genome editing. Wiley Interdiscip. Rev. RNA 9, e148 (2018).
Wrona, D., Siler, U. & Reichenbach, J. CRISPR/Cas9-generated p47 phox -deficient cell line for chronic granulomatous disease gene therapy vector development. Sci. Rep. 7, 6–11 (2017).
Wrona, D., Siler, U. & Reichenbach, J. Novel Diagnostic Tool for p47 phox -deficient chronic granulomatous disease patient and carrier detection. Mol. Ther. Methods Clin. Dev. 13, 274–278 (2019).
Dehé, P. M. & Gaillard, P. H. L. Control of structure-specific endonucleases to maintain genome stability. Nat. Rev. Mol. Cell Biol. 18, 315–330 (2017).
Maizels, N. & Davis, L. Initiation of homologous recombination at DNA nicks. Nucleic Acids Res. 46, 6962–6973 (2018).
Scully, R., Panday, A., Elango, R. & Willis, N. A. DNA double-strand break repair-pathway choice in somatic mammalian cells. Nat. Rev. Mol. Cell Biol. 20, 698–714 (2019).
Ling, A. K. et al. Double-stranded DNA break polarity skews repair pathway choice during intrachromosomal and interchromosomal recombination. Proc. Natl. Acad. Sci. USA. 115, 2800–2805 (2018).
Shen, B. et al. Efficient genome modification by CRISPR-Cas9 nickase with minimal off-target effects. Nat. Methods 11, 399–402 (2014).
Caldecott, K. W. Single-strand break repair and genetic disease. Nat. Rev. Genet. 9, 619–631 (2008).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. https://doi.org/10.1126/science.1225829 (2012).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Boutin, J. et al. CRISPR-Cas9 globin editing can induce megabase-scale copy-neutral losses of heterozygosity in hematopoietic cells. Nat. Commun. 12, 1–12 (2021).
Canver, M. C. et al. Characterization of genomic deletion efficiency mediated by clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 nuclease system in mammalian cells. J. Biol. Chem. 292, 2556 (2017).
Kraft, K. et al. Deletions, inversions, duplications: engineering of structural variants using CRISPR/Cas in mice. Cell Rep 10, 833–839 (2015).
Turchiano, G. et al. Quantitative evaluation of chromosomal rearrangements in gene-edited human stem cells by CAST-Seq. Cell Stem Cell 28, 1136–1147.e5 (2021).
Bayés, M., Magano, L. F., Rivera, N., Flores, R. & Pérez Jurado, L. A. Mutational mechanisms of williams-beuren syndrome deletions. Am. J. Hum. Genet. 73, 131–151 (2003).
Osborne, L. R. & Mervis, C. B. Rearrangements of the Williams–Beuren syndrome locus: molecular basis and implications for speech and language development. Expert Rev. Mol. Med. 9, 1 (2007).
Kimura, R. et al. Williams–Beuren syndrome as a potential risk factor for burkitt lymphoma. Front. Genet. 9, 368 (2018).
Decimi, V. et al. Williams syndrome and mature B-Leukemia: a random association? Eur. J. Med. Genet. https://doi.org/10.1016/j.ejmg.2016.10.007 (2016).
Liu, Y. et al. LncRNA ABHD11-AS1 promotes the development of endometrial carcinoma by targeting cyclin D1. J. Cell. Mol. Med. 22, 3955–3964 (2018).
Wu, D. D. et al. Role of the lncRNA ABHD11-AS1 in the tumorigenesis and progression of epithelial ovarian cancer through targeted regulation of RhoC. Mol. Cancer 16, 138 (2017).
Li, Y. et al. Multifaceted regulation and functions of replication factor C family in human cancers. Am. J. Cancer Res. 8, 1343 (2018).
N, V. et al. Fourteen new cases contribute to the characterization of the 7q11.23 microduplication syndrome. Eur. J. Med. Genet. 52, 94–100 (2009).
Jerez, A. et al. Loss of heterozygosity in 7q myeloid disorders: clinical associations and genomic pathogenesis. Blood 119, 6109 (2012).
Hosono, N. et al. Recurrent genetic defects on chromosome 7q in myeloid neoplasms. Leukemia 28, 1348 (2014).
Carvalho, C. M. B. & Lupski, J. R. Mechanisms underlying structural variant formation in genomic disorders. Nat. Rev. Genet. 17, 224 (2016).
Brunson, T., Wang, Q., Chambers, I. & Song, Q. A copy number variation in human NCF1 and its pseudogenes. BMC Genet. 11, 13 (2010).
Malkova, A. & Ira, G. Break-induced replication: functions and molecular mechanism. Curr. Opin. Genet. Dev. 23, 271–279 (2013).
Costantino, L. et al. Break-induced replication repair of damaged forks induces genomic duplications in human cells. Science 343, 88 (2014).
McCarty, N. S., Graham, A. E., Studená, L. & Ledesma-Amaro, R. Multiplexed CRISPR technologies for gene editing and transcriptional regulation. Nat. Commun. 11, 1–13 (2020).
Siow, K. M. et al. Targeted knock-in of NCF1 cDNA into the NCF2 locus leads to myeloid phenotypic correction of p47 phox -deficient chronic granulomatous disease. Mol. Ther. Nucleic Acids 35, 102229 (2024).
De Ravin, S. S. et al. Targeted gene addition in human CD34+ hematopoietic cells for correction of X-linked chronic granulomatous disease. Nat. Biotechnol. 34, 424–429 (2016).
Hayashi, H., Kubo, Y., Izumida, M. & Matsuyama, T. Efficient viral delivery of Cas9 into human safe harbor. Sci. Rep. 10, 1–14 (2020).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Heath, J. M. et al. Prime editing efficiently and precisely corrects causative mutation in chronic granulomatous disease, restoring myeloid function: toward development of a prime edited autologous hematopoietic stem cell therapy. Blood 142, 7129–7129 (2023).
Fiumara, M. et al. Genotoxic effects of base and prime editing in human hematopoietic stem cells. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01915-4 (2023).
Weisser, M. et al. Hyperinflammation in patients with chronic granulomatous disease leads to impairment of hematopoietic stem cell functions. J. Allergy Clin. Immunol. 138, 219–228.e9 (2016).
Essletzbichler, P. et al. Megabase-scale deletion using CRISPR/Cas9 to generate a fully haploid human cell line. Genome Res. 24, 2059–2065 (2014).
High-performance long-read assay enables contiguous data with N50 of 6–7 kb on existing Illumina platforms. Genomics Research Hub. Available at: https://www.illumina.com/science/genomics-research/articles/infinity-high-performance-long-read-assay.html (2022).
The HiFi difference — true long reads vs synthetic long reads. Products, Procedures + Protocols. Available at: https://www.pacb.com/blog/the-hifi-difference-true-long-reads-vs-synthetic-long-reads/. (2022).
Osborne, L. R. et al. A 1.5 million-base pair inversion polymorphism in families with Williams-Beuren syndrome. Nat. Genet. 29, 321–325 (2001).
Weissensteiner, M. H. et al. Combination of short-read, long-read, and optical mapping assemblies reveals large-scale tandem repeat arrays with population genetic implications. Genome Res. 27, 697–708 (2017).
Bothmer, A. et al. Characterization of the interplay between DNA repair and CRISPR/Cas9-induced DNA lesions at an endogenous locus. Nat. Commun. https://doi.org/10.1038/ncomms13905 (2017).
Schwarze, L. I., Głów, D., Sonntag, T., Uhde, A. & Fehse, B. Optimisation of a TALE nuclease targeting the HIV co-receptor CCR5 for clinical application. Gene Ther. 28, 588–601 (2021).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, 1247997 (2014).
Kissling, L., Monfort, A., Swarts, D. C., Wutz, A. & Jinek, M. Preparation and electroporation of Cas12a/Cpf1-guide RNA complexes for introducing large gene deletions in mouse embryonic stem cells. Methods Enzymol. 616, 241–263. https://doi.org/10.1016/bs.mie.2018.10.028 (2019).
Mohanraju, P., Van Der Oost, J., Jinek, M. & Swarts, D. C. Heterologous expression and purification of CRISPR-Cas12a/Cpf1. Bio Protoc. 8, e2842 (2018).
Brinkman, E. K. & van Steensel, B. Rapid quantitative evaluation of CRISPR genome editing by TIDE and TIDER. Methods Mol. Biol. https://doi.org/10.1007/978-1-4939-9170-9_3 (2019).
Klermund, J. et al. On- and off-target effects of paired CRISPR-Cas nickase in primary human cells. Mol. Ther. https://doi.org/10.1016/j.ymthe.2024.03.006 (2024).
Rhiel, M. et al. T-CAST: an optimized CAST-Seq pipeline for TALEN confirms superior safety and efficacy of obligate-heterodimeric scaffolds. Front.Genome Ed. 5, 1130736 (2023).
Acknowledgements
The study was supported by the second implementation phase of Highly Specialized Medicine (HSM2) of University of Zürich (awarded to J.R.), the Swiss National Science Foundation Grant 31003A_182567 (awarded to M.J.), and the Horizon Europe Pathfinder Challenge X-PAND consortium ID 101070950 (awarded to T.C.). F.R. and K.M.S. were supported by the Clinical Research Priority Program ImmuGene of University of Zürich (awarded to J.R. and M.J.). D.W. was funded by the Uniscientia Stiftung (awarded to J.R.). D.W. and O.P. received research grants from the University of Zurich (Forschungskredit; respectively FK-17-041 and FK-17-053). We thank the University Research Priority Program ITINERARE of the University of Zürich for funding our research on rare diseases. We thank Professor Dr. Ansgar Schulz, University Medical Center Ulm, Germany, for providing p47 CGD patient bone marrow HSCs for our experiments.
Author information
Authors and Affiliations
Contributions
F.R., K.M.S., D.W., C.F.G., T.C., U.M., M.J., U.S., and J.R. designed the experiments, analyzed the results, and wrote the manuscript; F.R., K.M.S, D.W., C.F.G., and O.P. performed experiments; F.R., K.M.S., and C.F.G. prepared the figures; M.S., K.B., L.K. and D.C.S. produced reagents used in the study; C.F.G., G.A., and T.C. analyzed the data generated from CAST-Seq. M.J. provided technical expertise related to design of CRISPR approaches.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Carsten Lederer, Philip Hublitz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Pei Hao and Mengtan Xing. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Raimondi, F., Siow, K.M., Wrona, D. et al. Gene editing of NCF1 loci is associated with homologous recombination and chromosomal rearrangements. Commun Biol 7, 1291 (2024). https://doi.org/10.1038/s42003-024-06959-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-024-06959-z