Abstract
Methylenetetrahydrofolate reductase (MTHFR) is a key enzyme in one-carbon (1C) metabolism, catalyzing the reduction of methylenetetrahydrofolate to methyltetrahydrofolate. Interestingly, Sphingobium lignivorans SYK-6, a model bacterium for the catabolism of lignin-derived aromatic compounds, possesses a unique MTHFR (S6MTHFR) that catalyzes the reverse reaction of typical MTHFRs—namely, the oxidation of methyltetrahydrofolate. However, no direct evidence supports this function. Here, we show that S6MTHFR catalyzes the oxidation of methyltetrahydrofolate and elucidate the molecular mechanism underlying the unique enzymatic properties of S6MTHFR based on its crystal structure. Furthermore, a database search reveals that a group of bacteria, including S. lignivorans SYK-6, utilize tetrahydrofolate-dependent demethylases to produce methyltetrahydrofolate, which is subsequently oxidized by an S6MTHFR-type enzyme. We propose that the combination of a demethylase with an S6MTHFR-type enzyme represents a distinct type of 1C metabolism that may regulate methionine biosynthesis.

Similar content being viewed by others
Introduction
One-carbon (1C) metabolism, a fundamental metabolic pathway for life, can be divided into folate and methionine cycles1. In the folate cycle, tetrahydrofolate (THF) accepts several types of 1C units, forming a series of intermediates1. Two of these intermediates, methylenetetrahydrofolate (CH2-THF) and methyltetrahydrofolate (CH3-THF), play pivotal roles in 1C metabolism. In most organisms, a 1C unit is introduced into the folate cycle by serine hydroxymethyltransferase or the glycine cleavage system as a form of methylene. The methylene group is transferred to THF to produce CH2-THF2,3,4, which is used to synthesise thymidylate. 10-Formyl-THF, used in the biosynthesis of fMet-tRNA and purine, is also generated from CH2-THF via methenyl-THF5. The remaining CH2-THF is converted by methylenetetrahydrofolate reductase (MTHFR; EC1.5.1.20) into CH3-THF, which transfers the 1C unit to homocysteine for methionine synthesis. The synthesised methionine can be converted to S-adenosylmethionine (Fig. 1a)6. MTHFR, which is a FAD-containing flavoenzyme, utilises NADH (NADPH in mammals) to reduce FAD, which in turn reduces CH2-THF to form CH3-THF (Fig. 1a)7,8. While CH3-THF can also reduce FAD, most MTHFRs have significantly higher FAD-reducing activity with NADH than with CH3-THF9. Additionally, the typical 1C metabolism synthesises CH2-THF prior to the MTHFR reaction. Therefore, MTHFR is generally thought to convert only CH2-THF to CH3-THF (i.e., forward reaction) in cells due to high concentrations of NADH/NADPH in physiologically conditions10,11.
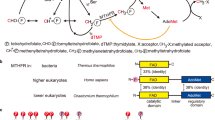
a In E. coli, MTHFR catalyses the reduction of CH2-THF, which is generated by the Gly cleavage system or serine hydroxymethyltransferase (SHMT). Electrons used in the reduction of CH2-THF were derived from NADH. b Predicted reaction scheme of S6MTHFR. S6MTHFR has been suggested to catalyse the oxidation of CH3-THF, which is generated by the methyl-transfer reaction from vanillate and syringate to THF by LigM and DesA, respectively. The reduced form of FAD (FADH2) must be oxidised by an electron acceptor (X).
Sphingobium lignivorans SYK-6 (formerly Sphingobium sp. SYK-6, hereafter SYK-6) is the most well-characterised bacterial degrader of lignin-derived aromatic compounds and is a promising platform for producing value-added chemicals from lignin12,13. Therefore, it is crucial to elucidate not only the catabolism of lignin-derived aromatic compounds in SYK-6 but also the metabolic system linked to it. SYK-6 can degrade various aromatic dimers and monomers with methoxy groups through vanillate and syringate. LigM and its homologue DesA, which are THF-dependent O-demethylases, convert vanillate to protocatechuate and syringate to 3-O-methylgallate by transferring a 1C unit from a methoxy group of the substrates to THF (Fig. 1b)14,15. Although LigM is structurally similar to an aminomethyltransferase of the Gly cleavage system, with 26% amino acid sequence identity, its product is not CH2-THF as in the typical glycine cleavage system, but CH3-THF15,16. Interestingly, SYK-6 exhibits methionine auxotrophy when growing on protocatechuate, which lacks a methoxy group, as a carbon source14. This fact suggests that CH2-THF and its derivatives, methenyl-THF and 10-formyl-THF, are produced during the growth of SYK-6 on protocatechuate, but CH2-THF cannot be converted to CH3-THF; thus, demethylation of the carbon source seems to directly affect methionine synthesis. Furthermore, 13C metabolic flux analysis of SYK-6 suggested that MTHFR of SYK-6 (S6MTHFR) catalyses only the oxidization of CH3-THF17. Thus, it is reasonable to consider that the S6MTHFR favours the reverse reaction of typical MTHFRs to produce CH2-THF from CH3-THF (Fig. 1b). Because of the unique characteristics of S6MTHFR, demethylation of the carbon source seems to directly affect methionine synthesis in SYK-6.
Although multiple studies have suggested that S6MTHFR has unique properties, no direct evidence of these properties has been obtained12,13,14,15,16,17; the exact biochemical and biological functions of S6MTHFR remain elusive. Since the four catalytically essential residues in Escherichia coli MTHFR (hereafter EcMTHFR)7,18,19,20,21—Glu28, Asp120, Gln183, and Phe223—are conserved in S6MTHFR, it is reasonable to assume that unknown key residues control the direction of the catalytic reaction of S6MTHFR. To investigate this possibility, we biochemically characterised S6MTHFR and determined its crystal structures. Our study revealed that S6MTHFR catalyses the reverse catalytic reaction of typical MTHFRs. It also elucidated the molecular mechanism underlying S6MTHFR based on those crystal structures. Moreover, structure-based amino acid sequence analysis showed that S6MTHFR can be classified into an unprecedented cluster in the MTHFR family. Some bacteria with methionine auxotrophy utilise THF-dependent demethylases to obtain a 1C unit and produce CH3-THF. In these bacteria, S6MTHFR-type enzymes appear to play an essential role in the oxidation of CH3-THF to CH2-THF, forming a unique folate cycle.
Results
Enzymatic characterization of S6MTHFR
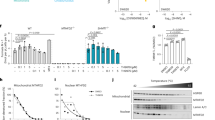
To elucidate the enzymatic properties of S6MTHFR, we evaluated its FAD reduction activity using NADH, with EcMTHFR serving as a control (Supplementary Fig. 1). As expected, S6MTHFR exhibited very low specific activity, reaching only 0.03 μmol·min⁻¹·mg⁻¹, compared to the significantly higher activity of EcMTHFR, which measured 125.4 μmol·min⁻¹·mg⁻¹, even in the presence of 250 μM NADH (Supplementary Table 1). In addition, no electron transfer from chemically reduced FAD (FADH2) to NAD+ was detected, suggesting that NADH and NAD+ hardly bind to S6MTHFR (Supplementary Fig. 2). Next, we analysed the CH3-THF oxidation activity in which FAD serves as the electron acceptor. The specific activity of the FAD reduction with CH3-THF (44.92 ± 0.71 μmol·min−1·mg−1) was significantly higher than that of EcMTHFR (1.88 ± 0.05 μmol·min−1·mg−1) (Supplementary Table 1). Additionally, unlike EcMTHFR, S6MTHFR showed no detectable CH₃-THF generation when NADH and CH₂-THF are used as substrates (Supplementary Fig. 3). An anaerobic titration experiment showed that the FAD in S6MTHFR was almost completely reduced by an equimolar amount of CH3-THF, suggesting that one hydride ion appears to be transferred from CH3-THF to FAD (Fig. 2a). High-performance liquid chromatography–mass spectrometry (HPLC–MS) analysis revealed that S6MTHFR converts CH3-THF to CH2-THF with kcat and Km values of 29.2 ± 0.412 s−1 and 0.0282 ± 0.00152 mM, respectively (Fig. 2b-e). The optimal pH and temperature of the catalytic reaction were 7.5 and 35°C, respectively (Fig. 2f, g). These results demonstrated that S6MTHFR has a different characteristic from typical MTHFRs: S6MTHFR catalyses the oxidation of CH3-THF as a substrate. Interestingly, despite the difference in enzymatic character, all critical residues for the catalytic reaction of EcMTHFR are conserved in S6MTHFR. We then determined the crystal structure of S6MTHFR to elucidate the molecular basis of the unique enzymatic properties of S6MTHFR.
a Anaerobic titration of S6MTHFR with CH3-THF. In an anaerobic chamber, 19 nmol of S6MTHFR (oxidised FAD) was titrated with 25 nmol of CH3-THF to reach saturation. The initial spectrum is shown by the blue line, and the final spectrum is black. The arrow indicates the maximum absorption wavelength, 450 nm, of FAD. Inset: Changes in the amount of oxidised FAD (measured at 450 nm) with the addition of CH3-THF. Reactions were performed once. b HPLC chromatograms of reaction mixtures using 20 nM S6MTHFR and 200 μM CH3-THF in the presence of 400 μM menadione at 30 °C. The retention times of CH3-THF and CH2-THF were 1.44 and 2.12 min, respectively. c, d ESI–MS (positive-ion mode) spectra of CH3-THF and CH2-THF observed at the start and after 5 min of the reaction, respectively. m/z values for the protonated molecular ions [M + H]+ of CH3-THF and CH2-THF were 460 and 458, respectively. e Steady-state kinetic analysis of S6MTHFR. Optimal pH and temperature were determined. Reactions were carried out in four independent experiments (n = 4). Data points marked with circles represent the means of the replicate measurements. Dots show individual data points. f, g Optimal pH and temperature were determined. Reactions were carried out in four independent experiments (n = 4). Data points marked with circles represent the means of the replicate measurements. Dots show individual data points.
Crystal structures of S6MTHFR and active site structure
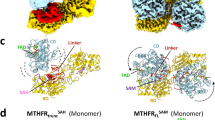
The crystal structures of S6MTHFR in the substrate-free and complex forms were determined at 1.5 Å and 1.85 Å resolution, respectively (Table 1). Although the asymmetric unit of the crystal contains two subunits, size exclusion chromatography with multiangle light scattering (SEC-MALS) analysis revealed that S6MTHFR exists as a monomer in solution (Supplementary Fig. 4). S6MTHFR adopts a TIM-barrel fold centred on eight parallel β-strands surrounded by eight α-helices (Fig. 3a). This fold is commonly observed in MTHFR structures22. The secondary structures are listed in Supplementary Fig. 5. Each loop region is designated using the names of the N- and C-terminal secondary structures.
a The crystal structure of the S6MTHFR‒CH3-THF complex is shown in a rainbow-coloured cartoon representation from the N to C termini (blue to red; PDB ID: 7XLF). FAD and the experimentally observed part of the CH3-THF molecule are shown by stick models in yellow and pink (for carbon atoms), respectively. The (|Fo | - |Fc | ) map, computed after annealing and refinement of a model with CH3-THF omitted, is shown as a mesh representation contoured at 3σ. Each secondary structure is labelled. b Active site structure of the substrate complex form. Carbon atoms of S6MTHFR, FAD, and CH3-THF are shown in green, yellow, and purple, respectively. c Comparison of the free and complex forms of S6MTHFR with CH3-THF. Helix α8 makes an induced fit upon substrate binding. The substrate-free (PDB ID: 7XG9) and complex forms are shown in light green and light orange, respectively. Phe215, Cys219, and helix α8 are shown in green and purple for the substrate-free and complex forms, respectively.
In the substrate (CH3-THF) complex, CH3-THF is located at the C-terminal edge of the TIM-barrel structure (Fig. 3a). While the glutamate moiety density of CH3-THF was not observed in the omit map, the rest of the CH3-THF was observed. The pterin ring of the CH3-THF is stacked on the isoalloxazine ring of FAD in a nearly parallel manner, with an average spacing of about 3.9 Å. The p-aminobenzoic acid (pABA) moiety of the CH3-THF is located in the cavity formed by Glu21, Thr23, Thr46, Leu48, Phe176, Phe215, Cys219, Tyr267, and Phe269 (Fig. 3b). Interestingly, the bound CH3-THF has a folded conformation (Fig. 3b), whereas CH3-THF bound to EcMTHFR adopts an extended conformation18.
Least-squares (LSQ) fitting of substrate-free and complex forms using Cα atoms (residues 7 to 288) yielded a root-mean-square deviation (rmsd) of 0.58 Å (Supplementary Table 2). Large differences were found at residues 210-224 (helix α8 to loop α8-α9) (Fig. 3c). Upon CH3-THF binding, helix α8 shifts approximately 1.5 Å towards the active site, and the side chains of Phe215 and Cys219, both located on helix α8, make van der Waals interactions with CH3-THF (Fig. 3b, c). Furthermore, the side chain of Phe215 forms a π‒π interaction with that of Phe269. The side chains of Phe215 and Phe269 interact with the bound CH3-THF, resulting in its folded conformation in S6MTHFR (Fig. 3b).
Structural comparison with EcMTHFR
Next, we analysed the structural differences between S6MTHFR and EcMTHFR. LSQ fitting of the CH3-THF complexes between S6MTHFR and EcMTHFR (PDB ID: 1ZP4) yielded an rmsd value of about 0.7 Å (58 Cα atoms in the eight parallel β-strands). The isoalloxazine ring of FAD and the pterin moiety of CH3-THF are located at nearly the same positions in both structures (Fig. 4a). Of the four catalytically essential residues of EcMTHFR (Glu28, Asp120, Gln183, and Phe2237,18,19,20,21), Glu28, Asp120, and Gln183 interact with the pterin moiety of CH3-THF. Their corresponding residues in S6MTHFR—Glu21, Asp107, and Gln175—showed nearly the same interactions with CH3-THF (Fig. 4a). However, the position of Phe215 in the S6MTHFR-CH3-THF complex is significantly different from that of Phe223, the corresponding residue of Phe215 in EcMTHFR (Fig. 4a).
a Comparison of the active site structures of the CH3-THF complex between S6MTHFR and EcMTHFR (PDB ID: 1ZP4). Cartoon models of S6MTHFR and EcMTHFR are shown in light green and cyan, respectively. Helix α8, loop β2-α3, and loop β8-α11 are highlighted in dark blue and green for EcMTHFR and S6MTHFR, respectively. CH3-THF in EcMTHFR and S6MTHFR is shown in cyan and yellow, respectively. S6 and Ec in parentheses represent residues of S6MTHFR and EcMTHFR, respectively. b CH3-THF oxidation activity of S6MTHFR and its mutants. Specific activities were calculated from the decreased amount of CH3-THF. Reactions were carried out in triplicate (n = 3). Dots show individual data points.
In EcMTHFR, the side chain of Phe223 underwent a large conformational change upon CH3-THF binding, resulting in the side chain of Phe223 interacting with the pterin moiety of CH3-THF. This interaction appears to contribute to the proper positioning of the pterin moiety for the catalytic reaction. Notably, the conformation of loop β8-α11 of EcMTHFR does not interfere with the accommodation of an extended conformer of CH3-THF (Fig. 4a). In S6MTHFR, Phe215, which corresponds to Phe223 of EcMTHFR, shifts by 1.5 Å and forms a π‒π interaction with Phe269 upon CH3-THF binding. Phe215 and Phe269 then make van der Waals interactions with the pABA moiety of CH3-THF, causing CH3-THF to adopt a folded conformation. Notably, CH3-THF in the extended conformation is unlikely to be accommodated in the active site of S6MTHFR due to a collision with loop β8-α11 (Fig. 4a).
S6MTHFR residues required for CH3-THF oxidation
To identify residues crucial for CH3-THF oxidation, we prepared some variants of S6MTHFR based on its crystal structures. Single amino acid substitutions of Cys219Ala and Phe269Gly reduced CH3-THF oxidation activity, whereas Pro49Ala, Leu48Gly-Pro49Gly, and Phe215Ala increased it (Fig. 4b). The negative and positive effects of the substitutions on CH3-THF oxidation can be explained by the crystal structures of S6MTHFR. Since Cys219 and Phe269 interact with CH3-THF, Cys219Ala, and Phe269Gly seem to change the shape of the active site and weaken the interaction with CH3-THF. On the other hand, substitutions of Pro49, Leu48, and Phe215 seem to reduce close contacts with the substrate. A combination of positive (Leu48Gly or Phe215Ala) and negative (Cys219Ala and Phe269Gly) mutations resulted in substantially reduced activity (Fig. 4b), suggesting the crucial roles of Cys219 and Phe269 in CH3-THF oxidation.
Creation of S6MTHFR variants with NADH oxidation activity
To further examine the difference between S6MTHFR and EcMTHFR, we analysed the interaction between S6MTHFR and NADH using the superimposed structures of S6MTHFR and the EcMTHFR‒NADH complex (PDB ID: 1ZPT). The complex’s crystal structure shows that NADH in EcMTHFR adopts a folded conformation18 (Fig. 5a). The superimposed structures suggested that S6MTHFR cannot accommodate the folded NADH due to multiple collisions with it. First, NADH’s adenine moiety seems to collide with the side chains of Phe215 and Phe269. The conformation of loop β8-α11 in S6MTHFR appears to cause the clash between Phe269 and NADH (Fig. 5a). In EcMTHFR, the corresponding loop is positioned away from the adenine ring, allowing it to accommodate NADH. Second, the nicotinamide mononucleotide moiety of NADH appears to collide with Leu48 and Pro49, presumably inhibiting NADH binding (Fig. 5a). Third, Cys219, located near the adenine ring of the NADH, also seems to clash with NADH (Fig. 5a). These structural features of S6MTHFR are likely to prevent NADH binding to the active site, thereby preventing the reduction of FAD by NADH.
a Superposition between the EcMTHFR‒NADH complex (cyan; PDB ID: 1ZPT) and S6MTHFR (green). The van der Waals surface of NADH is shown as a dot representation. b NADH oxidation activity of S6MTHFR and its mutants. LP: L48G-P49G variant, LPCF: L48G-P49G-C219A-F269G variant, LP-PFGGL: L48G-P49G-P268T-F269L-G270N-G271R-L272A variant. Specific activities were calculated from the decreased amount of NADH. c HPLC chromatograms (290 nm) of the reaction product by S6MTHFR (WT) and the LP-PFGGL variant in the presence or absence of NADH. A red arrow indicates CH3-THF. The retention times of NADH, CH2-THF derivative, CH3-THF, and CH2-THF were 0.65, 1.13, 1.50, and 2.11 min, respectively. The CH2-THF derivative seems to be THF, as methylenetetrahydrofolate is unstable and spontaneously decomposes to THF and formaldehyde44. d The ESI–MS spectrum (positive-ion mode) of CH3-THF produced from CH2-THF by the LP-PFGGL variant. The m/z-value for protonated molecular ions [M + H]+ of CH3-THF was 460. Reactions were carried out in triplicate (n = 3). Dots show individual data points.
With these results in hand, we then attempted to confer NADH oxidation activity on S6MTHFR by making amino acid substitutions. In the first variant, L48G-P49G (LP), the clashing amino acids were replaced with small side chains. This variant increased the specific activity for NADH oxidation approximately 38-fold compared to the wild type (Fig. 5b). The second variant, L48G-P49G-C219A-F269G (LPCF), exhibited no significant change in activity compared to the LP variant, highlighting the critical role of Leu48 and Pro49 in NADH binding. The third variant, L48G-P49G-P268T-F269L-G270N-G271R-L272A (LP-PFGGL), was designed to incorporate clash-avoiding substitutions at Leu48 and Pro49 along with five amino acid substitutions from loop β8-α11 to the respective residues of EcMTHFR. This design aimed to alter the conformation of loop β8-α11 to match that of EcMTHFR. The NADH oxidation activity of wild-type S6MTHFR did not saturate, even with NADH concentrations of up to 10 mM, and exhibited a turnover of 0.42 ± 0.02 s⁻¹. In contrast, the LP-PFGGL variant displayed significantly improved activity, with kcat and Km values for NADH oxidation of 21.4 ± 1.71 s⁻¹ and 2.96 ± 0.59 mM, respectively (Supplementary Fig. 6 and Supplementary Table 3). Moreover, this variant could convert CH2-THF into CH3-THF using electrons from NADH in the manner of typical MTHFRs (Fig. 5c,d). A structural change in loop β8-α11 of S6MTHFR may cause similar interactions not only with NADH but also with CH3-THF, as observed in typical MTHFRs.
S6MTHFR homologues form a distinct cluster in the MTHFR family
The amino acid sequence of S6MTHFR suggests that it has evolved from a typical MTHFR. However, S6MTHFR catalyses the reverse catalytic reaction of typical MTHFRs. This characteristic of S6MTHFR seems to be related to the unique auxotrophy of SYK-6. SYK-6 cannot utilise protocatechuate, a carbon source lacking methoxy groups, and instead needs a 1C unit from aromatic compounds containing a methoxy group, such as vanillate14,15. The methyl moiety of the methoxy group of vanillate is transferred to THF by LigM, producing CH3-THF. S6MTHFR then converts CH3-THF to CH2-THF (Fig. 1b). Due to the functional link between LigM and S6MTHFR, the metF (the S6MTHFR gene) and ligM genes are components of the ligM-metF-ligH operon23. However, it is unclear whether the combination of LigM and S6MTHFR is a unique example found only in SYK-6.
Previous studies revealed that Methylorubrum extorquens CM4 and Rhizorhabdus dicambivorans Ndbn-20 each have a THF-dependent demethylase: CmuAB24,25,26 (a different type of methyltransferase than LigM) and Dmt27, respectively. Since these enzymes produce CH3-THF, it is reasonable to consider that these bacteria have an S6MTHFR homologue. As expected, M. extorquens CM4 and R. dicambivorans Ndbn-20 have S6MTHFR homologues, designated as CM4MTHFR and N20MTHFR, respectively (Supplementary Fig. 5). Importantly, short amino acid fragments in loop β2-α3, helix α8, and loop β8-α11 of S6MTHFR, which are crucial for its unique enzymatic properties, are conserved in CM4MTHFR and N20MTHFR (Fig. 6a), suggesting that both likely have the same enzymatic properties as S6MTHFR. Indeed, N20MTHFR has been reported to exhibit CH3-THF oxidation activity and does not oxidise NADH28.
a Three conserved motifs of type 4 MTHFRs. Type 4 MTHFRs, including S6MTHFR, have conserved motifs at Loop β2-α3, Helix α8, and Loop β8-α11. These motifs play crucial roles in the oxidation of CH3-THF. The conserved residues are shown in red, orange, cyan, and green boxes. b The phylogenetic analysis of MTHFRs. The strain names corresponding to types 1, 2, 3, and 4 are shown in cyan, green, orange, and red, respectively. Circles: previously reported papers, blue star: structures already registered in the PDB, red star: structures published in this study. The MTHFRs are listed in Supplementary Data 4. c Overlay of structurally known MTHFRs and S6MTHFR. The LSQ fitting was performed using MTHFRs from the following species (PDB IDs are in parentheses): Saccharomyces cerevisiae (6FNU), Homo sapiens (6FCX), Thermus thermophilus HB8 (3APY), Haemophilus influenzae (5UME), Neisseria meningitidis (7RML), and E. coli (1ZP3). All MTHFRs except EcMTHFR are shown in white. EcMTHFR is in cyan. S6MTHFR in substrate-free and complex forms is shown in light green and light orange, respectively. Leu48, Arg78, Pro268, FAD, and CH3-THF in the S6MTHFR‒CH3-THF complex are shown in stick models.
On the basis of this result, we performed a database search for S6MTHFR homologues using a two-step search. First, we searched for bacteria with one or more THF-dependent demethylases similar to LigM, DesA, CmuAB, and Dmt, yielding more than 50 bacteria with their homologues. We then searched for S6MTHFR homologues in these bacteria and found 68 S6MTHFR homologues (Supplementary Data 1) that share the amino acid sequences in loop β2-α3, helix α8, and loop β8-α11 of S6MTHFR (Fig. 6a). The conservation of the three fragments suggests that they likely share with S6MTHFR the ability to catalyse CH3-THF oxidation.
We next performed phylogenetic analysis of MTHFRs. The obtained phylogenetic tree showed that MTHFRs can be categorised into four groups, labelled type 1 to type 4 (Fig. 6b). Type 1 MTHFRs consist of eukaryotic MTHFRs, which are represented by human MTHFR (HuMTHFR)29 and comprise the N-terminal catalytic and C-terminal regulatory domains. Type 2 MTHFRs include bacterial ordinary MTHFRs such as EcMTHFR, which are composed of only the catalytic domain22. Type 3 MTHFRs are heteromultimers with an electron transfer subunit containing an Fe-S-cluster30,31. While these three types of MTHFR have different domain and subunit compositions, they likely catalyse the reduction of CH2-THF. Indeed, they do not have the three S6MTHFR-specific motifs required for CH3-THF oxidation. In addition, all structurally known type 1 or 2 MTHFRs have structural features similar to those of EcMTHFR. First, the structure of a loop region corresponding to loop β8-α11 of S6MTHFR is similar to that of EcMTHFR (Fig. 6c). Second, these MTHFRs lack a protruding side chain that prevents NADH binding at the position of Leu48 (of S6MTHFR) (Fig. 6c). The residue corresponding to Leu48 is predominantly Gly. While HuMTHFR has a His residue at the corresponding position, the direction of the side chain is oriented away from the NADH binding site, allowing NADH binding.
On the other hand, type 4 MTHFRs, including S6MTHFR, catalyse CH3-THF oxidation due to the conservation of the three S6MTHFR-specific motifs in loop β2-α3, helix α8, and loop β8-α11. Type 4 MTHFRs constitute a distinct family of enzymes in combination with THF-dependent demethylases.
Discussion
Earlier reports showed that FAD in typical MTHFRs can be reduced by both NADH and CH3-THF; however, the reduction with the former is substantially faster than that with the latter7,18,32. Moreover, typically in 1C metabolism, CH₂-THF is synthesised before the MTHFR reaction occurs. Therefore, typical MTHFRs have been thought to receive a hydride ion from NADH and to convert CH2-THF to CH3-THF in cells. This is an essential reaction in the folate cycle of 1C metabolism1. In contrast, our studies suggested a unique enzymatic property of S6MTHFR. SYK-6 obtains a 1C unit from methoxylated aromatic compounds like vanillate and generates CH3-THF using a THF-dependent demethylase, LigM15,16. Earlier experimental results suggested that CH3-THF is converted into CH2-THF by S6MTHFR14. In this study, we demonstrated that S6MTHFR converts CH3-THF into CH2-THF and rarely reacts with NADH (Figs. 2b-d and 5b). Moreover, our structural analysis revealed that loop β2-α3, helix α8, and loop β8-α11 all play crucial roles in CH3-THF oxidation (Figs. 4a,5a, 6c).
Of the three fragments, loop β2-α3 and loop β8-α11 in S6MTHFR have different conformations from the corresponding loops of types 1 and 2 MTHFRs (Fig. 6c). This hinders the binding of NADH to the active site of S6MTHFR due to the collision with Leu48, Pro49, and Phe269. The structural comparison between S6MTHFR and other MTHFRs revealed that the conformational difference in loop β8-α11 occurs at the position of Pro268 (Fig. 6c), which is likely to change the conformation of the N-terminal part of loop β8-α11 of type 4 MTHFRs. While Pro268 is conserved in type 4 MTHFRs, its corresponding residue in types 1 and 2 MTHFRs is Thr (Fig. 6a and Supplementary Fig. 5). As described above, the substitution of five amino acids, including Pro268, significantly improved the NADH oxidation activity of S6MTHFR (Fig. 5b) and enabled the detectable generation of CH₃-THF (i.e., the forward reaction). This substitution appears to alter the loop conformation to be similar to the loop conformations found in types 1 and 2 MTHFRs.
Loop β2-α3 is also critical for inhibiting NADH binding to the active site. The side chain of Leu48 in S6MTHFR seems to collide with NADH. However, Leu48 is replaced by Gly in most types 1 and 2 MTHFRs, allowing NADH to be accommodated. The superposition of the structurally known types 1 and 2 MTHFRs reveals diversity in the conformation of this loop region (Fig. 6c). However, amino acid sequence analysis of type 4 MTHFRs suggests that this type has a conserved Leu residue in loop β2-α3 and inhibits NADH binding (Fig. 5a). Furthermore, the side chain conformation of Leu48 is constrained by van der Waals interactions with Pro49 and Arg78 and cannot move away from the NADH binding site. Since Arg78, Leu48, and Pro49 are conserved in type 4 MTHFRs, the conformation of Leu48 appears to be conserved in type 4 MTHFRs.
In addition to the NADH reactivity, the crystal structure of the S6MTHFR‒CH3-THF complex appears to explain the high CH3-THF oxidation reactivity of S6MTHFR relative to that of EcMTHFR. The superposition of S6MTHFR and EcMTHFR with folded and extended CH3-THF, respectively, shows that the distance between the C11 and N10 atoms in the folded CH3-THF is significantly shorter than in the extended form (Supplementary Fig. 7). Since the distance between the N10 and C11 atoms in S6MTHFR is 3.4 Å, it seems reasonable to predict that the lone pair of the N10 atom attacks the C11 atom of the pterin ring. The geometric arrangement of the lone pair of these atoms seems suitable for the catalytic reaction. On the other hand, the corresponding distance of the extended form of CH3-THF is rather long, 4.1 Å, for the catalytic reaction, and the direction of the lone pair of the N10 atom is inappropriate for the catalytic reaction.
Interestingly, since the conversion of CH2-THF to CH3-THF was not observed in S6MTHFR, the demethylation of the carbon source appears to be the sole pathway for CH3-THF generation. This suggests that the amount of synthesised CH3-THF reflects the environmental carbon source levels. Therefore, the amount of methionine seems to reflect the carbon source level. Since methionine is an essential amino acid for initiating protein synthesis as the first amino acid of proteins, the combination of S6MTHFR and demethylase may function as an environmental monitor to determine the growth of SYK-6 in relation to the cellular methionine level. In fact, the growth of SYK-6 on vanillate and syringate is markedly inhibited in the presence of excess methionine33. This observation indicates that the amount of methionine strictly regulates the catabolic system of lignin-derived aromatic compounds in SYK-6, which may support the above hypothesis.
In summary, we have elucidated the unique enzymatic properties of S6MTHFR. S6MTHFR favours the reverse reaction of the typical MTHFRs, converting CH3-THF to CH2-THF. The crystal structures of S6MTHFR revealed that structural differences in loop β2-α3, helix α8, and loop β8-α11 from those of EcMTHFR appear to facilitate CH3-THF oxidation. Additionally, the inhibition of NADH/NAD+ binding to S6MTHFR indicates the presence of alternative electron acceptors in SYK-6 cells. Since the conversion of CH2-THF to CH3-THF was not observed in S6MTHFR, the amount of methionine biosynthesis in SYK-6, which requires CH3-THF, is directly determined by the demethylation of the carbon source in the environment. A database search revealed many S6MTHFR homologues in bacteria, and these enzymes likely function with THF-dependent demethylase to transfer a 1C unit into the folate cycle for 1C metabolism. Our research has established a distinct family of MTHFRs that can function with a demethylase.
Methods
Plasmid construction
A double-stranded DNA encoding S6MTHFR (protein accession number BAK65950.1) of Sphingobium lignivorans SYK-6 (metF, SLG_12750) was synthesised (Eurofins) and amplified by PCR with a synthesised DNA fragment and the primer pairs (SYKmetF-5, 5ʹ-TATAGGATCCGCTACTGCAACGCTCGACAAG-3ʹ and SYKmetF-3, 5ʹ-TATAGTCGACTCAGATGCCCTGCTTCCCCTTG-3ʹ) using a KOD Plus kit (Toyobo, Osaka, Japan). The amplified DNA was inserted into the pET-44a(+) vector (Novagen, Pretoria, South Africa) between the BamHI and SalI sites using the Ligation High Ver.2 kit (Toyobo). We deleted the Nus-tag and inserted the HRV-3C cleavage site between the His-tag and the target gene sequence. The plasmid was cloned into E. coli DH5α. Point mutations of the S6MTHFR gene were introduced by the PCR method. The above plasmid was amplified with a set of outward-facing, overlapping primers containing nucleotide substitutions, then processed by DpnI and cloned into DH5α. All mutant gene sequences were verified. The primers and plasmids used in this study are shown in Supplementary Data 2 and 3. The pET22b-EcMTHFR plasmid, which contains the EcMTHFR (protein accession number AAC76923.1) gene with an HRV-3C cleavage site and His-tag, was synthesised by Eurofins.
Protein expression and purification of S6MTHFR
The above plasmid was transformed into E. coli BL21(DE3) (Novagen). To express recombinant proteins, the E. coli cells were grown in 1 L lysogeny broth (LB) at 37 °C, and isopropyl-D-1-thiogalactopyranoside (IPTG) was added at OD600 = 0.5 to a final concentration of 0.5 mM. After 20 h of incubation at 18 °C, the cells were harvested and resuspended in 40 ml of lysis buffer containing 50 mM Tris-HCl (pH 8.0), 300 mM NaCl, 1 mM dithiothreitol (DTT), and 10 mM imidazole (pH 8.0), and then disrupted by sonication using a UD-201 ultrasonic disruptor (Tomy, Tokyo, Japan). After removing the debris and insoluble fraction by centrifugation at 18,000 g for 30 min at 4 °C, the supernatants were collected and filtrated using a 0.22 μm filter (Millipore, Burlington, MA, USA). The filtrated supernatants were mixed with Ni-NTA resin (Qiagen, Hilden, Germany) and incubated on a rotary shaker for 2 h at 4 °C. They were then loaded onto an Econo-Column (Bio-Rad Laboratories, Hercules, CA, USA) filled with Ni-NTA resin and washed with wash buffer containing 50 mM Tris-HCl (pH 8.0), 25 mM NaCl, 1 mM DTT, and 30 mM imidazole (pH 8.0). The His-tag was cleaved on-column by adding PreScission protease (Cytiva, Marlborough, MA, USA) and incubated overnight at 4 °C, after which S6MTHFR was eluted by the wash buffer. Anion exchange chromatography (MonoQ 5/50 GL, Cytiva) was then performed on an Akta pure system (Cytiva). The MonoQ column was equilibrated with 50 mM Tris-HCl (pH 8.0), 25 mM NaCl, and 1 mM DTT (Buffer A) and eluted using a linear gradient of NaCl up to 1 M. The eluted fractions were collected and further purified by size-exclusion chromatography using a Superdex 200 increase 10/300 GL column (Cytiva) under the conditions of 20 mM HEPES (pH 7.5) and 100 mM NaCl. Finally, the purified proteins were concentrated by 30 K MWCO Amicon Ultra and Centricon centrifugal filter units (Millipore) and stored at −80 °C. The enzyme concentration was calculated using the molar extinction coefficient 14,300 at 450 nm, since FAD is a cofactor for MTHFR7. The purity of purified proteins was confirmed by SDS-PAGE (Supplementary Fig. 8).
Protein expression and purification of EcMTHFR
The pET22b-EcMTHFR plasmid was transformed into E. coli BL21(DE3) (Novagen). To express the recombinant protein, the E. coli cells were grown in 1 L lysogeny broth (LB) at 37 °C, and IPTG was added at OD600 = 0.6 to a final concentration of 0.5 mM. After overnight incubation at 18 °C, the cells were harvested and resuspended in 50 ml of lysis buffer containing 50 mM Tris-HCl (pH 7.4), 500 mM NaCl, 1 mM dithiothreitol (DTT), and 10 mM imidazole (pH 7.4), and then disrupted by sonication using a UD-201 ultrasonic disruptor (Tomy, Tokyo, Japan). The debris and insoluble fraction were removed by centrifugation at 18,000 g for 30 min at 4 °C. The supernatans was then collected and filtrated using a 0.22 μm filter (Millipore, Burlington, MA, USA). The filtered supernatant was mixed with Ni-NTA resin (Qiagen, Hilden, Germany) and incubated on a rotary shaker for 2 h at 4 °C. The supernatant was centrifuged at 20,000 rpm for 20 mins, resuspended with wash buffer [50 mM Tris-HCl (pH 7.4), 500 mM NaCl, 1 mM dithiothreitol (DTT), and 100 mM imidazole (pH 7.4)], and loaded onto an Econo-Column (Bio-Rad Laboratories, Hercules, CA, USA) filled with Ni-NTA resin. The resin was washed with the washed buffer five times. The His-tag was cleaved on-column by adding PreScission protease (Cytiva, Marlborough, MA, USA) and incubated overnight at 4 °C. After which Ec-MTHFR was eluted by elution buffer containing 50 mM Tris-HCl (pH 7.4), 500 mM NaCl, 1 mM dithiothreitol (DTT), and 300 mM imidazole (pH 7.4). The eluted fractions were collected and further purified by size-exclusion chromatography using a Superdex 200 increase 10/300 GL column (Cytiva) under the conditions of 50 mM Tris-HCl (pH 7.4), 100 mM NaCl and 1 mM DTT. Finally, the purified proteins were concentrated by 30 K MWCO Amicon Ultra and Centricon centrifugal filter units (Millipore) and stored at −80 °C. Enzyme concentration was calculated using the molar extinction coefficient 14,300 at 450 nm, since FAD is a cofactor for EcMTHFR. The purity of purified proteins was confirmed by SDS-PAGE.
Anaerobic titration
Experiments were conducted in an anaerobic chamber filled with a gas mixture (96% N2, 4% H2) to prevent the oxidation of reduced S6MTHFR34. S6MTHFR, CH3-THF (Sigma, St. Louis, MO, USA), and NAD+ (Millipore) were degassed in the chamber overnight before titration experiments. A 1.0 ml solution of 19 μM S6MTHFR containing 100 mM potassium phosphate (pH 7.2) was placed in a cuvette, and the enzyme solution was titrated with 0.5 ml of an 8, 10, or 50 mM solution of CH3-THF/NAD+ until saturation. After each addition of CH3-THF/NAD+, an absorbance spectrum between 300 and 800 nm was recorded by a DU730 spectrophotometer (Beckman Coulter, Brea, CA, USA) at room temperature.
Enzyme activity assay
The enzyme reactions of S6MTHFR and its mutants were carried out by incubating 20 nM purified enzymes with 100–1000 µM CH3-THF and 400 µM menadione in 50 mM KPO4 buffer (pH 7.2) for 3 min at 30 °C. The reaction was halted by mixing with an equal volume of 20% (v/v) formic acid. The supernatant separated by centrifugation (19,000 g, 10 min at 4 °C) was filtrated, and the CH3-THF concentration was measured by HPLC. Specific activities were calculated from the decreased amount of CH3-THF. To identify the reaction product, the reaction mixture was analysed directly by HPLC–MS without adding formic acid to prevent degradation of the reaction product due to the acidic pH. To determine the optimal pH, the reaction was performed with 50 mM GTA buffer (50 mM 3,3-dimethylglutarate, 50 mM Tris, and 50 mM 2-amino-2-methyl-1,3-propanediol; pH 5.0–9.0) at 30 °C. The optimal temperatures were determined in the range of 15–70 °C using 50 mM KPO4 buffer (pH 7.2) and 200 μM menadione. To determine the kinetic parameters for the conversion of CH3-THF by S6MTHFR, 5 nM and 20 nM of purified enzyme were used for 5–50 µM and 100–500 µM CH3-THF, respectively. For the reduction of CH2-THF, 500 nM of EcMTHFR, 20 nM or 5 µM of S6MTHFR and the LP-PFGGL mutant were incubated with 200 µM or 300 µM CH2-THF, along with 10 mM DTT and 2 mM NADH, in a 50 mM KPO4 buffer (pH 7.2) for 30 min at 30 °C. The reaction products were analysed by HPLC–MS. For NADH oxidation assays, 1 μM of S6MTHFR or 1 µM of each of its mutants was incubated with 0–10,000 µM NADH and 400 µM menadione in 50 mM KPO4 buffer (pH 7.2) at 30 °C. Reaction times were 30 min to determine specific activity and 1 min to determine the kinetic parameters. NADH oxidation was monitored by the decrease in absorbance at 343 nm (molar coefficient of NADH: ε343 = 6,220 M−1cm−1) using a BioTek Synergy LX plate reader (Agilent Technologies, Santa Clara, CA, USA)7. Specific activity was calculated from the decreased amount of NADH. Values of kcat and Km were determined using specific activities fitted to the Michaelis–Menten equation using GRAPHPAD PRISM software (version 10.1.2; GraphPad Software, La Jolla, CA, USA). The NADH oxidation assay was performed according to previously reported experimental conditions32. Reactions were performed in 50 mM KPO₄ buffer (pH 7.2) containing 0.3 mM EDTA, 10% glycerol, and 140 µM menadione. Each reaction mixture contained either 2.45 µM S6MTHFR or 30 nM EcMTHFR, with NADH concentrations ranging from 0 to 375 µM. Specific activity was calculated by monitoring the decrease in NADH absorbance at 343 nm32. An extinction coefficient of 6220 M−1 cm−1 was used for NADH.
HPLC and HPLC–MS analysis
HPLC analysis was performed with an ACQUITY UPLC system (Waters, Milford, MA, USA) and an ACQUITY UPLC I-Class system (Waters)35. All analyses were carried out using a BEH C18 column (2.1 × 100 mm, 1.7 µm particle size; Waters) and a mobile phase consisting of a mixture of solution A (acetonitrile containing 0.1% formic acid) and solution B (water containing 0.1% formic acid) at a flow rate of 0.5 mL/min and 30 °C under the following gradient conditions: 0–1.2 min, 6% solution A; 1.2–2.0 min, a linear gradient of 6% to 35% solution A; 2.0–3.0 min, 35% solution A; 3.0–3.1 min, a decreasing linear gradient of 35% to 6% solution A; 3.1–4.0 min, 6% solution A. The detection wavelength was 290 nm. HPLC–MS analysis was performed using the UPLC I-Class system coupled with an ACQUITY TQ detector (Waters). Mass spectra were obtained using the positive-ion mode with settings: capillary voltage, 3.0 kV; cone voltage, 10 to 40 V; source temperature, 120 °C; desolvation temperature, 350 °C; desolvation gas flow rate, 650 liter/h; and cone gas flow rate, 50 liter/h.35.
SEC-MALS analysis
Size-exclusion chromatography was performed with a Superdex 200 increase 10/300 GL column (Cytiva) using an Alliance 2695 system (Waters). Light scattering (LS) and the refractive index (RI) were measured using a DAWN HELEOS II detector (Wyatt Technology, Santa Barbara, CA, USA) and a 2414 RI detector (Waters), respectively. Before SEC-MALS analysis, the column was equilibrated at 20 °C with 10 mM HEPES-NaOH, pH 7.5, containing 100 mM NaCl. The purified S6MTHFR (5 mg/ml, 30 µl) was injected at a buffer flow rate of 0.5 ml/min. Data were processed with ASTRA 6.1 software (Wyatt Technology).
Crystallization
The initial crystallization screening was performed by using Crystal Screen 1 and 2 (Hampton Research, Aliso Viejo, CA, USA), Wizard Screens I and II (Rigaku, Tokyo, Japan), PEGsII (Qiagen), Index (Hampton Research), PEGIon/PEGIon2 (Hampton Research), and Protein Complex Suite (Qiagen) with the Protein Crystallization System 2 (PXS2) at the Structural Biology Research Center (KEK, Tsukuba, Japan)16. The sitting-drop vapour-diffusion method with crystallization drops consisting of 0.2 μl protein solutions (10 mg/ml) and 0.2 μl screening solutions at 20 °C and 4 °C was adopted. Twinned plate-shaped crystals were observed within a week under the conditions of 20% (w/v) PEG4000, 0.1 M sodium citrate at pH 4.5 (Protein Complex suite #26) at 20 °C. To obtain single crystals of S6MTHFR, the crystallization conditions were optimised with the hanging-drop vapour-diffusion method. Crystallization with microseeds gave single crystals of S6MTHFR using the same crystallization buffer at 4 °C. Crystals of the CH3-THF complex were prepared by the soaking method at 4 °C with soaking solution containing 10 mM CH3-THF, 22% (w/v) PEG4000, 0.1 M sodium citrate at pH 4.5. The soaking time was 1 h. Before diffraction data collection, all crystals were cryo-protected by soaking in a solution containing 30% (w/v) PEG4000, 0.1 M sodium citrate at pH 4.3 for 30 sec.
Data collection and structure determination
X-ray diffraction data of the substrate-free form of S6MTHFR were collected at 95 K using an Eiger X4M detector on BL-1A of the Photon Factory, KEK (Tsukuba, Japan). X-ray diffraction data of the S6MTHFR–CH3-THF complex were collected using a PILATUS 2M-F detector on X06DA (PXIII) at the Swiss Light Source (Villigen PSI, Switzerland). Diffraction data were processed and scaled by XDS and XSCALE, respectively36. The phases of the substrate-free S6MTHFR were determined using the programme Crank2 with the native SAD method37. The molecular replacement calculation by MOLREP was performed to obtain a higher resolution by using the initial model38. The crystal structure of the S6MTHFR–CH3-THF complex was determined by the molecular replacement method using the programme MOLREP. Crystallographic refinement and model building were performed using PHENIX.refine and Coot, respectively39,40.
Bioinformatic analysis
The MTHFR homologous sequences were collected using Protein BLAST and NCBI’s non-redundant protein database (https://www.ncbi.nlm.nih.gov/protein). Various bacteria and eukaryotes were searched for homologues of S6MTHFR, CM4MTHFR, N20MTHR, EcMTHFR, and HuMTHFR. A total of 60 MTHFR sequences were used for phylogenetic analysis (Supplementary Data 4). Substitution models were tested by MEGAX41, and the LG substitution model with “+G + I” was chosen as the best-fit model based on the Akaike information criterion42. The phylogenetic tree was reconstructed by the maximum likelihood method using RAxML-HPC2 on XSEDE under the LG + G model43, and bootstrap values were calculated from 1000 replications.
Statistics and reproducibility
The number of experimental replicates is indicated in the corresponding legend. Data points in each graph represent the means of the replicate measurements and individual data points are indicated by dots.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Any data generated or analysed during this study, associated protocols, primer sequences, plasmid maps, materials, and public databases (such as the PDB database) are included in the article and are also available from the corresponding authors upon request. Structural data from crystallographic analyses are available from the Protein Data Bank (https://www.rcsb.org/): the PDB codes for the crystal structures are 7XG9 (for S6MTHFR-Apo) and 7XLF (for S6MTHFR-Holo (CH3-THF)). The source data for the figures and tables in the main text are provided in Source Data 5, and those for the Supplementary Figs. and tables are available in Source Data 6, included with this paper.
References
Ducker, G. S. & Rabinowitz, J. D. One-carbon metabolism in health and disease. Cell Metab. 25, 27–42 (2017).
Kikuchi, G., Motokawa, Y., Yoshida, T. & Hiraga, K. Glycine cleavage system: reaction mechanism, physiological significance, and hyperglycinemia. Proc. Jpn. Acad. Ser. B 84, 246–263 (2008).
Locasale, J. W. Serine, glycine and one-carbon units: cancer metabolism in full circle. Nat. Rev. Cancer 13, 572–583 (2013).
Stover, P. & Schirch, V. Serine hydroxymethyltransferase catalyzes the hydrolysis of 5,10-methenyltetrahydrofolate to 5-formyltetrahydrofolate. J. Biol. Chem. 265, 14227–14233 (1990).
Sah, S. & Varshney, U. Methionyl-tRNA formyltransferase utilizes 10-formyldihydrofolate as an alternative substrate and impacts antifolate drug action. Microbiology 169, 001297 (2023).
Lu, S. C. S-Adenosylmethionine. Int. J. Biochem. Cell Biol. 32, 391–395 (2000).
Sheppard, C. A., Trimmer, E. E. & Matthews, R. G. Purification and Properties of NADH-dependent 5,10-methylenetetrahydrofolate reductase (MetF) from Escherichia coli. J. Bacteriol. 181, 718–725 (1999).
Igari, S. et al. Properties and crystal structure of methylenetetrahydrofolate reductase from Thermus thermophilus HB8. PLoS ONE 6, e23716 (2011).
Zhou, J., Kang, S. S., Wong, P. W., Fournier, B. & Rozen, R. Purification and characterization of methylenetetrahydrofolate reductase from human cadaver liver. Biochem. Med. Metab. Biol. 43, 234–242 (1990).
Matthews, R. G., Vanoni, M. A., Hainfeld, J. F. & Wall, J. Methylenetetrahydrofolate reductase. Evidence for spatially distinct subunit domains obtained by scanning transmission electron microscopy and limited proteolysis. J. Biol. Chem. 259, 11647–11650 (1984).
Bezerra, G. A. et al. Identification of small molecule allosteric modulators of 5,10-methylenetetrahydrofolate reductase (MTHFR) by targeting its unique regulatory domain. Biochimie 183, 100–107 (2021).
Masai, E., Katayama, Y. & Fukuda, M. Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. Biosci. Biotechnol. Biochem. 71, 1–15 (2007).
Kamimura, N. et al. Bacterial catabolism of lignin-derived aromatics: New findings in a recent decade: Update on bacterial lignin catabolism. Environ. Microbiol. Rep. 9, 679–705 (2017).
Masai, E. et al. A novel tetrahydrofolate-dependent O-demethylase gene Is essential for growth of Sphingomonas paucimobilis SYK-6 with syringate. J. Bacteriol. 186, 2757–2765 (2004).
Abe, T., Masai, E., Miyauchi, K., Katayama, Y. & Fukuda, M. A tetrahydrofolate-dependent O-demethylase, LigM, is crucial for catabolism of vanillate and syringate in Sphingomonas paucimobilis SYK-6. J. Bacteriol. 187, 2030–2037 (2005).
Harada, A. et al. The crystal structure of a new O-demethylase from Sphingobium sp. strain SYK-6. FEBS J. 284, 1855–1867 (2017).
Varman, A. M. et al. Decoding how a soil bacterium extracts building blocks and metabolic energy from ligninolysis provides road map for lignin valorization. Proc. Natl. Acad. Sci. 113, E5802–E5811 (2016).
Pejchal, R., Sargeant, R. & Ludwig, M. L. Structures of NADH and CH3-H4 folate complexes of Escherichia coli methylenetetrahydrofolate reductase reveal a spartan strategy for a ping-pong reaction. Biochemistry 44, 11447–11457 (2005).
Trimmer, E. E. et al. Aspartate 120 of Escherichia coli methylenetetrahydrofolate reductase: evidence for major roles in folate binding and catalysis and a minor role in flavin reactivityst. Biochemistry 44, 6809–6822 (2005).
Lee, M. N. et al. Functional role for the conformationally mobile phenylalanine 223 in the reaction of methylenetetrahydrofolate reductase from Escherichia coli. Biochemistry 48, 7673–7685 (2009).
Zuo, C. et al. A role for glutamine 183 in the folate oxidative half-reaction of methylenetetrahydrofolate reductase from Escherichia coli. Arch. Biochem. Biophys. 642, 63–74 (2018).
Matthews, R. G. et al. The structure and properties of methylenetetrahydrofolate reductase from Escherichia coli suggest how folate ameliorates human hyperhomocysteinemia. Nat. Struct. Biol. 6, 359–365 (1999).
Araki, T. et al. Regulation of vanillate and syringate catabolism by a MarR-type transcriptional regulator DesR in Sphingobium sp. SYK-6. Sci. Rep. 9, 18036 (2019).
Studer, A., Vuilleumier, S. & Leisinger, T. Properties of the methylcobalamin:H4 folate methyltransferase involved in chloromethane utilization by Methylobacterium sp. strain CM4. Eur. J. Biochem. 264, 242–249 (1999).
Vannelli, T., Messmer, M., Studer, A., Vuilleumier, S. & Leisinger, T. A corrinoid-dependent catabolic pathway for growth of a Methylobacterium strain with chloromethane. Proc. Natl. Acad. Sci. 96, 4615–4620 (1999).
Studer, A., McAnulla, C., Büchele, R., Leisinger, T. & Vuilleumier, S. Chloromethane-induced genes define a third C1 utilization pathway in Methylobacterium chloromethanicum CM4. J. Bacteriol. 184, 3476–3484 (2002).
Yao, L. et al. A tetrahydrofolate-dependent methyltransferase catalyzing the demethylation of dicamba in Sphingomonassp. strain Ndbn-20. Appl. Environ. Microbiol. 82, 5621–5630 (2016).
Yao, S. et al. The properties of 5-methyltetrahydrofolate dehydrogenase (MetF1) and its role in the tetrahydrofolate-dependent dicamba demethylation system in Rhizorhabdus dicambivorans Ndbn-20. J. Bacteriol. 201, e00096–19 (2019).
Froese, D. S. et al. Structural basis for the regulation of human 5,10-methylenetetrahydrofolate reductase by phosphorylation and S-adenosylmethionine inhibition. Nat. Commun. 9, 2261 (2018).
Bertsch, J., Öppinger, C., Hess, V., Langer, J. D. & Müller, V. Heterotrimeric NADH-oxidizing methylenetetrahydrofolate reductase from the acetogenic bacterium Acetobacterium woodii. J. Bacteriol. 197, 1681–1689 (2015).
Westphal, L., Wiechmann, A., Baker, J., Minton, N. P. & Müller, V. The Rnf complex is an energy-coupled transhydrogenase essential to reversibly link cellular NADH and ferredoxin pools in the acetogen Acetobacterium woodii. J. Bacteriol. 200, e00357–18 (2018).
Trimmer, E. E., Ballou, D. P. & Matthews, R. G. Methylenetetrahydrofolate reductase from Escherichia coli: elucidation of the kinetic mechanism by steady-state and rapid-reaction studies. Biochemistry 40, 6205–6215 (2001).
Bleem, A. et al. Multiplexed fitness profiling by RB-TnSeq elucidates pathways for lignin-related aromatic catabolism in Sphingobium sp. SYK-6. Cell Rep. 42, 112847 (2023).
Senda, M. & Senda, T. Anaerobic crystallization of proteins. Biophys. Rev. 10, 183–189 (2018).
Fukuhara, Y. et al. Characterization of the isophthalate degradation genes of Comamonas sp. strain E6. Appl. Environ. Microbiol. 76, 519–527 (2010).
Kabsch, W. XDS. Acta Crystallogr. D. Biol. Crystallogr. 66, 125–132 (2010).
Skubák, P. & Pannu, N. S. Automatic protein structure solution from weak X-ray data. Nat. Commun. 4, 2777 (2013).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D. Biol. Crystallogr. 68, 352–367 (2012).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Stecher, G., Tamura, K. & Kumar, S. Molecular evolutionary genetics analysis (MEGA) for macOS. Mol. Biol. Evol. 37, 1237–1239 (2020).
Akaike, H. Akaike’s information criterion. In International Encyclopedia of Statistical Science (ed. Lovric, M.) 25–25 https://doi.org/10.1007/978-3-642-04898-2_110 (Springer Berlin Heidelberg, Berlin, Heidelberg, 2011).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Osborn, M. J., Talbert, P. T. & Huennekens, F. M. The structure of “active formaldehyde” (N5, N10-methylene tetrahydrofolic acid). J. Am. Chem. Soc. 82, 4921–4927 (1960).
Acknowledgements
We thank the staff of the macromolecular X-ray crystallography beamlines of the Photon Factory at the Institute of Materials Structure Science of KEK for their technical assistance. We also thank Dr. Naoyuki Kuwabara for assistance with crystallization experiments, Dr. Kotaro Koiwai for helpful discussions, and Akane Saito for assistance with biochemical experiments. This research was supported by Research Support Project for Life Science and Drug Discovery (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED under grant number JP24ama121001(T.S.) and by JST grant number JPMJPF2104 (E.M.).
Author information
Authors and Affiliations
Contributions
HY.Y., E.M., and T.S. conceived the study. HY.Y., R.K., MJ. G., M.S. and N.K. performed the biochemical assays. HY.Y. and M.S. performed the crystallographic analyses. HY.Y. performed the bioinformatic analyses. HY.Y., N.K., E.M., and T.S. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Thomas McCorvie and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Laura Rodríguez Pérez.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yu, H., Kamimura, N., Kato, R. et al. Discovery of a distinct type of methylenetetrahydrofolate reductase family that couples with tetrahydrofolate-dependent demethylases. Commun Biol 8, 323 (2025). https://doi.org/10.1038/s42003-025-07762-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-07762-0