Abstract
Fish species of the family Scatophagidae possessing known candidate sex-determining genes (male-specific dmrt1Ys), offer suitable models for studying sex chromosome evolution. Here, we analyzed sex chromosome turnover events in three representative fish species of the family Scatophagidae, belonging to the genera Scatophagus and Selenotoca, which diverged 12.8 million years ago (Mya). Prior to the divergence of Sc. argus and Sc. tetracanthus 7.2 Mya, their dmrt1Y was differentiated from its locus, the truncated dmrt1ΔX, through allelic variation. The Y chromosome (Chr1) of Sc. tetracanthus is the result of the fusion of the original Y chromosome (Chr4) with an autosome (Chr13). The Se. multifasciata dmrt1Y arose from a duplication of dmrt1 on Chr4 and then translocated to the new Y chromosome (Chr18). The divergent evolutionary trajectories of the dmrt1Ys were accompanied by sex chromosome turnover in these three species. The sex chromosomes of the Scatophagidae family have evolved rapidly, albeit not randomly.
Similar content being viewed by others
Introduction
The evolution of sex chromosomes and the mechanisms of sex determination and differentiation are important issues for worldwide research in evolutionary biology1,2. Classical theory suggests that the sex chromosome originates from autosome(s)3. Initially, a new sex-determining (SD) gene emerges on an autosome, triggering its transformation into a proto-sex chromosome. Following this, sex-antagonistic genes or repetitive sequences progressively accumulate near the SD gene, inhibiting recombination of the sex chromosome and forming a differentiated sex-determining region (SDR). Through progressive evolutionary constraints, the homomorphic sex chromosomes undergo differentiation, ultimately resulting in the formation of heteromorphic sex chromosomes4. SD genes usually arise from those genes in the sex-determination and differentiation signaling network by gene duplication or allelic variation to gain SD function at the critical period of gonad development5,6. Regardless of how a new SD gene is generated, the sex chromosome sequence will change over time. Sex chromosome sequences change by various genetic mechanisms, including mutations7, rearrangements8, and gene loss9. These changes lead to recombination suppression and subsequently shape the evolution of sex chromosomes.
Most mammals have an XX-XY male heterogametic sex determination system, while most birds have a ZZ-ZW female heterogametic sex determination system, with their sex chromosomes showing a high degree of variation among different species1. Fish are the largest group of vertebrates with highly diverse reproductive strategies, including gonochorism, hermaphroditism, and unisexual gynogenesis10. The sex of fish is usually determined by genetic factors, environmental factors, or both11. Unlike mammals and birds, sex determination systems vary among fish species, even within closely related species12. Many fishes have relatively ‘young’ SD genes and sex chromosomes with low levels of sex chromosome differentiation13,14, making them real-time models for studying the early evolution of sex chromosomes in vertebrates.
The traditional view is that poorly differentiated sex chromosomes exist due to their youth, having had insufficient time to evolve, such as the 10 million year-old medaka Y chromosome15. In contrast, the ancient sex chromosomes of sturgeons, which evolved about 180 million years ago, have recently been found to be less differentiated16. Similarly, the bivalve sex chromosomes also have maintained a low level of differentiation for 350 million years. This may be related to the enrichment of sex-biased genes on their ancient homomorphic sex chromosomes17. The relationship between the degree of sex chromosome differentiation and the duration of formation is not yet fully understood. There is an urgent need for more comparative studies on the origin and differentiation of sex chromosomes in different species, and various fishes with diverse sex determination systems and genes provide excellent models.
The sex chromosome system can undergo changes during the evolution of a species or related species, which are thought to be associated with frequent sex chromosome turnover. Sex chromosome turnover is one possible outcome of sex chromosome evolution and involves the transformation of sex-determining mechanisms, potentially leading to the replacement of the original sex chromosomes with new ones, or the switching of roles between sex chromosomes and autosomes18,19. This phenomenon is relatively common in lower vertebrates20,21,22, including fish, reptiles, and amphibians, but it is rare in mammals and birds. Sex chromosome turnover can cause rapid changes in the sex chromosome(s), which has important biological significance in speciation23. For instance, stickleback fishes (family Gasterosteidae) exhibit remarkable diversity in sex determination among closely related species, which have experienced sex chromosome turnovers24, transitions between XY and ZW sex determination25, and fusions between sex chromosomes and autosomes26. Sex chromosome fusion and fission events are common methods of sex chromosome transformation, and they can then form a multiple-sex-chromosome system such as X1X1X2X2/X1X2Y, which has been reported in more than 30 fish species (16 cases of Perciformes, 9 cases of Cyprinidontiformes, 3 cases of Gymnotiformes, 2 cases of Gasterosteiformes, 2 cases of Tetraodontiformes, 1 cases of Siluriformes, etc.)27,28,29,30. In the earlier studies, there was mainly cytological evidence for the existence of such a multiple-sex-chromosome system, whereas current genomic technologies make it much easier to determine the origin and differentiation of multiple sex chromosomes in fishes. Recently, a sex chromosome fusion event in barred knifejaw (Oplegnathus fasciatus) was reported by sequencing both X1X1X2X2 female and X1X2Y male, revealing chromosome rearrangements in the fused Y chromosome31. Detailed comparisons of the fused chromosome and its precursors are therefore valuable for in-depth investigations into the roles of sex chromosomes in species differentiation and formation.
The family Scatophagidae belongs to the order Perciformes, and there are currently three reported species known to have evolved within 13 million years, all of which are either important aquaculture or aquarium fishes. Scatophagus (Sc.) argus (Linnaeus 1766) and Sc. tetracanthus (Lacepède 1802) belong to the same genus, while Selenotoca multifasciata (Richardson 1846) is the only reported relative in a different genus. The development of a sex-specific marker for Sc. argus revealed for the first time that Sc. argus has an XX-XY sex determination system32. Combined with genome and transcriptome sequencing, two copies of dmrt1s were identified in Sc. argus and Sc. tetracanthus, with one exhibiting a normal coding sequence (termed dmrt1Y/dmrt1) and the other a truncated coding sequence (termed dmrt1ΔX/ dmrt1b). dmrt1ΔX is present in both male and female individuals, whereas dmrt1Y appears only in male fish and is located on the Y chromosome with possible origin from allelic variation32,33. The male-specific dmrt1Y is a candidate SD gene in Sc. argus and Sc. tetracanthus, and the sex determination system of Se. multifasciata is also XX-XY type. Two previously identified dmrt1s in Se. multifasciata was assumed to be located on the X and Y chromosomes with a name of dmrt1X and dmrt1Y, respectively; dmrt1X and dmrt1Y encode 309 and 305 amino acids, respectively, with 85% sequence similarity34. In contrast to dmrt1ΔX in Sc. argus and Sc. tetracanthus, Se. multifasciata dmrt1X is normally coded with high expression in the adult testis, suggesting that it might still be functional. The male-specific dmrt1Y may determine sex in Se. multifasciata through its earlier expression than dmrt1X in the undifferentiated gonads34.
With a known sex determination system and candidate SD genes, the three representative species in the family Scatophagidae are good models for comparing the evolution of sex chromosome sequences within closely related species. The female genome of Sc. argus was previously reported by our group, and its X chromosome sequence was isolated with the sex-linked marker containing dmrt1ΔX35. Compared to the homologous X/Z chromosome, the Y/W chromosome sequences are more difficult to obtain due to their heterogeneity status in XY/ZW individuals. This situation would be harsher for assembling sex chromosome sequences in XY/ZW individuals when the sex chromosome remains young. At the early stages, partial Y chromosome sequences were determined mainly by constructing BAC or Fosmid libraries12,36,37. Fortunately, rapid advances in third-generation sequencing (TGS) technology have allowed researchers to better assemble sex chromosome(s) with even full length. In Northern pike (Esox lucius), researchers employed Nanopore technology to construct the genome of an XY individual and analyzed sequence differences between the undifferentiated X and Y chromosomes38. The male zigzag eel (Mastacembelus armatus) was sequenced using PacBio HiFi sequencing technology (TGS) and Hi-C assisted assembly method, and the haplotype sequence of the Y chromosome was obtained39. Here, we performed whole-genome sequencing of the three species including male Sc. argus, female and male Sc. tetracanthus and Se. multifasciata using TGS and Hi-C and then compared their sex chromosome sequences. Unexpectedly, dmrt1Y was found to originate from a gene duplication and translocation of the autosome dmrt1 (dmrt1a) in Se. multifasciata34. The dmrt1Ys in Sc. argus and Sc. tetracanthus were confirmed to originate from allelic variation. In this study, we showed for the first time that the Y chromosome in Sc. tetracanthus is derived from a fusion of the Y chromosome and an autosome, forming a multiple sex chromosome system (X1X1X2X2/X1X2Y). Our data support the evolutionary model of vertebrate sex chromosomes and provide a valuable genetic resource for in-depth comparative studies of sex determination and differentiation mechanisms within closely related species.
Results
Phenotypic and genotypic sex
A total of 186 specimens of Sc. argus (31 females and 31 males), Sc. tetracanthus (31 females and 31 males) and Se. multifasciata (31 females and 31 males) were collected for histological comparison (Supplementary Fig. 1A–C). Specimens containing oocytes were classified as females, while those with male germ cells (e.g. spermatozoa) were classified as males (Supplementary Fig. 1D, E). Thus far, no intersex individuals have been identified in any batch of the three fish species. The sex-linked markers previously developed by our research group, in conjunction with the findings of this study, have confirmed a 100% concordance between phenotypic and genotypic sexes in these examined individuals (Supplementary Fig. 1F–H).
Summary of chromosome-level genome assemblies of the three Scatophagidae species
In this study, the female genome of Sc. argus, previously sequenced using PacBio’s CLR technology in our previous study35, was employed for a comparative analysis. De novo genome assemblies were constructed for the three Scatophagidae species using long-read sequencing (PacBio’s CCS technology), yielding five primary genome assemblies (Table 1). The PacBio sequencing platform generated 39.3 ~ 55.8 Gb of raw data for each sample, as detailed in Supplementary table 1.
The six assembled genomes were subsequently scaffolded to a chromosome level using Hi-C reads. The resulting genetic maps revealed 24 chromosomes in both female and male Sc. argus and Se. multifasciata, consistent with the cytogenetic details reported by Suzuki et al.40. However, Sc. tetracanthus exhibited 24 chromosomes in female and 23 chromosomes in male (Supplementary Fig. 2 and Supplementary Table 2). Notably, the male Sc. tetracanthus genome contained an exceptionally large chromosome (52.59 Mb), approximately twice the size of other chromosomes, suggesting that certain chromosome fusion occurs in the male genome13. Moreover, the sex-linked markers of Sc. tetracanthus33 were localized to this large chromosome.
The BUSCO assessment revealed high completeness scores, ranging from 96.97% to 98.60% (Supplementary Table 3), indicating that all assemblies are high-fidelity and suitable for further comparative analysis. The number of predicted protein-coding genes in the three Scatophagidae species ranged from 21,512 to 24,256 (Table 1). The gene coding sequences and exon lengths were similar to those in other fishes’ genomes (Supplementary Fig. 3 and Supplementary Table 4). Additionally, repeat sequences account for 19.89% to 22.91% of these assembled genomes.
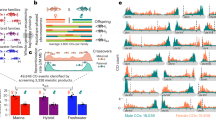
A total of 3016 single-copy orthologous genes were identified and used for phylogenetic analysis, encompassing the three Scatophagidae species and seven other fishes. Our results showed that the three Scatophagidae species diverged from Takifugu rubripes approximately 52.9 million years ago (Mya). Within the family of Scatophagidae, the genus Scatophagus diverged from the genus Selenotoca about 12.8 Mya. Furthermore, Sc. tetracanthus and Sc. argus were found to be closely related, with their divergence estimated at ~7.2 Mya (Fig. 1).
3016 single-copy orthologs were used for phylogenetic analysis by maximum likelihood. Female gene sequences of the three Scatophagidae species were used for this analysis. Blue numbers on the branches represent diverge times in Mya (million years ago). Red dots denote the fossil calibration nodes. Known sex determination genes or candidates were listed in the right panel for comparison. The red numbers mark the bootstrap support.
Haplotype-resolved genome assembly
Conventional diploid or polyploid assemblies disregards the distinctions between homologous chromosomes, while integrating the chimeric sequences of biparental allelic genomes. This presents a challenge when attempting to obtain Y or W chromosome sequences through the sequencing of XY or ZW individuals. To advance the study of sex chromosomes evolution, haplotype-resolved assembly techniques are essential39. In our present study, we utilized PacBio CCS technology to sequence the genomes of male individuals from the three Scatophagidae species, enabling haplotype-resolved assembly. The approach successfully generated two complete haploid genomes for each species, with genome sizes and scaffold N50 values comparable to those of the diploid assembly of both male and female (Table 2).
Following the generation of high-quality haploid genomes, sex-linked markers were used to distinguish X and Y haplotypes for each species. The presence of male-specific markers denote the Y haplotype genome (HapY) and vice versa as the X haplotype genome (HapX). It should be noted that the nomenclature of HapY and HapX is only applicable to the sex chromosomes in these haploid genomes. The number of chromosomes observed in the HapY of male Sc. argus and Se. multifasciata (n = 24) is consistent with those observed in their HapX of male and XX female genomes (n = 24). The chromosome lengths were found to be similar. Similarly, the HapX of the male and XX female genomes in Sc. tetracanthus comprises 24 chromosomes. Moreover, the length of the HapX chromosomes of Sc. tetracanthus was also comparable to that observed in Sc. argus and Se. multifasciata.
Consistent with the diploid genome of male Sc. tetracanthus, the HapY genome of the male consisted of 23 chromosomes, with one of the largest chromosomes being approximately twice the length of the others (Supplementary Table 2). For subsequent analyses, the male haploid genome (HapY) and the female XX genome were used as reference genomes.
Sex chromosome fusion in Sc. tetracanthus
The Hi-C data provide primary evidence for chromosome fusion in male Sc. tetracanthus. In the heatmap of interactions between 500 kbp genomic bins, there were 24 genomic blocks in female and male Sc. argus and Se. multifasciata, and female Sc. tetracanthus. However, the male genome of Sc. tetracanthus contained 23 genomic blocks, with the largest chromosome (Chr1 of the HapY genome) displaying strong interaction signals indicative of two merged genomic blocks (Fig. 2A, B). Furthermore, comparative synteny analysis between the HapY genome and the female XX genome revealed a predominantly one-to-one chromosomal correspondence in Sc. tetracanthus, with the exception of Chr4 and Chr13 (Fig. 2C). The Chr1 of the HapY genome exhibited strong conserved synteny with Chr4 and Chr13 of the female XX genome (Fig. 2D, E), supporting that Chr1 of the HapY genome probably originated from the fusion of the female Chr4 and Chr13.
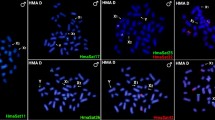
A The heatmap of interactions among haploid genome bins of 500 kbp along 23 chromosomes for male Sc. tetracanthus. B The heatmap of interactions among haploid genome bins of 500 kbp along Chr1 of male Sc. tetracanthus. C Comparative synteny analysis between the HapY genome and the female XX genome. D Genomic alignment of Chr1 from the HapY genome with Chr4/Chr13 from female genome in Sc. tetracanthus. E One-to-two synteny relationships between the chromosomal arms of Chr1 and Chr4/Chr13. F Karyotype of a female Sc. tetracanthus. G Karyotype of a male Sc. tetracanthus.
To further confirm the chromosome fusion in male Sc. tetracanthus, we performed additional karyotype analysis. The results validated that male Sc. tetracanthus contains 2n = 47 chromosomes (1m + 46a), whereas female possess 2n = 48 chromosomes (48a) (Fig. 2F, G). Comparative karyotypic examination in Sc. tetracanthus showed that males (n = 11) possess a largest metacentric chromosome, which is absent in females (n = 17) (Supplementary Fig. 4). According to karyotypes in other fish species with sex chromosome fusion, the distinctive largest metacentric chromosome in male Sc. tetracanthus should be the Y chromosome23,31,41. This observation was corroborated by in silico analysis, which precisely localized the Y-specific fragment to this metacentric chromosome. Moreover, homologous fragments of the sex-linked markers shared by male and female Sc. tetracanthus were situated on Chr4 of the XX and HapX genomes. This evidence suggests that Chr4 is the X1 chromosome, while Chr13 could be considered as the X2 chromosome. Taken together, this demonstrates the largest metacentric chromosome was formed by the fusion of the ancestral sex chromosome (Chr4) and an autosome (Chr13) during the evolution of Sc. tetracanthus, resulting in formation of the neo-Y chromosome (Chr1 of the HapY).
To identify the direction of fusion, the haploid genome of Sc. tetracanthus was searched for repetitive sequences to locate the centromeric and telomeric regions. Our results showed that there is considerable abundance of repetitive sequences present at the chromosomal termini. In particular, Chr1 of HapY exhibited a differential distribution, with repetitive sequences present at both ends and an additional cluster in the middle (Supplementary Fig. 5). Through annotation of satellite sequences within these repetitive sequences, several centromeric and telomere-associated repeat sequences were identified. The predicted centromeric repeat sequences were located at one end of the chromosomes, with the exception of Chr1, where they were centrally positioned. Similarly, the telomere-associated repeat sequences were restricted to one chromosomal end, except for Chr1, which contained these sequences at both termini (Fig. 2E). Combined with the synteny analysis, we propose that Chr4 (X1) and Chr13 (X2) underwent centromeric fusion, resulting in the formation of the metacentric chromosome (Chr1/ChrY) in male Sc. tetracanthus.
Characterization of sex chromosome differentiation in the three Scatophagidae species
Although we have identified sex-linked DNA markers and candidate SD genes in the three Scatophagidae species through bioinformatics analysis and PCR verification in our previous studies32,33,34,42, our understanding of their sex chromosomes remains in its infancy. For example, the sex chromosome fusion event in Sc. tetracanthus has only been revealed in our present study. Since we have detected our markers in XX and XY (X1X2Y) individuals, but not in YY individuals, it seems that the male-specific fragment was located on the Y chromosome. However, we are not sure that the ‘X’ specific fragment is actually located on the X chromosome.
In Sc. argus, both X- and Y-linked homologous sequences of the previous markers are located on Chr4 of female/HapX and HapY genome, corresponding to the X and Y chromosomes, respectively. In Sc. tetracanthus, the X- and Y-specific fragments of the sex-linked markers are located on Chr4 of the female/HapX genome and Chr1 of the HapY genome, respectively. Chr4 and Chr1 are homologous chromosomes, although they differ in length due to the chromosome fusion as mentioned above. Chr4 or its homologous chromosome serves as a shared sex chromosome in both Sc. argus and Sc. tetracanthus, indicating a similar origin of their sex chromosomes. The XX/XY sex determination system in Sc. argus could be transformed to X1X1X2X2/X1X2Y in Sc. tetracanthus by sex chromosome fusion. In Se. multifasciata, the Y-linked fragment of our previous sex-linked markers is located on Chr18 of the HapY, whereas the presumed ‘X’-specific fragment resides on Chr4 of the XX genome and the HapX genome of male fish34. This suggests that the previously characterized ‘X’-linked fragment was misassigned and should be reclassified as an autosomal fragment. This phenomenon may be due to the origin of the sex-specific fragment being duplicated from Chr4 and then translocated to the novel sex chromosome (Chr18).
Genome re-sequencing was performed to elucidate the sex-determining region (SDR) in the three Scatophagidae species. After filtering and quality control, 294.55-862.58 Gb of clean data were obtained in three Scatophagidae species. These clean data were subsequently mapped to the HapY genome and the female genome, with a mean mapping rate of 99.35–99.51% and a coverage depth of 7.35–39.87× (Supplementary Table 5). Based on the alignment, a total of 553,210 to 2,693,502 single-nucleotide polymorphisms (SNPs) were identified (Supplementary Data 1). The sex-specific SNPs that were heterozygous in males and homozygous in females were identified, confirming that these fish have an XX-XY or X1X1X2X2/X1X2Y system. In Sc. argus, a genomic region of approximately 0.70 Mb (from 25.39–26.09 Mb) on Chr4 of the HapY genome harbored significant SNPs associated with phenotypic sex (Supplementary Fig. 6A). The similar enrichment of SNPs was observed in the 25.04–25.72 Mb region (0.68 Mb) of Chr4 in the female genome (Supplementary Fig. 6B). Furthermore, this region showed a strong increase in the fixation index (FST) compared to other regions, and therefore it can be considered as a SDR (Supplementary Fig. 6C). In Sc. tetracanthus, the region with SNP genotypes showing 100% concordance with sex phenotypes spans from 16.79 to 31.31 Mb (14.52 Mb in length) on the Chr1 of the HapY genome. This 14.52 Mb region was here designated as a SDR in Sc. tetracanthus too (Supplementary Fig. 7, A and B). The similar enrichment of sex-linked SNPs was also observed in the 21.37-27.85 Mb region of Chr4 (X1) and 17.99–24.23 Mb region of Chr13 (X2) in the female genome (Supplementary Fig. 7C). In Se. multifasciata, a 2.94 Mb region (7.69–10.63 Mb) on Chr18 of the HapY was identified as being associated with sex and considered as its SDR (Fig. 3, A and B). A similar enrichment was observed in the 7.48–10.23 Mb region of Chr18 in the female genome (Supplementary Fig. 8). The size of the SDR in these three species elevates as follows: Sc. argus < Se. multifasciata < Sc. tetracanthus.
A Genome-wide association study for a sex determining locus based on 30 females and 30 males, where the genome-wide significant cutoff value after Bonferroni corrections is shown (red horizontal line). B Z-transformed FST scan for sex determining locus on Chr18. C Genomic comparisons between Chr18 from male and female genomes. D The upper panels display alignments of the PacBio reads from XX, and XY individuals and the Illumina reads from Female pool and Male pool to the HapY genome revealed a Y-specific insertion in the SDR which contained a male-specific duplicate of dmrt1 (named dmrt1Y). The lower panels display distribution of class I and class II transposable elements (TEs) near male-specific region.
To further demarcate the SDR, genetic mapping was performed for Sc. argus (Supplementary Table 6). Highly significant QTLs for sex trait (LOD > 5.6 at the whole genomic level) were mapped on LG7 corresponding to Chr4, spanning an interval of 1.45 cM (Supplementary Fig. 9A). Four sex-associated SNPs were found to be distributed in the 25.28–26.30 Mb (1.02 Mb) and 25.64–26.69 Mb (1.05 Mb) regions of Chr4 in the XX and HapY genome, respectively. 64.5% of the chromosome SDR obtained by re-sequencing overlapped with that determined by the QTL in Sc. argus (Supplementary Fig. 9B).
Sex chromosome alignment revealed sequence similarity of over 99% between the X/(X1 + X2) and Y chromosomes in the three Scatophagidae species (Supplementary Table 7). Further alignment of the SDRs revealed that their similarity was also greater than 98%, indicating limited sex chromosome differentiation in the three species (Supplementary Table 7). Notably, when comparing SNP density, we found that SDRs in both Sc. argus and Sc. tetracanthus exhibited significantly higher SNP densities compared to autosome. In contrast, the SNP density of SDRs in Se. multifasciata was not significantly different from that of autosome (Supplementary Fig. 10A). Furthermore, an examination of repetitive sequences within the genomes of Scatophagidae indicated that the SDRs exhibited a higher density of transposons relative to both the autosome and other segments of the sex chromosome (Supplementary Fig. 10B). Intriguingly, comparative genomic alignment of the SDR in Se. multifasciata exhibited intricate structural variance within a limited region (~0.15 Mb: 8.47–8.62 Mb), encompassing inversions and insertion/deletion (Fig. 3C). Based on the inverted sequence, our further analysis revealed that those genes on the X chromosome of Se. multifasciata were not collinear with Scatophagus genus autosomes on Chr18 (Supplementary Fig. 11).
Sequencing depth analysis of the sex chromosomes in Scatophagidae species revealed that the coverage of X and Y chromosome reads was essentially consistent and comparable to that of autosomal reads overall (Supplementary Fig. 12). Further detailed alignment of the reads to the SDR regions was performed to determine if there were any differences in the depth of coverage in these three fish species. Alignment of the PacBio HiFi reads from Se. multifasciata female and male individuals with the HapY genome revealed that a region from 8.59 to 8.62 Mb (30 kbp) on the SDR was mapped to reads from the male fish only. Consistently, the coverage of Illumina short reads around the 30 kbp fragment from pooled female samples is significantly lower than that of males (Fig. 3D). While the flanking regions of this 30 kbp fragment show no differences in coverage depth between the sexes, indicating that these regions are shared by X and Y chromosomes. These results suggest that this 30 kbp fragment should be specific to males. This male specific 30 kbp fragment is homologous to Chr4, indicating it may originate from duplication and translocation (Supplementary Fig. 13). Annotation results indicated that this 30 kbp duplicated fragment contained a single gene (from 8,590,644 bp to 8,602,581 bp), designated as dmrt1Y, which is a duplicate copy of the dmrt1 located on Chr4.
To validate the complex structural variations in the SDR of Se. multifasciata, particularly the male specificity of the gene dmrt1Y, a series of X- and Y-chromosome-specific primer pairs were designed at the boundary of the inversion and male-specific insertion (Supplementary Fig. 14A). Subsequently, seven female and seven male individuals were genotyped. Our results showed that the X-chromosome bands were present in all individuals, whereas the Y-chromosome bands were exclusively present in male individuals (Supplementary Fig. 14B). Moreover, one pairs of co-dominant sex-linked molecular markers were successfully obtained and validated in 60 fish (30 female and 30 male). These sex-linked markers developed for Se. multifasciata based on the third-generation sequencing should replace the markers developed from the next-generation sequencing in our previous study. This is because the previous ‘X’ specific fragment was existing in an autosome (Chr4) as mentioned above. This also indicate that the autosome dmrt1 linked with the ‘X’ specific fragment of our previous marker is wrongly defined as “dmrt1X” and should be corrected to dmrt1a. The qPCR results confirmed that the copy number of the X-specific fragment in XX individuals was almost twice that of XY individuals (Supplementary Fig. 14C). We can use the copy number difference of the X-specific fragment to discriminate the XX and XY individuals.
Identification of candidate SD genes in the three Scatophagidae species
A total of 21, 373, and 87 genes were identified within the SDRs of Sc. argus, Sc. tetracanthus and Se. multifasciata, respectively. Among them, the candidate SD gene identified in our previous study, dmrt1Y, was also located within these SDRs (Supplementary Data 2). The sex-linked markers of the three Scatophagidae species, all located within the dmrt1Y locus, support that dmrt1Y is the best candidate SD gene. The gonad transcriptome data of these Scatophagidae species revealed a significantly biased transcription profile for dmrt1Y between the ovary and testis (Fig. 4A–C). In addition, those genes closely surrounding the dmrt1Y also showed a male-biased expression pattern in the gonads (Supplementary Fig. 15), while only dmrt1Y showed the highest fold change and consistency across the three species.
The expression levels of those genes in the gonads were compared between females and males of Sc. argus (A), Sc. tetracanthus (B) and Se. multifasciata (C). D Venn diagram of significantly differentially expressed genes (DEGs) among three Scatophagidae species. DEGs correspond to log2(FoldChange) > 1 and p-value < 0.05.
Furthermore, the expression profile of SDR genes was examined in different tissues. We found that the dmrt1Y and its neighboring genes were transcribed higher in the testis than other tissues (Supplementary Fig. 16 and Supplementary Data 3). These results also suggest that dmrt1Y is an excellent candidate as a master SD gene in the three Scatophagidae species (Fig. 4D).
Comparative analyses of dmrt1s in the three Scatophagidae species
In order to ascertain whether dmrt1Ys of the three Scatophagidae species have a shared origin, we conducted a phylogenetic analysis. In the protein phylogeny, dmrt1s of Selenotoca were observed to cluster together at the root of the Scatophagus suborder, with significant support of high bootstrap values. Furthermore, Dmrt1Ys of Sc. argus and Sc. tetracanthus were identified to cluster together as a sister clade to their Dmrt1ΔX (Fig. 5B). These results indicate that dmrt1Y underwent independent lineage-specific duplication events between Scatophagus and Selenotoca species, with a common origin in Sc. argus and Sc. tetracanthus.
A Gene structure between dmrt1ΔX/dmrt1a and dmrt1Y. B Phylogenetic tree of dmrt1ΔX/dmrt1a and dmrt1Y proteins from different fishes. C Transposable element distribution and sequence identity of dmrt1 on X and Y chromosomes.
The previously cloned dmrt1a/dmrt1ΔX and dmrt1Y sequences32,33,34 were aligned with the assembled female and HapY genomes of the three Scatophagidae species using BLAST software to identify their exon and intron sequences. The dmrt1Ys of these Scatophagidae species exhibited a similar structure comprising five exons and four introns. Alignment of the dmrt1Y and dmrt1a/dmrt1ΔX genomic sequences demonstrated that the nucleotide identity between the exon sequences of these genes ranged from 75.8% to 98.7% (average: 87.5%; Fig. 5A). Conversely, the sequences and lengths of the introns presented considerable variation, as did the sequences in the upstream and downstream 5k bp regions (Fig. 5C).
A further analysis of repetitive sequences revealed accumulation of transposable elements (TEs) within the dmrt1Y of the three Scatophagidae species. These TEs identified within the dmrt1Y of Sc. argus, Sc. tetracanthus and Se. multifasciata were predominantly DNA transposons, long terminal repeat retrotransposons (LTRs), and long interspersed elements (LINEs) (Supplementary Data 4).
Discussion
HiFi and Hi-C are powerful technologies that can be utilized to sequence the sex chromosome in XY individuals39. Sex-linked markers are important for distinguishing the X and Y haplotpyes in XY individual sequencing. Alternatively, genomic information from parental fish is required to differentiate the X and Y haplotypes. Fortunately, in our previous research, we successfully developed sex-linked markers for the three Scatophagidae species32,33,34. The X and Y chromosomes assembled in this study are of high quality and will be valuable for future comparative studies. Our results confirmed that HiFi and Hi-C data, together with sex-linked markers, enable the direct assemble of the sex chromosomes in XY/ZW systems without the necessity of constructing Fosmid or BAC libraries. This approach also reduces the reliance on breeding YY or WW individuals, which have traditionally been used for sex chromosome sequencing in various fishes8,43,44. Re-sequencing of female and male individuals using NGS appears to be an effective strategy for identifying the SDR and even for pinpointing the candidate SD gene. In Sc. argus, the SDRs identified through re-sequencing and QTL mapping are of comparable length, both encompassing the candidate SD gene, dmrt1Y, which is male-specific. In addition to analyzing SNPs generated by NGS, coverage depth analysis has proven to be an efficient method for identifying the key SD region adjacent to the SD gene, as observed in Selenotoca multifasciata. Similar findings have been reported in golden pompano (Trachinotus ovatus) using coverage depth analysis45. Both NGS and TGS are powerful tools for studying sex chromosomes. In the future, the increasing application of these advanced sequencing approaches will contribute to a growing repository of fish sex chromosome sequences. This expanding dataset will enhance our understanding of the mechanisms underlying SD gene formation and differentiation. Furthermore, these data will provide empirical support for evolutionary theories related to sex chromosome evolution.
The expression profile of genes within the SDR supports our previous hypothesis that male-specific dmrt1Ys are candidate SD genes in the three Scatophagidae species, despite the notably low expression of dmrt1Y in ovarian (FPKM: 0-0.21). This low expression may attribute to the high similarity between the exon sequence of the dmrt1Y and dmrt1ΔX/dmrt1a, combined with the high sensitivity of the RNA-seq. Some dmrt1ΔX/dmrt1a reads in females may have been erroneously assigned to dmrt1Y when calculating their expression values. Therefore, the low expression level of dmrt1Y in females could be regarded as a false-positive signal and it does not impact its classification as a male-specific gene. One of the most important findings of this study is that the mechanisms underlying the formation of dmrt1Ys differ among species. Analysis of the X(X1X2) and Y chromosome sequences confirms that the dmrt1Ys in Sc. argus and Sc. tetracanthus originated through allelic variation. In Nile tilapia and zebrafish, mutations in dmrt1 can lead to male-to-female sex reversal, and the dmrt1-/- individuals cannot be rescued as males using aromatase inhibitors, indicating that dmrt1 is not only essential for male sex differentiation but also crucial for maintaining male sex in certain fish species46,47. The phenotypic effects of dmrt1-/- mimic the truncated mutant status of dmrt1ΔX in Sc. argus and Sc. tetracanthus. Consistently, treatment with an aromatase inhibitor also failed to induce female-to-male sex reversal in XX fish, suggesting that dmrt1ΔX is nonfunctional in Sc. argus32. In addition to being truncated and dysfunctional, the dmrt1ΔXs exhibit low expression level in the testes32,33. This reduced expression may result from mutations in critical promoter regions near the transcription start site (TSS) in both Sc. argus and Sc. tetracanthus (Supplementary Fig. 17). These findings suggest that the dmrt1ΔXs gradually become pseudogenes. To our knowledge, this represents the first reported case of XX individual lacking an intact dmrt1 gene in fish. In contrast, the dmrt1Y in Se. multifasciata originated through duplication and translocation. Similar to what have been observed in Japanese medaka, the autosomal (Chr9) dmrt1 was duplicated and translocated to a new chromosome, which subsequently evolved into the sex chromosome. In our previous study, we mistakenly concluded that Se. multifasciata dmrt1Y also arose from allelic variation and incorrectly defined autosomal dmrt1 as dmrt1X (dmrt1 on the X chromosome)34. The high-quality XX and XY genome assemblies in the present study allow us to correct this misclassification. According to our previous study, dmrt1Y is expressed at significantly lower levels than the autosomal dmrt1a in the testes of adult Se. multifasciata, suggesting that dmrt1Y primarily functions in sex determination, whereas dmrt1a plays a role in testicular maintenance34. Unlike Sc. argus, treatment with an aromatase inhibitor successfully induced female-to-male reversal in XX fish, indicating that dmrt1a should be functional in Se. multifasciata (Our unpublished data). This makes Se. multifasciata the second fish species in which dmrt1Y is responsible for sex determination while dmrt1a is involved in testicular maintenance, the first being Japanese medaka37. Further investigations are needed to elucidate the molecular mechanisms underlying these functional differences between dmrt1Y and dmrt1a.
dmrt1 and/or its duplicates are frequently selected as SD genes in diverse vertebrates. In the three species of the family Scatophagidae, dmrt1Y functions as their SD gene, providing further evidence for the “limited option” hypothesis48. According to the “limited option” hypothesis, certain genes are more likely to be reused or to evolve independently as SD genes. In Japanese medaka, the SD gene dmrt1by (or dmy) is a duplicate of the autosomal dmrt1, which is expressed earlier than dmrt1 in the male gonads and subsequently initiates male sex determination49. In African clawed frog (Xenopus laevis), a truncated duplicate of dmrt1, termed as dmW, determines the female sex by exerting a dominant-negative effects on the normal dmrt1 in ZW females50. In Chinese tongue sole (Cynoglossus semilaevis), Z-linked dmrt1 may determine the male sex via dose-dependent effects51. In the fighting fish (Betta splendens), a transposon insertion into the fourth intron of dmrt1 renders the gene susceptible to methylation modification, leading to its repression. This epigenetic regulation results in the differentiation of one homologous chromosome into the X chromosome and the other into the Y chromosome, with dmrt1 as the candidate SD gene52. Similarly, in Sc. argus, the promoter regions of dmrt1Y and dmrt1ΔX are hypomethylated and hypermethylated, respectively, which may contribute to their high and low expression levels in the testes53. The enrichment of TEs near the dmrt1Y and dmrt1ΔX may play a role in the allelic diversification of DNA epigenetic modification levels in the Sc. argus. The origin of dmrt1Y in Se. multifasciata is also likely to be mediated by TEs, similar to the duplication of dmy in Japanese medaka, where its expression is regulated by TEs within its promoter region54. However, the specific TEs involved in the dmrt1Y duplication and expression regulation require further confirmation in Se. multifasciata. Given the observed enrichment of TEs in dmrt1Y across the three Scatophagidae species, it is plausible that these elements contribute to the differentiation of dmrt1Y in this family.
The fish sex determination systems exhibit remarkable plasticity in the SD genes, as well as in structure of sex chromosomes. The turnover of sex chromosome within the family Scatophagidae provides an excellent example of sex chromosome evolution among closely related species, with a maximum divergence time of about 12.8 Mya. Chromosome fusion is one of the main routes of sex chromosome turnover55. Genome sequencing and karyotype analysis have revealed that the male Sc. tetracanthus has an X1X2Y system. Three potential mechanisms could account for the formation of this multiple sex chromosome system: (1) fusion of the Y chromosome with an autosome, (2) fission of the X chromosome in a species originally possessing an XX-XY system, or (3) reciprocal translocation between the X chromosome and an autosome in a species with an XX-XO system56. Previous studies suggest that X1X2Y systems in fish predominantly arise through chromosomal fusion rather than fission, resulting in the formation of large metacentric neo-Y chromosomes via Robertsonian fusion between the Y chromosome and an autosome31,56. In Sc. tetracanthus, the chromosome number in females (2n = 48) is identical to that of its closely related species, Sc. argus, whereas males possess one fewer chromosome (2n = 47), indicating that an independent chromosome fusion event has occurred. Comparative genomic analyses reveal that the large neo-Y chromosome (Chr1; 52.59 Mb) in Sc. tetracanthus males exhibits 99% sequence identity with the female chromosomes Chr4 (X1) and Chr13 (X2). Further chromosome comparisons show that Chr4 and Chr13 in Sc. tetracanthus females exhibit strong synteny with their counterparts in Sc. argus, suggesting that these chromosomes in Sc. argus correspond to the ancestral chromosomes that later fused in Sc. tetracanthus (Supplementary Fig. 18). Moreover, the ancestral chromosomes corresponding to Chr4 and Chr13 in unrelated species remain unlinked, confirming that this chromosomal fusion event is unique to Sc. tetracanthus (Supplementary Fig. 19). Based on genomic comparisons and karyotypic evidence, we propose that centric fusion between the Y chromosome (Chr4) and an autosome (Chr13) led to the emergence of the multiple X1X2Y sex determination system in male Sc. tetracanthus. Additionally, the sex-determining region (SDR) in Sc. tetracanthus is significantly larger than that in Sc. argus and Se. multifasciata. The fusion of a sex chromosome with an autosome is hypothesized to reduce recombination and accelerate the differentiation of sex chromosomes. Variations in SDR genotypes may play a critical role in the process of speciation.
The sex determination systems in the three Scatophagidae species can be categorized into two types based on their candidate SD genes. Type 1 corresponds to the sex determination system found in Sc. argus and Sc. tetracanthus, as their SD genes originated from allelic variation. However, they have also diverged due to a sex chromosome fusion event. Type 2 represents the sex determination system of Se. multifasciata, where its SD gene arose through a copy-and-paste mechanism. The first scientific question we face is determining which of the two systems is ancestral, or whether both evolved from other systems. One established fact is that a common allelic diversification process occurred on the sex chromosome of the ancestor shared by Sc. argus and Sc. tetracanthus. This suggests that the Type 1 system evolved in their common ancestor, meaning dmrt1Y and dmrt1ΔX have existed for more than 7.2 million years. Since dmrt1ΔX was not identified in Se. multifasciata, we infer that its truncation must have occurred after the divergence between the genera Scatophagus and Selenotoca, but before the split between Sc. argus and Sc. tetracanthus, approximately between 7.2 and 12.8 Mya. Alternatively, dmrt1ΔX may have been present in the common ancestor of both Scatophagus and Selenotoca, but was gradually lost during the evolution of Selenotoca, along with its allele dmrt1Y losing its SD function. If this hypothesis is correct, dmrt1Y and dmrt1ΔX have been present in the genus Scatophagus for more than 12.8 million years, while the duplication of dmrt1Y in Selenotoca occurred independently within the last 12.8 million years. The low SNP density in the SDR region of Se. multifasciata supports this hypothesis, suggesting that the duplication of dmrt1Y may have occurred more recently than the allelic variation of dmrt1 in Scatophagus (Supplementary Fig. 10). Nevertheless, we cannot entirely rule out the possibility that the ancestral sex determination system was Type 2 or belonged to an entirely different type. Furthermore, in more distantly related species within Perciformes, dmrt1 is also sex-linked in the large yellow croaker (Larimichthys crocea)57 and the yellow drum (Nibea albiflora)58 of the family Sciaenidae. Investigating whether dmrt1 has evolved through a shared evolutionary origin, independently, or via convergent evolution as a candidate SD gene in these species would be of significant interest. The evolution of the sex chromosome is driven by evolutionary forces such as sexual conflict and sexual selection59. Traditionally, it has been assumed that the degree of sex chromosome differentiation is time-dependent, with younger sex chromosomes being homomorphic and older ones being heteromorphic. It is therefore not surprising to identify the poorly differentiated sex chromosomes that have persisted for 30 million years in Takifugu7 and 40 million years in the Northern pike (Esox lucius)22. However, this assumption has been challenged by discoveries in sturgeons16 and bivalve species17, which suggest that even ancient sex chromosomes may exhibit low differentiation. Sex chromosome decay may not be the only evolutionary trajectory. In the three Scatophagidae species, sequence variation between X and Y chromosomes is low, which could be attributed to the relatively young age of these sex chromosomes. The rapid turnover of sex chromosomes in these species may act as an evolutionary force that maintains a relatively low level of differentiation. Conversely, it has been proposed that the accumulation of repetitive DNA on sex chromosomes contributes to the suppression of recombination. The suppressive effects of repetitive DNA have been reported in several species across different taxa, including vertebrates60,61, worms62 and plants63. Gene duplication and translocation mediated by TEs could also indirectly influence sex chromosome evolution. In this study, the inversion of the X chromosome in Se. multifasciata may have resulted from the insertion of dmrt1Y on the Y chromosome, leading to the rearrangement of neighboring genes and ultimately causing recombination suppression between X and Y chromosomes. These findings suggest that TEs have played a pivotal role in the evolution of sex chromosomes within the Scatophagidae family. In the future, comparative analyses of sex chromosome sequences across a broader range of vertebrate species could provide deeper insights into the fundamental principles governing sex chromosome differentiation.
In summary, the sex chromosomes within closely related species within the family Scatophagidae had undergone rapid but not random turnover events, as they still share some common features, such as the conserved SD genes, dmrt1Ys. The three Scatophagidae species with different sex chromosome systems provide excellent models for comparative studies of sex chromosome evolution between closely related species.
Methods
Ethics statement
All animal procedures and experiments were approved by the Guangdong Ocean University Animal Care and Use Committee (no. 201903004). We have complied with all relevant ethical regulations for animal use.
Sample collection, library construction and genome sequencing
For whole genome sequencing, one adult female and one adult male Sc. argus, Sc. tetracanthus and Se. multifasciata were collected from the Donghai Island Cultivation Base of Guangdong Ocean University (Zhanjiang, China), Liwan Ornamental Fish Market (Guangzhou, China), and Xiashan Market for Aquatic Products (Zhanjiang, China), respectively. The Sc. argus specimens were originally purchased from the Yucheng Fish Fry Culture Farm in Zhuhai (Guangdong, China), and transferred to the Donghai Island Cultivation Base of Guangdong Ocean University. The Sc. tetracanthus individuals were originally imported from South Africa (Supplementary Fig. 20) under a CITES Export Permit and China’s Biosafety Import Permit for Live Animals and Plants, with transport in accordance with the IATA Live Animals Regulations and veterinary health certification. In addition, 180 individuals (30 females and 30 males of Sc. argus, 30 females and 30 males of Se. multifasciata, 30 females and 30 males of Sc. tetracanthus) were also sampled for whole-genome re-sequencing. These Sc. argus and Se. multifasciata samples for whole-genome re-sequencing were obtained from the commercial fish market (Xiashan Market for Aquatic Products), whereas Sc. tetracanthus specimens were obtained from the ornamental fish market (Liwan Ornamental Fish Market).
Fish individuals were anaesthetized using 1% eugenol (E809010, Macklin Biochemical, Shanghai, China) before dissection. Their muscle tissues were collected and then immediately frozen in liquid nitrogen for storage at −80°C until use (for extraction of genomic DNA). Brain, heart, liver, spleen, kidney, muscle, and gonad tissues from the same fish were harvested for transcriptome sequencing.
Phenotypic sex identification
The phenotypic sex of all experimental fish were determined by histological observation of their gonads. Histological sectioning and staining were performed as reported in our previous study64. In brief, the collected gonads were fixed in Bouin’s solution overnight, dehydrated in alcohol, immersed in xylene for transparency, and then embedded in paraffin. Paraffin blocks were sectioned at 5-μm thickness and stained with hematoxylin and eosin (HE). A Nikon Eclipse Ti-E microscope (Tokyo, Japan) was used to capture images of the stained sections.
Genotypic sex identification
Sex linked markers were developed for Sc. argus, Sc. tetracanthus, and Se. multifasciata in our previous reports32,33,34. Genotypic sex was confirmed using these sex-linked markers for all the experimental fish in our present study. Briefly, a small piece of muscle was collected from each fish and stored in ethanol at −20°C for DNA extraction. Genomic DNA (gDNA), extracted using a DNA extraction kit (Cat#N1173, Dongsheng Biotecon, Guangzhou, China), was used for sex validation by PCR amplification. Sequences of each primer pair are provided (Supplementary Table 8).
Genome sequencing and assembly
High quality gDNA was extracted from muscle tissues using a nucleic acid purification kit according to the manufacturer’s instructions (Dongsheng Biotecon). Subsequently, it was used for MGI and PacBio HiFi sequencing. A library with an insert size of ~350 bp was prepared using the VAHTS Universal DNA Library Prep Kit from MGI (Shenzhen, China) and sequenced on a MGISEQ-2000 platform at Frasergen Bioinformatics Co. Ltd. (Wuhan, China). Raw sequencing reads were filtered using the Trimmomatic program v0.3365 to remove adapter sequences and low-quality reads. Clean reads were used to estimate the genome size and heterozygosity by a k-mer analysis (17-mer frequency distribution) using Jellyfish v2.066.
PacBio HiFi sequencing libraries were generated with an average insert size of ~15k bp, and sequenced on a PacBio Sequel II Sequencing System in Annoroad (Wuhan, China) using the CCS (circular consensus sequencing) algorithm. Raw sequencing reads were filtered using CCS 5.00 software (https://github.com/PacificBiosciences/ccs). HiFi clean reads were assembled using Hifiasm v0.1267 with the “-l 0” option to disable the purge duplication step of the software default settings, followed by a round of duplicate purging using Purge_Dups v1.2568. Genome completeness was estimated using the Benchmarking Universal Single-Copy Orthologs (BUSCO) v.5.2.269, based on orthologs derived from the Actinopterygii lineage.
Hi-C sequencing and chromosome assembly
Hi-C sequencing was performed to support the genome assembly at a chromosomal level. To construct the Hi-C library, each muscle sample was fixed in 1% formaldehyde for 10 min at room temperature to induce crosslinking. The reaction was quenched by the addition of 2.5 M glycine for 5 min. Nuclei DNA were digested with Mbo I, labeled with biotin-14-dCTP (Invitrogen, USA) and ligated with T4 DNA ligase. After an overnight incubating to reverse crosslink, the ligated DNA was sheared into 300–600 bp fragments. These DNA fragments were blunt-end repaired and A-tailed, followed by purification by biotin-streptomycin-mediated pulldown. The Hi-C libraries were then quantified and sequenced on an Illumina HiSeq X Ten platform (Illumina, USA) using a PE-150 module. Raw reads were filtered using the HTQC package70 to avoid the potential influence of low-quality reads in subsequent analyses.
Hi-C reads were aligned to the assembled contigs using Bowtie2 v2.3.271 with the end-to-end model (-very-sensitive -L 30). Only read pairs where both ends were uniquely aligned to the assembled contigs were selected. The number of Hi-C contacts between each contig was then calculated and normalized by standardizing the digestion sites of Mbo I in the genome sketch. Lachesis72 was then applied to cluster, order, and orient the contigs. Finally, each candidate assembly was manually verified using the Juicebox assembly tools73.
Genome re-sequencing
Genome re-sequencing was performed to compare the genomic differences between female and male. The gDNA from 180 tails of Sc. argus (30 females and 30 males), Sc. tetracanthus (30 females and 30 males) and Se. multifasciata (30 females and 30 males) was extracted for genome re-sequencing. Sequencing libraries were prepared using an Illumina TruSeq Nano DNA HT Library Prep Kit (Illumina) according to the manufacturer’s instructions. Library quality was assessed using a Fragment Analyzer (Advanced Analytical Technologies, Inc., USA). Libraries were quantified by qPCR using a Kapa Library Quantification Kit (KAPA Biosystems, USA). Sequencing was fulfilled on an Illumina HiSeq X platform using a paired-end read length of 2 × 150 nt with HiSeq X Reagent Kits (Illumina).
Transcriptome sequencing
To annotate protein-coding genes, transcriptome sequencing of multiple tissues was performed for those fishes used for whole genome sequencing. Total RNA was extracted from various tissues (including brain, heart, liver, spleen, kidney, muscle, and gonad) using TRIzol Reagent (Invitrogen, USA) according to the manufacturer’s instructions. RNA purity, degradation/contamination level, concentration, and integrity were measured using NanoDrop One (Thermo Fisher Scientific, USA), 1% agarose gels, Qubit RNA Assay Kit with a Qubit 3.0 Fluorometer (Life Technologies, USA), and RNA Nano 6000 Assay Kit with a Bioanalyzer 2100 system (Agilent Technologies, USA), respectively. The validated RNA samples were used for Illumina library construction. cDNA libraries were sequenced on an Illumina HiSeq 2000 system with a paired-end read length of 150 bp.
Annotation of repetitive elements
Tandem repeats (TRs) were analyzed using TRF v4.07b74 and GMATA v2.275. For transposable elements (TEs), the RepeatModeler v1.0.11, LTR_FINDER76, LTRharvest77, and MITE-Hunter78 were employed to construct a TE sequence library based on the principle of de novo methods and TE-specific architecture. These data were then combined with RepBase79. Finally, RepeatMasker v1.33180 was applied to predict TEs according to the final constructed TE database.
Gene prediction and function annotation
Putative ab initio, homology, and transcriptome-based methods were used to predict protein-coding genes. For the ab initio gene prediction, repeat regions in the assembled genome were first masked based on the result of the repeat sequence annotation, and then Augustus v3.2.281, which was trained to model parameters from homologous genes of BUSCO, was employed to generate gene structures. Homology-based prediction was performed using GeMoMa v1.682. Protein sequences of zebrafish (Danio rerio, GCA_008692375.1), large yellow croaker (Larimichthys crocea, GCF_000972845.3), Nile tilapia (Oreochromis niloticus, GCA_001858045.3), and Japanese medaka (Oryzias latipes, GCA_004347445.1) were downloaded from the NCBI database. For the transcriptome-assisted prediction, clean transcriptome (RNA-seq) data from the seven tissues were aligned to the assembled genome using HISAT2 v2.1.083 and assembled using StringTie v1.3.684. These predicted transcripts were then processed using PASA85. Finally, EVidenceModeler v1.186 was applied to integrate the above three respective annotation files to generate a consensus gene set. The TransposonPSI v08222010 (http://transposonpsi.sourceforge.net) alignment was conducted to remove sequences containing transposons to obtain the final results.
Functional annotation of the predicted genes was performed by alignment to the public SwissProt, NR, KOG, KEGG, and GO databases using Blastall and KAAS v2.187. These results were integrated to generate a final gene set.
Phylogenetic tree construction and divergence time estimation
To identify gene families in the assembled genomes of the three representative Scatophagidae fishes, the genome data of yellowfin seabream (Acanthopagrus latus, GCF_904848185.1), Japanese pufferfish (Takifugu rubripes, GCF_901000725.2), Nile tilapia (GCF_001858045.3), large yellow croaker (GCF_000972845.3), big head croaker (Collichthys lucidus, GCF_000002035.6), Japanese medaka (GCA_004347445.1) and coelacanth (Latimeria chalumnae, GCF_000225785.1) were downloaded from the NCBI. The OrthoMCL v2.0.9 pipeline88 was employed to identify gene families in the genomes of these species. Phylogenetic relationships between the three Scatophagidae species and other fish were established using the common single-copy orthologs. The protein sequences of these single-copy orthologs were concatenated into a supermatrix, which was aligned using MAFFT v7.46389, and phylogenetic analyses were conducted using the maximum likelihood method implemented RAxML v8.2.1163790, employing the GTR + GAMMA model and 100 bootstrap replicates. Based on the phylogenetic tree, the MCMCTREE software in the PAML v4.991 was applied to estimate the divergence times, using divergence times from the TimeTree database (http://timetree.org/) for calibration92.
Identification of the sex-linked region (SLR)
To identify the sex-linked regions of Scatophagidae species, genome-wide association analysis was conducted and the re-sequencing reads from both females and males were pooled separately and mapped to the assembled female and male genomes using BWA-mem v0.7.12 with default parameters93. After removing the duplicates generated from the PCR-based library construction using Picard v.2.18.2 (http://broadinstitute.github.io/picard/), single-nucleotide polymorphic (SNP) sites were called using the GATK v4.1.4.0 joint calling pipeline. The parameters used to filter the variants were “QD < 2.0 | | FS > 60.0 || MQRankSum < -12.5 || RedPosRankSum < -8.0 | | SOR > 3.0 | | MQ < 40.0”. The SNPeff v4.3t94 was employed to select those SNPs that are heterozygous in one sex but homozygous in the other sex. The EMMAX v8.2295 was applied to identify the SNPs that are potentially associated with sex. In addition, read mapping, sequencing depth, and coverage analysis of the TGS and pool NGS data were performed. Briefly, female and male HiFi reads were mapped to the assembled female and male genomes using Minimap2 v2.17-r94196 with default parameters. SAMtools v1.997 was employed to calculate the sequencing depth and coverage information for each mapping. The average coverage depth was extracted using a 10k bp non-overlapping sliding window. Any possible sex-specific region was identified according to the differences between these mappings. Integrative Genomics Viewer v2.5.298 was applied to search for sequence differences between males and females.
Furthermore, based on the constructed high-density SNP genetic linkage map of Sc. argus using two parents and their 200 offspring99, the Kruskal-Wallis test and interval mapping (IM) method in MapQTL v5.0 software were used to locate quantitative trait locus (QTL) for sex phenotypic traits in the mapping population.
Comparative genome analyses
Whole-genome alignment between the female and male Scatophagidae genome assemblies was performed using LAST v1452100. The X chromosome sequences were aligned to the Y chromosome sequences using Winnowmap2 with parameters: -c -x asm20 -eqx. The genomes of the three Scatophagidae species were employed to perform a chromosome collinear analysis using JCVI (MCscan Python version)101. The genome sequences of the three Scatophagidae species within the SLR were aligned using Progressive Cactus v2.7.1102. Subsequently, halSynteny v2.2103 was employed to identify synteny blocks among the aligned genomes, with a minimum block size of 1000 bp. Further, to obtain a higher resolution view of synteny and to check the identity between the three SLR regions, we then applied wfmash v0.15.0 (https://github.com/waveygang/wfmash) with parameters -p 50 -n 1 -4 to perform pairwise alignments.
The following procedures were conducted to measure sequence identity: the makewindows function of BEDTools v2.31.0104 was used to create 10k bp windows on the reference genome. Subsequently, the liftover function of the Rustybam tool v0.1.31 (https://github.com/mrvollger/rustybam) was employed to map intervals, while the stats function of rustybam was used to generate statistical data from the alignment. Sequence identity was calculated by dividing the number of matches by the sum of matches and mismatches.
Identification of putative centromeric and telomere-associated repeats
In order to identify satellite DNA sequences of centromeres based on the annotation results of genome tandem repeat sequences, the proportion of various types of satellite sequences in the assembled genomes was calculated and their distribution in the genomes was plotted. The region with the richest satellite sequences in each chromosome was defined as the putative centromere region39,105. Telomeric repeats usually were (TTAGGG)n, and based on this pattern their distribution in the genome was plotted.
Reconstruction of ancestral chromosomes
Protein sets of the S. argus, S. multifasciata and S. tetracanthus were aligned to the protein set of previously predicted ancestor chromosomes106, by employing BLASTp (e-value < 1 × 10−10). The reciprocal BLASTp best-hits between each selected species and the ancestor chromosomes were generated. Finally, chromosomal fissions and fusions were determined by using SVG in Perl.
Chromosome preparation, microscopy and image processing
Mitotic chromosomes were obtained from head kidney cells according to the protocol described by Bertollo et al.107. For conventional cytogenetic analysis, chromosomes were stained with 5% Giemsa solution (pH 6.8; Solarbio, China). At least 30 metaphase spreads per individual were checked to confirm the diploid number (2n) and karyotype structure. Images were captured using a Nikon Eclipse Ti-E microscope, and the images were processed using Image Pro Plus v4.1 software (Media Cybernetics, USA). The final images were optimized and arranged using Adobe Photoshop v7.0.
Statistics and Reproducibility
Statistical analysis of the data was performed using the R programming language v4.0.4 software. All values are expressed as mean ± standard deviation (SD). Statistical significance was analyzed using Student’s t-test, Bonferroni or Benjamini-Hochberg test. In the text and figures, significance levels are indicated by exact p values or asterisks (*p < 0.05; **p < 0.01). Sample sizes reported in the text were selected to satisfy statistical power, experimental robustness and availability. At least 3 biological replicates were used in an experiment.
Data availability
The data of final genome assembly and annotation in this study have been deposited in the figshare database (https://doi.org/10.6084/m9.figshare.28706396). Additionally, raw reads obtained by Illumina, PacBio, and Hi-C reads are available in the NCBI database under accession numbers PRJNA637812, PRJNA1243538, and PRJNA825131. The uncropped and unedited images for the gels in this study are shown in the Supplementary Figs. 21–34. The female Scatophagus argus genome in our previous study was deposited at the National Genomics Data Center (https://ngdc.cncb.ac.cn/gwh/) under accession number GWHAOSK00000000.1. and the raw genome sequencing data was deposited at NCBI under BioProject PRJNA637812.
References
Bachtrog, D. et al. Tree of sex consortium. Sex determination: Why so many ways of doing it?. PLoS. Biol. 12, e1001899 (2014).
Meisel, R. P. Ecology and the evolution of sex chromosomes. J. Evol. Biol. 35, 1601–1618 (2022).
Martínez, P. et al. Genetic architecture of sex determination in fish: applications to sex ratio control in aquaculture. Front. Genet. 5, 340 (2014).
Charlesworth, D., Charlesworth, B. & Marais, G. Steps in the evolution of heteromorphic sex chromosomes. Heredity 95, 118–128 (2005).
Kikuchi, K. & Hamaguchi, S. Novel sex-determining genes in fish and sex chromosome evolution: novel sex-determining genes in fish. Dev. Dyn. 242, 339–353 (2013).
Herpin, A. & Schartl, M. Plasticity of gene-regulatory networks controlling sex determination: of masters, slaves, usual suspects, newcomers, and usurpators. EMBO Rep. 16, 1260–1274 (2015).
Kamiya, T. et al. A trans-species missense SNP in Amhr2 is associated with sex determination in the tiger pufferfish. PLoS. Genet. 8, e1002798 (2012).
Gong, G. et al. Origin and chromatin remodeling of young X/Y sex chromosomes in catfish with sexual plasticity. Natl Sci. Rev. 10, nwac239 (2023).
Wang, T. et al. Genomic anatomy of homozygous XX females and YY males reveals early evolutionary trajectory of sex-determining gene and sex chromosomes in silurus fishes. Mol. Biol. Evol. 41, msae169 (2024).
Li, X. et al. Sex determination mechanisms and sex control approaches in aquaculture animals. Sci. China Life. Sci. 65, 1091–1122 (2022).
Nagahama, Y., Chakraborty, T., Paul-Prasanth, B., Ohta, K. & Nakamura, M. Sex determination, gonadal sex differentiation and plasticity in vertebrate species. Physiol. Rev. 101, 1237–1308 (2021).
Takehana et al. Co-option of Sox3 as the male-determining factor on the Y chromosome in the fish Oryzias dancena. Nat. Commun. 5, 4157 (2014).
Smith, S. H., Hsiung, K. & Bohne, A. Evaluating the role of sexual antagonism in the evolution of sex chromosomes: new data from fish. Curr. Opin. Genet. Dev. 81, 102078 (2023).
Kitano, J., Ansai, S., Takehana, Y. & Yamamoto, Y. Diversity and convergence of sex-determination mechanisms in teleost fish. Annu. Rev. Anim. Biosci. 12, 233e259 (2024).
Kondo, M., Nanda, I., Hornung, U., Schmid, M. & Schartl, M. Evolutionary origin of the medaka Y chromosome. Curr. Biol. 14, 1664–1669 (2004).
Kuhl, H. et al. A 180 Myr-old female-specific genome region in sturgeon reveals the oldest known vertebrate sex determining system with undifferentiated sex chromosomes. Philos. Trans. B. 376, 20200089 (2021).
Han, W. et al. Ancient homomorphy of molluscan sex chromosomes sustained by reversible sex-biased genes and sex determiner translocation. Nat. Ecol. Evol. 6, 1891–1906 (2022).
Fan, B. et al. A single intronic single nucleotide polymorphism in splicing site of steroidogenic enzyme hsd17b1 is associated with phenotypic sex in oyster pompano, Trachinotus anak. Proc. Biol. Sci. 288, 20212245 (2021).
Tao, W., Conte, M. A., Wang, D. & Kocher, T. D. Network architecture and sex chromosome turnovers: Do epistatic interactions shape patterns of sex chromosome replacement?. Bioessays 43, e2000161 (2021).
Meisel, R. P. Evolution of sex determination and sex chromosomes: a novel alternative paradigm. Bioessays 42, 1900212 (2020).
Graves, J. A. Evolution of vertebrate sex chromosomes and dosage compensation. Nat. Rev. Genet. 17, 33e46 (2016).
Vicoso, B. Molecular and evolutionary dynamics of animal sex-chromosome turnover. Nat. Ecol. Evol. 3, 1632e1641 (2019).
Kitano, J. et al. A role for a neo-sex chromosome in stickleback speciation. Nature 461, 1079–1083 (2009).
Ross, J. A., Urton, J. R., Boland, J., Shapiro, M. D. & Peichel, C. L. Turnover of sex chromosomes in the stickleback fishes (Gasterosteidae). PLoS Genet 5, e1000391 (2009).
Natri, H. M., Merilä, J. & Shikano, T. The evolution of sex determination associated with a chromosomal inversion. Nat. Commun. 10, 145 (2019).
Yoshida, K. et al. Sex chromosome turnover contributes to genomic divergence between incipient stickleback species. PLoS. Genet. 10, e1004223 (2014).
Bitencourt, J. A., Sampaio, I., Ramos, R. T. C., Vicari, M. R. & Affonso, P. R. A. M. First report of sex chromosomes in Achiridae (Teleostei: Pleuronectiformes) with inferences about the origin of the multiple X1X1X2X2/X1X2Y system and dispersal of ribosomal genes in Achirus achirus. Zebrafish 14, 90–95 (2016).
Krysanov, E. & Demidova, T. Extensive karyotype variability of African fish genus Nothobranchius (Cyprinodontiformes2). Comp. Cytogenet. 12, 387–402 (2018).
Zhang, S. et al. Cytogenetic characterization and description of an X1X1X2X2/X1X2Y sex chromosome system in Collichthys lucidus (Richardson, 1844). Acta Oceanol. Sin. 37, 34–39 (2018).
Marajó, L. et al. Chromosomal rearrangements and the first indication of an ♀X1X1X2X2/♂X1X2Y sex chromosome system in Rineloricaria fishes (Teleostei: Siluriformes). J. Fish. Biol. 102, 443–454 (2023).
Xiao, Y. et al. Chromosome-level genome reveals the origin of neo-y chromosome in the male barred knifejaw Oplegnathus fasciatus. iScience 23, 101039 (2020).
Mustapha, U. F. et al. Male-specific dmrt1 is a candidate sex determination gene in spotted scat (Scatophagus argus). Aquaculture 495, 351–358 (2018).
Peng, Y. X. et al. Identification of sex-linked marker and candidate sex determination gene in ornamental fish, African scat (Scatophagus tetracanthus). Aquaculture 563, 739023 (2023).
Jiang, D. N. et al. Establishment of the Y-linked dmrt1Y as the candidate sex determination gene in spotbanded scat (Selenotoca multifasciata). Aquacult. Rep. 23, 101085 (2022).
Huang, Y. Q. et al. A chromosome-level genome assembly of the spotted scat (Scatophagus argus). Genome. Biol. Evol. 13, evab092 (2021).
Li, M. H. et al. A tandem duplicate of anti-mullerian hormone with a missense SNP on the Y chromosome is essential for male sex determination in Nile tilapia, Oreochromis niloticus. PLoS. Genet. 11, e1005678 (2015).
Matsuda, M. et al. DMY is a Y-specific DM-domain gene required for male development in the medaka fish. Nature 417, 559–563 (2002).
Pan, Q. et al. Identification of the master sex determining gene in Northern pike (Esox lucius) reveals restricted sex chromosome differentiation. PLoS. Genet. 15, e1008013 (2019).
Xue, L. et al. Telomere-to-telomere assembly of a fish Y chromosome reveals the origin of a young sex chromosome pair. Genome Biol. 22, 203 (2021).
Suzuki, A. et al. Chromosomes of Scatophagus argus and Selenotoca multifasciata (Scatophagidae). Jpn. J. Ichthyo. 35, 102–104 (1988).
Blanco, D. R. et al. Origin of the X1X1X2X2/X1X2Y sex chromosome system of Harttia punctata (Siluriformes, Loricariidae) inferred from chromosome painting and FISH with ribosomal DNA markers. Genetica 142, 119–126 (2014).
Huang, Y. et al. A rapid method for genetic sex identification in the spotted scat (Scatophagus argus). J. Fish. Sci. China 4, 515–524 (2022).
Bao, L. et al. The Y chromosome sequence of the channel catfish suggests novel sex determination mechanisms in teleost fish. BMC Biol. 17, 6 (2019).
Tao, W. et al. High-quality chromosome-level genomes of two tilapia species reveal their evolution of repeat sequences and sex chromosomes. Mol. Ecol. Resour. 21, 543–560 (2021).
Dai, S. et al. Germline sexual fate is determined by the antagonistic action of dmrt1 and foxl3/foxl2 in tilapia. Development 148, dev199380 (2021).
Luo, H. et al. The male and female genomes of golden pompano (Trachinotus ovatus) provide insights into the sex chromosome evolution and rapid growth. J. Adv. Res. 65, 1–17 (2023).
Wu, K., Song, W., Zhang, Z., Ge, W. Disruption of dmrt1 rescues the all-male phenotype of the cyp19a1a mutant in zebrafish—a novel insight into the roles of aromatase/estrogens in gonadal differentiation and early folliculogenesis. Development 147, dev182758 (2020).
Marshall Graves, J. A. & Catherine, L. P. Are homologies in vertebrate sex determination due to shared ancestry or to limited options?. Genome Biol. 11, 4 (2010).
Kobayashi, T., Kajiura-Kobayashi, H., Guan, G. & Nagahama, Y. Sexual dimorphic expression of dmrt1 and Sox9a during gonadal differentiation and hormone-induced sex reversal in the teleost fish Nile tilapia (Oreochromis niloticus). Dev. Dyn. 237, 297–306 (2008).
Yoshimoto, S. et al. Opposite roles of dmrt1 and its W-linked paralogue, DM-W, in sexual dimorphism of Xenopus laevis: implications of a ZZ/ZW-type sex-determining system. Development 137, 2519–2526 (2010).
Cui, Z. et al. Genome editing reveals dmrt1 as an essential male sex-determining gene in Chinese tongue sole (Cynoglossus semilaevis). Sci. Rep. 7, 42213 (2017).
Wang, L. et al. Transposon-induced epigenetic silencing in the X chromosome as a novel form of dmrt1 expression regulation during sex determination in the fighting fish. BMC Biol. 7, 5 (2022).
Jiao, K. Z. et al. Integrated analysis of the gonadal methylome and transcriptome provides new insights into the expression regulation of sex determination and differentiation genes in spotted scat (Scatophagus argus). Aquaculture 589, 740974 (2024).
Herpin, A. et al. Transcriptional rewiring of the sex determining dmrt1 gene duplicate by transposable elements. PLoS. Genet. 6, e1000844 (2010).
Wang, J., Tao, W., Kocher, T. D. & Wang, D. Sex chromosome turnover and biodiversity in fishes. J. Genet. Genom. 2, 00222–00224 (2024).
Kitano, J. & Peichel, C. L. Turnover of sex chromosomes and speciation in fishes. Environ. Biol. Fishes. 94, 549–558 (2012).
Lin, A. et al. Identification of a male-specific DNA marker in the large yellow croaker (Larimichthys crocea). Aquaculture 480, 116–122 (2017).
Sun, S. et al. Study on sex-linked region and sex determination candidate gene using a high-quality genome assembly in yellow drum. Aquaculture 563, 738987 (2023).
Hayashi, S. et al. Sex chromosome cycle as a mechanism of sex determination. J. Biochem. 176, 81–95 (2024).
Peichel, C. L. et al. Assembly of the threespine stickleback Y chromosome reveals convergent signatures of sex chromosome evolution. Genome Biol. 21, 177 (2020).
Bachtrog, D. A dynamic view of sex chromosome evolution. Curr. Opin. Genet. Dev. 16, 578–585 (2006).
Sember, A. et al. Patterns of sex chromosome diferentiation in spiders: insights from comparative genomic hybridisation. Genes 11, 849 (2020).
Bergero, R., Forrest, A., Kamau, E. & Charlesworth, D. Evolutionary strata on the X chromosomes of the dioecious plant Silene latifolia: evidence from new sex-linked genes. Genetics 175, 1945–1954 (2007).
Cui, D., Liu, Z., Liu, N., Zhang, Y. & Zhang, J. Histological study on the gonadal development of Scatophagus argus. J. Fish. China 37, 696–704 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Garg, S. et al. Chromosome-scale, haplotype-resolved assembly of human genomes. Nat. Biotechnol. 39, 309–312 (2021).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898 (2020).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with singlecopy orthologs. Bioinformatics 31, 3210–3212 (2015).
Yang, X. et al. A fast quality control toolkit for Illumina sequencing data. BMC Bioinforma. 14, 33 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Durand, N. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell. Syst. 3, 99–101 (2016).
Benson, B. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Wang, X. W. & Wang, L. GMATA: an integrated software package for genome-scale SSR mining, marker development and viewing. Front. Plant Sci. 7, 1350 (2016).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, 265–268 (2007).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinforma. 9, 18 (2008).
Han, Y. J. & Wessler, S. R. MITE-hunter: a program for discovering miniature invertedrepeat transposable elements from genomic sequences. Nucleic Acids Res. 38, e199 (2010).
Bao, W., Kojima, K. K. & Kohany, O. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob. Dna. 6, 11 (2015).
Bedell, J. A., Korf, I. & Gish, W. MaskerAid: a performance enhancement to RepeatMasker,. Bioinformatics 16, 1040–1041 (2000).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19, ii215–ii225 (2003).
Keilwagen, J. et al. Using intron position conservation for homologybased gene prediction. Nucleic Acids Res. 44, e89 (2016).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Haas, B. J. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7 (2008).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, 182–185 (2007).
Li, L., Stoeckert, C. J. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 4, 772–780 (2013).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 9, 1312–1313 (2014).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Hedges, S. B., Dudley, J. & Kumar, S. TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22, 2971–2972 (2006).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly. (Austin) 6, 80–92 (2012).
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824 (2012).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Li, H. et al. 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinf. 14, 178–192 (2013).
Yang, W. et al. ddRADseq-assisted construction of a high-density SNP genetic map and QTL fine mapping for growth-related traits in the spotted scat (Scatophagus argus). BMC Genom. 21, 278 (2020).
Kielbasa, S. M., Wan, R., Sato, K., Horton, P. & Frith, M. C. Adaptive seeds tame genomic sequence comparison. Genome Res. 21, 487–493 (2011).
Tang, H. et al. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res. 18, 1944–1954 (2008).
Armstrong, J. et al. Progressive Cactus is a multiple-genome aligner for the thousand-genome era. Nature 587, 246–251 (2020).
Krasheninnikova, K. et al. halSynteny: a fast, easy-to-use conserved synteny block construction method for multiple whole-genome alignments. GigaScience 9, giaa047 (2020).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Miga, K. H. Centromeric satellite DNAs: hidden sequence variation in the human population. Genes 10, 352 (2019).
Kasahara, M. et al. The medaka draft genome and insights into vertebrate genome evolution. Nature 447, 714–719 (2007).
Bertollo, L. A. C., Born, G. G., Dergam, J. A., Fenocchio, A. S. & Moreira-Filho, O. A biodiversity approach in the neotropical Erythrinidae fish, Hoplias malabaricus. Karyotypic survey, geographic distribution of cytotypes and cytotaxonomic considerations. Chromosome Res. 8, 603–613 (2000).
Acknowledgements
All authors thank Professor De-Shou Wang, Professor Wen-Jing Tao and Dr. Shu-Qing Zheng from Southwest University of China for helpful discussions, Ms Gaganpreet Sidhu from York University of Canada and Dr. Umar Farouk Mustapha from University of Saskatchewan of Canada for language assistance, Yucheng Fish Fry Culture Farm in Zhuhai (Guangdong, China) for providing fish, and two anonymous reviewers for their constructive suggestions. This study was supported by grants from the National Natural Science Foundation of China [grant numbers: 32172971, 32273131 and 32473155]; Guangdong Basic and Applied Basic Research Foundation [grant number: 2024A1515010328]; Guangdong Provincial Science and Technology Programme [grant number: 2023B0202010005]; Youth science and technology innovation talent of guangdong TeZhi plan talent [grant number: 2023TQ07A888].
Author information
Authors and Affiliations
Contributions
Data curation, Huang Y.Q., Zhang X.H., and Bian C.; Formal analysis, Jiang D.N., Huang Y.Q., Shi Q., Zhang X.H., Bian C., Jiao K.Z., Huang Yu, and Zhang L.; Funding acquisition, Jiang D.N., Li G.L., and Chen H.P.; Investigation, Jiang D.N., Huang Y.Q., Jiao K.Z., Yang W and Li Y.; Methodology, Jiang D.N., Huang Y.Q., Zhang X.H., Bian C., Zhang L., and Tian C.X.; Project administration, Jiang D.N.; Resources, Zhu C.H., Deng S.P., Chen, H.P., Shi G., and Huang Yang; Supervision, Jiang D.N., Shi Q., and Li G.L.; Writing—original draft, Jiang D.N. and Huang Y.Q.; Writing—review and editing, Jiang D.N., Huang Y.Q., Shi Q., Bian C., and Li G.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: John Mulley and Johannes Stortz. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, YQ., Zhang, XH., Bian, C. et al. Allelic variation and duplication of the dmrt1 were associated with sex chromosome turnover in three representative Scatophagidae fish species. Commun Biol 8, 627 (2025). https://doi.org/10.1038/s42003-025-08056-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08056-1