Abstract
Although the connection between COVID-19 and coagulopathy has been clear since the beginning of SARS-CoV-2 pandemic, the underlying molecular mechanisms remain elusive. Available data support that the hyper-coagulant state is sustained by systemic inflammation. Here we show that the SARS-CoV-2 main protease (Mpro) can play a direct role in the activation of coagulation. Adding Mpro to human plasma increased clotting probability by 3-fold. Enzymatic assays and degradomics analysis indicate that Mpro cleaves and activates coagulation factors VII and XII. This activity is compatible with an extended secondary specificity of Mpro for R↓X that diverge from its well-established preference for LQ↓X. This finding is supported by HDX-MS characterization of the Mpro complex with an Arg-containing inhibitor, as well as the proteolytic cleavage of the peptide FTRLR↓SLEN by Mpro. Overall, integrating biochemical, proteomics and structural biology experiments, we unveil a novel mechanism linking SARS-CoV-2 infection to thrombotic complications in COVID-19.
Similar content being viewed by others
Introduction
Clinical evidence accumulated so far shows that systemic or localized microbial infections, of bacterial or viral origin, are independent risk factors for thromboembolic diseases (e.g., myocardial infarction and stroke)1,2. More recently, the outbreaking viral pneumonia (COVID-19), caused by the beta coronavirus SARS-CoV-2, has dramatically put forward the association between thrombotic complications and virus infection3.
While hypercoagulability stands as a hallmark for severe COVID-194,5, the detailed molecular mechanisms through which SARS-CoV-2 infections can activate blood coagulation have not been conclusively elucidated. Thus, there is still a large demand for the characterization of the biochemical pathways leading to thrombotic events in predisposed patients6,7,8. Filling this knowledge gap is key to understand the clinical phenotype of COVID-19 patients with severe coagulopathies and to identify novel, potential molecular targets for drug discovery9,10,11,12. Severe COVID-19 is associated with a potent activation of the innate immune response, which results into a massive release of pro-inflammatory cytokines (i.e., the “cytokine storm”)13,14 and high bradykinin levels (i.e., the “bradykinin storm”)3. For this reason, it has been proposed that inflammatory cytokines can lead to the activation of blood coagulation in COVID-19 patients by (i) enhancing the expression of pro-coagulant Tissue Factor, (ii) downregulating both anticoagulant and fibrinolytic pathways15, and (iii) stimulating the formation of new platelets with an increased sensitivity to thrombin activation16. In addition, the “bradykinin storm” further sustains coagulation by enhancing the expression of pro-inflammatory and pro-coagulant cytokines (IL-1β, IL-2, IL-6, and IL-8) in a positive feedback loop17.
SARS-CoV-2 is an RNA coronavirus with an unusually large genome (30 kb) that encodes at least 13 open reading frames18, including the main protease Mpro (also known as 3CL protease nsp5) and the papain-like PLpro nsp3 protease. Mpro is a cysteine protease that targets a Gln residue at P1 and a hydrophobic residue (Leu/Met) at the P2 position19,20,21,22,23,24,25,26,27,28. It contributes to viral maturation by cleaving the R1AB polyprotein, resulting in the envelope, membrane, spike, and nucleocapsid proteins18. During infection, Mpro cleaves other host substrates, resulting in the reorganization of the host proteome and, in some cases, enables the virus to evade cellular defense mechanisms27,29,30,31. Among the proteins encoded in the SARS-CoV-2 genome, Mpro has emerged as a potent antigen in infected individuals. Indeed, anti-Mpro antibodies were detected in the saliva and plasma of COVID-19 patients, with a positive correlation between antibody levels and disease severity32,33. Moreover, Mpro-encoding mRNA has been detected in extra-respiratory sites like blood and feces34,35, and the recent identification of Mpro in bronchoalveolar lavage fluid has led to the characterization of an unconventional secretion mechanism from infected cells, involving caspase and gasdermin pores36. Strikingly, extracellular Mpro retains most of its proteolytic activity in blood serum (~70%), indicating that it is not inhibited by physiological protein protease inhibitors, abundantly present in vivo36. Altogether, these findings indicate that Mpro, along with other viral constituents, circulates in the blood of COVID-19 patients, possibly interacting with host plasma proteins and coagulation factors. Nevertheless, the direct involvement of viral proteases in the activation of blood coagulation has not yet been demonstrated.
Blood coagulation is a complex, highly regulated, and evolutionary conserved process aimed at preventing blood loss from the circulating system17,37. It starts from the exposure of Tissue Factor and pro-coagulant surfaces (e.g., collagen), in the sub-endothelium of damaged blood vessels, and the activation of coagulation factors VII and XII. The latter are positioned at the beginning of the so-called “extrinsic” and “intrinsic” pathways, respectively. The coagulation cascade is characterized by the sequential proteolytic activation of protease zymogens at conserved sites, typically Arg↓Ile/Val bonds38, and culminates with the generation of α-thrombin, a serine protease playing a pivotal role in platelet activation and fibrin generation39.
In a previous work40, we have shown that subtilisin, a serine protease secreted from Bacillus subtilis (i.e., a non-pathogenic Gram-positive bacterium in the human gut microbiome), can clot human plasma. We demonstrated that it cleaves prothrombin into active thrombin species, thus bypassing the canonical activation pathway of the coagulation cascade. Building on these earlier results, here we explored the possibility that exogenous Mpro could directly activate the coagulation cascade.
In this study, we found that the addition of Mpro to human plasma samples enhances the clotting probability. Using a wide array of biochemical, biophysical, and proteomics techniques, we demonstrate that Mpro cleaves and activates factors VII and XII at their canonical activation sites. Furthermore, we discovered that (alongside the conventional preference for a Gln at P1 position) the protease can cleave, albeit with lower specificity, peptidyl substrates with an Arg-residue at P1 position. This secondary specificity, combined with the observed pro-coagulant effect of Mpro and its potential to circulate in the COVID-19 patient’s bloodstream, supports the hypothesis that Mpro can play a direct role in the activation of blood coagulation and provides an additional pathogenetic mechanism for the onset of thrombotic complications in SARS-CoV-2 infections.
Results
Exogenous Mpro triggers plasma clotting in vitro
To assess whether Mpro can trigger coagulation in human plasma, we conducted turbidimetric assays by adding to plasma samples the recombinant protease of commercial source (rcMpro) or in-house produced in E. coli cell cultures (rMpro) (Figs. 1A and S1A, B). Furthermore, we tested human plasma either freshly collected from healthy donors or of commercial origin, after re-calcification (Fig. S1C). From the resulting clotting curves, we extracted the clotting time (tc), calculated from the intersection point of the tangent line to the curve at the inflection point with the baseline, and the maximal change in the apparent absorbance at 350 nm (ΔAmax).
A Clotting curves obtained by adding rMpro, dmMpro, or a mock control to human plasma freshly collected from a healthy donor. Plasma samples were diluted 1:2 with HBS, re-calcified, and the increase of turbidity at 350 nm was measured over time at 37 ± 0.1 °C. Measurements were carried out in triplicates, with shaded areas corresponding to the standard deviation at each time point. B Survival curves (Kaplan–Meier) of time to clotting. The clotting probability for plasma incubated with and without Mpro (mock control) is compared at different time points. In the lower panel, the number at risk table reports non-clotted plasma. For this analysis, the lower Mpro concentration triggering coagulation (range from 50 to 500 nM) is considered. The statistical analysis is performed using plasma samples from a cohort of 21 healthy donors. For each sample, the clotting curve was manually evaluated for the presence of a sigmoidal trend indicating a clotting event and, accordingly, the clotting time was calculated (Fig. S2A, B).
The representative clotting curves, shown in Fig. 1A, provide clear-cut evidence of clot formation, whose fibrin origin was confirmed by SDS-PAGE (Fig. S1D). A tc value of ~20 min could be estimated, regardless of rMpro concentration. Furthermore, by doubling the protease concentration, the ΔAmax value was increased by about 50%, suggesting that more thrombin is generated at higher rMpro concentration (Fig. S1A)41. The same phenotype was observed when repeating the experiment using commercial recombinant Mpro (rcMpro) added to freshly prepared plasma (Fig. S1B), or recombinant in-house produced rMpro added to commercial plasma (Fig. S1C), indicating that the reagent source does not appreciably influence the results. An indication that clotting of plasma was caused solely by Mpro, rather than contaminating molecules, came from the observation that the inactive double mutant H41A/C145A of Mpro (dmMpro), in which the catalytic amino acids His41 and Cys145 were both replaced by Ala, failed to induce clotting (Fig. 1A).
These findings prompted us to validate the pro-coagulant effect of Mpro in a cohort of healthy blood donors (n = 21). To this aim, we applied non-parametric statistics, i.e., the Kaplan–Meier method42, to estimate at fixed time points the probability of clotting, i.e., the fraction of plasma samples that clotted in the presence and absence of rcMpro (Figs. 1B and S2). Despite the inter-individual biological variability, possibly affecting the plasma concentration of some coagulation factors43, after 60-min incubation with Mpro, a 3-fold increase in the clotting probability was detected for the Mpro-treated group compared to the control group, with a statistically significant p value of 0.003 (Fig. 1B, top panel). The greater tendency of plasma to clot in the presence of Mpro is shown in the table of Fig. 1B (bottom panel). At incubation times <20 min, all 21 plasma samples tested remained unclotted, regardless of the presence of Mpro. Conversely, at increasing incubation times, the number of unclotted samples progressively decreased to a greater extent in the Mpro-treated group, compared to the control group: after 1 h of incubation, only 8 of the 21 Mpro-treated samples remained unclotted, compared to 17 unclotted samples in the control (Mpro-untreated) group.
Mpro selectively activates coagulation factors VII and XII
Fibrin generation is the endpoint of the coagulation cascade and the result of a complex and precise balance existing between pro-coagulant and anti-coagulant mechanisms. To elucidate the mechanism of Mpro-induced plasma clotting, we screened in vitro the zymogens of pro-coagulant factors (Fibrinogen, Prothrombin, FVII, FX, FIX, FXI, and FXII) for their susceptibility to proteolytic activation by the exogenous protease. Since the observed pro-coagulant effect of Mpro might arise from selective proteolytic degradation/inactivation of anticoagulant factors, we explored the effect of rcMpro on antithrombin III (ATIII), a major irreversible inhibitor of α-thrombin44.
The formation of active coagulation factors was assessed in a simplified system by incubating Mpro with each zymogen, along with a para-nitroaniline (pNA)-releasing chromogenic substrate specific to the resulting active factor. Our results clearly show that the incubation of FVII and FXII zymogens with Mpro significantly increases the release of pNA (p value < 0.05, Kruskal–Wallis test) (Figs. 2 and 3A, B). Conversely, we did not observe any activation with prothrombin (FII), FIX, FX, and FXI zymogens (Fig. S3A–D), nor any change in the inhibition of α-thrombin (FIIa) by ATIII (Fig. S3E). Furthermore, turbidimetric analysis also shows that Mpro is not able to generate fibrin from isolated fibrinogen (Fbg) (Fig. S3F).
A Coagulation factor activation screening workflow. Coagulation factor zymogens were incubated at 37 °C with rcMpro in HBS-PEG buffer, pH 7.4, for 3 h at the specific E:S molar ratio and calcium chloride concentration (see the “Methods” section for experimental details). The generation of active proteases was monitored by enzymatic assays, using specific chromogenic substrates and monitoring the release of pNA at 405 nm. FIX activation was indirectly tested by measuring FIXa-mediated activation of FX. The ability of rcMpro to generate fibrin from fibrinogen was assessed by turbidimetric assay (see “Methods” section for experimental details). A positive (i.e., the active coagulation factor added to the chromogenic substrate solution at a concentration identical to that of the zymogen in the proteolysis mixture with rcMpro) and negative (i.e., the coagulation factor zymogen added to the chromogenic substrate solution at a concentration identical to that of the zymogen in the proteolysis mixture with rcMpro) control assay were always run for each test. B Screening assays for coagulation factor zymogens activation by Mpro. For each zymogen, the bar plot displays the initial rate of pNA release (v0) after the treatment with (red bars) or without (blue bars) rcMpro. Data are presented as mean ± standard deviation (SD), and the Kruskal–Wallis test was used for the comparison. Sample size: for FII, FVII, FIX, FXI, FXII n = 3, for Fbg, FX, FIIa+ATIII n = 2. *p ≤ 0.05. The curves are reported in Figs. 3A, B and S3A–F.
A Activation of FVII by rcMpro monitored by enzymatic assay. FVII zymogen (200 nM) was pre-incubated at 37°C with rcMpro (2 µM) in HBS-PEG buffer, pH 7.4, containing 3 mM CaCl2. After a 3-h reaction, an aliquot (5 µl) of the proteolysis mixture was taken and added to a solution of the FVIIa-specific chromogenic substrate MeSO2-D-CHA-But-Arg-pNA (300 µM) containing 100 nM recombinant tissue factor (TF). The time course of pNA release was monitored at 37 °C by recording the absorbance change at 405 nm obtained with FVII zymogen after 3-h pre-incubation with ( ) or without (▬) rcMpro. The positive control (
) or without (▬) rcMpro. The positive control ( ) corresponds to the kinetics of substrate hydrolysis by fully active FVIIa added at a concentration (10 nM) identical to that of the FVII zymogen in the proteolysis reaction with rcMpro. Data are presented as mean ± standard deviation. B Activation of FXII by rcMpro monitored by enzymatic assay. FXII zymogen (50 nM) was incubated at 37 °C with rcMpro (50 nM) in HBS-PEG buffer, pH 7.4, containing 5 mM CaCl2 and the chromogenic substrate S2302 (300 µM). The appearance of FXIIa activity was detected by recording the time course release of pNA at 405 nm (
) corresponds to the kinetics of substrate hydrolysis by fully active FVIIa added at a concentration (10 nM) identical to that of the FVII zymogen in the proteolysis reaction with rcMpro. Data are presented as mean ± standard deviation. B Activation of FXII by rcMpro monitored by enzymatic assay. FXII zymogen (50 nM) was incubated at 37 °C with rcMpro (50 nM) in HBS-PEG buffer, pH 7.4, containing 5 mM CaCl2 and the chromogenic substrate S2302 (300 µM). The appearance of FXIIa activity was detected by recording the time course release of pNA at 405 nm ( ). The negative and positive control experiments were carried out by incubating FXII zymogen with a S2302 solution in the absence (▬) or presence (
). The negative and positive control experiments were carried out by incubating FXII zymogen with a S2302 solution in the absence (▬) or presence ( ) of PCPS (40 μM) liposomes. A further control experiment was performed by incubating FXII zymogen with a S2302 solution in the presence of BSA (50 nM) (
) of PCPS (40 μM) liposomes. A further control experiment was performed by incubating FXII zymogen with a S2302 solution in the presence of BSA (50 nM) ( ), as a negatively charged protein. Data are presented as mean ± standard deviation. C Rate of FVIIa substrate hydrolysis at increasing pre-incubation times. The initial rate (v0) of pNA release from the FVIIa-specific substrate was determined under the same experimental conditions as those reported in (A), at increasing time of incubation of FVII zymogen with rcMpro in the time range 0–3 h (n = 1). D Quantitative determination of active FVIIa generated by rcMpro-catalyzed cleavage of FVII zymogen. The initial rate (v0) of substrate hydrolysis by active FVIIa is plotted as a function of increasing FVIIa concentrations (○). The calibration curve, resulting from the linear interpolation of the data points, was used to estimate the concentration/amount of active FIIa, which is generated after 3-h incubation of FVII zymogen with rcMpro.(
), as a negatively charged protein. Data are presented as mean ± standard deviation. C Rate of FVIIa substrate hydrolysis at increasing pre-incubation times. The initial rate (v0) of pNA release from the FVIIa-specific substrate was determined under the same experimental conditions as those reported in (A), at increasing time of incubation of FVII zymogen with rcMpro in the time range 0–3 h (n = 1). D Quantitative determination of active FVIIa generated by rcMpro-catalyzed cleavage of FVII zymogen. The initial rate (v0) of substrate hydrolysis by active FVIIa is plotted as a function of increasing FVIIa concentrations (○). The calibration curve, resulting from the linear interpolation of the data points, was used to estimate the concentration/amount of active FIIa, which is generated after 3-h incubation of FVII zymogen with rcMpro.( ).
).
The amount of newly generated FVIIa proportionally increases with the incubation time of FVII zymogen with Mpro (Figs. 3C and S4); after 3 h of reaction, we estimate ~14% of the total amount of FVII zymogen present in the activation reaction (Figs. 3D and S4). In the case of FXIIa generation, about 60% yield of active protease could be solely quantified with respect to the maximal activation induced by phosphatidylcholine/phosphatidylserine (PCPS) liposomes45.
These data are further corroborated by the ability of FVIIa and FXIIa generated by rMpro of activating their physiological substrates (Fig. S5). Indeed, enzymatic assays indicate that the rMpro-activated FVIIa and FXIIa are functionally active as they can activate FX and FXI, respectively.
At physiological pH conditions, rcMpro is negatively charged (pI = 5.9). To rule out the possibility that rcMpro could provide, per se, a negatively charged surface for the conformational activation of FXII, we compared the effect of rcMpro to that observed with bovine serum albumin (BSA, pI = 5.9). The data in Fig. 3B indicate that BSA does not activate FXII, further supporting the hypothesis that Mpro activates FXII via a proteolytic mechanism.
Overall, the results of this screening highlight FVII and FXII as potential substrates for rcMpro in human plasma, whereby the two zymogens are strategically positioned at the beginning of the extrinsic and intrinsic pathways, respectively, of the coagulation cascade.
Mpro displays secondary substrate specificity for Arg-X peptide bonds
Activation of FVII and FXII should involve cleavage at peptide bonds with Arg at P1 position (i.e., Arg152-Ile153 for FVII46 and Arg353-Val354 for FXII47,48). These substrate specificity requirements stand in contrast to the canonical cleavage specificity of Mpro reported so far, which instead mandates the presence of a Gln at P1 and a hydrophobic residue (Leu/Met) at P219,20,21,22,23,24,25,26,27,28,49,50. Hence, we decided to investigate Mpro substrate specificity by applying the high-throughput protease screen (HTPS)38 workflow (Fig. 4A) (see also the “Methods” section). This strategy allows to capture any potential secondary substrate specificity of Mpro by analyzing the identified cleavages of the protease with time resolution (5 min–24 h).
A HTPS workflow. Briefly, rcMpro is incubated with a native standardized cellular extract at different time points. The peptide fragments, generated under native conditions, are isolated by filter-aided sample preparation (FASP) and subsequently identified by data-dependent acquisition (DDA) mass spectrometry (see “Methods”). Significant enrichment (p value < 0.01) of amino acid frequency, compared to a random distribution, enables generating in triplicate an amino acid-position frequency matrix and the profiling of Mpro substrate specificity. B Bar plot of identified peptides/cleavages in HTPS assay. Peptides are generated by incubating rcMpro in the standardized cellular extract for 5 min, 30 min, 4 h, and 24 h. As a control, background peptides are identified in the absence of rcMpro after 30-min and 24-h incubation time (see “Methods”). Histograms refer to the average number of peptides generated in three independent assays (n = 3) at each time point, with error bars corresponding to the standard deviation. C Positional substrate preferences of Mpro (from P4 to P4′) determined by HTPS assay. The heatmaps report the amino acid enrichment (log2FC) compared to a random amino acid distribution. Significant (p value < 0.01) and positive enrichment are shown using a red color palette. Specificity of Mpro at each time point is calculated from three independent replicate experiments (n = 3). The respective natural amino acids are sorted alphabetically. D, E Substrate specificity of Mpro. IceLogo plot of substrate specificity for P3–P3′ positions of Mpro after 30-min reaction. The specificity is reported considering cleavages with Gln in P1 position (D) and for all identified cleavages (E). F Time-course analysis of peptide substrate hydrolysis by rcMpro. The synthetic peptide FTRLQ↓SLEN and its analog FTRLR↓SLEN were separately incubated (10 µM) in HBS at 37 °C with rcMpro (E:S molar ratio 1:40). The relative amount of residual intact peptides (full) and the generated N-terminal peptides (FTRLQ and FTRLR, N-term) were quantified by LC-MS at increasing time points (Fig. S7). Measurements were carried out in three independent experiments, with error bars corresponding to the standard deviation at each time point.
After incubating the native cell extract with rcMpro for 5 min, 30 min, 4 h, and 24 h, we identified 1105 ± 31, 1139 ± 42, 1173 ± 238, and 1442 ± 55 unique cleavages, respectively, while the number of background peptides in the absence of Mpro was negligible. (Fig. 4B, Supplementary Data S1). The substrate preference, calculated as log2-fold change enrichment of amino acids compared to a random distribution, confirmed the canonical specificity for Mpro, with Gln in P1, Leu and Val in P2, and Ser/Ala in P1’ position at all measured time points (Fig. 4C, D). Notably, even after 5-min incubation, there is a significant (p value < 0.01) log2FC enrichment >1.0 for Leu in P2 and Gln at P1 position (Fig. 4C). Noteworthy, the time resolution dimension in this analysis allows to probe the changes in Mpro substrate preferences. In fact, we observed that only after 30-min incubation, rcMpro displays an emerging secondary specificity for basic amino acid residues (Arg and Lys) at P1 position (with a log2FC = 0.6, p value < 0.01), which is maintained up to 24 h only for Arg (Fig. 4C, E and Supplementary Data S1). During the HTPS analysis, we identified >103 cleavage sites containing an Arg-residue in position P1, corresponding to about 10% of all the cleavage sites identified.
To rule out any possible contribution from trace amounts of contamination of proteases in the native cell lysate, we performed HTPS experiments in the absence of Mpro (mock control) after 30-min and 24-h incubation. As a result, less than 10 background peptides per condition were identified, thus excluding any bias generated from active proteases present in the native standardized cell lysate (Fig. 4B and Supplementary Data S1). The purity of rMpro preparations was confirmed by SDS-PAGE and intact mass analysis (Fig. S6). As a further control, we used bottom-up proteomics analysis with rcMpro preparations treated with Trypsin and Glu-C specific endoproteinase, searching the results against Uniprot Pan proteomes (https://ftp.uniprot.org/pub/databases/uniprot/current_release/knowledgebase/pan_proteomes/). At the best of the sensitivity of our MS system (see “Methods” section), we could not identify any protease sequence in the rcMpro preparations tested (Supplementary Data S2).
To validate HTPS findings obtained with Mpro, we synthesized the peptide FTRLQ↓SLEN, recapitulating the canonical substrate preference of the enzyme51, and its analog FTRLR↓SLEN in which Gln was replaced with an Arg-residue. The data shown in Figs. 4F and S7 indicate that, although the efficiency at which the analog is hydrolyzed by Mpro is much lower, the presence of an Arg-residue in P1 position does not impair cleavage by the protease.
Overall, the integration of the HTPS screening dataset with the biochemical characterization of synthetic peptidyl substrates puts forward the possibility that Mpro has an extended substrate specificity and can proteolytically activate both FVII and FXII to trigger plasma clotting.
Mpro interacts with Arg residues in the substrate primary specificity site
To validate on a structural level the secondary substrate specificity of Mpro, we challenged the protease with Arg-containing peptidyl-inhibitors of α-thrombin (Argatroban and PPACK). Even though Mpro and α-thrombin belong to different protease families (i.e., α-thrombin is a serine protease, whereas Mpro is a cysteine protease), they share a common α-chymotrypsin fold52,53 with Mpro active site (i.e., His41 and Cys145) positioned at the interface of two orthogonal β-barrels (i.e., domains I and II). The structural similarity highlighted above for α-thrombin and Mpro can suggest similar molecular recognition properties of the two proteases toward substrates and inhibitors.
Argatroban is a reversible thrombin inhibitor (Ki = 19 nM), whereas PPACK (D-Phe-Pro-Arg-CH2Cl) irreversibly inhibits the enzyme (Ki = 24 nM) by forming a covalent bond with the side chain of the catalytic His57 in the enzyme active site. Noteworthy, both inhibitors orient an Arg-side chain in the protease S1 substrate specificity site via electrostatic coupling with Asp18954. The results of enzyme inhibition assays (Fig. 5A) prove that both inhibitors can effectively block Mpro hydrolytic activity. After 2 h of incubation with excess inhibitor (1:140, E:I molar ratio), the rate of substrate hydrolysis by rcMpro is reduced by ~20% with Argatroban, whereas PPACK completely abrogates the catalytic activity, with an estimated inhibition constant (Ki) of 160 ± 2 nM (Fig. 5B), comparable to that earlier reported for thrombin40,54.
A Bar plot of Mpro inhibition by argatroban and PPACK. Argatroban or PPACK (7 μM) was incubated for 2 h in HBS-PEG, pH 7.4, at 37 ± 0.1 °C with rcMpro (50 nM). The reaction was started by the addition of the fluorogenic substrate (1.25 μM), and the rate of hydrolysis was determined by recording the fluorescence increase at 530 nm. The data are expressed as the relative rate (vi/v0) of substrate hydrolysis in the presence (vi) and absence (v0) of the inhibitor (n = 2). B Determination of Mpro inhibition constant by PPACK. Enzyme inhibition assays were carried out as in (A). The values of (vi/v0) are plotted as a function of inhibitor concentration. The data points were interpolated with Eq. 2, describing the tight binding inhibition model, to extract the apparent inhibition constant (Kiapp), from which a value of Ki = 160 ± 2 nM was estimated (see “Methods”) (n = 1).
In the next step, Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS)55,56,57,58 was used to directly map the binding region for PPACK on Mpro structure. HDX-MS analysis of PPACK binding to rcMpro (Fig. 6) was performed at 20 °C in PBS, pH 7.4, and 95% D2O, after 30-min incubation of Mpro with excess PPACK. Under these conditions, 50 different Mpro peptides were identified, with an amino acid sequence coverage >78% and a peptide redundancy of ~2 (Fig. S8A). The relative difference deuterium uptake (ΔD%) for each peptide, normalized for the number of exchangeable hydrogens, was determined at increasing incubation times (15 s–2 h) and reported in Fig. 6A (see also Supplementary Data S3). To better identify the regions that are conformationally altered upon inhibitor binding, a differential three-dimensional map of deuterium uptake was obtained by projecting ΔD% values, obtained after 2-h exchange with D2O, onto on the crystallographic structure of Mpro in the dimeric active form (6y2e.pdb)19 (Fig. 6B). At short incubation times (<5 min), regions that are embedded in or close to the dimerization interface (i.e., peptides 1–17 and 287–295), become more flexible/exposed in the presence of PPACK (Fig. 6B, regions depicted in red), likely because covalent inhibitor labeling might destabilize the dimeric structure causing exposure/flexibilization of the regions at the dimer interface. Conversely, peptides encompassing the catalytic His41 (e.g., peptide 38–44) are more buried/rigid in the PPACK-bound form even after 2h-incubation (Fig. 6B, regions depicted in blue). A similar pattern, though less pronounced, is observed for peptides near the active site (fragment 48–57). Furthermore, fragments 141–150 and 142–150, both encompassing the catalytic Cys145, could be identified solely in the free form of the enzyme, thus providing strong evidence that Cys145 is covalently derivatized by PPACK (Fig. S8B).
A HDX-MS analysis of Mpro-PPACK interaction. The difference deuteration (ΔD%), measured for peptide fragments deriving from rcMpro in the PPACK-bound and -unbound form, is reported in the time range of D2O labeling 15 s–2 h, as indicated (see “Methods”). Negative values of ΔD% correspond to regions that become more shielded/rigid upon inhibitor binding, whereas positive values correspond to regions that become more exposed/flexible. The active site is indicated. Peptides with a significant change in deuteration are highlighted in the red or blue boxes, depending on the direction of the change, positive or negative, respectively, and their deuterium uptake plots are reported in Fig. S8C. Experimental conditions were as follows: rcMpro (1.25 μM) incubated at 20 °C in 20 mM PBS in 95:5 D2O:H2O solution, pD 7.4, containing 150 mM NaCl. At each time point, HDX-MS measurements were conducted in triplicate. B Three-dimensional difference map of Mpro deuterium uptake after PPACK binding. The values of ΔHDX, as obtained from HDX-MS data of rcMpro in the PPACK-bound and -unbound state after 2-h exchange, are projected onto protomer A in the dimeric Mpro structure (6y2e), represented as a ribbon drawing. The regions that are significantly (adj p value < 0.05) more (ΔHDX < −0.25 Da) or less (ΔHDX > 0.25 Da) protected from H/D exchange after PPACK binding are colored blue or red, respectively. Regions without statistically significant changes are in white, while regions not covered are shown in gold. For the sake of clarity, protomer B is in gray. The relative deuterium uptake plots of relevant peptides in the active-site region (sequence 38–44), and at the N-terminal (sequence 1–17) and C-terminal (sequence 287–295) regions, in the PPACK-bound and -unbound states, are also reported.
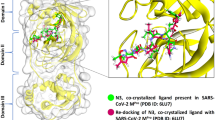
The results of enzyme inhibition assays and HDX-MS analysis provide structural support to the secondary substrate specificity of Mpro highlighted above and to the proposal that the protease can allocate an Arg-side chain in the S1 sub-site. This conclusion is further corroborated by molecular docking simulations (docking energy score: −135.571), showing that the positively charged Arg-side chain of PPACK could interact with the negative Glu166 in the protease S1 site, while Pro points toward His41 in the active site, and D-Phe contacts Thr25 and Thr26 in the S1′ sub-site on Mpro structure (Fig. 7A). Furthermore, the largely negative surface electrostatic potential of Mpro dimer (Fig. 7B) is expected to favor binding and cleavage of positive charged ligands and substrates, as shown in this study for FVII and FXII substrates, and as reported in the literature for other numerous electropositive interactors, such as histones, histone-related proteins, ribosomal and ribonuclear proteins31.
A Close-up view of D-Phe-Pro-Arg peptide (shown in stick) (i.e., the peptidyl moiety of PPACK D-Phe-Pro-Arg- CH2Cl) docked into Mpro active site (shown as van der Waals surface, light gray). Ligand atoms are color coded: nitrogen in blue, oxygen in red, carbon in dark gray. Relevant amino acid side-chains in Mpro are indicated. The Arg guanidyl-group is electrostatically coupled to Glu166 in the S1 site, whereas D-Phe points toward His41 in the active site (AS), whereas Pro interacts with Thr25 and Thr26 in the S1′ sub-site. HPEPDOCK software was used in docking simulations (see “Methods”). B Surface electrostatic potential of Mpro in the dimeric form. Calculations were carried out using the APBS software on the crystallographic structure of Mpro in the dimeric form (6y2e.pdb). The surface is colored according to the electrostatic potential (blue, positive; red, negative), as indicated, and expressed as kJ/(mol·q).
Mpro cleaves FVII and FXII zymogens at the physiological activation sites
The discovery of Mpro secondary preference for Arg at P1 position suggests that Mpro-mediated activation of FVII and FXII might occur via canonical proteolysis at Arg residues, as earlier reported for known physiological activators38,46. To verify our hypothesis, we identified the Mpro activation cleavage sites on purified FVII and FXII by exploiting a simplified version of the Terminal Amine Isotopic Labeling of Substrates (TAILS) protocol59. This method enables the identification of new N-termini and cleavage sites on a protein through reductive di-methylation and a bottom-up MS workflow. (Fig. 8A).
A TAILS workflow. Simplified version of TAILS workflow in which the substrates (FVII and FXII) are proteolyzed with Mpro (1:10 E:S molar ratio) and mock treatment. Generated cleavage fragments are subsequently di-methylated under reducing conditions. Di-methylated peptides are identified by mass spectrometry and subjected to differential mass analysis to identify Mpro cleavage sites. B Mpro-specific cleavages mapped in the FVII sequence. Results from the TAILS experiment using rcMpro and mock treatment. The experiment is performed with three independent replicates. Cleavages identified in at least two replicates (except 153IVGGKVCPKGECPWQVLLLVNG174) and not in the mock control are shown. Dashed lines indicate the cleavage site (Arg152-Ile153) for FVII physiological activators. The peptide 153IVGGKVCPKGECPWQVLLLVNG174 generated by the cleavage in the activation site is annotated in red. C Quantitative determination of 153[DimethNter]IVGGKVCPK[DimethLys]GECPWQVLLLVNG174 peptide. The peptide is quantified using a targeted mass spectrometry approach in both the mock control (FVII alone) and in the presence of rcMpro. The intensity reported is calculated from the sum of the top five peptide transitions. This experiment was conducted with three replicates, with each circle denoting a single measurement and the diamond indicating the mean value. The elution profiles of the peptide are reported in Fig. S9C. D Mpro-specific cleavages mapped in the FXII sequence. Results from the TAILS experiment using rcMpro and mock treatment. The experiment is performed with three independent replicates. Cleavages identified in at least two replicates and not in the mock controls (t = 0 and 3 h) are shown. Dashed lines indicate the cleavage sites for FXII physiological activation.
After 3 h incubation of FVII with rcMpro, we identified three N-terminal di-methylated peptides in at least two of three biological replicates and absent in the mock control, resulting from the cleavage at Gln366-Gly367, 392Arg-Ser393, and Val399-Leu400 bonds (Fig. 8B and Supplementary Data S4). Furthermore, in one of the three replicates (but not in the mock control), we could identify the cleavage of rcMpro at the Arg152-Ile153 bond (Fig. 8B, green peptide), matching with the canonical activation site on FVII by its physiological activators46. As the generation of 153IVGGKVCPKGECPWQVLLLVNG184 can be taken as a safe marker of FVII activation, we leveraged the sensitivity of Parallel Reaction Monitoring (PRM) to selectively measure the intensity of the peptide (Fig. 8C, Supplementary Data S4, Fig. S9). After 3 h incubation with rcMpro, trace amounts of the activation peptide were detected even in the mock control samples, consistent with the autoactivation of the FVII zymogen earlier reported60. Nevertheless, we found a significant (p value = 0.001, unpaired two-sided Student’s t test) two-fold increase in the intensity of the activation peptide in rcMpro-treated samples, compared to controls. Even though unspecific proteolysis at the FVII C-terminal region might in principle cause a loss of function in the resulting truncated species (i.e., degradation), the generation of active FVIIa in the proteolysis reaction of FVII with rcMpro, as experimentally observed in Fig. 3A, indicates that the cleavage at the physiological activation site (i.e., Arg152-Ile153) is prevalent (i.e., activation).
The same TAILS strategy, highlighted above for FVII, was exploited to identify the cleavage sites for rcMpro on FXII zymogen. After binding to negatively charged membranes, FXII zymogen is cleaved in vivo primarily at Arg353-Val354 to generate active FXIIa protease, and eventually at two other extra bonds, i.e., Arg343-Leu344, and Arg334-Asn335. After incubation of FXII in solution with rcMpro (in the absence of negatively charged liposomes), 19 N-terminal di-methylated peptides can be identified in at least two out of three replicates, that are absent in the mock controls (t = 0 or 3 h) (Fig. 8D). The variable nature of the amino acids at P1 position in the cleaved peptide bonds once more supports the extended specificity of Mpro, (Fig. 4C). Most of the cleavage sites that are mapped in Fig. 8D are localized outside the FXII protease domain and therefore not relevant to zymogen activation. Importantly, our TAILS analysis identifies in all three replicates, but not in the control experiment, the peptide 354VVGGLVALR362, generated from the cleavage at the physiological activation site (Arg353-Val354 bond). The latter finding clearly indicates that rcMpro can also proteolytically activate FXII.
Overall, by applying the TAILS workflow, we discovered that Mpro can directly activate FVII and FXII by proteolysis at their physiological activation sites, fully consistent with the results of enzymatic assays (Fig. 3A, B).
Discussion
Thrombotic diseases may arise on their own or as secondary manifestations of a wide range of pathologies with different etiologies and clinical phenotypes (e.g., cancer61, rheumatoid arthritis62, autoimmune diseases63,64, amyloidosis65,66, and infectious diseases both of bacterial and viral origin67,68). While the molecular mechanisms underlying secondary thrombotic complications remain elusive, non-canonical mechanisms involving aberrant proteolytic activation of blood coagulation have been proposed40,69.
The link between COVID-19 and thrombosis has been clear since the early days of the SARS-CoV-2 pandemic, with COVID-19-associated coagulopathy strongly contributing to morbidity and mortality in infected patients8,9,10,11,12,13. During SARS-CoV-2 infections, innate immune response is activated, along with the massive release of pro-inflammatory and pro-coagulant cytokines that cooperate to boost a pathological feedback loop between inflammation and thrombosis14. The results of our study indicate that, in addition to the “cytokine and bradykinin storm” mechanism, SARS-CoV-2 can directly activate blood coagulation by exploiting the extracellular proteolytic activity of the viral main protease, Mpro.
Using a wide array of biochemical, biophysical, and proteomics techniques, we have shown that exogenous Mpro can trigger human plasma clotting by proteolytically activating FVII and FXII. The pro-coagulant activity of Mpro was further confirmed with plasma samples from a cohort of 21 healthy donors, allowing to estimate a 3-fold increase in the clotting probability of Mpro-treated plasma samples compared to the control group (p value ~ 0.003). Notably, the pro-coagulant potential of Mpro varies considerably across the plasma samples tested, reflecting the inter-individual variability and the different severity of thrombotic complications observed in COVID-19 patients11.
To identify the mechanism generating a pro-coagulant phenotype, we employed in vitro enzymatic assays and found that Mpro cleaves FVII and FXII and that the pro-coagulant effect is inherently associated with the proteolytic activity of the enzyme, as evidenced by the inactive double mutant H41A/C145A, which exhibits no plasma clotting activity.
Cleavage of FVII and FXII at the physiological Arg-X activation sites could not be anticipated by the reported substrate specificity of Mpro, dominated by the preference for a Gln residue at P1 and a hydrophobic amino acid in P2 (Leu, Phe, Met)19,20,21,22,23,24,25,26,27,28. Using time-resolved HTPS profiling of Mpro, we identified an extended/secondary specificity of Mpro, whereby the protease, in addition to its canonical specificity (LQ↓), can cleave (albeit with lower specificity) substrates with an Arg-residue at the P1 position. Noteworthy, this secondary specificity for Arg-X bonds appears only at longer incubation times (i.e., >30 min), when the preferred substrate (LQ↓) is consumed or becomes scarce. This observation aligns with recent degradomics studies showing that Mpro can cleave substrates with amino acids other than Arg at position P127,28 (e.g., His and Met).
This key finding was further validated by enzymatic inhibition assays and docking simulations, showing that Arg-containing inhibitors (i.e., Argatroban and PPACK) can block Mpro hydrolytic activity. Furthermore, HDX-MS analysis provided direct structural evidence for PPACK binding to the Mpro active site. It is noteworthy that the replacement of Q with R in a consensus substrate peptide could still be cleaved by Mpro, even with lower efficiency. The clear-cut proof that Mpro can cleave and activate FII and FXII came from the results of TAILS analysis, showing that the two zymogens are hydrolyzed at their physiological cleavage sites, i.e., Arg152-Ile153 for FVII and Arg353-Val354 for FXII. These data indicate that proteolysis occurs at sites with canonical (Q↓) and non-canonical substrate specificity. However, these cleavage sites are mostly located outside the protease domain, likely resulting in active truncated forms. Thus, the proteolytic activity of Mpro tends to activate zymogens rather than degrade them.
At this point, the question arises as to whether the results of this study could have pathological implications in SARS-CoV-2 infections. A first prerequisite to address this point is the extracellular localization of Mpro. In fact, it is necessary that the protease can interact with coagulation factor zymogens in the bloodstream. Viral proteases have been characterized in the intracellular compartment of infected cells. However, recent data indicate that mRNA encoding for Mpro is present in extra-respiratory districts like blood and feces34,35. Even more recently, the active protease Mpro has been directly identified in bronchoalveolar lavage fluid in infected mice, with secretion mediated by caspase-activated gasdermin pores36. Furthermore, extracellular Mpro retains most of its hydrolytic activity in blood serum (~70%), suggesting that it is not inhibited by physiological protease inhibitors36. Additionally, the SARS-CoV-2 spike protein has been found to facilitate virus interaction with fibrin8. This interaction likely enhances Mpro release at local sites, thereby amplifying its pro-coagulant effect.
Based on these considerations, the results of this study allow us to propose that during viral infections, Mpro, released (either locally or systemically) from SARS-CoV-2-infected cells, can proteolytically activate FVII and FXII. FVII and FXII zymogens are strategically positioned at the beginning of the “intrinsic” and “extrinsic” pathways, respectively. Hence, the pro-coagulant activity induced by Mpro can amplify the coagulation cascade, where even small amounts of active FVIIa and FXIIa trigger a hypercoagulable state26,70. Furthermore, the ensuing FXIIa has a dual role, being both pro-coagulant and pro-inflammatory. Indeed, besides activating FXI downstream in the coagulation cascade, FXIIa converts prekallikrein to active kallikrein, which promotes bradykinin release71 and activates FXII, amplifying its formation47. The release of bradykinin, in turn, boosts inflammation downstream.
The experiments in this study suggest that Mpro directly contributes to blood coagulation activation, highlighting an additional pathogenetic mechanism behind thrombotic complications in severe SARS-CoV-2 infections.
Methods
Reagents
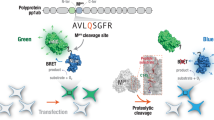
Human coagulation factors Prothrombin, Thrombin, FX, FXa, FVII, FVIIa, FIX, FIXa, FXI, FXIa, FXII, and recombinant TF (TF) were purchased from Haematologic Technologies Inc. (Essex Junction, VT, USA); fibrinogen from American Diagnostica (Stamford, CT, USA). Commercial Mpro (rcMpro) was purchased from Sigma-Aldrich (St. Louis, MO, USA). Chromogenic substrates S2238, S2765, S2366, and S2302 were purchased from Chromogenix (Milan, Italy), while MeSO2-D-CHA-But-Arg-pNA was from Sigma (Darmstadt, Germany). Mpro FRET-based substrate 5-FAM-AVLQSGFRK(DABCYL)K (ProteoGenix, Miami, FL, USA)- Commercial human plasma (STA-System Control N+P, Normal plasma) was purchased from STAGO (Milan, Italy). Egg phosphatidylcholine (PC) and brain phosphatidylserine (PS) were purchased from Avanti Polar Lipids, Inc. (Alabaster, AL, USA). Salts, solvents, and other reagents were of analytical grade and purchased from Merck LifeScience (Darmstadt, Germany).
Protein preparation
In-house produced recombinant Mpro (rMpro) and Mpro double-mutated to H14A and C145A (dmMpro) were purified and characterized as previously described72. Briefly, the sequence encoding for wild-type SARS-CoV-2 Mpro was inserted into a pET-SUMO expression vector with the restriction-free cloning method. Subsequently, to produce the catalytically inactive mutant, the resulting plasmid was double-mutated to H41A and C145A with a single site-directed mutagenesis reaction with primers 3′-AGAGGTGCAGATGACAGCACGAGGGCAATAGAC-5′ and 5′-TTCCTTAATGGCAGCGCTGGTTCGGTGGGCTTT-3′. The resulting plasmid was first sequenced to verify the insertion of the mutations and then used to transform an E. coli BL21(DE3) cell culture, pre-cultured in LB broth at 37 °C overnight in the presence of kanamycin (50 μg/mL). This culture was used to inoculate fresh LB medium supplemented with the antibiotic, and the cells were allowed to grow at 37 °C to an optical density at 600 nm of 0.6–0.8 A.U. Mpro expression was induced with 0.5 mM isopropyl β-D-1-thiogalactopyranoside (IPTG). The induction phase was allowed to proceed overnight at 20 °C. The cells were then harvested by centrifugation (5000 × g, 4 °C, 20 min) and the pellet was resuspended in buffer A (20 mM Tris, 150 mM NaCl, pH 7.8) containing DNAse I. After ultrasound treatment, the cell lysate was clarified by centrifugation at 12,000 × g and 4 °C for 30 min and loaded onto a HisTrap HP column (GE-Healthcare) equilibrated with 98% buffer A and 2% buffer B (20 mM Tris, 150 mM NaCl, 500 mM imidazole, pH 7.8). His-tagged Mpro was eluted with a linear imidazole gradient from 10 to 500 mM. The pooled fractions containing the target protein were subjected to buffer exchange with buffer A using a HiPrep 26/10 desalting column (GE-Healthcare). Next, ULP-1 protease was allowed to react (2 μg of ULP-1 per mL of Mpro solution) at 12 °C overnight to cleave the N-terminal HisTag-SUMO moiety. The reaction mixture was loaded onto a HisTrap HP column equilibrated in buffer A to separate the cleaved Mpro from the His-tagged ULP-1 protease, the HisTag-SUMO moiety, and the uncleaved protein. Mpro was finally purified using a Superdex 200 prep grade 16/600 SEC column (GE-Healthcare) equilibrated with buffer C (20 mM Tris, 150 mM NaCl, pH 7.8). The fractions containing the high-purity target protein were pooled, concentrated at 5 mg/ml, and flash-frozen in liquid nitrogen for storage in small aliquots at −80 °C.
Recombinant commercial Mpro (rcMpro) from SIGMA was resolubilised in water containing 1 mM DTT, to reach a final concentration of 1 mg/ml. Commercial human fibrinogen (America Diagnostica Inc) was desalted on a 5-ml HiTrap desalting column (GE Healthcare, IL, USA), eluted at a constant flow rate (1 ml/min) with 10 mM HEPES pH 7.4, 150 mM NaCl (HBS). All purifications were checked for potential proteases and LPS contamination. The material eluted in correspondence with the major chromatographic peak was collected and used for subsequent analyses. Fibrinogen concentration was determined spectrophotometrically at 280 nm, using a molar absorptivity of 513.400 M−1·cm−1.
Human plasma preparation
Platelet-poor plasma (PPP) was prepared from freshly withdrawn, citrated blood samples taken from 21 healthy donors (twelve men and nine women, 21–60 years of age). Donors gave their written informed consent to participate in this study, which was approved by the Institutional Ethics Committee of the Padua University Hospital. All ethical regulations relevant to human research participants were followed. PPP was prepared by centrifugation of whole blood at 1500 × g for 15 min, at room temperature, without brake, as recommended by the Clinical Laboratory Standards Institute guidelines73. For commercial plasma sample preparation, citrated lyophilized normal human plasma was reconstituted with distilled water following the manufacturer’s instructions.
Liposomes (PCPS) preparation
Liposomes containing phosphatidylcholine (PC) and phosphatidylserine (PS) were obtained by the extrusion method, as detailed elsewhere74. The final liposome solution (12 µM) was composed of PC and PS (50:50 molar ratio) unilamellar vesicles of 100 ± 30 nm diameter, as determined by Dynamic Light Scattering, using a Zetasizer Nano-S instrument (Malvern Instruments, UK).
Turbidimetric assays
Fibrin generation by rcMpro or dmMpro was probed on re-calcified citrated plasma diluted 1:2 with HBS or on a solution of freshly desalted fibrinogen (0.15 mg/ml). To assess the correct CaCl2 amount to be added to citrate plasma, a re-calcification scouting was run, and the maximum CaCl2 concentration that did not trigger plasma clotting within 1 h was selected. Plasma samples or fibrinogen were incubated with increasing concentration of rcMpro, at 37 ± 0.1 °C, or the correspondent higher amount of added DTT as a control, and the clot formation was followed by continuously recording the apparent absorbance of the solution at 350 nm, using a V-630 Jasco (Tokyo, Japan) spectrophotometer or a Victor Nivo Multiplate Reader (PerkinElmer, Waltham, MA, USA). The resulting clotting curves were analyzed to extract the values of tc and ΔAmax, where tc is the clotting time and is calculated from the intercept point of the tangent to the maximal slope of the curve with the time axis; ΔAmax is the maximal absorbance change when fibrin generation is complete40. Statistical difference in the probability of plasma clotting after rcMpro treatment was assessed by performing a turbidimetric assay on plasma samples from 21 healthy donors. Each clotting curve was evaluated for the presence of a sigmoidal curve indicating a clotting event, and the clotting time was accordingly calculated. The probability of clotting within 1 h of assay was estimated using the Kaplan–Meier method, and the survival curves were compared by log-rank test42 using the R-packages “survival” and “survminer”75,76.
Electrophoretic analysis of plasma clot
After 1-h incubation at 37 °C of 1:2 diluted plasma with rMpro (200 nM), the plasma clot was collected, washed with HBS buffer, and resuspended with GdnCl 8 M (incubated for 1 h at 37 °C). Albumin was then removed by a treatment with EtOH to a final concentration of 42% for 1 h at 4 °C and following centrifuging at 16,000 × g for 45 min at 4 °C. After removing the supernatant, the pellet was then resuspended with SDS sample loading buffer and analyzed by reducing SDS-PAGE (Bolt Bis-TRIS 4-12% precast gel) and Coomassie staining (Simply Blue SafeStain, Invitrogen). For comparison, commercial fibrinogen (Fbg) was also loaded. The typical α-, β-, and γ-chain of Fbg are indicated by arrows, along with the γ-chain dimer.
Coagulation factors activation assays
Activation of coagulation factors zymogens by Mpro was assessed by incubating the zymogen at 37 ± 0.1 °C in the absence (negative control) and presence of rcMpro and monitoring at the same temperature the release of p-nitroaniline (pNA) from the corresponding specific chromogenic substrate by measuring the absorbance increase at 405 nm (ɛM405nm = 9920 M−1·cm−1). For each zymogen, the experimental details are reported in the following. Prothrombin (10 nM) and FX (10 nM) were incubated with rcMpro at an E:S ratio of 10:1 (mol/mol) in HBS buffer, containing 0.1% (w/v) PEG8000 (HBS-PEG) and 5 mM CaCl2. After 3-h incubation, the reaction mixture was added with α-thrombin chromogenic substrate S2238 (20 μM) or with FXa chromogenic substrate S2765 (100 μM). Activation of prothrombin with 1 nM FXa in the presence of 100 μM PCPS (50:50) and activation of FX with 0.2 nM FVIIa in the presence of 2 nM TF were performed as positive controls. FVII (200 nM) was incubated for increasing periods of time in HBS-PEG with 5 mM CaCl2, at 37 ± 0.1 °C, alone or with rcMpro at an E:S ratio of 10:1 (mol/mol). At time points (i.e., 0, 45, 90, 180 min), aliquots of FVII from the reaction mixture (10 nM final concentration) were added to FVIIa-specific chromogenic substrate MeSO2-D-CHA-But-Arg-pNA (300 μM) in the presence of TF (100 nM). To calculate the concentration of the FVIIa generated by rcMpro, at each time point, the absorbance signal obtained from FVII in the presence of TF alone (reference signal) was subtracted from the absorbance signal obtained in the presence of rcMpro, and the initial rate (v0) of pNA release was calculated. The concentration of FVIIa was then estimated from a FVIIa titration curve in the presence of 100 nM TF. FIX (100 nM) was incubated with rcMpro at an E:S ratio of 1:1 (mol/mol) in HBS-PEG with 3 mM CaCl2. FIX activation was determined indirectly by activation of FX. Hence, after 3-h incubation, the reaction mixture was added with FX (10 nM) and FXa chromogenic substrate S2765 (100 μM). Activation of FX with 40 nM FIXa in the presence of 20 μM PCPS (50:50) was performed as a positive control. FXI (1 nM) was incubated with rcMpro at an E:S ratio of 1:1 (mol/mol) in HBS-PEG with 3 mM CaCl2. After 3-h incubation, the reaction mixture was added with FXIa chromogenic substrate S2366 (100 μM). Cleavage of S2366 by FXIa (0.1 nM) was performed as a positive control. FXII (100 nM) was treated with rcMpro at an E:S ratio of 1:1 (mol/mol) in HBS with 5 mM CaCl2 and added with FXIIa chromogenic substrate S2302 (300 μM) without previous incubation. Activation of FXII in the presence of 40 μM PCPS (50:50) was performed as a positive control. Activation of FXII by rcMpro was compared to BSA at the same E:S ratio.
The effect of rcMpro on the anticoagulant factor Antithrombin III (ATIII) was tested by incubating ATIII with rcMpro for 1 h at 37 °C in a 1:1 molar ratio (40 nM). This solution was further incubated with α-thrombin (1 nM) for 3 h at 37 °C in HBS-PEG, 3 mM CaCl2. α-Thrombin residual activity, after ATIII incubation, in the absence and presence of rcMpro, was tested using the S2238 chromogenic substrate.
HTPS analysis
In the high-throughput native microscale protease screen, a standardized native cell lysate is proteolyzed with the studied protease. The protease-generated peptides are collected, analyzed by MS, and the identified substrate peptides are analyzed to retrieve activity, specificity, and cleavage entropy data. The detailed workflow is described in Uliana et al.38.
Sample preparation
For HTPS characterization of rcMpro, a variation of the protocol for cysteine protease has been developed: native cell lysate in 20 mM Ammonium bicarbonate pH 7.8 (50 µg, 0.5 mg/ml), 5mM L-cysteine, 1 mM EDTA was incubated with rcMpro (5 µg, E:S 1:10) for 5 min, 30 min, 4 h and 24 h at 37 ± 0.1 °C under agitation (1400 × g) in a filtered 1.5 ml tube (10-kDa cutoff) from Sartorius (Göttingen, Germany), according to the filter-aided sample preparation (FASP) protocol for proteomic analysis77. Peptides were collected by 15-min centrifugation 10,000 × g followed by one wash with 100 µl of MS-grade water to increase peptide recovery. After cleanup using C18 StageTips, dried peptides were dissolved in 2% acetonitrile-0.1% formic acid before the analysis by MS in the data-dependent acquisition (DDA) mode. Samples were prepared in triplicate.
MS acquisition
LC-MS/MS was performed on an Orbitrap QExactive+ mass spectrometer (Thermo Fisher, Waltham, MA, USA) coupled to an EASY-nLC-1000 liquid chromatography system (Thermo Fisher). Peptides were separated using a reverse phase column (75 µm × 400 mm New Objective, in-house packed with ReproSil Gold 120 C18, 1.9 µm, Dr. Maisch GmbH) across 180 min gradient from 3% to 25% B in 160 min and from 25% to 40% B in 20 min (buffer A: 0.1% (v/v) formic acid; buffer B: 0.1% (v/v) formic acid, 95% (v/v) acetonitrile). The DDA acquisition mode was set to perform one MS1 scan followed by a maximum of 20 scans (TOP20) with MS1 scans (R = 70,000 at 400 m/z, AGC = 3e6 and maximum IT = 64 ms), HCD fragmentation (NCE = 25%), isolation windows (1.4 m/z), and MS2 scans (R = 35,000 at 400 m/z, AGC = 2e5 and maximum IT = 55 ms). A dynamic exclusion of 30 s was applied, and charge states lower than two and higher than seven were rejected for the isolation.
Data analysis
DDA data were searched with the MaxQuant software package (version 1.5.2.8) using HTPS_DB.fasta database (2557 entries). The search was performed with digestion mode set to unspecific, maximal peptide length to 40 AA, and only acetylation of N-termini and methionine oxidation as variable modifications. First search peptide mass tolerance was 20 ppm, and the main search peptide mass tolerance was 4/5 ppm. MS/MS match tolerance was set to 20 ppm. Target decoy approach was used to control FDR, which was set to 0.01 at the peptide and PSM levels. Protease substrate specificity analysis was performed in R (version 3.4.3) using the workflow deposited on Github (https://github.com/anfoss/HTPS_workflow, https://doi.org/10.5281/zenodo.4484341) under MIT license.
Mass spectrometry characterization of Mpro preparations
Bottom-up proteomics characterization
Five micrograms of rcMpro preparation was loaded on a centrifugal unit (Vivacon 500, 10MKCO, Sartorius) and centrifuged until dry (centrifugation 8000 × g, 15 min). After denaturation (8 M urea), reduction [5 mM tris(2-carboxyethyl)phosphine (TCEP)], and alkylation (10 mM iodoacetamide), the sample was washed with 25 mM ammonium bicarbonate (three centrifugation steps, 8000 × g, 15 min) and overnight proteolyzed [1 μg of trypsin (Promega) sequencing grade, 1 μg of endoproteinase Glu-C (Promega) sequencing grade]. Proteolysis was quenched by 0.1% formic acid, and dried peptides were resuspended in 10 μl of 0.1% formic acid and 2% acetonitrile. As control, only Trypsin and Glu-C were digested and analyzed by MS. Generated peptides were analyzed by LC-MS/MS on an LC-MS/MS was performed on an Orbitrap Q Exactive+ mass spectrometer (Thermo Fisher Scientific) coupled to an EASY-nLC-1000 liquid chromatography system (Thermo Fisher Scientific). Peptides were separated using a reverse phase column (75 μm ID × 400-mm new objective, in-house packed with ReproSil Gold 120 C18, 1.9 μm, Dr. Maisch GmbH) across 60-min linear gradient from 5 to 35% [buffer A, 0.1% (v/v) formic acid; buffer B, 0.1% (v/v) formic acid and 98% (v/v) acetonitrile]. The DDA mode was set to perform a maximum of 20 scans for the top 20 most intense peptides (TOP20) with MS1 identification (R = 140,000 at 400 m/z, AGC = 3 × 106, and maximum IT = 110 ms), HCD fragmentation (NCE = 25%), isolation windows (1.4 m/z), and MS2 identification (R = 35,000 at 400 m/z, AGC = 1 × 105, and maximum IT = 110 ms). Charge states lower than two and higher than seven were rejected. Acquired spectra were searched using the MaxQuant software version 1.5.2.8 embedded with the Andromeda search engine against the Pan proteome (https://ftp.uniprot.org/pub/databases/uniprot/current_release/knowledgebase/pan_proteomes/, singleton and non-singleton reference proteome clusters, 75% proteome similarity for Eukaryota and 55% proteome similarity for Archaea and Bacteria, downloaded 26.07.2021), extended with Mpro sequence and a contaminant list. The search parameters were set to include only fully tryptic peptides, carbamidomethylation as static peptide modification, and oxidation (M) as variable modification. The MS and MS/MS mass tolerances were set to 10 and 20 parts per million, respectively. False discovery rate of <1% was used at the peptide and protein levels.
Intact-MS characterization
One microgram of rMpro was diluted in 20 μl of 0.1% formic acid and directly injected into a Waters (Milford, MO, USA) Xevo G2-S Q-TOF mass spectrometer. The mass analysis was run in the positive ion mode, with the capillary potential set at 1.5 kV and source temperature at 100 °C. Monoisotopic mass values were determined at a resolution >35,000 and an accuracy <5 ppm. Spectra were acquired and deconvoluted with the Mass-Lynx MaxEnt 1 4.1 software (Waters).
Chemical synthesis
The peptide substrates (FTRLQSLEN and FTRLRSLEN) were synthesized by the manual solid-phase method using the 9-fluorenylmethyloxycarbonyl (Fmoc)/t-butyl (tBu) strategy55. After peptide chain assembly, resin cleavage and removal of side-chian protecting groups, the crude peptides were fractionated by RP-HPLC on a (4.6 × 250 mm, 5 μm granulometry, 300 Å pores size) C18 analytical column (Grace-Vydac, Hesperia, CA, USA), equilibrated with 0.1% (v/v) aqueous TFA and eluted with a linear acetonitrile-0.078% (v/v) TFA gradient. The peptide material eluted in correspondence with the major chromatographic peaks was collected, lyophilized, and analyzed by HR-MS, yielding mass values in agreement with the theoretical mass within 2 ppm accuracy.
Peptide samples (10 µM) in HBS were incubated at 37 °C in the presence of rcMpro, at a protease:peptide molar ratio of 1:40. At given time points (0 min, 1 h, 2 h, 4 h, 7 h, 16 h) aliquots (5 pmol) were taken, acid quenched with 0.1% (v/v) aqueous TFA, and then analyzed by LC-MS. Quantitative determination of intact peptides and proteolytic fragments (FTRLQSLEN, FTRLQ, SLEN, FTRLRSLEN, FTRLR, SLEN) was performed by integrating the area under the chromatographic peaks, using the Jasco ChromNav 2.0 software.
Inhibition of Mpro proteolytic activity by thrombin inhibitors Argatroban and PPACK
The inhibition of rcMpro by thrombin and FXa inhibitors Argatroban or PPACK was measured through an in vitro fluorogenic assay, using the fluorogenic rcMpro substrate 5-FAM-AVLQSGFRK(DABCYL)K. rcMpro (50 nM) was incubated with Argatroban or PPACK (7 μM) in HBS-PEG for 2 h, at 37 °C. After the addition of 5-FAM-AVLQSGFRK(DABCYL)K (1.25 μM), the emission of the released peptidyl 5-FAM fluorophore was measured at 530 nm. The residual proteolytic activity of rcMpro was determined as the ratio vi/v0, where vi and v0 are the initial velocities of Mpro-catalyzed substrate hydrolysis in the presence or in the absence of the inhibitor.
To calculate the inhibition constant (Ki) of PPACK, Mpro residual activity was measured at increasing concentrations of PPACK after 10-min incubation at 37 °C. The ratio vi/v0 was plotted as a function of PPACK concentration, and data points were fitted with Eq. (1), describing the slow-tight binding inhibition model78 to yield the apparent inhibition constant (Kiapp):
where E is the rcMpro concentration, and I is the PPACK concentration. The equilibrium inhibition constant (Ki) was then derived from Kiapp using Eq. (2) according to the competitive inhibition model:
where S is the substrate concentration, and KM is the Michaelis–Menten constant of rcMpro for 5-FAM-AVLQSGFRK(DABCYL)K, calculated as 20 μM.
Computational methods
Molecular docking
Molecular docking was performed with the HPEPDOCK web server79, starting from the inhibitor-free Mpro structure, after removal of water molecules (6y2e.pdb)19 and the (D)-Phe-Pro-Arg peptide, as the peptidyl moiety of the irreversible inhibitor PPACK. The software generated 3D structure models for the given sequences of peptides using the implemented MOPEP software, which considers peptide flexibility. Simulations were run with default parameters. One hundred poses were generated and ranked according to the CAPRI criteria80. The pose with the most favorable energy scores was selected and used for data analysis.
Electrostatics
Electrostatic potential calculations were performed using the APBS software run on the crystallographic structure of Mpro (6y2e.pdb)19. A solvent dielectric constant of 78.14 and a protein dielectric constant of 2.0 at 310 K in 150 mM NaCl were used. Final electrostatic maps were constructed by subtracting the protein self-energies from the calculated map using the DXMATH utility in APBS. Images were generated with PyMOL vs. 1.3 (DeLano Scientific, San Diego, CA, USA)
HDX-MS Local analysis
Sample preparation and LC-MS
HDX-MS experiment was performed following the procedure described by Acqusaliente et al.55. Briefly, samples were acquired in triplicate using a Waters Xevo G2-S mass spectrometer (Waters, Milford, MA, USA) equipped with an ACQUITY UPLC M-class chromatographic system (Waters). All sample handling was carried out using a Leap HDX PAL autosampler (Leap Technologies, Carrboro, NC, USA). rcMpro (25 μM) was incubated alone or with PPACK (2 mM) in PBS for 30 min at 20 °C. At each incubation time, deuterium labeling was initiated by a 20-fold dilution of the sample with D2O (95% D2O in 20 mM sodium phosphate, pH 7.16, 150 mM NaCl). At given time points (i.e., 15 s, 1 min, 5 min, 30 min, 2 h) the labeling reaction was quenched at 1°C by addition of an equal volume of quenching buffer (200 mM sodium phosphate, 1.5 M guanidine hydrochloride (Gdn-HCl), 1 mM Tris(2-carboxyethyl)phosphine hydrochloride (TCEP), adjusted to pH 1.99) to reach a final pH of 2.46. The pH and pD values were measured at 25 °C, using a Mettler-Toledo (Columbus, OH, USA) model. FiveEasy Plus pH-meter. Quenched samples were immediately injected into the LC system for online pepsin digestion using an in-house prepared immobilized pepsin column, thermostated at 15 °C and equilibrated with an isocratic flow of 0.23% (v/v) formic acid in water at a flow rate of 100μl/min. The resulting peptic fragments were online trapped on an Acquity UPLC BEH C18 VanGuard Pre-column (130 Å, 1.7 µm, 2.1 × 5 mm) and separated using an Acquity UPLC BEH C18 column (130 Å, 1.7 µm, 1 × 100 mm) with a linear acetonitrile-0.1% (v/v) formic acid gradient from 5% to 60% in 10 min, at a flow rate of 60 μl/min. Mass spectra were acquired in the resolution mode (m/z range 50–2000). Unlabeled proteins were prepared as reference samples in the same manner as those that were labeled with deuterium. Each sample was prepared in triplicate.
Data analysis
Peptides that were generated from online pepsin digestion were identified from the unlabeled protein samples using the Waters Protein Lynx Global Server 3.0. Only those fragments matching the following criteria were considered: (i) a length between 4 and 33 amino acids, (ii) at least 2 ion products identified, (iii) minimum products per amino acid of 0.2, (iv) maximum MH+ error tolerance of 6 ppm, and (v) the presence of the peptide in at least two of the three peptide identification runs. Those peptides meeting the filtering criteria were further processed by DynamX 3.0 (Waters) to calculate the relative amount of deuteration. Deuterium uptake was obtained by subtracting the centroid mass of the undeuterated form of each peptide from the deuterated form, at each time point, for each condition. Because the relative deuterium uptake of individual peptides was compared, no back-exchange correction was performed.
A Welch’s t-test was used to compare the mean uptake of each peptide between the two protein states ([Mpro + PPACK] vs [Mpro]) at each exposure time, and the p value was FDR-adjusted using Benjamini–Hochberg (BH) correction. A peptide is considered to have significant differences in deuterium uptake between the two states if the adjusted p value is <0.05.
Identification of Mpro cleavages on coagulation factors
The identification of cleavages on coagulation factors was performed using the TAILS protocol, which labels the neo-N-termini generated by the protease activity59.
Sample preparation
Purified commercial human coagulation factors FVII or FXII (4 μg) were incubated with rcMpro (1:10 E:S molar ratio) or with a mock treatment in HBS for 3 h at 37 ± 0.1 °C. After proteolysis, the reactions were terminated by heat-inactivation of the protease at 99°C for 5 min. Once cooled down, the samples were denatured with Gdn-HCl (4 M final concentration) for 15 min at 37 °C, reduced with TCEP for 30 min at 37 °C, and alkylated with iodoacetamide (IAA) for 30 min at room temperature in the dark. Next, we proceeded with reductive di-methylation of free N-termini by adapting the TAILS protocol59. Briefly, samples were incubated with 20 mM formaldehyde and 10 mM sodium cyanoborohydride (NaCNBH3) for 16 h at 37 °C. The reaction was quenched by 1:5 dilution in 50 mM ammonium bicarbonate, pH 7.8, for 4 h at r.t., and the labeled protein fragments were digested with trypsin (1:50 ratio (w/w)) for 16 h at 37 °C. Tryptic digestion was quenched by acidification with 0.5% formic acid, and the samples were desalted following the StageTips C18 protocol (washing step, 5% acetonitrile containing 0.1% formic acid; elution step, 50% acetonitrile containing 0.1% formic acid). Dried peptides were resuspended in 0.1% formic acid (15 μl). Identification and quantification of peptides were performed by liquid chromatography tandem mass spectrometry (LC-MS/MS) operating in DDA mode or in PRM. Samples were prepared in triplicate.
DDA analysis
LC-MS/MS was performed on an Orbitrap Eclipse Tribrid mass spectrometer (Thermo Fisher) coupled to an EASY-nLC2002 (Thermo Fisher) liquid chromatography system. Peptides were separated using a reverse phase column (75 μm × 40 mm New Objective, in-house packed with ReproSil Gold 120 C18, 1.9 μm, Dr. Maisch GmbH) across 60-min linear gradient from 3 to 30% (buffer A: 0.1% [v/v] formic acid; buffer B: 0.1% [v/v] formic acid, 80% [v/v] acetonitrile). The DDA data acquisition mode was set to perform one MS1 scan and multiple MS2 scans in a cycle time of 3 s with MS1 identification (R = 120,000 at 400 m/z, AGC = 200% and maximum IT = 100 ms, scan range 350–1400), HCD fragmentation (NCE = 30%), isolation windows (1.2 m/z) and MS2 identification (R = 30,000 at 400 m/z, AGC = 200% and maximum IT = 54 ms, scan range 150–1200). Charge states lower than two and higher than seven were rejected. Acquired spectra were searched using the SpectroMine software (Biognosys, Zurich, Switzerland) against the FVII sequence or the FXII sequence plus contaminants. The search was performed with the default parameters, changing the digestion specificity to semi-tryptic and adding Lys and N-terminal di-methylation (+29.039125) as a variable modification. The maximum number of modifications per peptide was increased to 3. False discovery rate of <1% was used at the peptide level. Only N-terminal di-methylated peptides not identified in any of the mock treatment replicates and identified in at least 2/3 replicates of the rcMpro-treated replicates were considered in the analysis.
PRM analysis
LC-MS/MS was performed on an Orbitrap Exploris 480 mass spectrometer (Thermo Fisher) coupled to a Vanquish Neo liquid chromatography system (Thermo Fisher). Peptides were separated using a reversed phase column (75 μm × 400 mm New Objective, in-house packed with ReproSil Gold 120 C18, 1.9 μm, Dr. Maisch GmbH) across 60 min linear gradient from 7 to 35% (buffer A: 0.1% [v/v] formic acid; buffer B: 0.1% [v/v] formic acid, 80% [v/v] acetonitrile). MS acquisition of targeted peptide was set up with the combination of one MS1 untargeted scan and MS2 scheduled targeted scan using an isolation window of 2.0 m/z and HCD fragmentation. Manual curated analysis of fragments was performed using Skyline daily81. Data analysis of PRM data was performed from the integration of the top five most intense fragments and transformed log2 values.
Statistics and reproducibility
The probability of clotting induced by Mpro was estimated on a total of 21 different plasma samples from healthy subjects, which were incubated both with and without Mpro. To assess if Mpro significantly increases the probability of a clotting event, we performed a survival analysis using the Kaplan–Meier method. A p values ≤ 0.05 was considered statistically significant.
The comparison between Mpro treatment and mock control in the enzymatic assays was performed using the Kruskal–Wallis test. Sample size: for FII, FVII, FIX, FXI, FXII n = 3, for Fbg, FX, FIIa+ATIII n = 2. P values ≤ 0.05 were considered statistically significant.
In the HTPS experiment, the specificity of Mpro and control proteases was assessed in three independent experiments. A two-sided paired t-test was employed to evaluate similarity between frequency matrices of different conditions.
In the HDX-MS analysis, the mean uptake of each peptide between the two protein states ([Mpro + PPACK] vs [Mpro]) at each exposure time was compared using a Welch’s t-test, and p values were FDR-adjusted using BH correction. An adjusted p value of ≤0.05 was considered statistically significant.
The abundance of the peptide generated by Mpro cleavage on FVII was compared across three independent experiments using a two-sided paired t-test.
Unless indicated, all experiments were carried out in three independent replicates.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw proteomics data scripts and generated table are deposited to ProteomeXchange Consortium via the PRIDE partner repository with identifiers PXD044733 (Characterization of Mpro substrate specificity by HTPS) and PXD046242 (Identification of direct cleavages on FVII and FXII by Mpro). HDX dataset is publicly available on the ETHZ research collection (Characterization of the binding of PPACK to Mpro by HDX-MS, https://www.research-collection.ethz.ch/handle/20.500.11850/637797). Source data are provided with this paper as Supplementary Data S5. All protein structures referred to in this study were obtained from PDB (https://www.rcsb.org/), with the accession codes [6y2e.pdb].
References
Beristain-Covarrubias, N. et al. Understanding infection-induced thrombosis: lessons learned from animal models. Front. Immunol. 10, 2569 (2019).
Nigade, P. P., Dhanagar, S. S. & Nikam, V. S. Venous thromboembolism in infectious diseases. Comp. Clin. Pathol. https://doi.org/10.1007/s00580-025-03657-y (2025).
Turnic, T. N. et al. Bradykinin and galectin-3 in survived and deceased patients with COVID-19 pneumonia: an increasingly promising biochemical target. Oxid. Med. Cell. Longev. 2022, 7920915 (2022).
Giannis, D., Douketis, J. D. & Spyropoulos, A. C. Anticoagulant therapy for COVID-19: What we have learned and what are the unanswered questions?. Eur. J. Intern. Med. 96, 13–16 (2022).
Gorog, D. A. et al. Current and novel biomarkers of thrombotic risk in COVID-19: a consensus statement from the International COVID-19 Thrombosis Biomarkers Colloquium. Nat. Rev. Cardiol. 19, 475–495 (2022).
Conway, E. M. et al. Understanding COVID-19-associated coagulopathy. Nat. Rev. Immunol. 22, 639–649 (2022).
Loo, J., Spittle, D. A. & Newnham, M. COVID-19, immunothrombosis and venous thromboembolism: biological mechanisms. Thorax 76, 412–420 (2021).
Ryu, J. K. et al. Fibrin drives thromboinflammation and neuropathology in COVID-19. Nature 633, 905–913 (2024).
Zheng, Y.-Y., Ma, Y.-T., Zhang, J.-Y. & Xie, X. COVID-19 and the cardiovascular system. Nat. Rev. Cardiol. 17, 259–260 (2020).
Lodigiani, C. et al. Venous and arterial thromboembolic complications in COVID-19 patients admitted to an academic hospital in Milan, Italy. Thromb. Res. 191, 9–14 (2020).
Helms, J. et al. High risk of thrombosis in patients with severe SARS-CoV-2 infection: a multicenter prospective cohort study. Intensive Care Med. 46, 1089–1098 (2020).
Llitjos, J.-F. et al. High incidence of venous thromboembolic events in anticoagulated severe COVID-19 patients. J. Thromb. Haemost.18, 1743–1746 (2020).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Jose, R. J. & Manuel, A. COVID-19 cytokine storm: the interplay between inflammation and coagulation. Lancet Respir. Med. 8, e46–e47 (2020).
Wolf, A. et al. The mechanistic basis linking cytokine storm to thrombosis in COVID-19. Thromb. Update 8, 100110 (2022).
Esmon, C. T. Possible involvement of cytokines in diffuse intravascular coagulation and thrombosis. Baillieres Best Pract. Res. Clin. Haematol. 12, 343–359 (1999).
Stojanovski, B. M. & Di Cera, E. Comparative sequence analysis of vitamin K-dependent coagulation factors. J. Thromb. Haemost. 20, 2837–2849 (2022).
Malone, B., Urakova, N., Snijder, E. J. & Campbell, E. A. Structures and functions of coronavirus replication–transcription complexes and their relevance for SARS-CoV-2 drug design. Nat. Rev. Mol. Cell Biol. 23, 21–39 (2022).
Zhang, L. et al. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science 368, 409–412 (2020).
Fang, S., Shen, H., Wang, J., Tay, F. P. L. & Liu, D. X. Functional and genetic studies of the substrate specificity of coronavirus infectious bronchitis virus 3C-like proteinase. J. Virol. 84, 7325–7336 (2010).
Shaqra, A. M. et al. Defining the substrate envelope of SARS-CoV-2 main protease to predict and avoid drug resistance. Nat. Commun. 13, 3556 (2022).
Zhao, Y. et al. Structural basis for replicase polyprotein cleavage and substrate specificity of main protease from SARS-CoV-2. Proc. Natl. Acad. Sci. USA 119, e2117142119 (2022).
Rut, W. et al. SARS-CoV-2 Mpro inhibitors and activity-based probes for patient-sample imaging. Nat. Chem. Biol. 17, 222–228 (2021).
Koudelka, T. et al. N-terminomics for the identification of in vitro substrates and cleavage site specificity of the SARS-CoV-2 main protease. Proteomics 21, e2000246 (2021).
Dai, W. et al. Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science 368, 1331–1335 (2020).
Owen, D. R. et al. An oral SARS-CoV-2 Mpro inhibitor clinical candidate for the treatment of COVID-19. Science 374, 1586–1593 (2021).
Pablos, I. et al. Mechanistic insights into COVID-19 by global analysis of the SARS-CoV-2 3CLpro substrate degradome. Cell Rep. 37, 109892 (2021).
Meyer, B. et al. Characterising proteolysis during SARS-CoV-2 infection identifies viral cleavage sites and cellular targets with therapeutic potential. Nat. Commun. 12, 5553 (2021).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Zhou, Y. et al. A comprehensive SARS-CoV-2–human protein–protein interactome reveals COVID-19 pathobiology and potential host therapeutic targets. Nat. Biotechnol. 41, 128–139 (2023).
Liu, X. et al. SARS-CoV-2–host proteome interactions for antiviral drug discovery. Mol. Syst. Biol. 17, e10396 (2021).
Martínez-Fleta, P. et al. SARS-CoV-2 cysteine-like protease antibodies can be detected in serum and saliva of COVID-19-seropositive individuals. J. Immunol.205, 3130–3140 (2020).
Jiang, H.-W. et al. SARS-CoV-2 proteome microarray for global profiling of COVID-19 specific IgG and IgM responses. Nat. Commun. 11, 3581 (2020).
Wang, W. et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 323, 1843–1844 (2020).
Andersson, M. I. et al. SARS-CoV-2 RNA detected in blood products from patients with COVID-19 is not associated with infectious virus. Wellcome Open Res. 5, 181 (2020).
Grin, P. M. et al. SARS-CoV-2 3CLpro (main protease) regulates caspase activation of gasdermin-D/E pores leading to secretion and extracellular activity of 3CLpro. Cell Rep. 43, 115080 (2024).
Coban, A., Bornberg-Bauer, E. & Kemena, C. Domain evolution of vertebrate blood coagulation cascade proteins. J. Mol. Evol. 90, 418–428 (2022).
Uliana, F. et al. Mapping specificity, cleavage entropy, allosteric changes and substrates of blood proteases in a high-throughput screen. Nat. Commun. 12, 1693 (2021).
Hoffman, M. Remodeling the blood coagulation cascade. J. Thromb. Thrombolysis 16, 17–20 (2003).
Pontarollo, G. et al. Non-canonical proteolytic activation of human prothrombin by subtilisin from Bacillus subtilis may shift the procoagulant–anticoagulant equilibrium toward thrombosis. J. Biol. Chem. 292, 15161–15179 (2017).
Farkas, ÁZ. et al. Structure, mechanical, and lytic stability of fibrin and plasma coagulum generated by Staphylocoagulase from Staphylococcus aureus. Front. Immunol. 10, 2967 (2019).
Bouck, E. G., Zunica, E. R. & Nieman, M. T. Optimizing the presentation of bleeding and thrombosis data: responding to censored data using Kaplan-Meier curves. Thromb. Res. 158, 154–156 (2017).
Bernardi, F. & Mariani, G. Biochemical, molecular and clinical aspects of coagulation factor VII and its role in hemostasis and thrombosis. Haematologica 106, 351–362 (2021).
Björk, I. & Lindahl, U. Mechanism of the anticoagulant action of heparin. Mol. Cell. Biochem. 48, 161–182 (1982).
Konrath, S., Mailer, R. K. & Renné, T. Mechanism, functions, and diagnostic relevance of FXII activation by foreign surfaces. Hamostaseologie 41, 489–501 (2021).
Ke, K. & Morrissey, J. The activation of factor VII by a variety of potential activators. FASEB J. 27, 6–995.6 (2013).
Naudin, C., Burillo, E., Blankenberg, S., Butler, L. & Renné, T. Factor XII contact activation. Semin. Thromb. Hemost. 43, 814–826 (2017).
Kenne, E. & Renné, T. Factor XII: a drug target for safe interference with thrombosis and inflammation. Drug Discov. Today 19, 1459–1464 (2014).
Cesar Ramos de Jesus, H. et al. Optimization of quenched fluorescent peptide substrates of SARS-CoV-2 3CLpro main protease (Mpro) from proteomic identification of P6—P6’ active site specificity. J. Virol. 98, e00049–24 (2024).
Luo, S. Y. et al. Identification of human host substrates of the SARS-CoV-2 Mpro and PLpro using subtiligase N-terminomics. ACS Infect. Dis. 9, 749–761 (2023).
Scott, B. M., Lacasse, V., Blom, D. G., Tonner, P. D. & Blom, N. S. Predicted coronavirus Nsp5 protease cleavage sites in the human proteome. BMC Genom. Data 23, 25 (2022).
Biembengut, ÍV., de Souza, T. & de, A. C. B. Coagulation modifiers targeting SARS-CoV-2 main protease Mpro for COVID-19 treatment: an in silico approach. Mem. Inst. Oswaldo Cruz 115, 1–4 (2020).
Anand, K. et al. Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain. EMBO J. 21, 3213–3224 (2002).
Kovach, I. M., Kelley, P., Eddy, C., Jordan, F. & Baykal, A. Proton Bridging in the Interactions of Thrombin with Small Inhibitors. Biochemistry 48, 7296–7304 (2009).
Acquasaliente, L., Pierangelini, A., Pagotto, A., Pozzi, N. & De Filippis, V. From haemadin to haemanorm: synthesis and characterization of full-length haemadin from the leech haemadipsa sylvestris and of a novel bivalent, highly potent thrombin inhibitor (haemanorm). Protein Sci. e4825. https://doi.org/10.1002/pro.4825 (2023).
Engen, J. R. & Komives, E. A. Complementarity of hydrogen/deuterium exchange mass spectrometry and cryo-electron microscopy. Trends Biochem. Sci. 45, 906–918 (2020).
Masson, G. R. et al. Recommendations for performing, interpreting and reporting hydrogen deuterium exchange mass spectrometry (HDX-MS) experiments. Nat. Methods 16, 595–602 (2019).
Konermann, L., Vahidi, S. & Sowole, M. A. Mass spectrometry methods for studying structure and dynamics of biological macromolecules. Anal. Chem. 86, 213–232 (2014).
Kleifeld, O. et al. Identifying and quantifying proteolytic events and the natural N terminome by Terminal Amine Isotopic Labeling of Substrates. Nat. Protoc. 6, 1578–1611 (2011).
Augustsson, C. & Persson, E. In vitro evidence of a tissue factor-independent mode of action of recombinant factor VIIa in hemophilia. Blood 124, 3172–3174 (2014).
Patmore, S., Dhami, S. P. S. & O’Sullivan, J. M. Von Willebrand factor and cancer; metastasis and coagulopathies. J. Thromb. Haemost. 18, 2444–2456 (2020).
Sokolov, A. V. et al. Thrombin inhibits the anti-myeloperoxidase and ferroxidase functions of ceruloplasmin: relevance in rheumatoid arthritis. Free Radic. Biol. Med. 86, 279–294 (2015).
Meroni, P. L., Borghi, M. O., Raschi, E. & Tedesco, F. Pathogenesis of antiphospholipid syndrome: understanding the antibodies. Nat. Rev. Rheumatol. 7, 330–339 (2011).
Pozzi, N. et al. β2-Glycoprotein I binds to thrombin and selectively inhibits the enzyme procoagulant functions. J. Thromb. Haemost. 11, 1093–1102 (2013).
Bever, K. M. et al. Risk factors for venous thromboembolism in immunoglobulin light chain amyloidosis. Haematologica 101, 86–90 (2016).
Acquasaliente, L. et al. Exogenous human α-Synuclein acts in vitro as a mild platelet antiaggregant inhibiting α-thrombin-induced platelet activation. Sci. Rep. 12, 9880 (2022).
Loof, T. G. et al. Staphylococcus aureus-induced clotting of plasma is an immune evasion mechanism for persistence within the fibrin network. Microbiology 161, 621–627 (2015).
Iba, T., Levy, J. H., Levi, M. & Thachil, J. Coagulopathy in COVID-19. J. Thromb. Haemost. 18, 2103–2109 (2020).
Lancellotti, S. et al. Oxidized von Willebrand factor is efficiently cleaved by serine proteases from primary granules of leukocytes: divergence from ADAMTS-13. J. Thromb. Haemost. 9, 1620–1627 (2011).
Yong, J. & Toh, C.-H. The convergent model of coagulation. J. Thromb. Haemost. 22, 2140–2146 (2024).
Hofman, Z., de Maat, S., Hack, C. E. & Maas, C. Bradykinin: inflammatory product of the coagulation system. Clin. Rev. Allergy Immunol. 51, 152–161 (2016).
Fornasier, E. et al. A new inactive conformation of SARS-CoV-2 main protease. Acta Crystallogr. Sect. Struct. Biol. 78, 363–378 (2022).
Adcock, D. M. et al. H21-A5 Collection, Transport, and Processing of Blood Specimens for Testing Plasma-Based Coagulation Assays and Molecular Hemostasis Assays: Approved Guideline 5th edn (Clinical and Laboratory Standards Institute, 2008).
Mui, B., Chow, L. & Hope, M. J. Extrusion technique to generate liposomes of defined size. Methods Enzymol. 367, 3–14 (2003).
Therneau, T. A package for survival analysis in R. R package version 3.8-3, https://CRAN.R-project.org/package=survival (2024).
Kassambara, A., Kosinski, M. & Biecek, P. survminer: drawing survival curves using ‘ggplot2’. 0.4.9. https://doi.org/10.32614/CRAN.package.survminer (2016).
Wiśniewski, J. R. Filter-aided sample preparation for proteome analysis. Methods Mol. Biol.1841, 3–10 (2018).
Copeland, A. R. Enzymes. A Practical Introduction to Structure, Mechanism, and Data Analysis. 2nd edn. 122–123 (John Wiley & Sons, USA, 2000).
Zhou, P., Jin, B., Li, H. & Huang, S.-Y. HPEPDOCK: a web server for blind peptide-protein docking based on a hierarchical algorithm. Nucleic Acids Res. 46, W443–W450 (2018).
Lensink, M. F. & Wodak, S. J. Score_set: a CAPRI benchmark for scoring protein complexes. Proteins 82, 3163–3169 (2014).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Acknowledgements
This work was supported by a Grant from the CaRiPaRo Foundation Excellence Research Project—BPiTA n. 52012 to V.D.F., Italian Ministry of University and Research (MIUR)—PRIN-2022 n. 2022ZSA2JP to P.S., Marie Curie Individual Fellowship (Grant agreement number 101207537, F.U.) We would like to thank Prof. Dr. Christofer M. Overall for sharing the N-terminomics data on Mpro specificity and for offering his expert insights on protease specificity characterization. The authors are grateful to Prof. Roberto Battistutta for the generous gift of the plasmid containing the DNA sequence of the inactive protease mutant.
Author information
Authors and Affiliations
Contributions
V.D.F. conceived and coordinated the work; A.P., G.N., A.P.I., E.C., and L.A. performed the biochemical work; F.U. and A.P. performed proteomic and mass spectrometry analyses; M.B., M.L.M., and B.G. provided in-house produced recombinant Mpro; G.D.S., P.M., and P.S. provided plasma samples; D.G. performed power analysis for the clotting experiments; P.P. supervised the TAILS tasks; V.D.F., A.P., and F.U. designed research and jointly wrote the manuscript; all authors analyzed and interpreted the data, reviewed and approved the final content of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Enrico Di Cera, Oliver Crook, and Leandro Neves for their contribution to the peer review of this work. Primary Handling Editors: Mark Collins and Laura Rodríguez Pérez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pagotto, A., Uliana, F., Cavedon, E. et al. The main protease (Mpro) from SARS-CoV-2 triggers plasma clotting in vitro by activating coagulation factors VII and FXII. Commun Biol 8, 1145 (2025). https://doi.org/10.1038/s42003-025-08570-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-08570-2