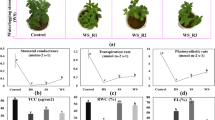
Abstract
Scenedesmus obliquus UTEX393 is a promising microalgal candidate for sustainable biomanufacturing but its limited halotolerance hinders large-scale cultivation in saline environments. To investigate the molecular basis of salt stress responses, we conducted a comprehensive multi-omic analysis integrating genomics, transcriptomics, proteomics, lipidomics, metabolomics, and DNA affinity purification sequencing (DAP-seq). An improved nuclear genome assembly and annotation yielded 19,017 gene models and a 97% BUSCO completeness score, enabling construction of a genome-scale metabolic model. Comparing 15 ppt salinity stress to 5 ppt control, growth and productivity were significantly reduced, accompanied by widespread transcriptomic and proteomic changes. Transcriptomic analysis revealed downregulation of photosynthetic machinery and energy conservation genes, and upregulation of stress-responsive elements such as expansins, flavodoxins, and osmoprotectants. Lipidomic profiling showed accumulation of triacylglycerols (TAGs) and degradation of galactosyl lipids, consistent with a shift toward lipid biosynthesis to mitigate redox imbalance. Depletion of key polar metabolites and branched-chain amino acids suggested a rerouting of central carbon metabolism under stress. DAP-seq identified key transcription factors, including LHY1 and SPL12, that target central metabolic enzymes involved in redox balancing, such as glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and malate dehydrogenase (MDH). These findings establish a regulatory-metabolic framework linking redox stress to lipid accumulation and reveal potential engineering targets to enhance salt tolerance. Overall, the multi-omic analysis supports the “overflow” hypothesis, where impaired photosynthesis results in excess reducing equivalents being diverted into TAG synthesis and highlights transcriptional regulators as candidates for improving algal robustness in brackish environments.
Similar content being viewed by others
Introduction
Green algae are being developed as chassis for biomanufacturing as they are useful as feedstocks to produce fuel and valuable chemical products1. These algae are also core components of bioremediation efforts for acid mine drainage2 and can be a source of high value nutraceutical and pharmaceutical compounds3. Given the promise of algae, bioprospecting and screening have identified numerous candidate production strains, including Scenedesmus4,5,6,7. The Scenedesmus genus contains coccoid green alga belonging to the Chlorophyceae, order Sphaeropleales. Recent phylogenetic assessments have resulted in the renaming of the genus Scenedesmus, thus this species is often synonymously referred to as Acutodesmus obliquus or Tetradesmus obliquus8,9. Overall, the Scenedesmus are highly productive with biomass compositions that are suitable as a source of both food and fuel, and are also effective at removing nutrients and other contaminants from wastewater10,11. One notable species; Scenedesmus obliquus UTEX393 (originally known as “D-3”), was a model organism used in early studies of photosynthesis, and in investigations that informed the discovery of the Calvin-Benson-Bassham cycle12.
Enhancing the salt tolerance of S. obliquus UTEX393 would significantly improve its industrial potential by enabling cultivation in non-potable, brackish water, thereby reducing resource costs and improving sustainability13. Although some halotolerant Scenedesmus strains exist, including Scenedesmus sp. NREL 46B-D314 and Scenedesmus sp. IITRIND215, most Scenedesmus species achieve optimal growth in freshwater or low salinity environments16,17,18 S. obliquus UTEX393 specifically demonstrates maximum growth at 5 parts per thousand (ppt) salinity, with growth rates declining by 35% at 15 ppt and growth is completely inhibited at 35 ppt (typical seawater salinity)11.
Previous studies have documented various molecular and physiological responses to salinity stress in some freshwater Scenedesmus and in more distantly related Chlorophytes. As reviewed in Shetty19, cyclophilins, photosynthetic ferredoxins, and glutathione peroxidases have been shown to increase salt tolerance in algae. In salt-stressed Chlamydomonas reinhardtii, electron transport and photochemical quenching were reduced, while non-photochemical quenching was increased20. In Dunaliella parva (a green halophilic algae) intracellular osmolytes, specifically glycerol are elevated and when salt shocked, Dunaliella salina induces the expression of carotenoid synthesis and reactive oxygen species scavenging enzymes21. Within the genus Scenedesmus, Scenedesmus obliquus HTB1 exhibited pigment deterioration22; while salinity stress induced lipid accumulation, Photosystem II disruption, and reactive oxygen species in Scenedesmus sp. BHU123
Despite these advances, the specific physiological responses and underlying systems-level ‘omics responses to saline environments remain largely unexplored for S. obliquus UTEX393. This knowledge gap creates a critical bottleneck in our ability to genetically enhance halotolerance in UTEX393 and in other industrially relevant alga. By systematically characterizing how this high-performing freshwater strain responds to salinity stress, we can lay the groundwork to genetically engineer strategies to maintain high levels of productivity in saline-rich cultivation schemes and potentially exploit salinity-induced responses to enhance valuable harvest end-products like triacylglyerol (TAG) biofuel precursors.
To understand (and potentially enhance) the physiological responses during salt stress in S. obliquus UTEX393, we improved the quality of the genome assembly and annotation and performed a comprehensive multi-omics study to characterize the transcriptomic, proteomic, and metabolomic responses under brackish conditions. Collectively, this study enabled the identification of specific physiological responses that limit halotolerance of UTEX393. Further, specific transcription factors were identified that may regulate UTEX393 environmental responses and their regulatory effects were characterized using DNA Affinity Purification and sequencing (DAP-seq)24. We report a significant disruption of photosynthetic components and indicators of redox dysregulation that may be causative of an “overflow” response wherein the excess reductive equivalents are used to fuel TAG synthesis. This redox imbalance and the resulting metabolic responses may be valuable to increase TAG content, however, the observed salinity response failed to allow for productive growth of S. obliquus UTEX393 under modest salt concentrations.
Results
Genome assembly, annotation, and comparative analysis
The genome assembly of S. obliquus UTEX393 was improved from the previously published version25 by merging multiple assemblies that originated from three independent Illumina assemblies and one PacBio assembly. The improved genome eliminated 103 gaps, resulting in a 3.9-fold reduction in contigs, a 13.8-fold improvement in scaffold L50, along with a concomitant increase in maximum scaffold sizes (Table 1). While the current assembly is similar in length (107.86 Mbp and 107.54 Mbp with and without organellar contigs, respectively), there is an increase in captured repetitive content with 20.9 Mbp of repeat coverage length, compared to 17.8 Mbp in the prior version. Additionally, organellar DNA content was removed to provide a standardized nuclear genome. Annotation of the new genome assembly resulted in 19,017 predicted genes and the completeness of the genome annotation based on BUSCO (chlorophyta_odb10) is 97%, a marked improvement compared to the prior version (93%) (Fig. 1A). The number of unique Pfams increased to 3508 domains compared to 3,402 domains and, of the ~3000 gene models that were identified as unique to the newer annotation based on MCL clustering of protein sequences26, over 85% were supported by RNA-seq, but only 9% were annotated with Pfam domains. 1229 gene models annotated as transposable elements or repeats in the previous annotation have been removed. To further enhance the strength of the genome, we leveraged a combination of homology-based approaches (see “Methods” for details) along with multiple rounds of manual curation to annotate a total of 4394 proteins with specific enzymatic activities. These proteins span a total of 1931 different Enzyme Commission numbers. After compartmentalization using the information about the predicted localization of enzymes into various cellular compartments and based on previous published algal models, the addition of transport and gap filling reactions, we successfully constructed a metabolic model that included a total of 3433 compounds and 5,592 reactions.
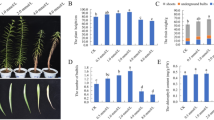
A BUSCO completeness assessed in protein mode using the chlorophyta_odb10 set. Percent of complete orthologs (single-copy and duplicated) is shown by genome. The first two rows show completeness of the latest annotation of S. obliquus UTEX393 compared to the annotation of the previously published version (Carreres et al., Genome Announcements, 2017; Carreres et al., Algal Research, 2019). B The phylogenetic tree inferred by the Orthology matrix analysis (OMA) based on hierarchical gene groups (HOGs). Orthologous matrix analysis (OMA) was performed using the filtered protein data for Micromonas pusilla CCMP1545 (MICPC), Micromonas commoda (MICCC), Ostreococcus lucimarinus (OSTLU), Ostreococcus tauri (OSTTA), Chlorella variabilis (CHLVA), Volvox carteri (VOLCA), Arabidopsis thaliana (ARATH), Chlamydomonas reinhardtii (CHLRE), Scenedesmus sp. 46B-D3 (SoblNREL46B3), Scenedesmus obliquus DOE152Z/DOE3031 (SoblDOE152Z), Scenedesmus obliquus UTEX393 (SoblUTEX393), Monoraphidium minutum 26BAM (Mmin26BAM), Raphidocelis subcapitata (RAPSUB35), Chromochloris zofingeiensis SAG211 (CzofSAG211), and Picochlorum renovo (Pren). C HOGs present in Scenedesmus obliquus UTEX393 that represent the number of inferred common ancestral genes shared within each set of species/strains. D the amino acid proportions of the NREL46B-D3 and UTEX393 proteins present in the 1464 HOGs present only in the Scenedesmus lineages and present in both strains (the second two bars of the upset plot). Significance is denoted by “*” on the plot (Bonferroni adjusted P-value ≤ 0.05).
Using the improved UTEX393 genome annotation, an orthologous protein matrix analysis (OMA)27 was performed to identify the conserved and unique functional gene groups within the Scenedesmaceae and other related species (Fig. 1C). OMA identified 27,066 hierarchical orthologous groups (HOGs) of genes—groups of genes that indicate a shared ancestral differentiation in protein sequence distinguishing them from orthologs in the broader tree. While not representative of a uniquely evolved function (given one HOG may be nested into a larger HOG along taxonomic lineages), they do catalog sets of proteins that are likely to differentiate from an inferred common ancestral gene. These HOGs indicate that Scenedesmus obliquus UTEX393 shares the most inferred common ancestor genes with Scenedesmus obliquus DOE152Z28 (UTEX3031) with an additional cohort of inferred common ancestor genes shared with the other members of the genus Scenedesmus. To our knowledge, amino acid utilization differences between halotolerant and non-halotolerant species have not been reported in green algae. However, the convergence of amino acid utilization in halotolerant eubacterial and archaebacterial lineages29 as well as the altered amino acid utilization of acidic/basic/hydrophobic amino acids in halophilic bacterium Salinibacter ruber indicates optimized amino acid utilization may be an adaptive advantage for halotolerant species30. Understanding the amino acid changes from the inferred common ancestor(s) with known halotolerance (Scenedesmus sp. NREL 46B-D3) may provide a basis for optimizing protein sequences in UTEX393 for increased osmotolerance. In fact, when examining the proportion of amino acids in the 1,464 HOGs present only in the Scenedesmus lineages and present in both strains (779 also present in DOE152Z), the differences from the inferred common ancestor as determined by comparing the UTEX393 and NREL46B-D3 protein sequences were significant. A significant increase in the levels of glycine, histidine, and arginine was observed in the halotolerant NREL 46B-D3 strain compared to the freshwater UTEX393, along with a significant decrease in serine levels (Fig. 1D).
Salinity perturbation study
Post genome construction, the physiological, systems level responses induced by salt stress were measured over time while grown in the presence of 15 ppt (“high salt”) or 5 ppt (“low”/control salt) sodium chloride media conditions. Throughout the culturing period, areal productivity (g/m3 day) and ash-free dry weight (AFDW) was reduced by 34% and 29% (average across three replicates at study end-point), respectively compared to the low salt condition (Fig. 2A, Supplementary Fig. 1). During this experiment, samples collected during an intensive 38-hour window (Fig. 2A) were subjected to a comprehensive multi-omics analysis. For each sampled time point, lipidomics, polar metabolomics, transcriptomics (RNA-seq), and proteomic (TMT-labeled) profiles were measured at each salt concentration. Additionally, deuterated water was added at the “0-h” sampling point to track newly synthesized metabolite(s) via isotopic labeling which defines the starting point of the perturbation study. Interestingly, the average drop in OD that occurs at night due to cellular respiration was reduced by 10% in 5 ppt to 2% in 15 ppt (P = 0.01, OD750 differences between day 4 peak sunlight and Day 5 pre-light).
A Optical density measurements (OD750nm) were taken throughout the growth experiment (error bars indicate the standard deviation across three replicates). Columns were inoculated at T = 0 days and intensive sampling was initiated when the 5 PPT (gray) medium columns dry weights approached 500 mg/L. The intensive sampling window is indicated by the rectangle where the 0-h time point was taken at midnight on the fourth day after inoculation. B Environmental conditions simulating the 30-year average light and temperature conditions of a modeled 20 cm deep pond in Mesa, AZ, on July 1 (blue = temperature, orange = Light intensity).
Transcriptomics analysis
The number of differentially expressed genes measured across the sampling window ranged from 2371 genes at 14 h after D2O addition, and 4744 genes at 4 h after D2O addition (Fig. 3, Supplementary Data 1). Of the genes differentially expressed over the time course, 1209 genes were significantly (FDR ≤ 0.05, |Log2FC | ≥ 1) differentially expressed at all time points. Interestingly, 763 (63%) of the genes were differentially expressed, but inconsistently across the time course (upregulated and downregulated at various time points).
Differential gene expression analysis of 15 ppt (high) salt compared to 5 ppt (low) control algal cultures during a 48-h time course. A Differentially expressed genes at each time point (FDR ≤ 0.05, |Log2FC | ≥ 1) (Gray=upregulated, black=downregulated, slashed = ambiguously differentially expressed across replicates). B Differential expression patterns of essential genes predicted through Flux variability analysis and screening gene-knockouts computationally using the metabolic model. Essential genes related to chloroplast function, amino acid and fatty acid metabolism are designated by a “*”. C Differentially expressed transcripts associated with photosynthesis. Yellow bar charts overlaid on axes indicate the light intensity at each time point with the orange trace line indicating the continuous program between the sampling time points.
Among the most differentially transcribed genes (Table 2) are 1707638, coding for a putative aldehyde dehydrogenase, and 1849939, a flavodoxin. Both genes are redox related as aldehyde dehydrogenases govern important redox enzymatic reactions preventing reactive oxygen species (ROS) buildup31 and flavodoxins are known to regulate redox potential as an electron acceptor32,33. Additionally, the transcripts encoding an expansin gene (1941549) were significantly increased (between 15- and 250-fold change in expression), potentially indicating an effort to regulate cell size and remodel the cell wall, a common response to salinity stress34,35. Notably, several genes (1994846, 1911475, 1789912, and 1883123) are upregulated at all but the 18 h sampling time point during the programmed decrease in light levels (moving toward dark in Mesa, AZ program). These four genes consisted of an alkaline peptidase (1883123), an oligopeptide transporter (1911475), a peptidase (1789912) and a NDP transferase (1994846).
To capture genes with dynamic responses to salt stress, we examined genes with significantly different levels of expression in at least half the sampled timepoints with the highest variability in differential expression occurring in response to circadian rhythms (Supplementary Table 1). Given cellular respiration was reduced by 10% in 5 ppt to 2% in 15 ppt in a dark-cycle specific manner, these genes are of interest as they may regulate the altered metabolism of UTEX393 during simulated night. Notably, the three genes that could be assigned a putative function and exhibited the strongest circadian pattern of differential expression were all septin domain-containing proteins. Modulation of septins (components of cytoskeletons) suggests that salt stress may induce a circadian entrained alteration of cell structure and cell division36,37. Additionally, genes responsible for modifying (glycoside hydrolases and glycosidases) or functionally defined by (fasciclins/FAS1-domain containing) covalent sugar modifications are differentially expressed in a circadian manner. This pattern of differential expression may indicate an altered glycation profile responsive to higher salt environments which can combat osmotic stress by replacing solvent/water interactions at molecular interfaces in both intracellular and extracellular environments38,39.
A large number (n = 81) of genes that were annotated to be involved in alteration of fatty acid synthesis and utilization pathways were also differentially expressed at varied timepoints. Leveraging gene functional annotations derived from the nearest BLAST-based gene orthologs in Arabidopsis we also observed an enrichment of differentially expressed genes (in dark cycle differential expression, RNA-seq timepoints T0–T4 and T24) that encode proteins involved in polyprenol metabolism (Supplementary Fig. 2), a key component of thylakoid membranes40. This alteration to polyprenol metabolism indicates a potential effort to alter thylakoid structural regulation to acclimate to the stress of higher salinity concentrations.
Broader examination indicated that these lipid and thylakoid regulatory shifts correlate within the broader landscape of altered chloroplast regulation; across the sampling time-course a systemic downregulation of Photosystems I/II, RuBisCO, ATP synthase, pigment, and B6F/redox components is prevalent (Fig. 3C) with some notable exceptions including a Chlorophyll a/b-binding protein domain-containing protein (1943860), Photosystem I assembly protein Ycf3 (1890725), probable chlorophyllide b reductase NYC1 (1799263), and Carotene biosynthesis-related protein CBR (1705857). The photosystem II-related gene was PPD4 (1740175) was consistently upregulated which has been shown to accumulate under high light conditions in pea plants41. Notably, several core enzymes were also differentially expressed. The plastid lipid processing enzymes Monogalactosyldiacylglycerol synthase (MGDG1, 1725787) and Plastid Galactoglycerolipid Degradation 1 (PGD1, 1835382) were both downregulated. Two digalactosyldiacylglycerol synthase (DGDG1, were differentially expressed with 1426071 downregulated and 1720584 demonstrating a circadian regulation where it was downregulated in dark time points and slightly upregulated during light timepoints. Additionally, Phosphoribulokinase (PRK, 1751426) was significantly downregulated over the time-course while 4 malate dehydrogenases (MDH, 1807543/1888054/1677471/1785663) and six Glyceraldehyde-3-phosphate dehydrogenase (GAPDH, 1887460/1771613/1733600/1965267/1881964/1888471) enzymes were more variable in their expression patterns. Of additional note, was the upregulation of the entry points to branched chain amino acid synthesis and acetate assimilation, Branched chain alpha-keto acid dehydrogenase (BCKDH, 1873871) and Acetate kinase (ACKA, 1714059), respectively. In the context of the broad metabolic shifts, the circadian differential expression (down in dark, up in light) of Adenosine kinase marks an alteration to a metabolic master regulator of metabolism controlling the phospho-adenosine pool. These central metabolic regulatory shifts indicate several alterations with the potential to enact altered metabolic states by modifying the availability of phospho-adenosine derivatives, branched chain amino acids, and the equilibrium of malate and oxaloacetate which controls the propensity for shuttling of carbon into and out of organellar spaces.
We further leveraged the transcriptome data and the metabolic model to study essential genes by simulating knockouts and conducting flux variability analysis (FVA). FVA analysis combined with the gene knockouts identified 15 genes as most necessary to organism function. Unsurprisingly, 8 out of the 15 detected genes are targeted to the chloroplast and are essential for autotrophic growth (Fig. 3B). When comparing the expression values of these enzymes across the stress conditions, a few trends become apparent. In keeping with their predicted role as essential genes, they are broadly expressed without exceptionally dynamic expression such as the enzyme Porphobilinogen synthase (1915403, EC 4.2.1.24), which is responsible for the second step of porphyrin biosynthesis, a key molecule in chlorophyll and photosynthesis. Four essential genes were involved in lipid metabolism and membrane lipid remodeling (1836679,1893280, 1840334). These genes are essential for many lipid precursors through various acetyl CoA transformations. This result further supports the importance of lipid metabolism not only as a storage strategy but also as a core metabolic process needed for cell growth and functioning in UTEX393 and likely an important part of the stress response42,43. Another major category of essential genes is associated with nitrogen metabolism and amino acid biosynthesis (1807962, 1817514, 1862262, 1676407, and 1700826).
Proteomics
Using TMT-labeled mass spectrometry, the differences in protein abundances were measured at each sampled interval. At the 14- and 18-h time points, there was a large cohort of significantly (adjusted P-value ≤ 0.05) differentially abundant proteins (Fig. 4) suggesting a more cohesive regulatory shift at these time points. One of the most differentially abundant proteins was another expansin (1741660). As seen in the transcriptomic data, many of the differentially abundant proteins are chloroplast components involved in photosynthesis. An increase in Chlorophyll binding proteins (1687918, 1900501, 1863588) was observed concurrent with a reduced abundance of Photosystem II repair protein (1809586), Oxygen-evolving enhancer protein 3 (1818388), Photosystem I reaction center subunit II (1773891), and Plastocyanin-like domain-containing protein (1882176) (Fig. 4, Supplementary Data 2). In addition to these photosynthetic alterations, we observed a major shift in redox machinery including one upregulated (633438) and two downregulated (1908791, 1758594) superoxide dismutase proteins as well as numerous oxidoreductases, dehydrogenases, and peroxidases (Supplementary Data 2).
Differentially abundant proteins from TMT-labeled tandem mass spectrometry between high (15ppt) and low (5ppt) salt conditions (FDR ≤ 0.05) (Gray = upregulated, black = downregulated). While variation in protein abundances left a subset of the proteins as differentially abundant (A), the general trend of the differentially abundant proteins appears mostly consistent across time points with the differential abundance (Log2FC) consistent despite only limited timepoints with significant differential abundance (asterisk indicates FDR ≤ 0.05) (B). Time points 14 and 18 showed the greatest number of differentially abundant proteins with the largest cohort of differentially abundant proteins occurring at the 18-h time point where the light program is trending toward dark (sunset). Yellow bar charts overlaid on axes indicate the light intensity at each time point with the orange trace line indicating the continuous program between the sampling time points.
Metabolomics
To identify changes in central metabolism, the allocation of carbon/energy, and other alterations of metabolites, polar metabolomics and lipidomics were performed across the time course. In general, triacylglycerols (TAGs) were more abundant in UTEX393 under high salt compared to the low salt conditions, concomitant with a shift toward higher relative abundances of saturated TAGs (Fig. 5A). Under normal conditions, TAG abundance increases in the light/daytime conditions (timepoints −10, 8, 14, 38) and are consumed during dark periods (timepoints 0, 4, 24). While the low salt controls do exhibit this pattern, UTEX393 grown in 15 ppt salt exhibits peak TAG abundance in the dark (time point 0) with a continuous reduction of TAGs throughout the intensive sampling window. Most notably, the drop in TAG abundance from T8 (simulated sunrise) to T14 (simulated midday) is counter to typical lipid production tendencies44,45. This consumption of TAGs during a period of photosynthetic activity may indicate a need to supplement energy production to perform cellular functions and may be linked to the decreased loss in OD during dark time points in high salt environments.
A The relative abundance of TAG species under increased salt conditions (relative to 5 ppt controls). Left bar denotes TAG species with ≤5 degrees of saturation (orange) and ≥6 degrees of saturation (gray). B Polar metabolites. C Sum of all metabolites identified as diacyl-, monoacyl-, and triacyl-glycerols (Black = 15ppt, green = 5 ppt, summed values were scaled to the highest abundance in each lipid subtype). Yellow bar charts overlaid on axes indicate the light intensity at each time point with the orange trace line indicating the continuous program between the sampling time points. * = significant at P ≤ 0.05 between 15 ppt and 5 ppt conditions.
Among the identified polar metabolites, standing pools of glyceric acid, 2-phosphoglycerate (2-PGA), pyruvic acid, fumaric acid, and adenine were reduced, suggesting a reduced flux through central metabolism (e.g., TCA) and production of new ATP building blocks (Fig. 5B). Furthermore, in the presence of high salt, the relative amount of branched chain amino acids (BCAAs) valine, leucine, and isoleucine and the amino acids serine, alanine and glycine were reduced. In addition to the evident shift toward TAG production, polar metabolomics revealed a marked increase in the osmoprotectants proline and betaine, along with a smaller but significant increase in trehalose. Additionally, there is an increase in ADP in the presence of 15 ppt salt, which trends higher over the intensive sampling window. While the patterns among deuterium-labeled metabolites (D2O addition at T0) revealed few clear trends in deuterium integration and, thus, de novo synthesis, some insights were gleaned from these data. While the increase in isocitrate appears to be resultant of de novo synthesis based on deuterium labeling ratios, the similar increase in betaine is not supported as a de novo synthesis event by labeling ratios (Supplementary Fig. 3). As betaine lipids are a building block of ER membranes this may indicate that membrane recycling/degradation occurs under salt stress.
Lipidomic characterization across the time-course provided some of the most intriguing evidence to corroborate the significant alterations in chloroplast regulation under salt stress. Monogalactosyl diacylglycerols (MGDGs) and digalactosyl diacylglycerols (DGDGs) are the primary constituents of the plastid membrane and their relative abundances shifted significantly across the time course experiment. MGDGs and DGDGs were less abundant in high salt conditions compared to the low salt condition (Fig. 6). The trend of decreased galactosyl diacylglycerols (GDGs) is most prominent in lipids containing at least one 16 carbon chain (32:X, and 34:X lipids). Notably, by summing across untargeted metabolomics peaks, all identified sub-classes of carotenoids were reduced for all measurements made in the intensive sampling window (Supplementary Fig. 4). While there was a broad overall reduction in GDGs, there was an observed increase in the longer 36 carbon MGDGs with medium (2 or 3 degrees) of unsaturation as well as the 34 carbon DGDGs with 5 or 6 degrees of unsaturation.
Chloroplast lipidomics indicated a significant alteration to the relative abundances of galactosyl diacylglycerols. A monogalactosyl diacylglycerols and B digalactosyl diacylglycerols are both indicated with “*” indicating significant (P-value ≤ 0.05). Yellow bar charts overlaid on axes indicate the light intensity at each time point with the orange trace line indicating the continuous program between the sampling time points.
DAP-seq
To investigate regulatory response(s), we used DNA affinity purification sequencing (DAP-seq)24 to identify the gene targets of key transcription factors. DAP-seq uses in vitro-synthesized transcription factors that are affinity-purified to capture binding events within sheared native genomic DNA (gDNA). High-throughput sequencing of the bound oligonucleotides and subsequent sequence analysis enables identification of the transcription factor binding sites and potential target genes under regulation. DAP-seq candidates were selected based on phylogenetic significance as well as differential expression across the salt stress time course (Supplementary Fig. 5). For 8 of the 24 transcription factors that were synthesized in vitro, at least 20 DAP-seq peaks were identified over the background, indicating TF binding (Supplementary Table 2). The number of predicted target genes of each TF, which were determined based on locations of intergenic peaks, varied from 1 to over 500 target genes. The full dataset is accessioned as PRJNA880555 including sequencing for all 24 DAP-seq experiments and 3 controls (Supplementary Table 3). These data can be interrogated in the DAPSeqViewer within the PhycoCosm46 genome portal (Supplementary Fig. 7).
Consensus binding motifs were constructed based on the DAP-seq binding sites. The two GATA TFs (JGI protein IDs 82029 and 1799829) had similar GATC-containing binding motifs. Comparisons to the binding site database JASPAR revealed that the binding motifs show similarity to motifs of GATA transcription factors in A. thaliana, also with the GATC motif (Franco-Zorrilla 2014) (E-value = 5.25e-3, https://jaspar2022.genereg.net/matrix/MA1396.1/). The binding motifs were identified for two SBP TFs, which both included the conserved GTAC motif that is essential for DNA-binding by SBP TFs across algae and plants (Birkenbihl47).
We discovered that the Myb-like TF (JGI protein ID 1730603) has a conserved motif that is similar to other Myb-like transcription factors including those in A. thaliana that are involved in regulation of the circadian clock, such as CCA1 and LHY1 (E-value = 1.04e-6, https://jaspar2022.genereg.net/matrix/MA1185.1/). Salinity stress induced modulation of the expression of the Myb-like TF in UTEX393, which correlated with the diurnal cycle, with higher expression during the night-time samples and lower expression during the day. A predicted target gene, a diacylglycerol transferase (JGI protein ID 691482), showed a similar expression pattern correlated with the diurnal cycle. Based on the similarity of the binding motif, sequence similarity of the DNA binding domains (68% sequence identity with the A. thaliana LHY1 gene), and an observed diurnal pattern of gene expression, the Myb-like TF may be circadian entrained in a manner similar to LHY1.
Of potential interest are the targets of several transcription factors identified as having binding motifs similar to LHY1 and SPL12. The LHY1 homolog was predicted to regulate two major central metabolic enzymes, including the mitochondrial malate dehydrogenase (MDH, 1785663) and a chloroplastic glyceraldehyde 3-phosphate dehydrogenase (GAPDH). Additionally, the SPL12 homolog was found to act on two cytosolic GAPDHs among its other targets. As a result, these two transcription factors may provide promising targets to alter some of the redox imbalances identified in UTEX393 under salt stress.
Metabolic modeling and gene interaction network analysis
Using the omics information, a metabolic model was constructed to provide a networked system wherein genes/proteins can be linked based on their reactions to metabolic substrate(s) and product(s). Integrating the DAP-seq TF-gene target interactions with the enzyme-molecule interactions from the metabolic model, we were able to construct a network defining the potential functional regulation of the characterized TFs (Fig. 7). Four transcription factors were predicted to bind upstream of a total of 66 enzyme-coding genes (287871 SBP/SPL12, 1730603 Myb-like/LHY1, 1008053 SBP, and 214616 SBP). Interestingly, 3 out of the 10 total TFs studied herein belonged to the SBP family. While this is a small sample size, it might point to a specific role of the SBP family of transcription factors in metabolic regulation48. The metabolic regulatory functions of some TFs show similarities to established functions of orthologs in model systems. TF 214616, which belongs to the SBP family, is predicted by this integrated network analysis to regulate an enzyme involved in copper metabolism, a process that is associated with the SBP family of transcription factors in Arabidopsis49,50. In addition, the network analysis suggests that SPL12-like TF (287871) regulates enzymes involved in phosphate metabolism, another well-documented metabolic process linked to the SBP family48.
A Comparison between binding motifs of LHY1 A. thaliana TF and Myb-like TF (1730603) in S. obliquus UTEX393, B comparison between binding motifs of GATA6 A. thaliana TF and C2C2-GATA family TF (1799829) in S. obliquus UTEX393, and C comparison between binding motifs of SPL12 A. thaliana TF and SBP family TF (287871) in S. obliquus UTEX393. D Combined network of the metabolic model with the Dap-seq enzyme targets. The full network shows that all the TFs (purple MYB-Like, turquoise SBP), their targets (orange) and metabolites (black) form a single connected component. While the individual TF-target networks form singular components when augmented with the metabolic model, they all form a single component linked through metabolite-enzyme interactions. Key enzymes found in the transcriptomic and transcription factor analysis are also highlighted, such as GAPDH, MDH, and DAGT.
Discussion
The improved genome assembly and annotation of Scenedesmus obliquus UTEX393 enabled a more accurate and comprehensive understanding of the physiology and regulatory mechanisms in this industry-relevant lineage. A comparative analysis of protein sequences derived from inferred common ancestors between the halotolerant NREL 46B-D3 strain and UTEX393 indicated a significant shift toward glycine, histidine, and arginine levels in the more halotolerant Scenedesmus species. This shift in the amino acid profiles may indicate an alteration to protein chemistry requisite for optimizing halotolerance traits, however, this remains an understudied topic with only a few studies examining halotolerance and amino acid utilization29,30.
Integration of the multi-omics measurements provided insight into the metabolic and regulatory landscape that is altered and responsive to higher salt concentrations (Fig. 8). Several observed differences between the salt-stressed and control samples represent known adaptations that allow for increased productivity under high salinity conditions. For example, the increase in proline levels as well as the increase in the protein abundance of proline synthesis gene, P5CR, are known responses to salt stress51. The consistent increase in transcription and/or expression of expansins (1941549 and 1741660) further indicate attempts to adapt to the salt stress. Despite these physiological alterations, the high-salt cultures did not successfully acclimate to 15ppt salt concentrations during the study period, as no noticeable improvement in growth was observed during the experiment.
Red = increased in salt stressed (15ppt) Scenedesmus relative to control (5 ppt), blue = decreased in salt stressed Scenedesmus relative to control. Bold arrows represent enzymes upregulated in salt stress conditions relative to controls and thin dashed arrows represent enzymes downregulated in salt stress conditions relative to controls. Long dashed arrows indicate collapsed metabolic pathways representing multiple reactions. Superscripts “LHY1” and “SPL12” indicate the enzyme was identified as a target of the given transcription factor by DAP-seq and superscript “P” indicates the differential expression identified in the figure was identified through proteomic analyses.
The differential expression patterns of transcripts which encode for central enzymes that regulate and utilize photosynethetically generated reducing equivalents (and thus the redox balance) is one of the most noteworthy elements of Scenedesmus obliquus UTEX393’s response to elevated salinity. The superoxide dismutases (in particular, superoxide dismutase 633438, the most significantly upregulated protein under many sampling timepoints) were upregulated, indicating a cellular redox imbalance. The increased transcript abundance of cytosolic malate dehydrogenase (MDH), along with the decreased levels of chloroplast and mitochondrial MDH paralogs also suggest a major alteration of redox balancing in the cell52,53,54,55. GAPDH paralogs, generally considered to be a regulatory hub for oxidative metabolic reactions56,57, are similarly altered with a shift in predominant paralog from 1888471 to 1881964 in the chloroplast, along with a shift from 1733600 and 1965267 to 1887460 in the cytosol. Alteration of MDH-GAPDH combined with the reduced expression of PRK which forms an additional regulatory bienzyme complex with GAPDH indicates a significantly altered metabolic potential regulated by redox signaling57,58. Importantly, an LHY1 protein identified in the DAPseq measurements was found to regulate the expression of mitochondrial MDH and chloroplastic GAPDH while SPL12 was found to regulate cytosolic GAPDH paralogs (1733600 and 1965267). Thus, genetic alternation of the expression of these transcription factors could be an effective strategy to modulate or improve the redox-driven regulatory shifts of UTEX393 under salt stress conditions.
The observed redox stress concomitant with the induction of TAG synthesis/storage indicates that the salinity stress response of UTEX393 is consistent with “overflow biology.” According to the overflow hypothesis, stress induces a shift in lipid production due to a decreased capacity to process reductive products of photosynthesis59,60,61,62,63. To offset this electron overflow, cells use TAG synthesis as an electron sink to store energy and balance the redox equilibrium. A reduced photosynthetic rate would have the benefit of decreasing the production of reducing equivalents while the overflow of reducing equivalents is shunted to TAG synthesis60. Indeed, an increased abundance of TAG species was observed. While the observed metabolic shifts would reduce electron ‘overflow’, the total growth rate would, as observed across the time-course, be reduced proportionally to the decreased photosynthetic productivity. Patterns exhibited by UTEX393 support the overflow hypothesis proposed in nitrogen deprivation studies59, yet the study presented herein occurs under nitrogen replete conditions (Supplementary Fig. 6). These responses to redox stress and the concomitant production of lipids appear to be an ubiquitous yet exploitable response of Scenedesmus to salt stress14,15,23. One of the most broad-reaching regulatory shifts that was observed is the circadian differential expression pattern of Adenosine Kinase (ADK). As the enzyme responsible for the conversion of Adenosine and ATP to AMP and ADP, the alteration to phosphorylated adenosine would alternately reduce the AMP pool in the dark (downregulated) and increase it in the light (upregulated). This AMP responsive signaling could act as a significant metabolic regulator64 which could be responsible for the altered TAG utilization and production pattern observed under salt stress.
Notably, the multiomics data indicate a marked reduction in chloroplast and thylakoid components concurrent with the redox imbalance. Reduced levels of MGDG and DGDG lipids (chloroplast membrane components)–despite the reduction in MGDG degrading enzyme PGD1–suggests a loss of chloroplasts and thus reduced photosynthetic capacity. Thylakoid-localized components of photosystem I and II, along with b6f and thylakoid ATPase components, were also systematically downregulated. Combined with the observed decrease in ATP production and NADPH-reducing equivalents, energy conservation is reduced, helping to explain the decreased productivity of UTEX393 under salt stress. As the intensive sampling time course was initiated approximately 4 days into the salt stress, we interpret the data as indicative of lipid remodeling that predates the sampling regime, especially in the case of galactosyl lipid components of the chloroplast membranes. The decreased abundance of MGDGs and DGDGs along with reduced expression of synthesis components MGD1, and DGD1 and the reduced expression of the galactosyl lipid degradation enzyme PGD1, indicates a steady state of reduced synthesis and degradation of chloroplast membrane components. Considering the increased TAG abundance, UTEX393 appears to slow photosynthesis to reduce electron overflow and redox stress. These regulatory and metabolic shifts provide context and insight into recent studies examining the salinity stress response in Scenedesmus sp. BHU123. Most notably, Singh et al. found a decrease in PSII-driven linear electron flow, which agrees with our interpretations of the molecular results of this study. The agreement of these two studies strongly indicates that similar regulatory mechanisms govern BHU1 and the freshwater UTEX393 strain. D2O chase data indicating increased betaine levels are not the product of de novo synthesis highlight one essential knowledge gap that remains in the Scenedesmus salt stress response. While primarily found in extraplastidial membranes (e.g., ER)65, betaine lipids like DGTS can occur in the chloroplast envelope membranes in certain Chlorophytes66 Thus, the increase in the concentration of betaine could be an indirect measure of chlorophagy in UTEX393 but the source of betaine under salt stress remains an outstanding question. Experiments examining the interaction of salt stress and high light conditions may provide important insights into the underlying signals that initiate “overflow” responses by reducing the photosynthetic production of reducing equivalents and the subsequent redox stress induced by increased salinity.
In addition to the redox-regulated components, additional “master regulators” of metabolism were altered. Linked to the observed overflow response, is the systematic upregulation of branched-chain ketoacid dehydrogenase (BCKDH). BCKDH is the first committed step of branched chain amino acid (BCAA) synthesis acting on pyruvate to produce the precursor molecular of isoleucine, leucine, and valine. Perhaps one of the most noteworthy points of integration in this multiomics assessment is the interface of BCAA production and utilization for TAG synthesis. After this initial committed step, BCAA synthetic pathways bifurcate with one pathway producing valine and leucine and the other producing isoleucine—all of which can be used as precursors for TAG synthesis. The observed depletion of BCAAs, an important building block for TAG synthesis, is consistent with a previous study that reported mutants deficient in BCAAs (BCKDH knockouts) have significantly reduced metabolic potential (dark oxygen consumption reduced by 20–35%)67. Despite the increased expression of BCKDH, the abundance of BCAAs is diminished likely due to the use of BCAAs as building blocks for TAG synthesis67 which further accounts for the decreased growth phenotype of UTEX393 under increased salinity. The depletion of the mitochondrial pyruvate pool toward BCAA synthesis to satisfy TAG production requirements may be responsible for a significant metabolic reduction as indicated by BCAA deficient Chlamydomonas mutants67. Concurrent with the increased enzyme expression levels and the implied increase in carbon flux toward BCAA synthesis in the mitochondria, the TCA cycle enzymes leading to 2-oxoglutarate (alpha-ketoglutarate) are ubiquitously upregulated. Importantly, 2-oxoglutarate is the precursor for proline biosynthesis where protein levels of pyrroline-5-carboxylate reductase (P5CR), the halotolerance-associated enzyme15,68,69 governing the final step in proline synthesis, are significantly upregulated.
The observed responses of certain TFs in Scenedesmus warrant a deeper investigation into the conservation of regulatory mechanisms across the Viridiplantae and provide new regulatory targets to alter physiological response(s) to salt stress. By combining metabolic modeling with DAP-seq data, we were able to gain insight into the transcriptional regulation of key metabolic processes. The Myb-like TF (1730603) in Scenedesmus was observed to control enzymes involved in all-trans-retinal metabolism, which is a key chromophore needed for the functioning of phototransduction in algae70. This response is similar to the Myb-like family of transcription factors, which are well known to be involved in various phytochrome and light receptor signaling pathways in plants71. Further, the circadian differential expression pattern of ADK, the enzyme responsible for the conversion of Adenosine and ATP to AMP and ADP, the alteration to phosphorylated adenosine would alternately reduce the AMP pool in the dark (downregulated) and increase it in the light (upregulated). This AMP-responsive signaling could also act as a significant metabolic regulator64 which could be responsible for the altered TAG utilization and production pattern observed under salt stress. The identification of these TFs specifically regulating central metabolic genes/proteins involved in redox balancing reactions provides specific next steps to improve Scenedesmus for increased salt tolerance. Genetic studies examining alterations to strain tolerance when Scenedesmus orthologs of SPL12 and LHY1 are modified is hypothesized to directly impact salinity tolerance. Taken wholistically, the TFs characterized by DAP-seq, the differential expression of rate-limiting enzymes, the modulation of chloroplast degradation and photosystem II all provide a roadmap for future intelligent engineering of Scenedesmus to alter salinity stress response.
Methods
Genome assembly and annotation
Multiple pieces of sequencing data were used to improve the assembly and annotation. All data were produced from the UTEX culture collection (accession UTEX393) for the haploid alga where the culture is non-axenic. First, contaminating bacterial reads/contigs were removed from previously published Illumina-based draft assembly25 using Deconseq (version 0.4.3) by mapping to the bacterial database generated by the script designers72, and the reference genomes of fungal species selected based on fungal sequences detected in the Carreres et al. assembly. A second Illumina-based genome assembly had been provided by David Nelson (NYU Abu Dhabi). The third Illumina-based genome had been assembled from sequence reads produced from the US Joint Genome Institute (JGI) Community Sequencing Project (CSP) (ID#1191226). The fourth genome assembly was based on PacBio data from the JGI CSP ID#1191226. The Illumina and PacBio-based genomes were assembled in house at Brooklyn College of the City University of New York with the resource available via the CUNY High Performance Computing Center. Each of the four separate assemblies was polished with Pilon 1.22 using the JGI Illumina reads. Following, all genome assemblies were merged using a sequential process using SPAdes 3.9.0, integrating each Illumina assembly, and finally with the PacBio assembly, which produced the new genome of 329 scaffolds with >5000 bp.
The nuclear genome assembly of UTEX393 was annotated using the JGI Annotation Pipeline73,74, which combines different gene model predictions based on ab initio75,76, homology-based77,78, and transcriptome-based79 methods. An automated step selects the best model at each locus based on homology and transcriptome support. Protein-coding genes are functionally annotated using InterProScan80 and alignments to databases, including NCBI NR, SwissProt81, KEGG82, and KOG83. RNAseq datasets that were used for the annotation were downloaded from NCBI (SRA ERR2699840, ERR2699873, ERR2699886, ERR2699865, ERR2699875, ERR2699881) and reads were assembled into consensus sequences using Trinity v2.3.284. In addition, high-quality IsoSeq transcripts sequenced by PacBio from a related Scenedesmus obliquus (strain EN0004) were used for the annotation. Genome assembly and annotation statistics were performed on the nuclear assembly after removal of potential organellar scaffolds. Using Blast+ v2.12.0 against the NCBI nr/nt database, a 235.6 Kbp scaffold (scaffold_72) was identified as a chloroplast scaffold, and an 82 Kbp scaffold (scaffold_154) was identified as a mitochondrial scaffold. Within scaffold_72, alignment identified 60 genes with exact matches to gene content found in the de Cambriaire et al. chloroplastic assembly and annotation85 confirming this previous chloroplast assembly as strain UTEX393 and indicating compatibility of the previous chloroplast resource with this genome release. For comparison to the previously published genome of UTEX39325, the genome sequence and gene models were downloaded from the European Nucleotide Archive (ENA) associated with Project Accession PRJEB15865. A genome portal and accompanying files are accessible in PhycoCosm46 under the name “Scenedesmus obliquus UTEX 393 v2.0.” Unique gene models were identified by MCL clustering the protein sequences from the two annotated gene sets. The completeness of the genome annotation was evaluated using BUSCO v5.0.0 in protein mode on the Chlorophyta ortholog dataset (chlorophyta_odb; 8-5-2020). The protein sequences used for running the BUSCO assessment were downloaded from the following PhycoCosm portals: Sceobl393_2, Sobl393_1 (ExternalModels gene set), Sceobl1, Scesp_1, Monmin1, Monneg1, Chrzof1, and Rapsub1. Orthology analysis was performed using the Orthologous Matrix standalone tool27 with the protein databases for Micromonas pusilla CCMP1545 (MICPC), Micromonas commoda (MICCC), Ostreococcus lucimarinus (OSTLU), Ostreococcus tauri (OSTTA), Chlorella variabilis (CHLVA), Volvox carteri (VOLCA), Arabidopsis thaliana (ARATH), Chlamydomonas reinhardtii (CHLRE), Scenedesmus sp. 46B-D3 (SoblNREL46B3), Scenedesmus obliquus DOE152Z/DOE3031 (SoblDOE152Z), Scenedesmus obliquus UTEX393 (SoblUTEX393), Monoraphidium minutum 26BAM (Mmin26BAM), Raphidocelis subcapitata (RAPSUB35), chromochloris zofingeiensis SAG211 (CzofSAG211), and Picochlorum renovo (Pren) with Arabidopsis thaliana as the outgroup. Analysis of hierarchical orthologous groups was performed using custom scripts and implementation of the “ape” package86 in R where the amino acid content for each amino acid sequence was calculated for both the NREL 46B-D3 and UTEX393 strains of Scenedesmus.
Growth medium
DISCOVR medium11 at 5ppt and 15 ppt salinities were used to perform the salt perturbation study. Vitamin B12 was added at 0.5 µg/L (per Guillard’s f/2 recipe), due to previously observed vitamin deficiencies in marine cultures. It is assumed in large-scale cultivation conditions, that bacterial consortia will provide this vitamin to the algal culture87.
Inoculum preparation
S. obliquus UTEX393 was obtained from the University of Texas Culture Collection and maintained in 5 ppt DISCOVR medium in 125 ml Erlenmeyer flasks under 450 µmol/m2/s light set on a 12:12 (h) cycle and pH was at 7 ± 0.2by continuously sparging with sterile-filtered 0.5% CO2-enriched air. The same culture was used for all experiments and is non-axenic. Cultures were scaled to 800 mL in a Roux bottle under similar light and sparging conditions to produce the inoculum for
PNNL Laboratory Environmental Algae Pond Simulator (LEAPS)88. Three LEAPS columns with 5 PPT medium and three with 15 PPT medium were inoculated and maintained following previously described conditions for summer conditions (July 1) in Mesa, AZ88. Samples were collected in parallel for multi-omics studies at 0, 4, 8, 14, 18, 24, and 38 h into the intensive sampling window (beginning at 108 h post-inoculation) along with a baseline sampling 10 h prior. OD750 measurements were used to calculate Ash-free dry weights (AFDW) at each time point (see Supplemental Fig. 1 for AFDW vs. OD750 calibration). Areal biomass productivities are calculated using the AFDW values. Immediately following the 0-hour timepoint, 10% v/v deuterium oxide (ACROS Organics) was added to each biological replicate. Prior to adding deuterium, the appropriate amount of salt was dissolved into the deuterium to reach either 5 or 15 PPT (5.6 or 16.8 g/L Crystal Sea Marine Mix, respectively), to avoid diluting the culture. For the ‘omics studies detailed in subsequent methods, 48 samples (triplicate for 8 timepoints under two conditions) were processed for analyses.
Transcriptomics
The concentration and quality of total RNA were obtained using the Tapestation RNA Assay Reagents and RNA ScreenTape (Agilent, Cat. #5067-5577 and 5067-5576). Polyadenylated RNA was selected using the TruSeq stranded mRNA Library Prep Kit (Illumina, Cat.#20020595). RNA was converted to cDNA, and adapters and indexes were added onto the ends of the fragments to generate Illumina libraries that can be sequenced on an Illumina sequencer. Illumina libraries are eluted in DNA Elution Buffer (Zymo Research, Cat. #D3004-4-10). The concentration of the libraries was obtained using the Qubit dsDNA HS Assay (ThermoFisher Scientific, Cat. #Q32854). The average size of the library was determined by the Agilent High Sensitivity DNA Kit (Agilent, Cat. #5067-4626). An accurate library quantification was determined using the Library Quantification Kit – Illumina/Universal Kit (KAPA Biosystems, KK4824).
Libraries were normalized to the same concentration based on the qPCR results. Each library was sequenced on approximately 5% of a NextSeq High Output flow cell to generate paired-end 151 bp reads using the NextSeq 500/550 High Output Kit v2.5 Kit (300 cycles) (Illumina, Cat. #20024908). The library was run using four flow cells to yield a minimum of 8,684,423 150 bp reads resulting in a minimum of 31.7X transcriptome coverage across all replicates. Raw read data (fastq) were aligned to the Scenedesmus obliquus UTEX393 genome (JGI) assembled and annotated herein which is filtered to exclude organellar components. Alignments were performed using STAR (2.7.1a)89 according to encode standard parameters with one exception. Specifically, STAR was implemented using the basic two-pass mode to identify novel splice junctions/events given the genomic resources are draft-level and quant mode was used to directly produce gene-level counts data. All gene counts were imported into the R/Bioconductor package EdgeR90, filtered to include only genes with at least 5 counts per million (CPM) in at least 3 replicates reducing the genes considered for analysis to 12,382 prior to the calculation of trimmed mean of M values (TMM) normalization size factors were calculated to adjust for sample-specific differences. The TMM size factors and the matrix of counts were then analyzed using the R package LIMMA (Ritchie et al.91). Weighted likelihoods based on the observed mean-variance relationships were then calculated for all samples and genes using voomWithQualityWeights92 prior to the calculation of differential expression profiles between each sample group using LIMMA. Gene counts and differentially expressed genes for each organism were annotated with entrez gene IDs, descriptions, and symbols using BiomaRt93 using the best amino acid blast hit to the Arabidopsis thaliana. Additional functional annotations were generated for genes by taking the highest percent identity match among blast matches (E-value ≤ 1e-2) to the Landmark, Swissprot94, and PDB94 blast databases. Enrichment and pathway analyses were performed using custom R scripts implementing the gene enrichment package GAGE95 (Luo et al. 2009) and Pathview96.
Proteomics
Samples were lysed cells using a BeadRuptor in 1 mL lysis buffer (4% SDS + 25 mM Tris-Hcl pH7.6) and transferred into a protein LoBind tube. Lysates were clarifed to remove debris by centrifugation at 21,000 RCF for 10 minutes at 4 C and supernatant transferred to fresh LoBind tube. Protein concentrations were quantified using a Pierce BCA Assay Kit (Thermo). Pre-wash of 50 mL 10 K MWCO filters was performed with 4 mL UA buffer (8 M urea + 100 mM Tris-HCl pH 8.6) and spun down at 5000 RCF for 20 minutes at RT prior to addition of ~200 ug of protein lysate reduced with DTT at a final concentration 10 mM at 90 °C for 15 minutes with gentle shaking. Reduced lysates were loaded onto MWCO filters with 10 mL UA buffer prior to acetylation using 0.5 M iodoacetamide in UA buffer (20 min, RT, dark). Filters were washed twice with 4 mL UA buffer and twice with 4 mL 100 mM TEAB prior to on filter digestion using trypsin/lysC (Thermo) at a 1:50 enzyme to protein ratio (overnight, 37°C). Digests were eluted twice with 4 mL 50 mM TEAB followed by a single wash with 4 mL 0.5 M NaCl. Elutions were pooled, acidified with 10% formic acid and desalted using 1 cc 30 mg SPE cartridges (Waters Oasis). Digests were quantified with a Pierce Quantitative Colorimetric Peptide Assay Kit before labeling with TMT-10plex (Thermo) according to standard protocols. TMT plexes were run on an acetonitrile:water gradient (Neo Vanquish UHPLC) prior to detection using a nanospray ion source (Vpositive = 1900, Vnegative = 600) prior to detection on a Thermo Q Exactive Orbitrap (resolution = 120,000, data dependent acquisition mode = Cycle time). Data processing and protein assignments were performed with Proteome Discoverer 3.0 using Sequest HQ and Percolator using the gene catalog proteins available on PhycoCosm using default parameters for analysis with phosphorylation of serine, threonine, and tyrosine included as variable modifications.
Metabolomics
Metabolite extraction
Prior to extraction, algae biomass (between 15-30 mg) was first lyophilized dry (FreeZone 2.5 Plus, Labconco) then powderized by bead-beating with a 3.2 mm stainless steel bead in a bead-beater (Mini-Beadbeater-96, BioSpec Products) for 5 s (2x).
Lipid extraction
For extraction of lipids, a chloroform-based lipid extraction was performed using a modified Bligh-Dyer approach97. Here, 300 μL methanol (MeOH), 150 μL CH3Cl and 120 μL water was added to the powderized pellet (final ratio of 2:1:0.8 MeOH:CH3Cl:H2O), vortexed and sonicated in a water bath for 10 min, then an additional 150 μL each of CH3Cl and MeOH was added (final ratio of 1:1:0.9 MeOH:CH3Cl:H2O), which was followed by a brief vortex and incubated for 10 min in a sonic water bath. Samples were centrifuged for 5 minutes at 5000 rpm, and the bottom lipid-enriched chloroform phase was transferred to a new tube. Then, 300 μL of chloroform was added, followed by another vortex, sonication, and centrifugation. The bottom chloroform phase was combined with the previously collected extract. The chloroform extracts of lipid were dried in a SpeedVac (SPD111V, Thermo Scientific) and stored at −20 °C.
Polar metabolite extraction
For extraction of polar metabolites, 450 μL MeOH was added to powderized pellets, vortexed and sonicated in a water bath for 10 minutes. After centrifuging for 5 min at 5000 rpm, 200 μL of supernatant was transferred to a separate tube. Extracts were then dried in a SpeedVac (SPD111V, Thermo Scientific) and stored at −20 °C.
LC-MS/MS
Liquid chromatography-tandem mass spectrometry (LC-MS/MS) was performed on extracts of algae biomass using both normal and reverse phase chromatography for polar and lipid metabolites, respectively. Chromatography was performed using an Agilent 1290 LC stack, with MS and MS/MS data collected using a Q Exactive HF Orbitrap MS (Thermo Scientific, San Jose, CA). Mass spectrometer source settings included a sheath gas flow rate of 55 (au), auxiliary gas flow of 20 (au), sweep gas flow of 2 (au), spray voltage of 3 kV and capillary temperature of 400 °C.
Reverse phase chromatography
Extracted lipids from wet algae biomass were resuspended in a 3:3:4 ratio of isopropyl alcohol (IPA), acetonitrile (ACN), and MeOH, with the resuspension volume varied for each sample to normalize by biomass weight. One hundred fifty microliters of this was centrifuge-filtered through a 0.22 μM hydrophilic PVDF membrane (#UFC30GV00, Millipore) and transferred to a glass LC-MS vial. To each of these, 8 μL of a lipid internal standard mixture (SPLASH LIPIDOMIX Mass Spec Standard, #330707, Avanti Polar Lipids) was added to each vial. Full MS spectra were collected from m/z 132-1500 at 60,000 resolution in both positive and negative ion mode, with MS/MS fragmentation data acquired using stepped 10, 20, and 40 eV collision energies at 15,000 resolution. Chromatography was performed using a C18 column (Agilent ZORBAX Eclipse Plus C18, Rapid Resolution HD, 2.1 × 50 mm, 1.8 μm) at a flow rate of 0.4 mL/min with a 2 μL injection volume. To detect lipids, samples were run on the C18 column at 55 °C equilibrated with 100% buffer A (60:40 H2O:ACN with 5 mM ammonium acetate and 0.1% formic acid) for 1 min, diluting buffer A down to 45% with buffer B (90:10 IPA:ACN w/ 5 mM ammonium acetate and 0.1% formic acid) over 2 min, down to 20% A over 8 minutes, then down to 0% A over 1.5 min, followed by isocratic elution in 100% buffer B for 1.5 min. Samples consisted of three biological replicates each and three extraction controls, with sample injection order randomized and an injection blank (2 μL of 3:3:4 IPA:ACN:MeOH) run between each sample.
Normal phase chromatography
Extracted polar metabolites from wet algae biomass were resuspended in 100% methanol containing 10 μM internal standard (5–50 μM of 13C, 15N Cell Free Amino Acid Mixture, #767964, Sigma), with the resuspension volume varied for each sample to normalize by biomass weight. One hundred fifty microliters of this was centrifuge-filtered through a 0.22 μM hydrophilic PVDF membrane (#UFC30GV00, Millipore) and transferred to a glass LC-MS vial. Full MS spectra were collected from m/z 70-1050 at 60,000 resolution in both positive and negative mode, with MS/MS fragmentation data acquired using stepped 10, 20, and 40 eV collision energies at 15,000 resolution. Chromatography was performed using a HILIC column (Agilent InfinityLab Poroshell 120 HILIC-Z, 2.1 × 150 mm, 2.7 μm, #673775-924) at a flow rate of 0.45 mL/min with a 2 μL injection volume. To detect metabolites, samples were run on the HILIC column at 40 °C equilibrated with 100% buffer B (95:5 ACN:H2O with 5 mM ammonium acetate) for 1 minute, diluting buffer B down to 89% with buffer A (100% H2O with 5 mM ammonium acetate and 5 μM methylenediphosphonic acid) over 10 min, down to 70% B over 4.75 min, then down to 20% B over 0.5 min, followed by isocratic elution in 80% buffer A for 2.25 minutes. Samples consisted of three biological replicates each and three extraction controls, with sample injection order randomized and an injection blank (2 μL MeOH) run between each sample.
Metabolomic identification
Metabolites were identified based on exact mass and comparing retention time (RT) and MS/MS fragmentation spectra to that of standards run using the same chromatography and MS/MS method. LC-MS data was analyzed using custom Python code98. For samples run on polar/HILIC chromatography, each detected feature (unique m/z coupled with RT) was assigned a score (0–3) representing the level of confidence in the metabolite identification. Metabolites given a positive identification had detected m/z ≤ 5 ppm or 0.001 Da from theoretical as well as RT ≤ 0.5 minutes compared to a pure standard run using the same LC-MS method. A compound with the highest level of positive identification (score of 3) also had matching MS/MS fragmentation spectra compared to either an outside database (METLIN)99 or internal database generated from standards run and collected on a Q Exactive HF Orbitrap MS. MS/MS mismatches to the standard invalidated an identification. For a detected lipid compound, lipid class was determined based on characteristic fragment ions or neutral loss, and coupled with exact mass to determine specific lipid identity (number of carbons in fatty acid tails and degree of unsaturation). In POS mode, MGDG and DGDG lipids ionized as [M + NH4]+ with a neutral loss of 179 and 341100, respectively, and TAGs as [M + NH4]+ with fatty acid tails detected in the MS/MS spectra101. Lipid standards run separately (DGDG (18:3/18:3)—#840524 P, MGDG (18:3/16:3)—#840523 P; Avanti Polar Lipids, Inc.) and deuterated internal standards of DG and TAG were used to verify the fragmentation pattern and elution times of each lipid class.
Untargeted analysis was performed according to previous work14. In brief, Feature-Based Molecular Networking workflow110 (version release 18) using MZmine2102 and Global Natural Products Social Molecular Networking (GNPS)103 were employed and using a filtered feature set absent isotopes, adducts and features without MS/MS data and selecting for the most intense fragmentation pattern.
Metabolic reconstruction and flux balance analysis
PRIAM104, eggNOGmapper105, KOFAM106, and the BioCyc database were used to assign enzyme commission numbers (EC numbers)107. To build a thorough consensus gene to EC number table, the EC numbers were carefully inspected and curated. The model was carried out using the COBRApy toolkit108, in accordance with the standard process detailed in ref. 109. The localization of different enzymes was based on predictions made by the PredAlgo pipeline in addition to information from previously published algal models. An effort was made to fill in the gaps in the metabolic model by using MENECO110 and by following the methodology provided in ref. 111. Iterations of manual curation were carried out to fill in any remaining gaps and guarantee the biosynthesis of essential metabolic precursors. This was accomplished with the use of knowledge on the composition of biomass gleaned from previously published models of algae and plants112. The model was saved in SBML format113 and analyzed with COBRApy and the Sybil package114. These analyses included phenotypic phase plane analysis, flux balance analysis, and flux variability analysis. To check for mistakes and provide assistance in the process of curating the model, the MEMOTE package was used to identify any remaining problems115.
DAP-seq
DAP-seq experiments were conducted as described previously in O’Malley et al.116, with minor modifications as described by Baumgart et al.117. The conditions are summarized here in brief. S. obliquus UTEX393 genomic DNA library was shared to an average size of 150 bp with a Covaris LE220-Plus focused-ultrasonicator (Covaris), followed by library preparation using the KAPA HyperPrep kit (Roche) following the manufacturer’s recommendations. In parallel, ligated Halo-tagged transcription factor proteins were cloned into a plasmid backbone, sequence-verified, and used for PCR amplification to produce linear PCR fragments. PCR product was purified using SPRI beads and the correct size of each was verified using a Fragment Analyzer (Agilent Technologies). At least 500 ng of each linear PCR product was used as protein expression template in the TnT T7 Quick for PCR DNA in vitro protein expression kit (Promega) following the manufacturer’s recommendations. Each DAP-seq reaction was run with 50 uL expressed protein, 50 ng of the previously prepared fragment library, and 250 ng salmon sperm DNA to reduce non-specific binding. The final DAP-seq libraries were pooled for sequencing on a NovaSeq using the S4 flowcell (Illumina), targeting 10 million 2×150 reads per sample.
Transcription factors were selected based on coexpression analysis, prioritizing candidates that showed significant differential expression under salt stress. The structural (i.e., intro-exon junctions) and functional (presence of a DNA-binding domain) annotations of the transcription factors were manually verified by comparison to transcriptomics data and previously curated protein domains. Once confirmed, the sequences were codon harmonized to match the Escherichia coli codon frequency table with BOOST using the balanced strategy of refactoring to overcome synthesis violations. Synthetic DNA building blocks were obtained from Twist Biosciences and assembled into a DAP-seq compatible vector containing an in-frame N-terminus Halo tag and 3’ poly-A tail by Gibson assembly. Assembled constructs were sequence verified (Pacific Biosciences).
Illumina reads were adapter trimmed and quality filtered using BBTools version 38.90 (https://sourceforge.net/projects/bbmap/) using the following options: k = 21 mink=11 ktrim=r tbo tpe qtrim=r trimq=6 maq=10. Filtered reads were aligned to the S. obliquus UTEX393 reference genome using bowtie2 version 2.4.2118 with the options --no-mixed --no-discordant. Peaks were called using MACS3 version 3.0.0a6119, using negative control samples with mock protein expression as the background file, and the following options: --gsize 107859680 --call-summits --keep-dup 1. Sequences representing up to the 100 most significant peaks, as scored by signal value in column 7 of the narrowPeak files, were extracted using a custom Python script.
These peak sequences were used to generate motifs using MEME (MEME Suite v.5.3.0) (Bailey 1994) with a zero-order background file generated from the reference genome, and the following additional options: -dna -revcomp -mod anr -nmotifs 2 -minw 8 -maxw 32. Peaks were assigned to putatively regulated genes with a custom script using BEDTools version 2.30.0120. The bedtools subtract function was used to filter for intergenic peaks, followed by bedtools closest to assign each peak to adjacent downstream genes. Motif calling was run twice for each dataset, once with the entire peak regions and once with only the regions ±50 bp flanking the summit location of each peak. Peaks located on chloroplast or mitochondrial scaffolds were excluded from motif calling. Significant motifs (e-value < 0.1) were manually inspected and compared to known motifs in JASPAR Non-redundant core database 2022121 using the Tomtom web server122 (MEME Suite v5.4.1). Alignments with e-value < 0.1 were considered significant. A DAPSeqViewer portal is available to review the targets and the inferred regulatory network of the interrogated TFs.
Data availability
The current genome assembly and annotations are publicly available at the JGI Algal Genome Portal PhycoCosm: https://phycocosm.jgi.doe.gov/Sceobl393_2. This Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession JARQZH000000000. Previous assemblies used are available under NIH accession NCBI GCA_900108755.1. The version described in this paper is version JARQZH010000000. Raw transcriptomic sequencing data are accessioned under SRA bioproject ID PRJNA952197. The DAP-seq libraries are available on SRA under the BioProject ID PRJNA880555 and SRA accession SRP398093. Raw mass spectrometry data are publicly available under MASSIVE accession MSV000099195. Source values underlying Fig. 3 and Fig. 4 are available in Supplementary Data 1 and 2, respectively, while both supplementary data files contributed to the generation of Fig. 8.
References
Chisti, Y. Biodiesel from microalgae. Biotechnol. Adv. 25, 294–306 (2007).
Rambabu, K. et al. Biological remediation of acid mine drainage: review of past trends and current outlook. Environ. Sci. Ecotechnol. 2, 100024 (2020).
Chew, K. W. et al. Microalgae biorefinery: high value products perspectives. Bioresour. Technol. 229, 53–62 (2017).
Coons, J. E., Kalb, D. M., Dale, T. & Marrone, B. L. Getting to low-cost algal biofuels: a monograph on conventional and cutting-edge harvesting and extraction technologies. Algal Res. 6, 250–270 (2014).
Dahlin, L. R. et al. Development of a high-productivity, halophilic, thermotolerant microalga Picochlorum renovo. Commun. Biol. 2, 388 (2019).
Dahlin, L. R. et al. Down-selection and outdoor evaluation of novel, halotolerant algal strains for winter cultivation. Front. Plant Sci. 9, 1513 (2018).
Unkefer, C. J. et al. Review of the algal biology program within the National Alliance for Advanced Biofuels and Bioproducts. Algal Res. 22, 187–215 (2017).
Cho, H. S. & Lee, J. Taxonomic reinvestigation of the genus Tetradesmus (Scenedesmaceae; Sphaeropleales) based on morphological characteristics and chloroplast genomes. Front. Plant Sci. 15, 1303175 (2024).
Hegewald, E. & Wolf, M. Phylogenetic relationships of Scenedesmus and Acutodesmus (Chlorophyta, Chlorophyceae) as inferred from 18S rDNA and ITS-2 sequence comparisons. Plant Syst. Evol. 241, 185–191 (2003).
Breuer, G., Lamers, P. P., Martens, D. E., Draaisma, R. B. & Wijffels, R. H. Effect of light intensity, pH, and temperature on triacylglycerol (TAG) accumulation induced by nitrogen starvation in Scenedesmus obliquus. Bioresour. Technol. 143, 1–9 (2013).
Huesemann, M. et al. DISCOVR strain pipeline screening – Part I: maximum specific growth rate as a function of temperature and salinity for 38 candidate microalgae for biofuels production. Algal Res. 71, 102996 (2023).
Stepka, W., Benson, A. A. & Calvin, M. The path of carbon in photosynthesis: II. Amino acids. Science 108, 304–304 (1948).
Hannon, M., Gimpel, J., Tran, M., Rasala, B. & Mayfield, S. Biofuels from algae: challenges and potential. Biofuels 1, 763–784 (2010).
Calhoun, S. et al. A multi-omic characterization of temperature stress in a halotolerant Scenedesmus strain for algal biotechnology. Commun. Biol. 4, 333 (2021).
Arora, N. et al. Elucidating the unique physiological responses of halotolerant Scenedesmus sp. cultivated in sea water for biofuel production. Algal Res. 37, 260–268 (2019).
Ji, X. et al. The effect of NaCl stress on photosynthetic efficiency and lipid production in freshwater microalga—Scenedesmus obliquus XJ002. Sci. Total Environ. 633, 593–599 (2018).
Kaewkannetra, P., Enmak, P. & Chiu, T. The effect of CO2 and salinity on the cultivation of Scenedesmus obliquus for biodiesel production. Biotechnol. Bioproc. E 17, 591–597 (2012).
Pandit, P. R., Fulekar, M. H. & Karuna, M. S. L. Effect of salinity stress on growth, lipid productivity, fatty acid composition, and biodiesel properties in Acutodesmus obliquus and Chlorella vulgaris. Environ. Sci. Pollut. Res 24, 13437–13451 (2017).
Shetty, P., Gitau, M. M. & Maróti, G. Salinity stress responses and adaptation mechanisms in eukaryotic green microalgae. Cells 8, 1657 (2019).
Zuo, Z., Chen, Z., Zhu, Y., Bai, Y. & Wang, Y. Effects of NaCl and Na2CO3 stresses on photosynthetic ability of Chlamydomonas reinhardtii. Biologia 69, 1314–1322 (2014).
Alkayal, F. et al. Expressed sequence tag (EST) profiling in hyper-saline-shocked Dunaliella salina reveals high expression of protein synthetic apparatus components. Plant Sci. 179, 437–449 (2010).
Jiao, F. et al. Impact of salinity on morphology, growth, and pigment profiles of Scenedesmus obliquus HTB1 under ambient air and elevated CO2 (10%) conditions. Algal Res. 88, 104027 (2025).
Singh, R. P. et al. Unlocking the adaptation mechanisms of the oleaginous microalga Scenedesmus sp. BHU1 under elevated salt stress: a physiochemical, lipidomics and transcriptomics approach. Front. Microbiol. 15, 1475410 (2024).
Bartlett, A. et al. Mapping genome-wide transcription-factor binding sites using DAP-seq. Nat. Protoc. 12, 1659–1672 (2017).
Carreres, B. M. et al. Draft genome sequence of the Oleaginous green alga Tetradesmus obliquus UTEX 393. Genome Announc. 5, e01449-16 (2017).
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Train, C.-M., Glover, N. M., Gonnet, G. H., Altenhoff, A. M. & Dessimoz, C. Orthologous Matrix (OMA) algorithm 2.0: more robust to asymmetric evolutionary rates and more scalable hierarchical orthologous group inference. Bioinformatics 33, i75–i82 (2017).
Biondi, T. et al. The telomere-to-telomere, gapless, phased diploid genome and methylome of the green alga Scenedesmus obliquus UTEX 3031 reveals significant heterozygosity and functional separation of the haplotypes. Algal Res. 79, 103431 (2022).
Gandbhir, M., Rasched, I., Marlière, P. & Mutzel, R. Convergent evolution of amino acid usage in archaebacterial and eubacterial lineages adapted to high salt. Res. Microbiol. 146, 113–120 (1995).
Oren, A. & Mana, L. Amino acid composition of bulk protein and salt relationships of selected enzymes of Salinibacter ruber, an extremely halophilic bacterium. Extremophiles 6, 217–223 (2002).
Choi, H., Tostes, R. C. & Webb, R. C. Mitochondrial aldehyde dehydrogenase prevents ROS-induced vascular contraction in angiotensin-II hypertensive mice. J. Am. Soc. Hypertens. 5, 154–160 (2011).
Buckel, W. & Thauer, R. K. Flavin-based electron bifurcation, ferredoxin, flavodoxin, and anaerobic respiration with protons (Ech) or NAD+ (Rnf) as electron acceptors: a historical review. Front. Microbiol. 9, (2018).
Lodeyro, A. F., Ceccoli, R. D., Pierella Karlusich, J. J. & Carrillo, N. The importance of flavodoxin for environmental stress tolerance in photosynthetic microorganisms and transgenic plants. Mechanism, evolution and biotechnological potential. FEBS Lett. 586, 2917–2924 (2012).
Jadamba, C., Kang, K., Paek, N.-C., Lee, S. I. & Yoo, S.-C. Overexpression of rice Expansin7 (Osexpa7) confers enhanced tolerance to salt stress in rice. Int J. Mol. Sci. 21, 454 (2020).
Le Gall, H. et al. Cell wall metabolism in response to abiotic stress. Plants 4, 112–166 (2015).
Bridges, A. A. & Gladfelter, A. S. Septin form and function at the cell cortex. J. Biol. Chem. 290, 17173 (2015).
Yamazaki, T. et al. Localization and evolution of septins in algae. Plant J. 74, 605–614 (2013).
Oliver, A. E. et al. Non-disaccharide-based mechanisms of protection during drying. Cryobiology 43, 151–167 (2001).
Bhattacharya, S. & Kundu, A. Sugars and sugar polyols in overcoming environmental stresses. In Protective Chemical Agents in the Amelioration of Plant Abiotic Stress 71–101 https://doi.org/10.1002/9781119552154.ch4 (John Wiley & Sons Ltd, 2020).
Akhtar, T. A. et al. Polyprenols are synthesized by a plastidial cis-prenyltransferase and influence photosynthetic performance. Plant Cell 29, 1709–1725 (2017).
Albanese, P. et al. Thylakoid proteome modulation in pea plants grown at different irradiances: quantitative proteomic profiling in a non-model organism aided by transcriptomic data integration. Plant J. 96, 786–800 (2018).
Avidan, O., Brandis, A., Rogachev, I. & Pick, U. Enhanced acetyl-CoA production is associated with increased triglyceride accumulation in the green alga Chlorella desiccata. J. Exp. Bot. 66, 3725–3735 (2015).
Kato, Y. et al. Light/dark cycling causes delayed lipid accumulation and increased photoperiod-based biomass yield by altering metabolic flux in oleaginous Chlamydomonas sp. Biotechnol. Biofuels 12, 39 (2019).
Singh, H., Shukla, M. R., Chary, K. V. R. & Rao, B. J. Acetate and bicarbonate assimilation and metabolite formation in Chlamydomonas reinhardtii: a 13C-NMR study. PLOS ONE 9, e106457 (2014).
Sweetlove, L. J., Beard, K. F. M., Nunes-Nesi, A., Fernie, A. R. & Ratcliffe, R. G. Not just a circle: flux modes in the plant TCA cycle. Trends Plant Sci. 15, 462–470 (2010).
Grigoriev, I. V. et al. PhycoCosm, a comparative algal genomics resource. Nucleic Acids Res. 49, D1004–D1011 (2021).
Birkenbihl, R. P., Jach, G., Saedler, H. & Huijser, P. Functional dissection of the plant-specific SBP-domain: overlap of the DNA-binding and nuclear localization domains. J. Mol. Biol. 352, 585–596 (2005).
Lei, K.-J. et al. Modulation of the phosphate-deficient responses by microRNA156 and its targeted SQUAMOSA promoter binding protein-like 3 in Arabidopsis. Plant Cell Physiol. 57, 192–203 (2016).
Mermod, M. et al. SQUAMOSA promoter-binding protein-like 7 mediates copper deficiency response in the presence of high nitrogen in Arabidopsis thaliana. Plant Cell Rep. 38, 835–846 (2019).
Yamasaki, H., Hayashi, M., Fukazawa, M., Kobayashi, Y. & Shikanai, T. SQUAMOSA promoter binding protein-like7 is a central regulator for copper homeostasis in Arabidopsis. Plant Cell 21, 347–361 (2009).
Jaeger et al. Neochloris oleoabundans is worth its salt: transcriptomic analysis under salt and nitrogen stress. PLOS ONE 13, e0194834 (2018).
Oesterhelt, C. et al. Redox regulation of chloroplast enzymes in galdieria sulphuraria in view of eukaryotic evolution. Plant Cell Physiol. 48, 1359–1373 (2007).
Khan, Z. et al. Cadmium induces GAPDH- and- MDH mediated delayed cell aging and dysfunction in Candida tropicalis 3Aer. Environ. Monit. Assess. 191, 490 (2019).
Liszka, A. et al. Three cytosolic NAD-malate dehydrogenase isoforms of Arabidopsis thaliana: on the crossroad between energy fluxes and redox signaling. Biochem. J. 477, 3673–3693 (2020).
Backhausen, J. E. et al. Transgenic potato plants with altered expression levels of chloroplast NADP-malate dehydrogenase: interactions between photosynthetic electron transport and malate metabolism in leaves and in isolated intact chloroplasts. Planta. 207, 105–114 (1998).
Hildebrandt, T., Knuesting, J., Berndt, C., Morgan, B. & Scheibe, R. Cytosolic thiol switches regulating basic cellular functions: GAPDH as an information hub?. Biol. Chem. 396, 523–537 (2015).
Lebreton, S., Gontero, B., Avilan, L. & Ricard, J. Memory and imprinting effects in multienzyme complexes–II. Kinetics of the bienzyme complex from Chlamydomonas reinhardtii and hysteretic activation of chloroplast oxidized phosphoribulokinase. Eur. J. Biochem 246, 85–91 (1997).
López-Calcagno, P. E., Howard, T. P. & Raines, C. A. The CP12 protein family: a thioredoxin-mediated metabolic switch?. Front. Plant Sci. 5, 9 (2014).
Juergens, M. T., Disbrow, B. & Shachar-Hill, Y. The relationship of triacylglycerol and starch accumulation to carbon and energy flows during nutrient deprivation in Chlamydomonas reinhardtii1. Plant Physiol. 171, 2445–2457 (2016).
Du, Z.-Y. & Benning, C. Triacylglycerol accumulation in photosynthetic cells in plants and algae. in Lipids in Plant and Algae Development (eds. Nakamura, Y. & Li-Beisson, Y.) 179–205 (Springer International Publishing, 2016). https://doi.org/10.1007/978-3-319-25979-6_8.
Weise, S. E., van Wijk, K. J. & Sharkey, T. D. The role of transitory starch in C(3), CAM, and C(4) metabolism and opportunities for engineering leaf starch accumulation. J. Exp. Bot. 62, 3109–3118 (2011).
Hays, S. G. & Ducat, D. C. Engineering cyanobacteria as photosynthetic feedstock factories. Photosynth Res. 123, 285–295 (2015).
Gründel, G., Scheunemann, R., Lockau, W., Zilliges, Y. Impaired glycogen synthesis causes metabolic overflow reactions and affects stress responses in the cyanobacterium Synechocystis sp. PCC 6803. Microbiology 158, (2012).
Boison, D. Adenosine kinase: exploitation for therapeutic gain. Pharm. Rev. 65, 906–943 (2013).
Salomon, S. et al. Betaine lipids: biosynthesis, functional diversity and evolutionary perspectives. Prog. Lipid Res. 97, 101320 (2025).
Jouhet, J. et al. Plant and algal lipidomes: analysis, composition, and their societal significance. Prog. Lipid Res. 96, 101290 (2024).
Liang, Y. et al. Branched-chain amino acid catabolism impacts triacylglycerol homeostasis in Chlamydomonas reinhardtii1[OPEN]. Plant Physiol. 179, 1502–1514 (2019).
Kishor, P. B. K., Hong, Z., Miao, G. H., Hu, C. A. A. & Verma, D. P. S. Overexpression of [delta]-pyrroline-5-carboxylate synthetase increases proline production and confers osmotolerance in transgenic plants. Plant Physiol. 108, 1387–1394 (1995).
Pérez-Arellano, I., Carmona-Álvarez, F., Martínez, A. I., Rodríguez-Díaz, J. & Cervera, J. Pyrroline-5-carboxylate synthase and proline biosynthesis: From osmotolerance to rare metabolic disease. Protein Sci. 19, 372–382 (2010).
Derguini, F. et al. All-trans-retinal is the chromophore bound to the photoreceptor of the alga Chlamydomonas reinhardtii. Photochem. Photobiol. 54, 1017–1021 (1991).
Ballesteros, M. L. et al. LAF1, a MYB transcription activator for phytochrome A signaling. Genes Dev. 15, 2613–2625 (2001).
Schmieder, R. & Edwards, R. Fast identification and removal of sequence contamination from genomic and metagenomic datasets. PLoS One 6, e17288 (2011).
Grigoriev, I. V. et al. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res. 42, D699–D704 (2014).
Kuo, A., Kohler, A., Martin, F. M. & Grigoriev, I. V. Expanding genomics of mycorrhizal symbiosis. Front. Microbiol. 5, (2014).
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. O. & Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 18, 1979–1990 (2008).
Salamov, A. A. & Solovyev, V. V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000).
Birney, E. & Durbin, R. Using GeneWise in the Drosophila annotation experiment. Genome Res. 10, 547–548 (2000).
Solovyev, V., Kosarev, P., Seledsov, I. & Vorobyev, D. Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol. 7, S10 (2006).
Zhou, S.-M., Chen, L.-M., Liu, S.-Q., Wang, X.-F. & Sun, X.-D. De novo assembly and annotation of the Chinese Chive (Allium tuberosum Rottler ex Spr.) transcriptome using the Illumina platform. PLOS ONE 10, e0133312 (2015).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–W120 (2005).
The UniProt Consortium UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Kanehisa, M., Sato, Y., Furumichi, M., Morishima, K. & Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 47, D590–D595 (2019).
Koonin, E. V. et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 5, R7 (2004).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
de Cambiaire, J.-C., Otis, C., Lemieux, C. & Turmel, M. The complete chloroplast genome sequence of the chlorophycean green alga Scenedesmus obliquus reveals a compact gene organization and a biased distribution of genes on the two DNA strands. BMC Evol. Biol. 6, 37 (2006).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Croft, M. T., Lawrence, A. D., Raux-Deery, E., Warren, M. J. & Smith, A. G. Algae acquire vitamin B12 through a symbiotic relationship with bacteria. Nature 438, 90–93 (2005).
Huesemann, M. et al. DISCOVR strain pipeline screening – Part II: Winter and summer season areal productivities and biomass compositional shifts in climate-simulation photobioreactor cultures. Algal Res. 70, 102948 (2023).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Bairoch, A. & Apweiler, R. The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 28, 45–48 (2000).
Luo, W., Friedman, M. S., Shedden, K., Hankenson, K. D. & Woolf, P. J. GAGE: generally applicable gene set enrichment for pathway analysis. BMC Bioinforma. 10, 161 (2009).
Luo, W. & Brouwer, C. Pathview: an R/bioconductor package for pathway-based data integration and visualization. Bioinformatics 29, 1830–1831 (2013).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Yao, Y. et al. Analysis of metabolomics datasets with high-performance computing and metabolite atlases. Metabolites 5, 431–442 (2015).
Smith, C. A. et al. METLIN: a metabolite mass spectral database. Ther. Drug Monit. 27, 747–751 (2005).
Botté, C. Y. et al. Identification of plant-like galactolipids in Chromera velia, a photosynthetic relative of malaria parasites. J. Biol. Chem. 286, 29893–29903 (2011).
McAnoy, A. M., Wu, C. C. & Murphy, R. C. Direct qualitative analysis of triacylglycerols by electrospray mass spectrometry using a linear ion trap. J. Am. Soc. Mass Spectrom. 16, 1498–1509 (2005).
Pluskal, T., Castillo, S., Villar-Briones, A. & Oresic, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinforma. 11, 395 (2010).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Claudel-Renard, C., Chevalet, C., Faraut, T. & Kahn, D. Enzyme-specific profiles for genome annotation: PRIAM. Nucleic Acids Res. 31, 6633–6639 (2003).
Cantalapiedra, C. P., Hernández-Plaza, A., Letunic, I., Bork, P. & Huerta-Cepas, J. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Aramaki, T. et al. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
Karp, P. D. et al. The BioCyc collection of microbial genomes and metabolic pathways. Brief. Bioinform. 20, 1085–1093 (2019).
Ebrahim, A., Lerman, J. A., Palsson, B. O. & Hyduke, D. R. COBRApy: Constraints-based reconstruction and analysis for Python. BMC Syst. Biol. 7, 74 (2013).
Thiele, I. & Palsson, B. Ø A protocol for generating a high-quality genome-scale metabolic reconstruction. Nat. Protoc. 5, 93–121 (2010).
Prigent, S. et al. Meneco, a topology-based gap-filling tool applicable to degraded genome-wide metabolic networks. PLOS Comput. Biol. 13, e1005276 (2017).
Satish Kumar, V., Dasika, M. S. & Maranas, C. D. Optimization based automated curation of metabolic reconstructions. BMC Bioinforma. 8, 212 (2007).
Dev Sarkar, R., Singh, H. B. & Chandra Kalita, M. Enhanced lipid accumulation in microalgae through nanoparticle-mediated approach, for biodiesel production: a mini-review. Heliyon 7, e08057 (2021).
Hucka, M. et al. The systems biology markup language (SBML): a medium for representation and exchange of biochemical network models. Bioinformatics 19, 524–531 (2003).
Gelius-Dietrich, G., Desouki, A. A., Fritzemeier, C. J. & Lercher, M. J. Sybil–efficient constraint-based modelling in R. BMC Syst. Biol. 7, 125 (2013).
Lieven, C. et al. MEMOTE for standardized genome-scale metabolic model testing. Nat. Biotechnol. 38, 272–276 (2020).
O’Malley, R. C. et al. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165, 1280–1292 (2016).
Baumgart, L. A. et al. Persistence and plasticity in bacterial gene regulation. Nat. Methods 18, 1499–1505 (2021).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Castro-Mondragon, J. A. et al. JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 50, D165–D173 (2022).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Acknowledgements
The work (proposals: https://doi.org/10.46936/10.25585/60001010 and https://doi.org/10.46936/10.25585/60001129) conducted by the U.S. Department of Energy Joint Genome Institute (https://ror.org/04xm1d337), a DOE Office of Science User Facility, is supported by the Office of Science of the U.S. Department of Energy, operated under Contract No. DE-AC02-05CH11231. This research was supported by the Bioenergy Technology Office (BETO) within the Department of Energy’s Office of Energy Efficiency and Renewable Energy (EERE) under Agreements NL0032355 and NL0032266. This manuscript has been reviewed and approved for unlimited release (LA-UR-23-25543).
Author information
Authors and Affiliations
Contributions
C.P.S.K.: Methodology, Validation, Formal analysis, Investigation, Data Curation, Writing—Original Draft, Writing - Review & eEditing, Visualization, Project administration. S.C.: Validation, Formal analysis, Investigation, Data Curation, Writing— Review & Editing, Visualization. B.K.: Formal analysis, Writing—Original Draft, Visualization. S.J.E.: Perturbation experiments. OH: Supervision. PMM: Investigation, Formal analysis, Nethodology. T.G.G.: Formal analysis, Methodology, Supervision. Y.K.: Investigation, Resources. Z.M.K.: formal analysis. L.B.: Investigation. I.K.B.: Investigation. B.B.: Investigation. C.C.: Investigation. J.G.: Investigation. A.L.: Investigation. K.L.: Investigation. R.M.: Investigation. T.N.: Investigation. C.D.G.: Investigation, Resources. J.T.: DNA preparation. D.T.: Metabolomics. Y.Z.: Investigation. A.G.: Perturbation experiments. R.C.O.: Investigation, Supervision. J.P.: Conceptualization. J.U.: Conceptualization. MH: Investigation, conceptualization. I.V.G.: Conceptualization, Supervision, Project administration, Funding acquisition, Resources, Writing—Review & Editing. S.R.S.: Conceptualization, Supervision, Project administration, Funding acquisition, Resources, Writing—Review & Editing, Writing—Original Draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Martin Sperfeld, Changhua Shang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Tobias Goris. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kruse, C.P.S., Calhoun, S., Kamel, B. et al. A multi-omic characterization of the physiological responses to salt stress in Scenedesmus obliquus UTEX393. Commun Biol 8, 1632 (2025). https://doi.org/10.1038/s42003-025-09016-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09016-5