Abstract
Urm1 from yeast is a unique ubiquitin-like protein with dual functionality. It has been shown to operate in tRNA thiolation and protein urmylation, combining features typical of bacterial sulfur carriers and classical ubiquitin-like modifiers. Hence, in evolutionary terms, Urm1 may be placed at the crossroad of prokaryotic sulfur transfer and eukaryotic protein conjugation pathways. Prompted by Urm1-like proteins identified in Archaea, we examined Urm1 functional conservation using URM1 gene shuffle from Sulfolobus acidocaldarius to Saccharomyces cerevisiae. We find that archaeal Urm1 conjugates to peroxiredoxin Ahp1, a bona fide urmylation target in yeast, but cannot support tRNA thiolation. Ahp1 conjugation requires sulfur transfer onto the archaeal Urm1 modifier from Uba4, the E1-like urmylation activator in yeast. Thus, thioactivation of archaeal Urm1 and urmylation-like conjugation are conserved and exchangeable processes between Sulfolobus and Saccharomyces. Our survey underlines that Urm1 likely occupies a key role in the evolution of the ubiquitin-like protein family.

Similar content being viewed by others
Introduction
Ubiquitin is an essential protein in eukaryotes1 that conjugates to other proteins in a process known as ubiquitylation. This post-translational protein modification is well-characterized and critical for key biological processes including cell cycle control and proteostasis2. Ubiquitylation is spread among eukaryotes and involves a cascade of E1–E3 enzymes3,4. It starts with a specific E1 activating enzyme that adenylates ubiquitin at its C-terminal glycine. Subsequently, the activated ubiquitin forms a thioester with an E2 conjugating enzyme and conjugates to a target protein with assistance of a substrate-specific E3 ligase5,6.
Genome wide searches in all domains of life have identified proteins with structural features typical of ubiquitin, collectively termed the ubiquitin-like (Ubl) protein family (Fig. 1A). Characteristic structural features of Ubl members are conserved throughout evolution and include a β-grasp fold (β-GF), composed of five β-sheets with a central α-helix, and a C-terminal di-glycine motif or more rarely, a single glycine residue (Fig. 1B)7. Some Ubl members (e.g., SUMO) are even expressed in inactive precursor forms, where the glycine motif must be exposed through proteolytic maturation8. Remarkably, bacterial members of the Ubl family (i.e., ThiS, MoaD) are known as sulfur carrier proteins (SCP) required for the synthesis of thio-cofactors (e.g. thiamine, molybdopterin)9. These proteins are also activated by E1-like enzymes but require subsequent sulfur incorporation at their C-termini in the form of thiocarboxylates for SCP function.
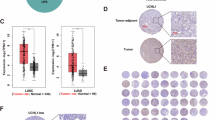
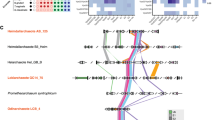
A Ribbon diagrams of ThiS from E. coli (PDB: 1f0z) and Urm1 from S. acidocaldarius (AlphaFold58) are shown next to Urm1 from S. cerevisiae (PDB: 2qjl) and human ubiquitin (PDB: 1ubq). Highlighted are conserved β-GFs with a typical sheet of five antiparallel β-strands (marine), a central α-helix (orange) and C-terminal di-glycine motifs (red). B Sequence alignments of the proteins in (A) using T-Coffee (Expresso)59,60 visualised by ESPript 3.061. Blue framed boxes denote groups of conserved residues (red characters) while identical residues of the C-terminal di-glycine motifs are boxed in red. C Unrooted phylogenetic tree of ubiquitin-like β-GF proteins. Multiple sequence alignment of Urm1 from eukaryotes (M. musculus, H. sapiens, A. thaliana, and D. discoideum) and Archaea (S. solfataricus, S. islandicus, and S. acidocaldarius) as well as eukaryotic Ubl modifiers (SUMO, ATG8, ATG12, and UFM1) and sulfur carrier families (prokaryotic MoaD, ThiS, and eukaryotic MOCS2A) was trimmed (Noisy v1.5.12.146,47) and used in maximum likelihood analyses (RAxML v8.2.1248 implemented in raxmlGUI2.0.1349).
Among eukaryotic Ubl modifiers, the ubiquitin-related modifier 1 (Urm1) uniquely functions as an SCP for tRNA thiolation and urmylation, a Ubl protein conjugation10,11,12,13. Like bacterial SCPs, Urm1 activity requires formation of a thiocarboxylate on its C-terminal di-glycine motif8. Thus, in evolutionary terms, Urm1 is likely placed at the crossroads of eukaryotic conjugation and prokaryotic sulfur transfer pathways14.
In Saccharomyces cerevisiae, Urm1 thiocarboxylation is required for tRNA thiolation and urmylation functions. It involves a network of enzymes with dedicated desulfurase (Nfs1), sulfurtransferase (Tum1) and E1-like (Uba4) activities for sulfur mobilization and transfer onto Urm1 (Supplementary Fig. 1)15. Interestingly, Urm1 conjugation to target proteins differs from ubiquitylation as it appears to be mechanistically simpler and without the need for E2/E3 enzymes16. Recent studies highlight the importance of Urm1 in the maintenance of proteostasis under conditions of cellular stress by inducing phase separation17 or protein persulfidation16. The latter post-translational protein thiolation was shown to be coupled to in vitro urmylation of target protein peroxiredoxin Ahp116,18.
Sulfur-dependence of Urm1 functions in tRNA thiolation and urmylation is conserved in eukaryotes19,20, and proteins related to Urm1 have also been identified in Archaea21,22,23. One such Urm1-like protein from Sulfolobus acidocaldarius (Saci_0669) was shown to conjugate to proteins depending on an E1-like enzyme22. However, whether this modifier also needs sulfur activation or operates in sulfur transfer, such as tRNA thiolation, has yet to be demonstrated and could provide valuable insights into the evolution of prokaryotic and eukaryotic members of the Ubl protein family.
Hence, we studied Urm1 functional conservation using URM1 gene shuffle and complementation between S. acidocaldarius and S. cerevisiae. We show that when expressed in yeast, the archaeal protein is thioactivated by Uba4 and attaches to Ahp1, analogous to S. cerevisiae Urm1. This shows that despite phylogenetic transition between Sulfolobus and Saccharomyces, Urm1-like conjugation has remained largely unchanged over time. Thus, the archaeal modifier qualifies as a candidate stepping-stone for studying the roots of eukaryotic members of the Ubl protein family.
Results and discussion
Urm1 from yeast and Sulfolobus cluster in a monophyletic clade
Structural and sequence comparison of ubiquitin-like β-GF proteins confirm highly conserved motifs and amino acids including the C-terminal di-glycine in all domains of life (Fig. 1A, B). Particularly, Urm1 from S. cerevisiae (ScUrm1) and S. acidocaldarius (SaciUrm1), the proteins of our interest, show striking similarities in their sequences and overall structures (Supplementary Fig. 2). However, structural analysis alone is not sufficient to deduce information on their evolutionary relationships. Therefore, we conducted phylogenetic analyses of various Ubl proteins (i.e., ubiquitin, SUMO, NEDD8, UFM1, ATG8, ATG12, MOCS2A, MoaD, ThiS, and Urm1) that uncover a close relationship between archaeal and eukaryotic Urm1 proteins (Fig. 1C).
In our phylogenetic tree, eukaryotic Ubl proteins (SUMO, ATG8, ATG12, and UFM1) are monophyletic, while sulfur carrier families (prokaryotic MoaD, ThiS, and eukaryotic MOCS2A) are paraphyletic. Urm1 proteins occur as monophyletic group with solid bootstrap support (BS = 80; Supplementary Fig. 3). Clustering of SaciUrm1 (Saci_0669) in the Urm1 clade, which is the most conserved clade of the Ubl family24,25, suggests that they share a common ancestor and might have functional similarities (Fig. 1C and Supplementary Fig. 3).
Sulfolobus URM1 gene shuffle reveals urmylation-like conjugation in yeast
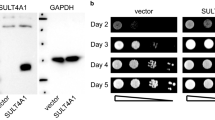
For complementation analysis in trans, we shuffled the S. acidocaldarius gene (Saci_0669) into a yeast URM1 deletion (urm1∆) strain in order to allow for expression of an HA-tagged SaciUrm1 variant. Based on anti-HA Western blots, the archaeal gene was robustly expressed (Supplementary Fig. 4A). To study archaeal Urm1-like conjugation in yeast, we analyzed protein extracts under conditions previously shown to detect protein urmylation11,20. In the presence of isopeptidase inhibitor N-ethylmaleimide (NEM) and based on electrophoretic mobility shift assays (EMSA), SaciUrm1 predominantly formed a candidate conjugate at ~40 kDa (Fig. 2A).
A, B EMSA under reducing and conjugating conditions. Protein extracts from urm1∆, urm1∆AHP1-c-MYC or urm1∆ahp1∆ strains expressing HA-SaciURM1 (A) or HA-ScURM1 (B) were subjected to anti-HA Western blots (top panels) to detect free HA-SaciUrm1 (~20 kDa) or HA-ScUrm1 (~23 kDa) and urmylated Ahp1 (~40 kDa) or Ahp1-c-Myc (~50 kDa) conjugates. Unconjugated Ahp1 (~20 kDa) or Ahp1-c-Myc (~25 kDa) is monitored in anti-Ahp1 Western blots (middle panels). C Urmylation analysis of Ahp1 mutants by EMSA. Protein extracts from urm1∆ahp1∆ strains co-expressing HA-SaciURM1 and indicated AHP1 substitutions or carrying empty vector (ev) were subjected to anti-HA Western blots (top panel). Free forms of HA-SaciUrm1 (~20 kDa) and urmylated Ahp1 (~40 kDa) conjugates are marked. Free Ahp1 (~20 kDa) is detected by an anti-Ahp1 blot (middle panel). Protein loading was controlled with anti-Cdc19 Western blots (bottom panels in A–C).
This shifted species is similar to the one formed between ScUrm1 and yeast peroxiredoxin Ahp1 (Fig. 2B)11,26. Accordingly, this conjugate was not detected in extracts from an ahp1∆ null-mutant but became even further shifted in size (~50 kDa) when Ahp1 was c-Myc-tagged (Fig. 2A). Thus, archaeal SaciUrm1 attaches to Ahp1 in a fashion that is analogous to bona fide Ahp1 urmylation in yeast (Fig. 2A, B)20. The latter requires Lys-32 on Ahp1 as major Urm1 acceptor site and two cysteine residues (Cys-31, Cys-62). While Cys-62 is critical for peroxiredoxin activity and attachment of Urm1 at Lys-32, substitution of Cys-31 with serine shows residual activity and Ahp1 urmylation in vivo26. We examined whether these molecular Urm1 acceptor criteria are conserved for urmylation-like conjugation of Ahp1 by archaeal SaciUrm1. EMSA data obtained from AHP1 substitution mutants clearly show that SaciUrm1 follows the above principles (Fig. 2C).
Hence, Urm1-like conjugation of Ahp1 by SaciUrm1 was unaltered, drastically decreased, or entirely abolished in Ahp1 variants with respective substitutions at Cys-31 (C31S), Lys-32 (K32R), or Cys-62 (C62S) alone and in tandem with Cys-31 (C31,62S) (Fig. 2C). Summing up, our data reveal that substrate specific requirements for attachment to Ahp1 are shared between an archaeal Urm1-like modifier and yeast Urm1. Thus, in evolutionary terms, Urm1-like conjugation by prokaryotic and eukaryotic modifiers of the Ubl family appears to be conserved.
Sulfur-dependence of urmylation is conserved from archaea to yeast
Prompted by the exchangeability between archaeal and yeast Urm1 for Ahp1 conjugation, we next examined whether sulfur dependence of Urm1 functions (i.e., urmylation, tRNA thiolation) known from yeast (Fig. 3A and Supplementary Fig. 1) is a feature conserved with SaciUrm1, too. Sulfur requirement for conjugation by SaciUrm1 was studied with sulfur transfer mutants previously shown to compromise Ahp1 urmylation in yeast15. The data show decreased or fully abolished Ahp1 conjugation by SaciUrm1 in the absence of Tum1 sulfurtransferase (tum1∆) or E1-like Uba4 thioactivator (uba4∆), respectively (Fig. 3B). Thus, Urm1-like conjugation to Ahp1 by SaciUrm1 appears to be sulfur-dependent and requires Uba4, the yeast enzyme generating the crucial Urm1 thiocarboxylate12,13.
A Scheme of the sulfur transfer in the yeast Urm1 pathway. Sulfur (red) flow required for Urm1 activation starts with sulfur extraction from cysteine through desulfurase Nfs1, followed by direct (Nfs1) or indirect (Tum1) sulfur transfer to the catalytic active cysteine (C397) of E1-like enzyme Uba4 that thiocarboxylates the C-terminus of Urm1. B Western blot analysis for sulfur dependence of SaciUrm1 urmylation. Protein extracts from urm1∆uba4∆, urm1∆uba4∆tum1∆ or urm1∆uba4∆ahp1∆ strains expressing HA-SaciURM1 with (+) or without (-) FLAG-UBA4 or FLAG-UBA4-C397S were subjected to an anti-HA Western blot (top panel) to detect free HA-SaciUrm1 (~20 kDa) and urmylated Ahp1 (~40 kDa) conjugates. FLAG-Uba4 or FLAG-Uba4-C397S (~55 kDa) and Ahp1 (~20 kDa) were detected by anti-FLAG and anti-Ahp1 antibodies (middle panels). Protein loading control involved an anti-Cdc19 Western blot (bottom panel).
Since thiocarboxylation of Urm1 requires a catalytic Cys residue (C397) in the rhodanese domain of Uba4 for sulfur transfer (Fig. 3A and Supplementary Fig. 1)13,15,27, we next studied SaciUrm1 conjugation to Ahp1 in yeast expressing a Uba4 mutant (C397S) defective in sulfur-dependent urmylation. Our EMSA data show a drastic effect, with Ahp1 conjugation by SaciUrm1 almost being abolished in the C397S mutant (Fig. 3B). This strongly suggests thioactivation of the archaeal modifier by Uba4, a scenario analogous to ScUrm1 in yeast. Likewise, we found that the conserved C-terminal di-glycine motifs in ScUrm1 and SaciUrm1 (Fig. 1B and Supplementary Fig. 2B) are required for urmylation (Supplementary Fig. 4B)10.
Thus, the archaeal modifier failed to form Ahp1 conjugates when its C-terminal Gly-84 was removed (∆G) (Supplementary Fig. 4B), strongly suggesting that a thiocarboxylated C-terminus plays a crucial and universally conserved role for modifications by Urm1-like proteins. To further investigate thiocarboxylation of SaciUrm1 by Uba4, we used SDS-PAGE containing APM ([(N-acryloylamino)phenyl]mercuric chloride)28. Sulfur-specific adduction of thiolated SaciUrm1 to APM induces mobility shifts that are detectable by Western blot (Fig. 4A)29,30. Our data show that APM indeed forms adducts with SaciUrm1 provided both functional Uba4 and a proper C-terminal di-glycine motif are present (Fig. 4B). These are requirements for APM adduction to SaciUrm1 that are very similar to APM retardation seen with ScUrm1 (Fig. 4C).
A Scheme illustrating APM usage in SDS-PAGE to diagnose Urm1 thiocarboxylation. Urm1-COOH is thio-activated at its C-terminal glycine by Uba4 resulting in Urm1-COSH. Adduct formation with thiocarboxylated Urm1 through the thiophilic character of APM results in retarded migration indicating the presence of sulfur in the thiocarboxylate (-COSH). B, C EMSA diagnostic for thiocarboxylation of SaciUrm1 (B) and ScUrm1 (C) by Uba4. Protein extracts from urm1∆uba4∆ strains coexpressing ScUBA4 (+) or not (-) with HA-tagged SaciUrm1 or HA-tagged ScUrm1 or their terminal glycine deletion (∆G) were separated by SDS-PAGE containing a layer of APM as specified (+/-). APM dependent retardation of anti-HA signals (right panels) distinguishes sulfur-free (-COOH) forms of HA-SaciUrm1 (B) or HA-ScUrm1 (C) from thiocarboxylated (-COSH) forms.
In sum, our data indicate that thiocarboxylation and urmylation principles known from yeast also apply to the archaeal Urm1-like modifier and that sulfur-dependent performance of SaciUrm1 in urmylation-like conjugation is conserved from prokaryotes to eukaryotes.
SaciUrm1 fails to mediate tRNA thiolation in yeast despite being thioactivated
With the confirmation of SaciUrm1 thiocarboxylation, we next asked whether the archaeal modifier can support sulfur transfer for tRNA thiolation. In yeast, a thiolase heterodimer (Ncs2-Ncs6) incorporates sulfur from ScUrm1 into tRNA anticodons at wobble uridines (U34). Together with other tRNA modifiers, this forms composite 5-methoxycarbonyl-methyl-2-thio (mcm5s2) groups on tRNAGln, tRNALys, and tRNAGlu (tQKE) (Supplementary Fig. 1)12,13,31. Direct comparison of tRNA thiolation capacities between SaciUrm1 and ScUrm1 by liquid chromatography-tandem mass spectrometry (LC-MS/MS) revealed that in contrast to ScUrm1, mcm5s2U34 formation is absent from urm1∆ cells expressing SaciUrm1 (Fig. 5A).
A LC-MS/MS-based tRNA modification analysis of U34 nucleosides (ncm5U34, mcm5U34, mcm5s2U34) from urm1∆ strains expressing ScURM1, SaciURM1 or empty vector (ev). The red arrow marks sulfur-dependent mcm5s2U34 modification. Each measured modified nucleoside signal was normalized to total uridine content. Graphs represent the mean ± SD from n = 3 biologically independent samples; individual data points displayed as dots. Statistical significance was performed between ev control and ScURM1 or SaciURM1 expressing cells with two-sided Student’s t-test. p-values < 0.005 are indicated as ** and not significant as ns. Data values are available in Supplementary Table 4 and the Supplementary Data Excel file. B Phenotypic growth tests of urm1∆ strains expressing ScURM1, SaciURM1 alone or with overexpression of tRNAGln, tRNALys, and tRNAGlu (tQKE) from multi-copy plasmids on medium with DNA damaging reagent zeocin (top panel, right) or without (control). Similarly, thermo-sensitive growth was analyzed at 30 °C and 37 °C (bottom panel) in an urm1∆ mutant that lacks tRNA pseudouridylation gene DEG1 (urm1∆deg1∆).
Similarly, expression of SaciUrm1, rather than ScUrm1, associates with excess levels of 5-methoxycarbonyl-methyl (mcm5) groups that are typical of a thiolation defect (Fig. 5A). Furthermore, LC-MS/MS revealed no impact of SaciUrm1 on sulfur-independent 5-carbamoyl-methyl (ncm5) groups (Fig. 5A), thus further indicating a thiolation-specific block of SaciUrm1 in mcm5s2U34 synthesis. We also confirmed this defect in vivo by showing that SaciUrm1 expression cannot rescue phenotypes that result from U34 thiolation loss in urm1∆ cells (Fig. 5B). In yeast, lack of tRNA thiolation at the wobble position U34 does not result in a reduced abundance of hypomodified tRNAs, but has been shown to reduce the binding affinity of the hypomodified anticodon to the cognate codon in the ribosomal A-site32,33. The defect results in inefficient decoding and error-prone mRNA translation, which is why the growth and viability of URM1 gene deletion (urm1∆) strains becomes significantly compromised, particularly upon cultivation at elevated temperatures or in the presence of chemical stressors such as zeocin (Fig. 5B). Intriguingly, overexpression of hypothiolated tRNAs (i.e., tQKE) compensates for the decoding inefficiency of urm1∆ cells and partially rescues the thermo and drug sensitivity phenotypes (Fig. 5B). Unlike with ScUrm1 or partial rescue by tRNA overexpression (tQKE, see above), SaciUrm1 expression, however, could not complement zeocin sensitivity or synthetic lethality in tandem with tRNA pseudouridylation mutant deg1∆ (Fig. 5B)13,20,34.
Hence, our data show that despite being thiocarboxylated by Uba4 for sulfur-dependent Urm1-like conjugation (Fig. 4), SaciUrm1 cannot transfer the sulfur for tRNA thiolation in yeast (Fig. 5). Such incompatibility may be explained by SaciUrm1 failing to properly interact with the thiolase complex in yeast. In contrast to eukaryotic thiolase heterodimers, archaeal homologues rather form homodimers, indicating an evolutionary diversification from homo- to heterodimeric tRNA thiolation enzymes30,35,36. This could reflect an evolutionary barrier that may have resulted in functional specification of the eukaryotic thiolase complex that is unable to recognize and use the archaeal Urm1 protein as sulfur donor36,37.
To the best of our knowledge there is no data linking SaciUrm1 directly to sulfur transfer associated with tRNA thiolation in Sulfolobus. However, archaeal tRNA thiomodifications do exist as has been recently uncovered by comprehensive LC-MS/MS profiles on tRNAs isolated from several Archaea including S. acidocaldarius38. Thus, a sulfur donor function of SaciUrm1 for tRNA thiolation likely exists in its host organism S. acidocaldarius and needs further investigations. These should also include the two open reading frames from S. acidocaldarius, predicted to encode Ubl proteins (Saci_0952 and Saci_1652) (Supplementary Fig. 3). They may represent potential SCPs involved in different sulfur transfer reactions, as is known from other organisms such as Haloferax volcanii21 and Thermococcus kodakarensis23 that use several SCPs. To conclude, our gene shuffle approach has uncovered that the conserved Urm1-like conjugation function of SaciUrm1 can be separated in yeast from the tRNA thiolation function of ScUrm1.
Conclusions
Our gene shuffle experiments demonstrate that stable expression of an archaeal Urm1 homolog (SaciUrm1) in yeast can pave the way for evolutionary studies on prokaryotic and eukaryotic members of the ubiquitin-like protein family. We show SaciUrm1 cooperates with yeast Urm1 pathway components, in particular Uba4, for thioactivation and engages in sulfur-dependent Urm1-like conjugation to Ahp1. The latter peroxiredoxin is a bona fide Urm1 target in yeast, indicating that ubiquitin-like urmylation is conserved among prokaryotes and eukaryotes. Despite detectable thioactivation in vivo, SaciUrm1 fails to support sulfur transfer for tRNA thiolation in yeast. Hence, dual-functional sulfur carrier roles for Urm1 in tRNA thiolation and protein urmylation can be separated from one another by expressing an archaeal Urm1 homolog in yeast. Such scenario may be beneficial for further research that focuses on urmylation mechanisms and relevance, particularly when tRNA thiolation is to be avoided. Alternatively, in an attempt to rescue the tRNA thiolation incompatibility and overcome cross-species barriers, such separation of function scenarios may provide new evolutionary insights into Urm1 dual-functionality. Hence, we consider the Urm1-like modifier from Sulfolobus at a phylogenetic position suited to further investigate evolutionary roots of the ubiquitin-like protein family.
Methods
Yeast strains and plasmid constructions
S. cerevisiae strains used in this study (Supplementary Table 1) were grown on complete (YPD) or synthetic (SC) media according to standard methods39. Thermosensitivity was analyzed on SC medium at 30 °C or 37 °C, while drug sensitivity was analyzed on YPD medium at 30 °C using 0.02 mg/ml zeocin, after 2–3 days. Yeast cell transformations with PCR products or gene shuffle plasmids (Supplementary Table 2) were done according to Gietz & Woods40. Genomic deletions were generated by PCR41,42 with gene-specific oligonucleotides and confirmed with oligonucleotides binding outside the target loci (Supplementary Table 3).
For multi-copy ScURM1 expression, pAJ45 (Supplementary Table 2) was amplified with primers M13_FW_-40 and M13_RV_-27 (Supplementary Table 3) to obtain HA-tagged ScURM1 with ADH1 promoter and CYC1 terminator. The amplified product was cut with HindIII and SalI and ligated into YEplac195, resulting in pKZ25 (Supplementary Table 2). SaciURM1 was amplified directly from archaeal genomic DNA with primers SaciUrm1_Fw_NotI and SaciUrm1_Rv_MlsI (Supplementary Table 3) and cloned into pKZ25 (Supplementary Table 2), replacing ScURM1 with SaciURM1, resulting in pKZ6 (Supplementary Table 2). Similarly, SaciURM1 and ScURM1 were amplified from pKZ6 and pKZ25 (Supplementary Table 2) with reverse primers SaciURM1dG_MlsI_RV or ScURM1dG_MlsI_RV, deleting the C-terminal glycine residue (∆G), generating plasmids pKZ48 and pKZ50 (Supplementary Table 2), respectively. For single-copy SaciURM1 expression, pKZ6 (Supplementary Table 2) was amplified with primers M13_FW_-40 and M13_RV_-27 (Supplementary Table 3) to obtain PADH1-3xHA-SaciURM1-TCYC1 that was cloned into YCplac33 via HindIII and SalI, resulting in pKZ2 (Supplementary Table 2). To purify ScUrm1 and SaciUrm1 with the Strep-Tactin®XT 4Flow® high capacity resin (IBA Lifesciences, 2-5030-002), a construct of PADH1-8xHis-TwinStrep-TCYC1 was synthesized by GenScript Biotech (Netherlands) B.V. and cloned into YEplac195, resulting in pLK247 (Supplementary Table 2). The HA-tagged URM1 genes were amplified from pKZ25 and pKZ6 (Supplementary Table 2) and cloned into pLK247 (Supplementary Table 2) via NotI and MlsI, resulting in pKZ76 and pKZ77 (Supplementary Table 2), respectively. Truncated ∆G mutants thereof were generated as described above, resulting in plasmids pKZ82 and pKZ83 (Supplementary Table 2).
For expression of FLAG-tagged UBA4 and a C397S mutant, the PADH1-UBA4-TCYC1 construct from pAJ16 (Supplementary Table 2) was cloned via BamHI and EcoRI in pRS313 (Supplementary Table 2) and N-terminal FLAG tagging was performed with primers N-terminal FLAG FW and N-terminal FLAG RV (Supplementary Table 3), resulting in pKZ60 (Supplementary Table 2). Site-directed mutagenesis was done with primers UBA4_C397S_FW and UBA4_C397S_RV (Supplementary Table 3) to generate pKZ65 (Supplementary Table 2). All generated plasmids were checked by Sanger sequencing (Microsynth Seqlab GmbH).
Phylogenetic analysis
Phylogenetic trees of ubiquitin-like proteins from Eukaryota, Archaea and Bacteria were inferred using amino acid sequences obtained from the publicly available UniProt database43. Multiple sequence alignment was done by MAFFT version 7 using L-large-INS-1 as tree-based progressive method with default parameters44,45. Alignment was improved by removing homoplastic and random-like positions with Noisy (v1.5.12.1) and default parameters46,47. Maximum likelihood analyses were conducted using RAxML v8.2.1248 implemented in raxmlGUI2.0.1349. The best tree was obtained from 20 independent runs under empirical LG + G model, which was selected with ModelTest-NG v0.1.750 implemented in raxmlGUI. Bootstrap support values were calculated from 100 pseudoreplicates of the alignment. The resulting best tree was visualized by iTol version 651 and edited with CorelDRAW Suite X5.
Urmylation studies
Protein urmylation analysis used a lysis buffer for mechanical yeast cell disruption (10 mM K-HEPES pH 7.0, 10 mM KCl, 1.5 mM MgCl2, 0.5 mM PMSF, 2 mM benzamidine) containing complete protease inhibitors and 2.5 mM NEM. To detect unconjugated Urm1, NEM was excluded. Samples for Western blot analysis were prepared with equal protein concentrations according to Bradford52 in SDS sample buffer (62.5 mM Tris-HCl pH 6.8, 2% SDS, 10% glycerol, 0.002% bromophenol blue, 5% β-mercaptoethanol) after Laemmli53. Samples were run by SDS-PAGE and proteins transferred to PVDF membranes. For protein detection, primary anti-HA (1:5000; F7, Santa Cruz Biotechnology or 2-2.2.14, Invitrogen) and anti-FLAG (1:5000; FG4R, Invitrogen) antibodies were used. Expression of Ahp1 was verified with anti-Ahp1 serum54 (1:4000). Protein loading was controlled with anti-Cdc19 antibody (1:400000). Western blot signals were visualized using the LiCor Odyssey FC Imager System and Image Studio software (v.5.2.5, Li-COR Biosciences). All Western blots were performed with a minimum of three biological replicates.
Confirmation of Urm1 thiocarboxylation
To confirm thiocarboxylation of Urm1 in vivo, an additional Twin-Strep-tag® was used for protein purification using the Strep-Tactin®XT 4Flow® high capacity resin (IBA Lifesciences, 2-5030-002). Yeast cells grown to logarithmic phase were mechanically lysed in wash buffer W (100 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1 mM EDTA) containing complete protease inhibitors and 0.1% (v/v) NP-40. Proteins were purified according to manufacturer’s instructions by eluting in buffer BXT (100 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1 mM EDTA, 50 mM biotin). Western blot analysis was performed as described above with slight modifications. Specifically, SDS-PAGE gels were supplemented with 20 μg/ml APM28, and sample buffer was used without β-mercaptoethanol.
Yeast tRNA isolation and wobble uridine modification profiling
Yeast tRNA was isolated as described55, using NucleoZOL reagent (Macherey-Nagel). The pelleted tRNA was washed once with 75% ethanol and stored in 100% ethanol. LC-MS/MS analysis of wobble uridine modifications (ncm5U34, mcm5U34, mcm5s2U34) was performed according to Biedenbänder et al56. with the analyzed amount increased to 1 µg of digested RNA sample spiked with 100 ng of internal standard. Absolute quantification of biological triplicates (n = 3) used internal and external calibration as described57 with the total amount of modified nucleosides being normalized to the amount of uridines.
Statistics and reproducibility
All Western blots and phenotypic growth tests were performed with a minimum of three biological replicates. Graphical illustration and statistical analysis were performed with GraphPad Prism (version 8.0.2). LC-MS/MS data in Fig. 5A represent the mean ± SD from n = 3 biologically independent samples and individual data points are displayed as dots. Data values are available in Supplementary Table 4 and the Supplementary Data Excel file. Statistics were performed with two-sided Student’s t-tests with p-values < 0.005 indicated by ** and non-significant changes with ns.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Complete data for Fig. 5A is available in Supplementary Table 4 and the Supplementary Data Excel file. Full Western blot and phenotypic growth test images are shown in the Supplementary Raw Data section of the Supplementary Information file. The Multiple sequence alignment for phylogenetic analysis can be provided in FASTA format upon reasonable request. All other data are incorporated into the article and its Supplementary Information as well as the Supplementary Data Excel file.
References
Goldstein, G. et al. Isolation of a polypeptide that has lymphocyte-differentiating properties and is probably represented universally in living cells. Proc. Natl. Acad. Sci. USA 72, 11–15 (1975).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Ciechanover, A., Elias, S., Heller, H. & Hershko, A. ‘Covalent affinity’ purification of ubiquitin-activating enzyme. J. Biol. Chem. 257, 2537–2542 (1982).
Ciechanover, A., Heller, H., Elias, S., Haas, A. L. & Hershko, A. ATP-dependent conjugation of reticulocyte proteins with the polypeptide required for protein degradation. Proc. Natl. Acad. Sci. USA 77, 1365–1368 (1980).
Pickart, C. M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Hershko, A. Ubiquitin: roles in protein modification and breakdown. Cell 34, 11–12 (1983).
Burroughs, A. M., Iyer, L. M. & Aravind, L. The natural history of ubiquitin and ubiquitin-related domains. Front. Biosci. 17, 1433–1460 (2012).
Sengupta, S. & Pick, E. The ubiquitin-like proteins of Saccharomyces cerevisiae. Biomolecules 13, 734 (2023).
Cappadocia, L. & Lima, C. D. Ubiquitin-like Protein Conjugation: Structures, Chemistry, and Mechanism. Chem. Rev. 118, 889–918 (2018).
Furukawa, K., Mizushima, N., Noda, T. & Ohsumi, Y. A protein conjugation system in yeast with homology to biosynthetic enzyme reaction of prokaryotes. J. Biol. Chem. 275, 7462–7465 (2000).
Goehring, A. S., Rivers, D. M. & Sprague, G. F. Attachment of the ubiquitin-related protein Urm1p to the antioxidant protein Ahp1p. Eukaryot. Cell 2, 930–936 (2003).
Nakai, Y., Nakai, M. & Hayashi, H. Thio-modification of yeast cytosolic tRNA requires a ubiquitin-related system that resembles bacterial sulfur transfer systems. J. Biol. Chem. 283, 27469–27476 (2008).
Leidel, S. et al. Ubiquitin-related modifier Urm1 acts as a sulphur carrier in thiolation of eukaryotic transfer RNA. Nature 458, 228–232 (2009).
Pedrioli, P. G. A., Leidel, S. & Hofmann, K. Urm1 at the crossroad of modifications. ‘Protein modifications: beyond the usual suspects’ review series. EMBO Rep. 9, 1196–1202 (2008).
Jüdes, A., Bruch, A., Klassen, R., Helm, M. & Schaffrath, R. Sulfur transfer and activation by ubiquitin-like modifier system Uba4•Urm1 link protein urmylation and tRNA thiolation in yeast. Micro. Cell 3, 554–564 (2016).
Ravichandran, K. E. et al. E2/E3-independent ubiquitin-like protein conjugation by Urm1 is directly coupled to cysteine persulfidation. EMBO J. 41, e111318 (2022).
Cairo, L. V. et al. Stress-dependent condensate formation regulated by the ubiquitin-related modifier Urm1. Cell 187, 4656–4673 (2024).
Kaduhr, L. et al. Urm1, not quite a ubiquitin-like modifier?. Micro. Cell 8, 256–261 (2021).
Nakai, Y., Harada, A., Hashiguchi, Y., Nakai, M. & Hayashi, H. Arabidopsis molybdopterin biosynthesis protein Cnx5 collaborates with the ubiquitin-like protein Urm11 in the thio-modification of tRNA. J. Biol. Chem. 287, 30874–30884 (2012).
Jüdes, A. et al. Urmylation and tRNA thiolation functions of ubiquitin-like Uba4·Urm1 systems are conserved from yeast to man. FEBS Lett. 589, 904–909 (2015).
Humbard, M. A. et al. Ubiquitin-like small archaeal modifier proteins (SAMPs) in Haloferax volcanii. Nature 463, 54–60 (2010).
Anjum, R. S. et al. Involvement of a eukaryotic-like ubiquitin-related modifier in the proteasome pathway of the archaeon Sulfolobus acidocaldarius. Nat. Commun. 6, 8163 (2015).
Hidese, R. et al. Functional redundancy of ubiquitin-like sulfur-carrier proteins facilitates flexible, efficient sulfur utilization in the primordial archaeon Thermococcus kodakarensis. mBio 15, e0053424 (2024).
Yang, Z. et al. A phylogenetic analysis of the ubiquitin superfamily based on sequence and structural information. Mol. Biol. Rep. 41, 6083–6088 (2014).
Xu, J. et al. Solution structure of Urm1 and its implications for the origin of protein modifiers. Proc. Natl. Acad. Sci. USA 103, 11625–11630 (2006).
Brachmann, C. et al. Redox requirements for ubiquitin-like urmylation of Ahp1, a 2-Cys peroxiredoxin from yeast. Redox Biol. 30, 101438 (2020).
Sokołowski, M. et al. Molecular basis for thiocarboxylation and release of Urm1 by its E1-activating enzyme Uba4. Nucleic Acids Res. 52, 13980–13995 (2024).
Igloi, G. L. Interaction of tRNAs and of phosphorothioate-substituted nucleic acids with an organomercurial. Probing the chemical environment of thiolated residues by affinity electrophoresis. Biochemistry 27, 3842–3849 (1988).
Van der Veen, A. G. et al. Role of the ubiquitin-like protein Urm1 as a noncanonical lysine-directed protein modifier. Proc. Natl. Acad. Sci. USA 108, 1763–1770 (2011).
Chen, M. et al. The [4Fe-4S] cluster of sulfurtransferase TtuA desulfurizes TtuB during tRNA modification in Thermus thermophilus. Commun. Biol. 3, 1–9 (2020).
Noma, A., Sakaguchi, Y. & Suzuki, T. Mechanistic characterization of the sulfur-relay system for eukaryotic 2-thiouridine biogenesis at tRNA wobble positions. Nucleic Acids Res 37, 1335–1352 (2009).
Ranjan, N. & Rodnina, M. V. Thio-Modification of tRNA at the Wobble Position as Regulator of the Kinetics of Decoding and Translocation on the Ribosome. J. Am. Chem. Soc. 139, 5857–5864 (2017).
Johansson, M. J. O., Esberg, A., Huang, B., Björk, G. R. & Byström, A. S. Eukaryotic wobble uridine modifications promote a functionally redundant decoding system. Mol. Cell Biol. 28, 3301–3312 (2008).
Lin, S. Role of the ubiquitin-like protein Urm1 as a protein modifier. https://doi.org/10.5282/edoc.21608. (Ludwig-Maximilians-Universität München, 2017).
Chavarria, N. E. et al. Archaeal Tuc1/Ncs6 Homolog Required for Wobble Uridine tRNA Thiolation Is Associated with Ubiquitin-Proteasome, Translation, and RNA Processing System Homologs. PLoS One 9, e99104 (2014).
Bimai, O. et al. The thiolation of uridine 34 in tRNA, which controls protein translation, depends on a [4Fe-4S] cluster in the archaeum Methanococcus maripaludis. Sci. Rep. 13, 5351 (2023).
Guy, M. P. & Phizicky, E. M. Two-subunit enzymes involved in eukaryotic post-transcriptional tRNA modification. RNA Biol. 11, 1608–1618 (2014).
Wolff, P. et al. Comparative patterns of modified nucleotides in individual tRNA species from a mesophilic and two thermophilic archaea. RNA 26, 1957 (2020).
Sherman, F. Getting started with yeast. Methods Enzymol. 194, 3–21 (1991).
Gietz, R. D. & Woods, R. A. Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol. 350, 87–96 (2002).
Gueldener, U., Heinisch, J., Koehler, G. J., Voss, D. & Hegemann, J. H. A second set of loxP marker cassettes for Cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res 30, e23 (2002).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
The UniProt Consortium et al UniProt: the Universal Protein Knowledgebase in 2025. Nucleic Acids Res. 53, D609–D617 (2025).
Kuraku, S., Zmasek, C. M., Nishimura, O. & Katoh, K. aLeaves facilitates on-demand exploration of metazoan gene family trees on MAFFT sequence alignment server with enhanced interactivity. Nucleic Acids Res 41, W22–W28 (2013).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform 20, 1160–1166 (2019).
Dress, A. W. M. et al. Noisy: identification of problematic columns in multiple sequence alignments. Algorithms Mol. Biol. 3, 7 (2008).
Grünewald, S., Forslund, K., Dress, A. & Moulton, V. QNet: an agglomerative method for the construction of phylogenetic networks from weighted quartets. Mol. Biol. Evol. 24, 532–538 (2007).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Edler, D., Klein, J., Antonelli, A. & Silvestro, D. raxmlGUI 2.0: A graphical interface and toolkit for phylogenetic analyses using RAxML. Methods Ecol. Evolut. 12, 373–377 (2021).
Darriba, D. et al. ModelTest-NG: A New and Scalable Tool for the Selection of DNA and Protein Evolutionary Models. Mol. Biol. Evol. 37, 291–294 (2020).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Laemmli, U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
Iwai, K., Naganuma, A. & Kuge, S. Peroxiredoxin Ahp1 acts as a receptor for alkylhydroperoxides to induce disulfide bond formation in the Cad1 transcription factor. J. Biol. Chem. 285, 10597–10604 (2010).
Krutyhołowa, R. et al. Kti12, a PSTK-like tRNA dependent ATPase essential for tRNA modification by Elongator. Nucleic Acids Res. 47, 4814–4830 (2019).
Biedenbänder, T. et al. RNA modifications stabilize the tertiary structure of tRNAfMet by locally increasing conformational dynamics. Nucleic Acids Res 50, 2334–2349 (2022).
Thüring, K., Schmid, K., Keller, P. & Helm, M. Analysis of RNA modifications by liquid chromatography-tandem mass spectrometry. Methods 107, 48–56 (2016).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Di Tommaso, P. et al. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39, W13–W17 (2011).
Armougom, F. et al. Expresso: automatic incorporation of structural information in multiple sequence alignments using 3D-Coffee. Nucleic Acids Res. 34, W604–W608 (2006).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Acknowledgements
The authors thank Dr Kuge (Tohoku Pharmaceutical University, Japan) and Dr Thorner (University of California, USA) for providing anti-Ahp1 and anti-Cdc19 antibodies, respectively, and Dr Ohsumi (Tokyo Institute of Technology, Japan) and Dr Robinson (University of Lancaster, United Kingdom) for the HA-Urm1 expression plasmid and SaciURM1 DNA, respectively. We also thank Drs Igloi and Albers (University of Freiburg, Germany) for kindly donating APM and reading the manuscript and Drs West (The College of Wooster, USA), Hering, Mayer, and Klassen, as well as Niklas Metzendorf (University of Kassel, Germany) for their technical assistance and valuable comments on the manuscript. We acknowledge Urm1 project support by Deutsche Forschungsgemeinschaft (DFG), Germany, to R.S. (SCHA750/15-2) and their SPP1784 program Chemical Biology of Native Nucleic Acid Modifications to R.S. (SCHA750/20-2). Work in the laboratory of M.H. was supported by DFG Collaborative Research Center RMaP (TRR 319, TP C03, Project Id 439669440). L.K. has been awarded a PhD scholarship by the Otto Braun-Fonds (B. Braun, Melsungen AG, Germany). L.K. and K.Z. received support by ZFF-Pilot grants #2887 and #2620 (Zentraler Forschungsfonds, University of Kassel, Germany) awarded to R.S.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
K.Z., L.K., and R.S. designed the experiments; K.Z. and L.K. performed the phylogenetic analyses and the molecular biological experiments; L.B. carried out the LC-MS/MS; K.Z., L.K., L.B., M.H., and R.S. analyzed the data; K.Z., L.K., and R.S. wrote the paper; M.H. and R.S. secured funding support (for details, see above).
Corresponding author
Ethics declarations
Competing interests
Author M.H. declares the following competing interest: consultant for Moderna Inc. All other authors declare no competing interests.
Peer review
Peer review information
Communications Biology Marcin Suskiewicz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Elah Pick and Mengtan Xing. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zupfer, K., Kaduhr, L., Bessler, L. et al. Evolutionary conservation of ubiquitin-like protein urmylation as revealed by URM1 gene shuffle from archaea to yeast. Commun Biol 8, 1637 (2025). https://doi.org/10.1038/s42003-025-09212-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09212-3