Abstract
The protein disulfide isomerase (PDI) is a conserved redox protein that plays an essential role during host cell adhesion and invasion in apicomplexan parasites. PDI is also expressed in Cryptosporidium parvum, but its role remains unknown. Here we show that C. parvum PDI (CpPDI, encoded by cgd6_120) contains two conserved thioredoxin domains with catalytic domains (CGHC), is highly expressed early in development in C. parvum, discharged from the invading sporozoites into host cells, and is mostly distributed in endoplasmic reticulum and the cytosol of free sporozoites. In addition, the PDI inhibitor bacitracin, DNTB, anti-CpPDI antibodies and recombinant CpPDI protein could effectively with interfere the invasion of sporozoites into host cells. LC-MS/MS and Co-immunoprecipitation analysis identified Keratin, type I cytoskeletal 18 protein (KRT-18), as a PDI interactor that is not redox-dependent and whose interaction is not related to the catalytic activity of disulfide bond formation. Downregulation in the expression of KRT-18 impairs C. parvum infection. Collectively, these results demonstrate that CpPDI is essential across invasive stages of the C. parvum parasite, and identify a non-redox dependent PDI interactor--KRT-18 in host cells, these data provide molecular insights into how Cryptosporidium interacts with its intestinal host.
Similar content being viewed by others
Introduction
Cryptosporidium spp. are among the main enteric protozoan parasites that cause diarrheal diseases in humans and animals worldwide. Cryptosporidium is transmitted via the fecal-oral route, and sources of infection include water and food contaminated with oocysts1. Most human diseases are caused by Cryptosporidium hominis, which only infects humans, and Cryptosporidium parvum, which can be zoonotically transmitted2,3. More than half of the waterborne disease outbreaks in the United States are caused by this parasite4. Cryptosporidium has recently been recognized as a leading global cause of severe diarrheal disease in young children and an important contributor to under-2 childhood mortality, especially in poor-resource settings5,6,7. It is estimated that cryptosporidiosis-associated acute infection caused more than 48,000 deaths as well as over 4.2 million disability-adjusted life-years lost in children younger than 5 years in 20168. Nitazoxanide, the only United States Food and Drug Administration-approved drug for cryptosporidiosis, is ineffective in immunocompromised or malnourished individuals and is not approved for use in children younger than 2 years9. Currently, there are no effective vaccines for C. parvum, and fundamental studies on parasite biology and host–pathogen interactions are needed to identify potential therapeutic and vaccine targets.
The life cycle of Cryptosporidium alternates between asexual and sexual reproduction. In contrast to most other apicomplexans, the entire cycle occurs within a single host10. Cryptosporidium infection occurs by ingestion of oocysts containing four infectious sporozoites. Then, sporozoites are released and invade host intestinal epithelial cells, which live in a unique intracellular but extracytoplasmic niche11,12. Although the formation mechanism of this niche is unclear, it is generally believed to be related to the rearrangement of the host actin cytoskeleton, a dense band, and other structures12. In apicomplexans, proteins secreted from micronemes, rhoptries, and dense granules have emerged as key mediators of invasion and pathogenesis. Among these, micronemes are related to adhesion and motility, rhoptry is related to invasion and initial immune evasion, and dense granules are related to host cell modifications12. Previous studies have shown that during invasion, Cryptosporidium sporozoites export proteins, such as the ROP1 and MEDLE protein families, into the cytosol of infected host cell13,14. Due to limitations in vitro culture and genetic manipulation techniques in the early stages, cryptosporidium invasion and host cell remodeling still need to be better understood mechanistically, and very few parasite proteins have been identified thus far.
Protein disulfide isomerase (PDI) is expressed in virtually all eukaryotic cells, including several protozoa15,16. These proteins generally contain at least one TRX-like domain, which is either catalytically active (a or a′) or non-catalytic (b or b′)17. The a and a’ domains contain the redox active site motif CXXC, whereas the b and b′ domain mostly serves as major sites for substrate binding and those responsible for chaperone activity, with a large multivalent hydrophobic surface18,19. In apicomplexan parasites, PDIs are located on the cell surface and are related to the adhesion and invasion of parasites18,20. For instance, PDI was identified and localized on the surface of Toxoplasma gondii tachyzoites and may activate the host cell membrane integrin15,21. Neospora caninum PDI is associated with the parasite surface membrane and is involved in tachyzoite–host cell interaction16; it has also been identified and partially characterized in other apicomplexans such as Eimeria tenella and Plasmodium20,22. However, the essentiality of PDI for C. parvum and its biological functions in the parasite are unknown. A previous proteomic analysis found that the PDI protein (encoded by the cgd6_120 gene) of C. parvum (CpPDI) was significantly increased during excystation and released in the process of invasion23. In the present study, CpPDI is a member of the thioredoxin superfamily of redox proteins. It contains two thioredoxin-like domains that include the CGHC active site, the cysteines of which are essential for its catalytic activity. Their findings prompted us to investigate their potential role in C. parvum (e.g., involvement in parasite invasion and identifying interacting substrates).
Here, we characterized the dynamic distribution of CpPDI during parasite invasion and intracellular development. We observed that CpPDI is mainly concentrated in endoplasmic reticulum and the cytoplasm of sporozoites using CRISPR/Cas9 technology and is highly expressed and released into host cells in the early stages of parasite infection. The inhibition of sporozoites invasion into host cells by anti-CpPDI antibodies and PDI inhibitors further supported the involvement of CpPDI in parasite invasion. Through pull down and mass spectrometry analysis, we discovered a non-redox dependent CpPDI interactor, the keratin, type I cytoskeletal 18 protein (KRT-18). By substituting selected amino acids in the CpPDI active sites, we further demonstrated the non–redox interaction between CpPDI and KRT-18. Inhibiting the expression of KRT-18 in host cells inhibits parasite adhesion and invasion. In summary, our study revealed that CpPDI participates in the invasion of Cryptosporidium by interacting with the KRT-18 protein.
Results
CpPDI is localized in sporozoites and is secreted during invasion
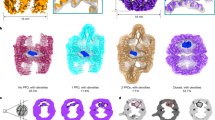
We compared the sequences of the 6 members of PDI family present in C. parvum using a neighbor-joining phylogenetic analysis. For comparison, we also included the Parasites PDIs and human PDI. Phylogenetic analysis indicated that CpPDI, encoded by the cgd6_120 gene, is most similar to the PDIs reported in other apicomplexan related to parasite adhesion and invasion, such as Plasmodium20, T. gondii15, and N. caninum16 (Fig. 1A). CpPDI contains a signal peptide with two conserved thioredoxin domains with catalytic domains (CGHC) (Fig. 1C). The CpPDI tertiary structure forms a “U” shape, forming hydrophobic pockets that facilitate binding with substrates. The two catalytic domains (CGHC) are highlighted in the form of green spheres (Supplementary Fig. 1A). To investigate the secretion and cellular localization of CpPDI, a rabbit polyclonal antibody against CpPDI was produced. Western blot analysis using this antibody detected single bands over 50 kDa from sporozoites, which agreed with the theoretical masses of native proteins (~53 kDa). Furthermore, pre-immune serum did not react with any protein from the lysates of C. parvum sporozoites (Supplementary Fig. 1B). To explore the localization of CpPDI in sporozoites, we generated epitope-tagged lines in widetype C. parvum parasites with a C-terminal 3 × HA tag (Fig. 1B, D). Immunofluorescence microscopy revealed that CpPDI showed perinuclear labeling that partially overlapped with BiP (endoplasmic reticulum maker) and anti-Sporozoites antibody (Fig. 1E, F). To assess whether CpPDI is secreted during invasion and the localization of CpPDI following invasion, we collected the supernatant of infected cells and cultures were fixed for immunofluorescence assay. We detected CpPDI in the supernatant of HCT-8 cells infected with Cryptosporidium sporozoites (Supplementary Fig. 1C). The IFA results showed that fluorescent signals were detected in the host cells during the invasion of Cryptosporidium sporozoites in 1 h (Supplementary Fig. 1D) and 6 h (Supplementary Fig. 1E). Overall, CpPDI may be secreted by the ER and released during the invasion of host cells by C. parvum.
A Phylogenetic relationship of, other apicomplexan PDIs and human PDI. The tree was constructed by a maximum likelihood analysis with 1000 replications for bootstrapping. B Schematic for endogenous tagging of cgd6_120 gene locus with 3 × HA epitope tag. The locations of the single guide RNA (sgRNA) in the 3′UTR, the Cas9 cut site (yellow arrow), and the repair template for homologous recombination are shown. Nluc, nanoluciferase; Neo, neomycin resistance marker; pEno, enolase promoter. C Domain architecture of C. parvum PDIs family showing the presence of one or two TRX-like active domain containing the active site CGHC. D PCR mapping of the correct integration events in the CpPDI-HA transgenic parasite. Fecal genomic DNA extracted from wild-type (WT) and CpPDI-HA transgenic parasites were used for PCRs. The sequences of primers for checking 5′ and 3′ integrations are provided in Supplementary Table 1. E Schematic representation of the subcellular structure of Cryptosporidium sporozoite. F Localization of CpPDI expression in sporozoites as revealed by endogenously tagging the gene. CpPDI is predominantly located near the parasite nucleus and cytoplasm. Anti-BiP antibodies served as an ER marker, anti-sporozoites antibodies serves as an cytoplasm maker. Scale bar: 5 μm.
CpPDI is highly expressed in the early and developmental stages of infection, both in vitro and in vivo
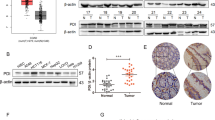
To examine the expression of CpPDI during the intracellular development of C. parvum in vitro, we infected HCT-8 cells with C. parvum oocysts. We tested its expression using reverse transcription-quantitative PCR (RT-qPCR) at different time points. After infection of the cells with sporozoites, low expression of CpPDI was observed at 24, 36, 48, and 72 h. The highest expression of the gene was observed at 2, 6, and 12 h post-infection. This observation was supported by published proteomic data at CryptoDB (http://cryptodb.org/cryptodb/) (Fig. 2A)24,25,26. Since the use of excysted sporozoites infected with HCT-8 cells in this study, there were differences in expression at some time points. To visualize the stage-specific expression and localization of CpPDI, we performed immunofluorescence assays (IFAs). Indirect immunofluorescence microscopy (IFM) detected signals of the CpPDI protein during the trophozoites, meront stages and microgamonts stage (Fig. 2B), although RT-qPCR data showed lower expression in these stages. We also assessed the CpPDI localization in vivo. The small intestines of IFN-γ−/− mice infected with C. parvum for 7 days were processed for histological analysis. Parasites were observed on the microvillar brush border, with the ileum showing a more intense infection (Fig. 2D). When imaged using IFM, CpPDI coincided with Vicia villosa lectin (VVL) (Fig. 2C).
A Relative transcription level of the CpPDI gene (cgd6_120) at 2, 6, 12, 24, 36, 48, and 72 h infection time points, as determined by reverse transcription-quantitative PCR (Blue, this study) and median transcript expression of cgd6_120 gene. Data shown here have been replotted from Mauzy et al. (Red). HCT-8 cells were infected with C. parvum sporozoites and cultured for specific time points, and RNA was collected from three wells per time point. Data from the Cryptosporidium 18S rRNA gene were used for normalization. Values are plotted as means ± standard deviation (SD). B IFA detection of CpPDI protein at different life stages of C. parvum. HCT-8 cells were infected with C. parvum sporozoites. The coverslips were fixed and stained with rat anti-CpPDI followed by goat anti-rat IgG Alexa Fluor 594, and DAPI for nuclear staining. Scale bars, 5 μm. C Immunofluorescence analysis of a histological section of the small intestine of a mouse infected with C. parvum. CpPDI is shown in red, parasites in green (VVL), and nuclei in blue (DAPI). Scale bar, 10 μm. D Burdens of intracellular parasites on the ileal villi in mice infected with C. parvum visualized by H&E-stained tissue sections. Arrows indicate intracellular parasites in the infected epithelium. Bar = 20 μm.
CpPDI affected C. parvum sporozoites invasion by influencing parasites–host interaction
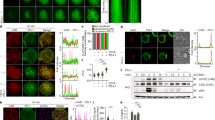
We further confirmed that anti-CpPDI antibodies and PDI inhibitors (bacitracin and 5,5′-Dithiobis(2-nitrobenzoic acid) (DNTB)) could interfere with sporozoites invasion using the invasion/growth inhibition assay. Compared with the control culture, significant (p < 0.01) reductions in C. parvum loads were observed in the cell cultures treated with antibodies against CpPDI (Fig. 3A, left). Preincubation of sporozoites with the sulfhydryl blocker DNTB and the PDI inhibitor bacitracin significantly reduced the C. parvum loads at 24 h after invasion (Fig. 3A, right). This assay indicated that interference in the function of surface associated CpPDI has a profound negative impact on C. parvum sporozoite–host cell interaction.
A Neutralization efficiency of C. parvum invasion by post-immune serum against CpPDI and PDI inhibitors (bacitracin and DNTB). Data are presented as the mean ± SD from three independent assays. B Enzymatic activities of recombinant CpPDI and CpPDI-AAAA. Recombinant CpPDI and CpPDI-AAAA were assayed for catalytic activity using the insulin reduction assay. C Dose-dependent inhibition of sporozoite invasion by recombinant CpPDI and CpPDI-AAAA as determined by fixation for microscopic counting in the 3-h invasion + 21-h infection assay.
To further confirm whether CpPDI can interfere with the invasion of sporozoites, we recombined and expressed CpPDI and CpPDI-AAAA proteins (mutation of the two catalytic domains from CGHC to AGHA) (Supplementary Fig. 2). To ensure that they maintained the expected level of catalytic activity and could be reversibly reduced and oxidized in the presence of intervening sequence variants, we measured the reductase activity of recombinant CpPDI and CpPDI-AAAA using an insulin reduction assay. Compared to recombinant CpPDI, CpPDI-AAAA did not exhibit reductase activity (Fig. 3B). Next, recombinant CpPDI and CpPDI-AAAA proteins were incubated with sporozoites and HCT-8 cells for 3 h, followed by the removal of excess recombinant proteins and uninvaded parasites and continuous culture of the invaded parasites for 21 h. In this assay, we detected a 25.3% increase in invasion by recombinant CpPDI but a 13.9% reduction by CpPDI-AAAA (each at 50 μg/mL) in a dose-dependent manner (Fig. 3C). We used recombinant proteins to incubate sporozoites separately, washed away excess proteins, and infected HCT-8 cells as described above. There was no significant reduction in parasite load in cultures treated with the recombinant proteins compared with those treated with normal infection (Fig. 3C). This further confirmed that CpPDI affects parasite invasion by influencing parasite–host interactions.
Identification of CpPDI substrates via pull down and LC-MS/MS analyses
To gain insight into the host pathway targeted by CpPDI, we conducted pull down to identify binding partners. The specific substrate of CpPDI is largely elusive, partly due to the transient nature of CpPDI substrate interactions. To overcome this challenge, we generated CpPDI variants by replacing selected amino acids on the active site that formed stable complexes with their substrates for subsequent isolation and identification (Fig. 4A). The activity of the variant was evaluated by monitoring the aggregation of reduced insulin through using turbidity measurements, as described above. Compared with CpPDI, the lag time of CpPDI-CACA (mutation of two catalytic domains from CGHC to CGHA) was significantly increased, while CpPDI-AAAA completely loses reductase activity (Fig. 4B). CpPDI-CACA trapping variants displayed a slower aggregation of insulin than wild-type CpPDI, indicative of a slower turnover of the insulin substrate due to the accumulation of stable intermediates.
A Structural diagram of CpPDI and its variants. B Enzymatic activities of recombinant CpPDI, CpPDI-CACA, and CpPDI-AAAA. The recombinant CpPDI, CpPDI-CACA, and CpPDI-AAAA were assayed for catalytic activity using the insulin reduction assay. C Silver-stained one-dimensional SDS-PAGE of GST-fusion protein coupled beads (GST-CpPDI). Left: Line 1: marker. Line 2: GST-CpPDI. Line 3:GST-CpPDI-CACA.Line 4: GST negative control group. Right: Line 1: marker. Line 2: GST-CpPDI + HCT-8 cells membrane proteins. Line 3: GST-CpPDI-CACA + HCT-8 cell membrane protein. Line 4: GST + HCT-8 cell membrane protein negative control group. D Heat map of interactors of CpPDI identified by mass spectrometry (credibility of ≥95% and ≥2 peptides).
To further determine the substrate of CpPDI during Cryptosporidium invasion, we extracted intact HCT-8 cell proteins. To verify that the CpPDI and CpPDI capture variants formed a stable complex with the substrates, the sample was subjected to SDS-PAGE followed by silver staining. Using the GST label as a blank control, we observed the substrates that bind to PDI and its variants (Fig. 4C). Next, the proteins in the three samples were excised and examined by mass spectrometry, of which 50 proteins were identified only in the CpPDI and CpPDI-CACA groups (Supplementary Fig. 3, Supplementary data 1). To further identify the most probable interactors of CpPDI, we filtered the candidates using the following criteria: (1) credibility of more than 95%;(2) at least 2 peptides, and ultimately obtained a total of 17 potential CpPDI interacting substrates (Fig. 4D).
CpPDI interacts with the host protein KRT-18 in vitro and in vivo
Since the invasion of Cryptosporidium is closely related to cell membrane and cytoskeleton proteins12,14, we selected proteins targeted at cell membrane and cytoskeleton to be included in further analysis. After analyzing the subcellular localization using bioinformatics data, two highly reliable proteins were identified in the CpPDI and CpPDI-CACA samples: a 38-kDa protein named Annexin A2 (ANXA2) located on the extracellular matrix and basement membrane (UniProt number P07355) and a 48-kDa protein named Keratin, type I cytoskeletal 18 protein (KRT-18) located on extracellular matrix and cytoskeleton-related (UniProt number P05783). We first tested the validity of the interaction between CpPDI and ANXA2 or KRT-18 using immunoprecipitation (IP). Using the CpPDI::flag knock-in strain with the expression of ANXA2::HA or KRT-18::HA, we performed an in vivo co-IP assay. We concluded that CpPDI::flag interacted with KRT-18::HA in vivo but did not interact with ANXA2::HA (Fig. 5A, Supplementary Fig. 4). Surprisingly, the addition of the reducing agent dithiothreitol (DTT) cannot eliminate the interaction between CpPDI::flag and KRT-18::HA (Fig. 5A), indicating that this interaction is largely independent on the intramolecular or intermolecular disulfide bonds between CpPDI::flag and KRT-18::HA.
A Western blot of a Flag immunoprecipitation experiment using HCT8 cells expressing CpPDI-Flag and KRT-18-HA in the presence or absence of DTT. KRT-18-HA was recovered both in the presence of CpPDI and in the absence of DTT. B Western blot of a Flag immunoprecipitation experiment using HCT8 cells expressing CpPDI-Flag and KRT-18-HA, CpPDI-CACA-Flag and KRT-18-HA, CpPDI-AAAA-Flag, and KRT-18-HA, respectively. C Immunofluorescence analysis of HCT8 expressing CpPDI-Flag (red), KRT-18 (green), and dapi (blue). Note the co-localization of KRT-18 with CpPDI-Flag, scale bar 5 μm. D Protein structure and molecular docking maps of CpPDI and KRT-18. E immunofluorescence of HCT8 cells infected with C. parvum. CpPDI in red, KRT-18 in green, VVL in green, and DAPI in blue. Scale bar 5 μm. F Immunofluorescence of histological sections of the ileum of C. parvum infected mice. VVL in green, KRT-18 in yellow, and CpPDI in red, colocalize at the apical brush of enterocytes. DAPI is in blue; scale bar, 5 μm.
To further understand whether CpPDI::flag interacts with KRT-18::HA through intermolecular disulfide bonds, we constructed a CpPDI::flag substrate-trapping mutant expression vectors: (CpPDI-CACA::flag and CpPDI-AAAA::flag)27,28. Consistent with the CO-IP results, Both CpPDI::flag and its mutants interact with KRT-18::HA, which is not affected by intermolecular disulfide bonds (Fig. 5B), indicating that KRT-18 is a non-redox dependent PDI interactor. Furthermore, we performed molecular docking studies to predict the binding sites of CpPDI and KRT-18 (Fig. 5D), indicating that their binding ability is not related to the active sites of CpPDI.
Previous studies have shown that in the early stages of Cryptosporidium infection, KRT-18 gene expression in host cells is upregulated29. Keratin is a specific component of the cytoskeleton of epithelial cells and a specific heteropolymer. Other studies have shown that loss or mutation of keratin leads to increased apoptosis30. In HCT8 cells, we found that KRT-18 co-localized with the ectopically expressed CpPDI-flag (Fig. 5C), and during parasite infection, KRT-18 accumulated at the infection site alongside CpPDI (Fig. 5E). In Cryptosporidium-infected mice, CpPDI and KRT-18 co-localized at the apical surface of the infected cells (Fig. 5F). We designed three siRNAs that can silence KRT-18 gene expression and transinfect HCT-8 cells to test the effect of reduced KRT-18 expression on C. parvum infection. Compared with the control, the level of KRT-18 mRNA was reduced by approximately 40–60% at siRNA concentration of 50 nM, whereas no or few changes were observed in the negative control by qRT-PCR (Supplementary Fig. 5A). Western blot analysis further validated gene silencing, in which KRT-18 protein expression was decreased (Supplementary Fig. 5B). Next, we briefly evaluated the relative impact of SiRNA2 inhibition of KRT-18 on parasite burden in vitro. Surprisingly, we detected a significantly lower parasite burden in cells transfected with siKRT-18 at 3 and 24 hpi, which was confirmed by qPCR (Supplementary Fig. 5C). These data indicate that the suppressed expression of KRT-18 mainly hinders the adhesion and invasion of parasites, and growth and development defects may be the downstream effects of adhesion and invasion.
Discussion
Over the last decade, the important role of PDIs in apicomplexan parasites has become increasingly apparent, with studies finding PDIs at the cell surface during the motile and invasive life cycle stages, which have been implicated in host cell adhesion and invasion, which are essential processes in their obligate intracellular life cycles15,16,31. The C. parvum genome contains six PDI isozyme, most of which have unknown functions. Here, we focused on CpPDI (encoded by the cgd6_120 gene), which increases significantly during excystation (26) and is secreted during invasion. CpPDI contains a signal peptide and two conserved thioredoxin domains with catalytic domains (CGHC). Like other Apicomplexa protozoa, CpPDI is highly expressed early in development in C. parvum and is mostly distributed in endoplasmic reticulum and the cytosol of free sporozoites. Our experiments, employing membrane impermeant reagents that act solely on the parasite surface (thioredoxin inhibitor DNTB, PDI inhibitor bacitracin, and anti-CpPDI antibodies), indicated that interference in the function of surface-associated CpPDI has a profound negative impact on C. parvum sporozoite–host cell interaction. By constructing a mutant PDI at the active site, we showed that CpPDI may interact with the host cell KRT-18, which is conducive to the invasion of Cryptosporidium.
Apicomplexan parasites contain numerous PDI isozymes, however, their roles have only been partially investigated. Recent bioinformatics studies have revealed the presence of classical thioredoxin domains in malarial parasites, with a total of nine PDI-like molecules identified18, of which only two of these proteins (PDI-8 and PDI trans) have been experimentally shown to exhibit PDI activity31. PDI-trans is male-specific, surface-expressed, essential for fertilization/transmission and a feasible target for antimalarial drugs and vaccines31. T. gondii PDI promotes adhesion and invasion by combining with the cell membrane in the early stages of infection in human cells and has been proposed as a vaccine candidate15. Previous work on N. caninum PDI protein indicated that the parasite surface membrane was associated physical contact between the thiol groups of PDI of the parasite and the host cell membrane. When PDI inhibitors or anti-NcPDI antibodies are used to incubate N. caninum tachyzoites, the adhesion between parasites and host cells can be reduced16. However, little is known regarding the expression and function of PDI proteins in Cryptosporidium spp. In this study, CpPDI had the same location and function as PDI of other apicomplexan parasites, and played an active role in the adhesion and invasion of Cryptosporidium. Antibodies against these proteins have been shown to inhibit parasite invasion in vitro, suggesting that they play a role in the processing or maturation of substrates involved in host cell recognition.
Previous studies have suggested the formation of electron-dense bands and intense remodeling of epithelial cells during Cryptosporidium infection using electron microscopy11,12. However, the underlying molecular mechanisms behind these changes are still unclear32. Apicomplexan parasites have evolved highly specialized pathogenic factors that allow them to invade and control a diverse range of host cells, such as the C. parvum ROP1 and MEDLE family12,14. Alternatively, surface contact activates the host integrin signal and causes actin polymerization in the host33. Here, we report the functions of C. parvum PDI and its ability to target various locations. Using the IFA, we found that this secreted protein is expressed in sporozoites and may be injected into the host cell; hence, it is likely to interact with the host cell during invasion. However, the mechanism of CpPDI externalization is still unclear. By constructing the fusion protein GST-CpPDI and GST-CpPDI-CACA as a probe, proteins derived from HCT-8 cells were incubated and initially analyzed for screening of candidate proteins potentially interacting with CpPDI. Proteins binding with GST only were eliminated through setting up the GST control, and the remaining proteins were determined as candidate receptors for CpPDI. KRT18 was identified as a potential non-redox dependent CpPDI-interacting partner, and this interaction was rigorously supported by co-IP and cellular co-localization in vitro and in vivo. Finally, interference with the expression of the KRT-18 gene leads to limitations in the growth and development of C. parvum, further indicating that it plays a positive role in the growth and development of Cryptosporidium.
KRT-18, a cytoskeletal protein, belongs to the family of intermediate filament proteins widely expressed by epithelial and endothelial cells; it plays an important role in tissue integrity34. KRT-18 is overexpressed in many different types of cancer, including colorectal cancer, and is associated with tumor stage, cancer migration, and invasion35. In addition, KRT-18 has been proven to be associated with cell apoptosis. During apoptosis, KRT-18 is cleaved by multiple caspase molecules and released during late apoptosis following the disintegration of the plasma membrane34. Loss or mutation of keratin can lead to increased cell apoptosis, which may be related to its interaction with TNF or Fas receptors30. Interestingly, in previous studies, it was reported that C. parvum infection causes upregulation of KRT-18 expression in the host cell29, indicating that the invasion of C. parvum may cause changes in the cytoskeleton and anti-apoptotic effects through changes in keratin. This suggests a broader role for KRT-18 during infection with enteropathogens. Further studies are required to understand the significance of these results.
In summary, we show here that CpPDI is located in the cytoplasm and surface of parasites and is secreted during invasion, playing an essential role across invasive stages of the C. parvum parasite, similar in localization and function to PDI in T. gondii15. However, the mechanism of PDI externalization is unclear. We also identified a non-redox dependent PDI interactor–KRT-18 in host cells. CpPDI-specific antibodies and inhibitors can also inhibit cell invasion. Thus, CpPDI represents a potential new target for anti-cryptosporidial strategies. Using recently developed genetic tools to study CpPDI might provide a deeper understanding of its role. In addition, it is worthwhile to develop an assay for high-throughput screening of selective anti-CpPDI inhibitors for potential anti-cryptosporidial drug development.
Methods
Animal studies
Studies on mice were reviewed and approved by the Research Ethics Committee of Henan Agricultural University. According to the Animal Ethics Procedures and Guidelines of the People’s Republic of China, all animals and experiments were handled strictly in accordance with good laboratory animal practice. We have complied with all relevant ethical regulations for animal use. All efforts were made to minimize animal suffering. IFN γ−/− (GKO) mice embryos were purchased from Jackson Laboratory (USA, product code: 002287) and resuscitated by the Medical Laboratory Animal Research Institute of Peking Union Medical College Hospital before being bred in-house at Henan Agricultural University. All mice were reared in a specific-pathogen-free facility and provided non-medicated feed and water ad libitum throughout the experiment. For C. parvum infection, the mice were gavaged with 5 × 105 C. parvum oocysts suspended in 200 mL of distilled water, and the mice in the negative-control group were inoculated with the same volume of distilled water.
Cell culture and infection
The HCT-8 human colon adenocarcinoma cell (HCT-8) line was purchased from ATCC (CCL-224TM) and maintained in RPMI 1640 medium (Biological Industries, Israel) supplemented with 10% fetal bovine serum (FBS), and 100 U/mL penicillin, 100 U/mL. For HCT-8 infection medium, serum concentrations were reduced to 2%. The cells were then inoculated with C. parvum sporozoites in RPMI 1640 supplemented with 2% FBS. Invasion stages of parasites were obtained by infecting HCT-8 cells for 1–6 h, while intracellular stages of parasites were obtained by infecting HCT-8 cells for 12–48 h. Monolayers were harvested at different times to observe trophozoites (12 h), meronts (24 and 36 h), and male and female gametes (48 h) using the IFA assay.
Cryptosporidium parvum parasites and excystation
C. parvum oocysts (subtype ⅡdA19G1 at the gp60 locus) were maintained in the laboratory (International Joint Research Laboratory for Zoonotic Diseases of Henan) and passaged in newborn Holstein calves via collection of feces after infection. Samples were purified as previously described36. To determine the viability of the oocysts and the accuracy of later experiments, the purified oocysts were stored at 4 °C in 2.5% potassium dichromate until use. Before the experiments, the oocysts were treated with 10% Clorox on ice for 10 min and washed three times with sterile phosphate-buffered saline (PBS). For some experiments, the oocysts were excysted before infection by incubating them with 0.75% sodium taurocholate hydrate (Sigma) and 0.5% BSA-PBS (Sigma) at 37 °C for 1 h.
Gene expression analysis
HCT-8 cells were grown in 12-well culture plates and incubated for 24 h before infection. Monolayers were infected with excysted sporozoites, washed twice with PBS at 2 hpi, and fresh HCT-8 medium was added. RNA was collected from three wells at each time point using RNA-easy Isolation Reagent (Vazyme, Nanjing, China), and then stored at –80 °C until further processing. First-strand cDNA was synthesized using HiScript® III All-in-One RT SuperMix Perfect for qPCR (Vazyme, Nanjing, China). Reverse transcription quantitative PCR (RT-qPCR) was performed using a CFX384TM Real-Time PCR system (Bio-Rad, Hercules, CA, USA) with ChanQ Universal SYBR qPCR Master Mix (Vazyme) using the primers listed in Supplementary Table 1. The following conditions were used for RT-qPCR: priming at 95 °C for 30 s, followed by 40 cycles of denaturation at 95 °C for 10 s, annealing at 60 °C for 30 s, and extension at 60 °C for 15 s, followed by melt curve analysis to detect nonspecific amplification. The relative expression of the CpPDI gene was calculated using the 2–ΔΔCT method37 using C. parvum 18S rRNA as the reference gene. Data are presented as the mean and standard error of the mean.
Indirect immunofluorescence microscopy
Sporozoites resuspended in PBS were dried on microscope slides, and the intracellular stages of C. parvum in HCT-8 cells were grown on coverslips for 1, 6, 12, 24, and 48 h. The slides and coverslips were fixed at 25 °C for 20 min with 4% paraformaldehyde. After three washes in PBS, the slides and coverslips were permeabilized with 0.5% Triton X-100 in PBS for 15 min and then incubated for 1 h with 5% BSA in PBS (BSA-PBS), followed by incubation with primary antibodies in 5% BSA for 2 h, washes in PBS, and incubation with secondary antibodies in 5% BSA for 1 h. Finally, the cells were incubated for 10 min with DAPI and washed three times with PBS. A drop of Fluoromount-G™ (Yeasen Biotech) was added to set the coverslip, which was sealed with nail polish38.
Intestines of infected mice were collected and “swiss-rolled” before fixation overnight in formalin, and immunofluorescence was performed as described previously39. And counterstained with VVL, which recognizes glycan epitopes found at all C. parvum stages.
Bioinformatics analysis, molecular modeling, and model evaluation of CpPDI
SignalP Server (version 4.1) was used to predict the cleavage sites in the signal peptide. A molecular model of CpPDI was generated using AlphaFold 2 as described previously40,41. The 3D structure and quality of the models were evaluated and verified using SAVES v5.0.
Purifying CpPDI protein and generat rat polyclonal antiserum
The full CpPDI sequence was amplified from the genomic DNA of the C. parvum IIdA19G1 isolate using the primers listed in Supplementary Table 1 and was inserted into the pGEX-4T-1 vector (Novagen, Madison, WI, USA). The plasmid was transformed into Escherichia coli Rosetta (DE3) cells (Tiangen Biotechnology Co., Ltd., Beijing, China), after which 0.1 mM of isopropyl-β-D-thiogalactopyranoside (IPTG) was added at 37 °C for 3 h to induce protein expression. For the purification of CpPDI, cultured E. coli were centrifuged and lysed, and the supernatant was filtered through a 0.45 μm cellulose acetate membrane filter (Millipore, Billerica, MA, USA) and loaded onto glutathione sepharose 4B beads (GE Healthcare, Pittsburgh, USA) at 4 °C for 3 h. After washing the beads with six volumes of PBS, CpPDI was eluted from the beads using elution buffer (50 mM Tris-HCl, 10 mM reduced glutathione, pH 8.0). The GST tag of the target was removed from protein by cutting it with thrombin (Solarbio, Beijing, China) at 25 °C for 12 h.
The antigen for CpPDI polyclonal antiserum was generated in pathogen-free New Zealand white rabbits. The priming antigen was mixed with 750 mg antigen and an equal volume of Freund’s Complete Adjuvant (FCA), emulsified, and then boosted three times at 21-day intervals with 500 mg antigen in Freund’s Incomplete Adjuvant (FIA). On the 7th day after the fourth immunization, the anti-serum titer of the ELISA test met the requirements, and whole blood was collected from the arteries and purified. All immunofluorescent staining experiments used terminally bled CpPDI antiserum at a 1:1000 dilution.
Western blot analysis of native CpPDI
For western blot analysis of native CpPDI, the released sporozoites and HCT-8 cells (2 × 106) were collected by centrifugation and resuspended in RIPA lysis buffer (Epizyme, Shanghai, China) containing 1% protease and phosphatase inhibitor cocktail (NCM, Suzhou, China). For the secretion of CpPDI, HCT-8 cells were infected with sporozoites for 1 h, and the supernatant (containing secreted proteins) and precipitate (sporozoites and HCT-8 cells) were collected. Similarly, the precipitate was lysed using the above method. The supernatant was collected into a 50-mL centrifuge tube, and 1% protease and phosphatase inhibitor cocktail was added, and the tube was centrifuged at 4 °C for 10 min at 12,000 × g, to remove the sporozoites and cell fragments. The supernatant was filtered through a 0.22 μm membrane and transferred to an ultrafiltration centrifuge tube (Merck Millipore; Amicon Ultra-50, Ultrael-3k) and centrifuged at 4000 × g at 4 °C and concentrated to 500 μL. Western blot analyses were performed as described previously42 using anti-CpPDI (1:1,000 dilution) and GAPDH (1:10,000 dilution) primary antibodies and horseradish peroxidase (HRP)-conjugated secondary antibodies at 1:10,000 dilution. Finally, the membrane was washed three times with PBST, and reactive protein bands in the membrane were detected using Immobilon Crescendo Western HRP substrate (Merck Millipore, MA, USA) and analyzed with Amersham ImageQuant 800 (GE, CT, USA).
Generating the pan Cp polyclonal antiserum
The antigen for the pan C. parvum sporozoites polyclonal antiserum was prepared in our laboratory, as previously reported43.
Transfection of C. parvum sporozoites
The C. parvum Cas9/guide vector and p3HA-Nluc-P2A-neo plasmid skeletons were both gifted by Boris Striepen Laboratory. Guide RNAs for the CpPDI gene (cgd6_120) were designed and cloned into BbsI sites of the C. parvum Cas9/guide vector. The tagging plasmid pCpPDI-3HA-Nluc-P2A-neo was generated by Gibson assembly using the pUC-HA3-pEno-Nluc-Neo-Eno vector as a template and ~500 bp flanks on both sides for double homologous recombination. Primers used for guide cloning and overhang PCRs for generating tagged strains are provided in Supplementary Table 1.
C. parvum oocysts (2 × 107) were excysted as described above, and the released sporozoites were resuspended in 80 μL complete SF buffer (consisting of 65.5 μL Cell Line Solution and 14.5 μL Supplementary 1). Afterwards, 20 μL of plasmid suspension (50 μg Cas9/gRNA expression plasmid and 50 μg homologous repair plasmid) was then added to the mixture, which was transferred to a cuvette (Lonza Cologne, Cologne, Germany) and electroporated using an AMAXA 4D nucleofector system (Lonza Cologne) with program EH10039,40. The transfected sporozoites were diluted in PBS and inoculated into GKO mice by oral gavage. Prior to infection, the mice were given 100 μL of 8% NaHCO3 solution to neutralize gastric acid. The next day, the drinking water in the animal cages was replaced with a solution containing 16 g/L paromomycin (Yuaye, Shanghai, China) for in vivo selection. Parasite shedding was monitored by measuring nanoluciferase activity in the feces of infected mice.
Luciferase assay
Luciferase assays were performed using the Nano-Glo Luciferase Assay Kit (Promega, Madison, USA) as previously described39. For mouse fecal samples, one fecal pellet was collected in a 1.5 mL microcentrifuge tube to which ten 3 mm glass beads (Fisher Scientific, Hanover, USA) and 0.8 mL fecal lysis buffer (50 mM Tris pH 7.6, 2 mM DTT, 2 mM EDTA pH 8.0, 10% glycerol, 1% Triton X-100 prepared in ddH2O) were added. The tube was vortexed for 60 Hz, 45 s, stopping for 15 s, repeated twice using homogenizer (Servicebio, Wuhan, China). Then incubate the supernatant with a mixture of the Nano-Glo Luciferase Substrate Mix in the dark for 3 min.
HCT-8 cells cultured for 24 h were co-cultured with the C. parvum, and luminescence was measured using the Nano-Glo Luciferase Assay kit on a Synergy HTX imaging multimode reader (BioTek, Winooski, USA).
In vitro evaluation of the effects of CpPDI on parasite invasion
The effect of polyclonal anti-CpPDI antibodies and PDI inhibitors (bacitracin and DNTB) on C. parvum infection in HCT-8 cells was examined using an in vitro neutralization assay. PDI inhibitors only act on the surface of sporozoites and prevent the formation of disulfide bonds by covalently modifying the surface thiol groups16,44. Briefly, HCT-8 cells were grown in 12-well plates and maintained in RPMI 1640 medium supplemented with 10% FBS at 37 °C in a humidified 5% CO2 incubator. Excysted sporozoites (2 × 106) were resuspended in RPMI 1640 medium supplemented with 10% FBS and added to a separate 12-well plate. Antibodies (1:50, 1:100, and 1:500) or varying concentrations of PDI inhibitors bacitracin and DNTB, ranging from 0.5 to 5 mM were added to the 12-well plate. Sporozoite-antibody or PDI inhibitor solutions were incubated at 37 °C for 30 min and then added to confluent HCT-8 cells in the 12-well plate. After a 2.5 h incubation, free sporozoites were washed off and incubated for an additional 18 h. The method for assessing of antibodies against C. parvum infection of HCT-8 cells in vitro was based on the quantitative real-time reverse transcription-PCR (qRT-PCR) technique, as described by Cai et al.45.
In another assay, recombinant CpPDI and CpPDI-AAAA (50, 100, and 500 μg/mL) were mixed with oocysts (1 × 106) in RPMI 1640 medium containing BSA (50 μg/mL) for 10 min. Oocysts and recombinant protein suspensions were then added to host cells cultured on 15-mm coverslips in 24-well plates at 70–80% confluence (oocyst/host cell ratio at 1:2). After incubation at 37 °C for 3 h to allow parasite excystation and invasion, uninfected parasites were removed by medium exchange and washed three times with PBS. After cultivation for an additional 21 h in the absence of proteins (total infection time: 24 h), the developmental stages of C. parvum in cells were stained with the A600FLR-20X Sporo-Glo antibody (Waterborne) and examined under a BX53 immunofluorescence microscope (Olympus, Tokyo, Japan). For each slide, images of 50 random fields were captured under 1000×. Parasite load was calculated as previously described46. Data from cells treated with pre-immune serum at the corresponding dilutions were used as controls. All experiments were performed in triplicate.
Generation of PDI variants and disulfide reductase activity assays
Classical PDI variants were designed by substituting the second cysteine residue in the CGHC active site with alanine as previously described27.
The cDNA for CpPDI was inserted into the pGEX-4T-1 expression vector. Using the wild-type cDNA vector as a template, we performed multiple rounds of PCR site-directed mutagenesis and sequencing to prepare plasmids encoding PDI variants; the primers used are listed in Supplementary Table 1. These included trapping variants with active sites in the CGHA (CpPDI-CACA) configuration and enzymatically inactive PDI with active-site motifs of AGHA (CpPDI-AAAA) to as controls (Supplementary Fig. 1). After sequence verification, the plasmids were transformed into BL21 competent E. coli cells to generate strains that expressed all PDI variants.
The ability of CpPDI to reduce disulfide bonds was assessed by a standard turbidimetric assay using insulin as a substrate. Natural insulin is composed of two chains: A and B. When natural insulin is reduced, the A-B chain separates, and the B chain forms a precipitate, causing the solution to become cloudy. The reduction of this disulfide bridge causes the separation of the two insulin chains and their precipitation, with a consequent increase in the turbidity of the solution, which can be monitored by measuring the absorbance at 650 nm using a SynergyTM HTX multi-mode reader (BioTek). The assay was performed by mixing 100 mM sodium hydrogen phosphate buffer (pH 7.0), 2 mM EDTA, and insulin to prepare 1 mg/mL insulin buffer and adding varying concentrations of CpPDI, CpPDI-CACA, and CpPDI-AAAA, ranging from 100 to 1000 μg, and 2 mM DTT. The absorbance values of the reaction solution were recorded at 650 nm at constant temperature treatment times of 0, 10, 20, 30, 40, 50, and 60 min at 37 °C.
GST-pull down and LC-MS/MS analysis
HCT-8 cell protein extraction was performed using a Minute™ Plasma Membrane Protein Isolation and Cell Fractionation Kit (Invent Biotechnologies) according to the manufacturer’s instructions. GST-CpPDI, GST-CpPDI-CACA, and GST proteins were incubated with 20 mM DTT at 0 °C for 20 min, desalted then added to glutathione sepharose 4B beads at 4 °C for 3 h in Pierce™Spin Columns-Screw Cap (Thermo Fisher Scientific, MA, USA), and the beads were washed several times using PBS buffer. The beads were then incubated with HCT-8 cell plasma membrane proteins in PBS at 4 °C for 3 h before adding 20 mM N-ethylmaleimide, and washed several times with PBS to remove unbound proteins. Each sample was examined by SDS-PAGE on a 10% gel, followed by silver staining using a Quicksilver staining 156 kit (Beyotime, China)27,47. The samples obtained by silver staining were sent to GeneCreate Biological Engineering (Wuhan, China), digested with trypsin overnight, added to the peptide extract (ACN/formic acid), desalinated using C18 desalting columns, and subjected to LC-MS/MS analysis according to standard protocols. Data were retrieved using ProteinPilot (version 4.5) software. For protein identification, the Paragon algorithm in ProteinPilot was used to search the UniProt Human Database. Certain filtering criteria were selected for the identified protein results, and peptides with an unused score >1.3 (credibility of more than 95%) were considered credible peptides, and proteins containing at least one unique peptide were retained.
Molecular docking
Molecular docking was performed using HDOCK. The structure of KRT-18 and CpPDI (predicated by AlphaFold using Uniprot ID P05783 and Q5CPJ7) was employed for model building. Structures and maps in the figures were rendered with PyMOL (The PyMOL Molecular Graphics System, v.4.6).
Statistics and reproducibility
All statistical analyses were performed with GraphPad Prism 8.0.2 (GraphPad Software, San Diego, CA, USA). Differences between the two groups were analyzed with an unpaired t-test with Bonferroni’s correction, and differences among multiple groups were analyzed with nonparametric one-way ANOVA. All data are expressed as means ± standard deviations (SD). All experiments were performed with at least three biological replicates. For all analyses, P < 0.05 was considered significant. *P < 0.05, **P < 0.01, and ***P < 0.001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All relevant data are available in the manuscript and the Supporting Information files. The source data behind the graphs in the paper can be found in Supplementary Data 2. All other data, including the plasmids used in this study, can be obtained from Zhang Longxian upon reasonable request.
References
Guo, Y., Ryan, U., Feng, Y. & Xiao, L. Emergence of zoonotic Cryptosporidium parvum in China. Trends Parasitol. 38, 335–343 (2022).
Nader, J. L. et al. Evolutionary genomics of anthroponosis in Cryptosporidium. Nat. Microbiol. 4, 826–836 (2019).
Ryan, U., Zahedi, A., Feng, Y. & Xiao, L. An update on zoonotic Cryptosporidium species and genotypes in humans. Animals 11, 3307 (2021).
Hlavsa, M. C. et al. Outbreaks associated with treated recreational water—united states, 2000-2014. MMWR Morb. Mortal. Wkly Rep. 67, 547–551 (2018).
Kotloff, K. L. et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case-control study. Lancet 382, 209–222 (2013).
Yang, X., Guo, Y., Xiao, L. & Feng, Y. Molecular epidemiology of human cryptosporidiosis in low- and middle-income countries. Clin. Microbiol. Rev. 34, e00087–19 (2021).
Kotloff, K. L. et al. The incidence, aetiology, and adverse clinical consequences of less severe diarrhoeal episodes among infants and children residing in low-income and middle-income countries: a 12-month case-control study as a follow-on to the Global Enteric Multicenter Study (GEMS). Lancet Glob. Health 7, e568–e584 (2019).
Khalil, I. A. et al. Morbidity, mortality, and long-term consequences associated with diarrhoea from Cryptosporidium infection in children younger than 5 years: a meta-analyses study. Lancet Glob. Health 6, e758–e768 (2018).
Amadi, B. et al. High dose prolonged treatment with nitazoxanide is not effective for cryptosporidiosis in HIV positive Zambian children: a randomised controlled trial. BMC Infect. Dis. 9, 195 (2009).
Tandel, J. et al. Life cycle progression and sexual development of the apicomplexan parasite Cryptosporidium parvum. Nat. Microbiol. 4, 2226–2236 (2019).
Elliott, D. A. & Clark, D. P. Cryptosporidium parvum induces host cell actin accumulation at the host-parasite interface. Infect. Immun. 68, 2315–2322 (2000).
Guérin, A. & Striepen, B. The biology of the intestinal intracellular parasite Cryptosporidium. Cell Host Microbe 28, 509–515 (2020).
Dumaine, J. E. et al. The enteric pathogen Cryptosporidium parvum exports proteins into the cytosol of the infected host cell. Elife 10, e70451 (2021).
Guérin, A. et al. Cryptosporidium rhoptry effector protein ROP1 injected during invasion targets the host cytoskeletal modulator LMO7. Cell Host Microbe 29, 1407–1420.e5 (2021).
Moncada, D. et al. Role of the 52 kDa thioredoxin protein disulfide isomerase of Toxoplasma gondii during infection to human cells. Exp. Parasitol. 164, 36–42 (2016).
Naguleswaran, A. et al. Neospora caninum protein disulfide isomerase is involved in tachyzoite-host cell interaction. Int. J. Parasitol. 35, 1459–1472 (2005).
Cuervo, N. Z. & Grandvaux, N. Redox proteomics and structural analyses provide insightful implications for additional non-catalytic thiol-disulfide motifs in PDIs. Redox Biol. 59, 102583 (2023).
Ali Khan, H. & Mutus, B. Protein disulfide isomerase a multifunctional protein with multiple physiological roles. Front. Chem. 2, 70 (2014).
Kozlov, G., Määttänen, P., Thomas, D. Y. & Gehring, K. A structural overview of the PDI family of proteins. FEBS J. 277, 3924–3936 (2010).
Angrisano, F., Ford, A., Blagborough, A. M. & Bullen, H. E. Protein disulfide isomerases—a way to tackle malaria. Trends Parasitol. 39, 622–625 (2023).
Meek, B., Back, J. W., Klaren, V. N., Speijer, D. & Peek, R. Conserved regions of protein disulfide isomerase are targeted by natural IgA antibodies in humans. Int. Immunol. 14, 1291–1301 (2002).
Han, H. et al. Molecular characterization and analysis of a novel protein disulfide isomerase-like protein of Eimeria tenella. PLoS ONE 9, e99914 (2014).
Snelling, W. J. et al. Proteomics analysis and protein expression during sporozoite excystation of Cryptosporidium parvum (Coccidia, Apicomplexa). Mol. Cell Proteom. 6, 346–355 (2007).
Mauzy, M. J., Enomoto, S., Lancto, C. A., Abrahamsen, M. S. & Rutherford, M. S. The Cryptosporidium parvum transcriptome during in vitro development. PLoS ONE 7, e31715 (2012).
Mirhashemi, M. E. et al. Transcriptome analysis of pig intestinal cell monolayers infected with Cryptosporidium parvum asexual stages. Parasit. Vectors 11, 176 (2018).
Estimates of global, regional, and national morbidity, mortality, and aetiologies of diarrhoeal diseases: a systematic analysis for the Global Burden of Disease Study 2015. Lancet. Infect. Dis. 17, 909–948 (2017).
Eriksson, O., Stopa, J. & Furie, B. Identification of PDI substrates by mechanism-based kinetic trapping. Methods Mol. Biol. 1967, 165–182 (2019).
Jessop, C. E., Watkins, R. H., Simmons, J. J., Tasab, M. & Bulleid, N. J. Protein disulphide isomerase family members show distinct substrate specificity: P5 is targeted to BiP client proteins. J. Cell Sci. 122, 4287–4295 (2009).
Wang, L. et al. Comparative proteomics reveals Cryptosporidium parvum infection disrupts cellular barriers. J. Proteom. 287, 104969 (2023).
Coulombe, P. A. & Omary, M. B. ‘Hard’ and ‘soft’ principles defining the structure, function and regulation of keratin intermediate filaments. Curr. Opin. Cell Biol. 14, 110–122 (2002).
Angrisano, F., Sala, K. A., Tapanelli, S., Christophides, G. K. & Blagborough, A. M. Male-specific protein disulphide isomerase function is essential for Plasmodium transmission and a vulnerable target for intervention. Sci. Rep. 9, 18300 (2019).
Lumb, R., Smith, K., O’Donoghue, P. J. & Lanser, J. A. Ultrastructure of the attachment of Cryptosporidium sporozoites to tissue culture cells. Parasitol. Res 74, 531–536 (1988).
Zhang, H., Guo, F. & Zhu, G. Involvement of host cell integrin α2 in Cryptosporidium parvum infection. Infect. Immun. 80, 1753–1758 (2012).
Lebherz-Eichinger, D., Krenn, C. G. & Roth, G. A. Keratin 18 and heat-shock protein in chronic kidney disease. Adv. Clin. Chem. 62, 123–149 (2013).
Kilic-Baygutalp, N. et al. Evaluation of serum HGF and CK18 levels in patients with esophageal cancer. Genet. Mol. Res. 15, 15038583 (2016).
Pawlowic, M. C., Vinayak, S., Sateriale, A., Brooks, C. F. & Striepen, B. Generating and maintaining transgenic Cryptosporidium parvum parasites. Curr. Protoc. Microbiol. 46, 20b.2.1–20b.2.32 (2017).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Xu, R., Feng, Y., Xiao, L. & Sibley, L. D. Insulinase-like protease 1 contributes to macrogamont formation in Cryptosporidium parvum. mBio 12, e03405–20 (2021).
Sateriale, A., Pawlowic, M., Vinayak, S., Brooks, C. & Striepen, B. Genetic manipulation of Cryptosporidium parvum with CRISPR/Cas9. Methods Mol. Biol. 2052, 219–228 (2020).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Tunyasuvunakool, K. et al. Highly accurate protein structure prediction for the human proteome. Nature 596, 590–596 (2021).
Cui, Z. et al. Cryptosporidium parvum gp40/15 is associated with the parasitophorous vacuole membrane and is a potential vaccine target. Microorganisms 8, 363 (2020).
Wu, S. et al. MiR-199a-3p regulates HCT-8 cell autophagy and apoptosis in response to Cryptosporidium parvum infection by targeting MTOR. Commun. Biol. 7, 924 (2024).
Benedikter, B. J. et al. Cigarette smoke extract induced exosome release is mediated by depletion of exofacial thiols and can be inhibited by thiol-antioxidants. Free Radic. Biol. Med. 108, 334–344 (2017).
Cai, X., Woods, K. M., Upton, S. J. & Zhu, G. Application of quantitative real-time reverse transcription-PCR in assessing drug efficacy against the intracellular pathogen Cryptosporidium parvum in vitro. Antimicrob. Agents Chemother. 49, 4437–4442 (2005).
Zhang, Q. et al. Characterization of calcium-dependent protein kinases 3, a protein involved in growth of Cryptosporidium parvum. Front. Microbiol. 11, 907 (2020).
Stopa, J. D., Baker, K. M., Grover, S. P., Flaumenhaft, R. & Furie, B. Kinetic-based trapping by intervening sequence variants of the active sites of protein-disulfide isomerase identifies platelet protein substrates. J. Biol. Chem. 292, 9063–9074 (2017).
Acknowledgements
We thank Wuhan GeneCreate Biological Engineering Co., Ltd. for their technical assistance. We thank LetPub (www.letpub.com) for linguistic assistance while preparing this manuscript. This work was partially supported by NSFC grants (32172882), the National Key Research and Development Plan Project (2022YFD1800200), NSFC-Henan Joint Fund Key Project (U1904203), the Key Program of the National Natural Science Foundation of China (31330079), and the Leading Talents of the Thousand Talents Program of Central China (19CZ0122). These funding bodies were solely involved in funding and had no role in the design of the study, the collection, analysis and interpretation of the data or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
L.X. Zhang and S.M. Zhang conceived and designed the experiments. L.Y. Wang, N. Li, and X.T. Zhang performed the experiments and analyzed the data. L.Y. Wang wrote the paper. L.Y. Wang, Z.H. Cui, and Y.X. Wang took the photographs. Y.Y., W.U., X.Y., Li, Y.Y. Yang, J.F. Zhao, S.M. Zhang, and L.X. Zhang contributed to the revision of the manuscript. All the authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Anna Feix and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ophelia Bu.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, L., Li, N., Zhang, X. et al. Host cell KRT-18 as a non-redox dependent interactor of Cryptosporidium parvum PDI enables parasite infection. Commun Biol 8, 1840 (2025). https://doi.org/10.1038/s42003-025-09238-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-025-09238-7