Abstract
Glycophagy, a selective form of autophagy critical for glycogen homeostasis, relies on the glycogen cargo receptor called starch-binding domain-containing protein 1 (STBD1), yet its evolutionary origins remain elusive. Here, we provide evidence that the Pacific oyster Crassostrea gigas utilizes glycophagy to manage glycogen mobilization during periods of energy deprivation. We identify an oyster STBD1 protein, and trace its origins through phylogenetic and comparative genomic analysis of the carbohydrate binding module family 20 (CBM20) domain within this protein across metazoans. Oyster STBD1 and those in other invertebrates contain an N-terminal CBM20, contrasting the C-terminal location of CBM20 in vertebrate STBD1. N-terminal CBM20 STBD1 proteins have a deep origin in bilaterians, with the vertebrate structural arrangement arising at the chordate root. Structural modelling and functional studies reveal that the N-terminal organization of the CBM20 domain in STBD1 enhances glycogen binding, with subsequent anchoring by GABARAPL2, facilitating an increased glycogen flux into autophagosomes for lysosomal degradation. We conclude that glycophagy is deeply conserved in bilaterians and that STBD1 structural evolution underlies potentially adaptive variation in metabolic strategies across distinct animal clades.
Similar content being viewed by others
Introduction
All animals rely on conserved metabolic processes to maintain energy balance and cellular homeostasis, essential for survival and fitness in changing environments1,2, including by managing the use of available energy reserves3,4. Autophagy is an ancient catabolic pathway that maintains cellular homeostasis when energy is limiting5. Unlike canonical metabolic pathways, autophagy operates at low baseline levels under normal conditions, but is activated in response to stressors including starvation6,7. Autophagy selectively targets intracellular nutrient macromolecules for degradation to supply a source of energy, including glycogen (glycophagy)8 and lipid droplets (lipophagy)9. Importantly, different metazoan clades are thought to primarily rely on distinct energy sources. Vertebrates and insects, for instance, use lipid as a predominant energy store, owing to its high energy density per unit mass10,11,12. A number of studies have shown that lipophagy is crucial for energy homeostasis in vertebrates13,14,15, while inhibiting autophagy in Drosophila diminished adipocyte differentiation and lipid droplet size without impacting glycogen mobilization16. On the other hand, studies on oysters indicate that glycogen rather than lipid is the primary energy source mobilized for gametogenesis17,18, strongly associated with an increase of autophagic vesicles19. These findings suggest that nutrient mobilization strategies via autophagy vary across the Metazoa, and this may be associated with lineage-specific adaptations in metabolism.
Glycophagy is a selective form of autophagy for transporting and degrading glycogen20. In vertebrates, starch-binding domain-containing protein 1 (STBD1) acts as the primary cargo receptor in this process21,22,23. Under nutrient deprivation, STBD1 recruits glycogen to form STBD1-glycogen complexes, binding with autophagy-related protein 8 (Atg8) family members to assemble glycophagosomes. These glycophagosomes fuse with lysosomes for glycogen degradation, mediated by lysosomal acid α-glucosidase (GAA)8. Underpinning this functional role is its structural domain composition. Mammalian STBD1 contains the hydrophobic N-terminal domain and Atg8-interacting motifs (AIMs), as well as a carbohydrate binding module family 20 (CBM20) domain at the C-terminus24. Among them, mutations in CBM20 diminish the stability of STBD1 and its ability to bind glycogen, while attenuating its interactions with glycogen-associated proteins25. Protein structural modelling revealed that CBM20 forms an immunoglobulin-like β-sandwich architecture, originally identified in bacterial amylolytic enzymes where it is localized to the C-terminus26,27. However, non-amylolytic CBM20-containing proteins such as LAF and GPCPD families exhibit an inverted topology with N-terminal CBM20 domains, potentially indicating an evolutionary adaptation of this domain for alternative metabolic pathways28,29. Interestingly, a recent enzyme engineering study on α-amylase (a member of the CBM20 family) demonstrated that relocating CBM20 to the N-terminus significantly enhanced its starch binding and degradation capabilities30. Current understanding remains insufficient to resolve the evolutionary significance of the N-terminal or C-terminal localization of CBM20 domain. That includes uncertainty surrounding the evolutionary origins of STBD1 and whether its potential role in glycophagy has shaped metabolic adaptability across diverse animal taxa.
Bivalve oysters exhibit distinctive glycogen metabolism strategies compared to most metazoans31,32,33. For instance, glycogen content in oysters is around 7-13-fold higher than that of human, fish, and Drosophila (Supplementary Fig. S1), while glycogen mobilization is highly regulated by the environment and both nutritional and reproductive status, which is not the case for lipid or protein stores33,34,35. As oysters have evolved to use glycogen metabolism as a primary strategy for energy homeostasis during stressful conditions, they provide an important system to explore potential adaptations in glycophagy. Here, we establish a key role for glycophagy in metabolic regulation in the Pacific oyster, Crassostrea gigas. STBD1 is present in oysters, but has an N-terminal CBM20 domain, and stably binds glycogen with high efficiency. N-terminal CBM20 represents the ancestral STBD1 organization, arising in the common ancestor of bilaterians and currently encoded in a range of lophotrochozoan genomes. We propose that STBD1-mediated glycophagy, as an ancestral strategy for maintaining cellular homeostasis during stress, likely supported metabolic adaptations during early bilaterian evolution.
Results
STBD1-mediated glycophagy plays a crucial role in energy metabolism in oysters
To investigate the role of autophagy in the response of oysters to stress, we manipulated nutritional status in C. gigas, using a 14-day fasting period followed by a refeeding phase (Fig. 1A). Immunohistochemistry showed that the autophagy marker LC3 was significantly enhanced by fasting, with signal weakening after refeeding in the labial palp, the adult tissue with the highest glycogen and lipid content (Fig. 1B, Supplementary Fig. S2 and S3A). In contrast, glycogen signals markedly diminished after fasting and subsequently reaccumulated upon refeeding, whereas lipid signals exhibited no changes (Fig. 1B, Supplementary Fig. S3B and S3C). Notably, LC3 signal was concentrated in the areas where glycogen accumulation, such as outer fold of mantle, interval of gill filament, ciliated epithelium (CE) and vesicular connective tissue (VCT) cells of labial palp, digestive diverticula and fringe of visceral ganglion, while showing minimal correlation with lipid signaling (Supplementary Fig. S3 and Supplementary Data 1). The high correlation between LC3 and glycogen signals, and the low correlation with lipid signals, implicates a specific role for autophagy in glycogen depletion under nutrient stress.
A Nutritional status in C. gigas was manipulated through a 14-day (d) period of fasting followed by a refeeding period lasting 48 hours (h). Created in BioRender. Ren, L. (2025) https://BioRender.com/l41ddy1. Parts of this figure were created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International license. B Signals for the autophagy marker LC3 (brown), glycogen (pink), lipid (red) in labial palp were observed after 14 d of fasting followed by 12 h of refeeding. Arrows indicate dense distribution of signals in the CE (black) and VCT (red) cells within the labial palp. Scale bars: 50 μm. Positive signals was quantified using ImageJ, as displayed in graphs on the right. C Expression levels of STBD1 and p62 in the labial palp of oysters after 14 d fasting followed by 48 h refeeding (n = 5). D Protein levels of STBD1 and p62 in the labial palp after 14 d fasting followed by 48 h refeeding (n = 3). E Expression levels of STBD1 in oyster individuals showing high and low glycogen content (n = 6). Data are represented as mean ± SD in (B, C, E), where statistical significance was calculated using one-way ANOVA in (B, C) and Student’s t-test in (E). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Numerical source data for the plots in this figure can be found in Supplementary Data 1.
To provide further evidence for the role of autophagy, we assessed the expression of candidate C. gigas orthologs for genes encoding the glycophagy receptor STBD1 (GenBank: LOC105333128) and the lipophagy receptor p62 (GenBank: LOC105345634)36. STBD1 mRNA expression increased significantly following 5-days fasting, reached maximum levels after 7 days, then declined, and exhibited a significant increase 6 h post-refeeding before returning to control levels (Fig. 1C). In contrast, p62 exhibited no significant alterations in expression across the same experiment. At the protein level, STBD1 protein abundance gradually decreased during fasting and significantly accumulated after refeeding, while p62 protein levels showed no statistically significant differences (Fig. 1D and Supplementary Data 1). These findings indicate that STBD1, rather than p62, functions as the substrate for utilization in the regulation of autophagy-related energy metabolism in oysters. Moreover, STBD1 showed increased mRNA expression in oysters with high levels of glycogen content (Fig. 1E and Supplementary Fig. S4), suggesting a close association between STBD1 and glycogen metabolism, implicating glycophagy as a crucial strategy used by oysters to maintain homeostasis during nutritional deficiency.
STBD1 mainly interacts with GABARAPL2 in oysters to drive glycophagy
The STBD1-glycogen complex binds to the Atg8 family of autophagy-modifying proteins through the AIM motif of STBD1, serving as another essential requirement for promoting the autophagic sequestration of glycogen24,37,38. GABARAPL1, an Atg8 family member, interacts with the STBD1-glycogen complex to trigger autophagosome maturation in vertebrates21,24. We screened the C. gigas genome for all known Atg8 family members to establish if GABARAPL1 is conserved in oysters. Phylogenetic analysis revealed C. gigas orthologs for LC3A, LC3C, GABARAP and GABARAPL2 (Fig. 2A), but not GABARAPL1, nor LC3B, an Atg8 family member involved in lipophagy through its interaction with p6213,14. These two Atg8 family members were also absent from other molluscan genomes we examined (Fig. 2A). The expression of the four identified oyster Atg8 family members increased significantly during fasting and decreased upon refeeding (Fig. 2B). We also identified a clear mRNA co-localization signal for STBD1 with Atg8 family members in the CE and VCT cells of labial palp, which was strongest for GABARAPL2, followed by GABARAPL and LC3A, while no signal was detected with LC3C in the VCT cells (Fig. 2C). To identify physical interactions between C. gigas STBD1 and Atg8 family members, each protein was co-expressed and their interactions were assessed using co-immunoprecipitation (co-IP) assays. The strongest interaction was between STBD1 and GABARAPL2, with an intensity about 4.4-fold higher than between STBD1 and the other three Atg8 family members (Fig. 2D and Supplementary Data 2). Therefore, STBD1 shows a preference for binding to GABARAPL2 member in oysters, which likely allows the entry of STBD1-glycogen complexes into autophagosomes. The potential loss of LC3B in oysters may also inhibit the autophagic degradation of lipid droplets in oysters (Fig. 2E). Overall, these results support the existence of a complete glycophagy pathway in oysters, analogous to that in vertebrates, whereas there is little evidence for an equivalent role for lipophagy.
A Phylogenetic tree of Atg8 family in C. gigas in relation to the other metazoans. C. gigas sequences are highlighted in red letters. The alpha-cyclodextrin glycosyltransferase from Thermoanaerobacterium thermosulfurigenes (Accession: AAB00845.1) was used as an outgroup. The corresponding abbreviations of species names see Supplementary Data 6. B Expression levels of Atg8 family-related genes, LC3A, LC3C, GABARAP and GABARAPL2, in the labial palp of oysters after 14 d fasting followed by 48 h refeeding (n = 5). C RNA co-localization of STBD1 with either LC3A, LC3C, GABARAP or GABARAPL2 in the labial palp of oysters. Signals from nucleus (blue), LC3A, LC3C, GABARAP and GABARAPL2 (green), STBD1 (red), and merge (orange). STBD1 co-localized with LC3A, LC3C, GABARAP or GABARAPL2 signals are indicated with white arrows. Scale bars: 75 μm. D Protein interaction between STBD1 and either LC3A, LC3C, GABARAP or GABARAPL2 using co-immunoprecipitation assays. Western blot detected bands from STBD1 (HA-conjugated), and LC3A, LC3C, GABARAP and GABARAPL2 (Flag-conjugated) (n = 2). E Schematic model of glycophagy in oysters: STBD1 interacts with GABARAPL2 directing glycogen into the autophagic degradation pathway, while p62-mediated lipophagy appears absent, possibly due to loss of LC3B. Created in BioRender. Ren, L. (2025) https://BioRender.com/lg0jx9r. Parts of this figure were created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International license. Data are represented as mean ± SD in (B), where statistical significance was calculated using one-way ANOVA. *p < 0.05, **p < 0.01. Numerical source data for the plots in this figure can be found in Supplementary Data 2.
N-terminal CBM20 STBD1 domain has higher predicted affinity for glycogen
Given the established role of conserved protein domains in STBD1-mediated glycogen binding and autophagosome degradation, we compared the domain composition of STBD1 from C. gigas alongside other mollusks with its orthologs in representative vertebrates. The most striking difference between molluscan STBD1 and vertebrate STBD1 was that the CBM20 domain was positioned at the N- and C-terminus, respectively (Fig. 3A). Moreover, the N-terminal hydrophobic domain of vertebrate STBD1 was absent in C. gigas (Fig. 3A). Potential AIM motifs were identified in C. gigas STBD1 as Y110-L113, Y145-V148, and F177-I180 (Fig. 3A). However, the characteristic W-X-X-V AIM motif found in vertebrate STBD1 proteins was absent in C. gigas STBD1.
A Comparative domain analysis of STBD1 in representative vertebrate and molluscan species. Blue and orange boxes represent vertebrate and mollusk STBD1 protein structures, respectively. Green rectangles indicate N-terminal hydrophobic domains, pink rectangles indicate AIM motifs, and blue and orange rectangles mark the CBM20 domains of vertebrates and mollusks, respectively. B–E Molecular docking analysis of mouse, zebrafish, wild-type C. gigas STBD1 and in silico-mutated C. gigas STBD1 with CBM20 moved to the C-terminus (90-194 aa) with branched maltotetraose. All protein models were generated using the AlphaFold 3 server. The Site 1 and Site 2 oligosaccharide-binding pockets of mouse and zebrafish are indicated by light blue and dark blue bars, respectively, while those of C. gigas and the C. gigas_mutant are indicated by yellow and orange bars, respectively. Branched maltotetraose is indicated by pink bars. Hydrogen bonds are indicated by green dashed lines and labeled with bond lengths. F–I Per-residue binding energy contributions from molecular dynamics simulations of mouse, zebrafish, wild-type C. gigas, and C. gigas_mutant STBD1 with oligosaccharides.
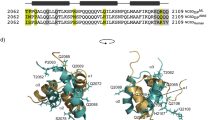
To elucidate the impact of the CBM20 domain’s location on STBD1 function, we predicted and obtained three-dimensional structures of C. gigas STBD1 (N-terminal CBM20, wild-type), vertebrate STBD1 proteins (C-terminal CBM20) and an in silico re-arranged C. gigas STBD1 (C-terminal CBM20) (Fig. 3B–E), followed by a detailed analysis of their binding kinetics with branched maltotetraose. For all STBD1 proteins, CBM20 was modelled as a distorted β-barrel composed of eight β-strands (Fig. 3B–E), showing high structural similarity to the recently resolved human CBM20 (PDB ID: 8X8K) (Supplementary Fig. S5A). Docking results revealed that both oyster and vertebrate STBD1 possess two oligosaccharide-binding pockets. Within the Site 1 binding pocket, C. gigas STBD1 W36, K69, W83, and N88 residues are highly conserved at the equivalent binding positions in mouse and zebrafish C-terminal CBM20 proteins (Fig. 3B–D and Supplementary Fig. S5B). However, in the Site 2 pocket of C. gigas STBD1, beyond the vertebrate-equivalent residues (Q14, R82, W104), the polar side chains of E85, N105, and E106 form additional hydrogen-bond interactions with maltotetraose (Fig. 3B–D), indicating a distinctive oligosaccharide-recognition mode. Further molecular dynamic (MD) analyses revealed that while all complexes converged on a similar timescale ( ~ 30 ns) (Supplementary Fig. S6), their binding energy profiles were distinct. Site 1 was the primary contributor in the mouse and zebrafish complexes (Fig. 3F, G), whereas in C. gigas, Site 2 contributed more prominently within a distributed energy profile across both sites (Fig. 3H). Together, these residues in the N-terminal CBM20 of C. gigas STBD1 yielded a predicted binding energy of −36.48 kcal/mol with maltotetraose, markedly stronger than that of the C-terminal CBM20 in mouse and zebrafish STBD1 (−23.67 to −24.99 kcal/mol) (Supplementary Data 3). Per-residue energy decomposition analysis revealed that binding at Site 1 of the C. gigas STBD1–maltotetraose complex was dominated by Van der Waals (W36, W83) and polar solvation interactions (N88), whereas Site 2 binding resulted from the synergistic contributions of electrostatic (Q14, E85, E106), Van der Waals (W104) and non-polar solvation interactions (F15) (Supplementary Data 4). In addition, comparison with the C. gigas STBD1_mutant-maltotetraose complex further confirmed that relocating CBM20 from the N- to the C-terminus markedly reduced the Site 2 residue contributions (Fig. 3H, I), indicating that positioning CBM20 at the N-terminus significantly enhances the oligosaccharide-binding affinity of Site 2 residues.
Oyster STBD1 possesses enhanced binding affinity that expedites glycogen breakdown
Next, we sought to experimentally validate the predicted enhanced glycogen-binding affinity conferred to oyster STBD1 by its distinctive N-terminal CBM20. We successfully isolated recombinant proteins for wild-type STBD1 proteins in zebrafish, mouse and C. gigas, along with a mutant C. gigas STBD1 protein with CBM20 moved to the C-terminus (Fig. 4A and Supplementary Fig. S7). Glycogen co-sedimentation assays showed that all isolated STBD1 proteins bound glycogen in vitro, with the strongest binding for wild-type C. gigas (Fig. 4B), approximately 1.5-fold higher than mutant C. gigas STBD1, and 2.1–2.7-fold higher than the wild-type vertebrate STBD1 proteins (Fig. 4B and Supplementary Data 5). Thus, the N-terminal position of CBM20 in oyster STBD1 markedly enhances affinity for glycogen compared to both the same protein, as well as evolutionarily distant STBD1 orthologs, containing a C-terminal CBM20 domain.
A Schematic diagram summarizing the construction and isolation of a C. gigas STBD1 protein with CBM20 moved to the C-terminus, which was purified along with mouse, zebrafish and wild-type C. gigas STBD1. Created in BioRender. Ren, L. (2025) https://BioRender.com/yr3ifov. Parts of this figure were created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International license. B Binding of the four recombinant STBD1 proteins with glycogen was measured. Western blot assessed the protein levels in inputs, supernatants, and the pellet after ultracentrifugation. PET32a vector protein was used as a negative control (n = 3). C Changes in protein levels of each STBD1 protein overexpressed in 293 T cells after transfection for 24 h, followed by treatment with or without EBSS for 6 h. Western blot was used to verify protein levels (n = 3). D Glycogen content in 293 T cells after overexpression of each STBD1 protein for 24 h, followed by treatment with or without EBSS for 6 h (n = 4). Data are represented as mean ± SD in (D), where statistical significance was calculated using two-way ANOVA. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Numerical source data for the plots in this figure can be found in Supplementary Data 5.
We next tested the hypothesis that the relatively stronger affinity between glycogen and STBD1 from C. gigas supports enhanced glycogen degradation. The same four STBD1 proteins were overexpressed in human 293 T cells and treated with the autophagy inducer EBSS for 6 h, before changes in STBD1 protein abundance and glycogen content were determined by comparison to controls. Western blot analysis showed that C. gigas STBD1 protein abundance was markedly higher than for vertebrate STBD1 (Fig. 4C). Moreover, an approximate 5.5-fold reduction in wild-type C. gigas STBD1 abundance was observed following 6 h autophagy induction, compared to a respective 1.3-fold, 2.0-fold and 2.7-fold reduction for mouse, zebrafish and the mutant C. gigas STBD1 (Fig. 4C and Supplementary Data 5), indicating that C. gigas STBD1 protein is utilized more efficiently than the three STBD1 proteins with C-terminal CBM20 domains. Importantly, the glycogen content of 293 T cells overexpressing wild-type C. gigas STBD1 was significantly lower after autophagy induction than for the other three STBD1 proteins (Fig. 4D). Nonetheless, 293 T cells overexpressing all four STBD1 proteins showed significant reductions in glycogen content following autophagy induction compared to controls (Fig. 4D). As these experiments were conducted in a heterologous human cell background, the observed differences should be interpreted as evidence of intrinsic, protein-level functional divergence under matched cellular conditions rather than direct measurements of organism-level metabolic flux across lineages. Nonetheless, the results support the interpretation that an N-terminal CBM20 enhances glycogen engagement with the autophagy pathway, while organism-level implications should be drawn with caution and validated in species-appropriate systems.
Resolving the evolutionary origins of STBD1 in metazoans
Our results show that glycophagy is conserved between oysters and vertebrates, revealing the significance of the CBM20 domain’s position for STBD1 glycogen binding function. Considering the vital role of CBM20 in glycophagy, we investigated the phylogenetic origin of genes encoding CBM20 domain-containing proteins across metazoan lineages, with the aim of tracing the evolutionary history of STBD1, including its CBM20 domain arrangement. We conducted a comprehensive genome-wide scan of CBM20 domain containing proteins in 74 metazoan species from 14 phyla (Supplementary Data 6) and compared their domain organizations. Proteins containing CBM20 were present in all examined phyla except Ctenophora (comb jellies), representing seven major domain architectures (Fig. 5 and Supplementary Fig. S8). To reconstruct their evolutionary relationships, we performed maximum-likelihood phylogenetic analyses using amino acid alignments filtered specifically to resolve CBM20 domain evolution, incorporating sequences from metazoans and 11 additional non-metazoan eukaryotes (Supplementary Fig. S9, Supplementary Data 7). In metazoans, CBM20-containing proteins form three clades: glycerophosphocholine phosphodiesterase 1 (GPCPD1), laforin (LAF), and STBD1, each including previously characterized vertebrate representatives (Fig. 6A and Supplementary Fig. S10), consistent with previous reports in mammals that CBM20 is restricted to GPCPD1, LAF, and STBD124. The presence of poriferan and placozoan sequences in the GPCPD1 clade, alongside the observation that GPCPD1, LAF and STBD1 groupings predate branching events within each of these clades, suggests that all three CBM20-containing clades may have been present in the metazoan ancestor, with subsequent lineage-specific patterns of loss and expansion. This includes evidence for prevalent losses of LAF proteins and widespread retention of GPCPD1 across different metazoan lineages (Fig. 6A). Moreover, cnidarian proteins within the LAF clade exhibit strong sequence and structural similarity to chordate LAF proteins. Multiple sequence alignments confirmed conservation of functional domain sequences, and structural superpositions of representative cnidarian and chordate LAF proteins revealed highly conserved overall folds (RMSD = 0.806 ~ 1.187 Å; mean MSA LDDT = 0.802) (Supplementary Fig. S11). Taken together, these results suggest that LAF was already present before the divergence of cnidarians and bilaterians, despite its apparent absence in protostomes and echinoderms.
The schematic tree indicates evolutionary relationships based on NCBI Taxonomy. CBM20 domain-containing proteins were categorized into seven types based on domain structures (Supplementary Fig. S8), with schematic diagrams in the upper-right corner. Other domains refer to EF-hand (IPR002048), peptidase M12B (IPR001590), IPT (IPR002909), and glycosyl hydrolase family 13 (IPR006047). CBM20 Carbohydrate binding module family 20 domain (IPR002044), GP-PDE Glycerophosphodiester phosphodiesterase domain (IPR030395), DSP Dual specificity phosphatase domain (IPR000340). The silhouette images of the animals in this figure were modified from PhyloPic (https://www.phylopic.org/), which is an open database of free silhouette images of animals, plants, and other life forms, available for reuse under Creative Commons licenses.
A Phylogenetic clade of eukaryotic STBD1, LAF, and GPCPD1 proteins. Proteins are colored according to the lineages. Red dots mark nodes with bootstrap support >50%. The corresponding abbreviations of species names are listed in Supplementary Data 6 and 7. The complete tree with bootstrap support values is in Supplementary Fig. S10. B Hypothesis for STBD1 evolution: a speculative model for the evolution of STBD1. The common ancestor of metazoans harbored GPCPD1 and two CBM20-2 proteins. One CBM20-2 protein likely gave rise to LAF in the common ancestor of parahoxozoans through domain gain, while the other preserved the ancestral N-terminal CBM20 and evolved into STBD1α in the common ancestor of bilaterians. GPCPD1 and CBM20-2 were lost in Cnidaria, while STBD1α was independently lost in Cnidaria, Ecdysozoa, and Echinodermata. At the root of Chordata, STBD1β evolved from STBD1α through N- to C-terminal CBM20 domain shuffling. Protostomes and echinoderms independently lost LAF. The yellow, blue, dark green, and purple branches indicate the presence of STBD1α, STBD1β, LAF, and GPCPD1 genes during evolution, respectively.
Notably, STBD1 proteins with a single N-terminal CBM20 domain (hereafter: STBD1α) were exclusively found in lophotrochozoans, while C-terminal CBM20 (hereafter: STBD1β) proteins were restricted to Chordata (Figs. 5 and 6A, Supplementary Fig. S10). A marked STBD1α expansion was observed in mollusks, especially the bivalves Sinonovacula constricta and Ostrea edulis. In addition, we identified STBD1 proteins with two CBM20 domains (STBD1γ) in S. constricta and some gastropods, and multiple CBM20 domains (STBD1δ) in nemerteans. These findings provide robust evidence for lineage-specific expansions and structural diversification of STBD1 in lophotrochozoans and chordates, suggesting a diverse repertoire of proteins in lophotrochozoans and chordates that may underlie functional divergence and specialization.
STBD1 and LAF were supported as sister clades using midpoint rooting39 of our phylogenetic tree (Fig. 6A). Given that STBD1 is present only in bilaterians, it is challenging to distinguish loss scenarios in other metazoan lineages from an origin specific to early bilaterian evolution. By incorporating CBM20-containing proteins from non-metazoan species, we found that metazoan LAFs cluster with diverse non-metazoan CBM20 proteins (e.g., DPE2, GLAA, and LAF), whereas bilaterian STBD1 proteins form a distinct clade. Notably, LAF appears to have originated independently in metazoans and in the protist Tetrahymena thermophila. These findings suggest that the common ancestor of animals may not have possessed LAF, but instead contained a protein with a single N-terminal CBM20 domain (hereafter: CBM20-2). Importantly, STBD1α exhibits a similar N-terminal CBM20 domain structure, consistent with CBM20-2 representing the ancestral domain architecture of both LAF and STBD1. Therefore, we propose a speculative model in which GPCPD1 and two CBM20-2 proteins were present in the common ancestor of metazoans (Fig. 6B). LAF likely arose from one of these CBM20-2 proteins through domain gain in the common ancestor of parahoxozoans, while the other CBM20-2 preserved the ancestral N-terminal CBM20 domain in the common ancestor of bilaterians and subsequently evolved into STBD1α. After the divergence from other deuterostomes, chordate-specific STBD1β may derive from domain shuffling of CBM20 from the N-terminus to the C-terminus. Moreover, lineage-specific domain duplication events in nemerteans and mollusks have resulted in the generation of STBD1γ and STBD1δ with two or more CBM20 domains. In lophotrochozoans, species-specific gene duplications have expanded the STBD1 repertoire. Overall, gene expansion, domain shuffling and duplication have shaped STBD1 evolution to the present various degrees of conservation and divergence observed across bilaterians.
Discussion
Glycogen, the primary reservoir for surplus glucose, facilitates rapid energy mobilization during periods of energy deficit through its highly branched architecture, enabling simultaneous enzymatic cleavage at multiple sites40,41. Although canonical glycogen metabolism pathways are highly conserved across metazoans, the efficiency of glycogen storage and utilization varies remarkably among lineages10,42,43,44. Glycophagy has been reported as a non-canonical energy producing pathway in vertebrates20,21,22. A recent study revealed that oysters undergoing energetically demanding gametogenesis showed an increased number of autophagic vesicles, serving as large reservoirs of glycogen19, suggesting a potential role for glycophagy in energy homeostasis, and hinting that this pathway may considerably predate vertebrates. Using oysters as a study system, this study aimed to comprehensively resolves the evolutionary origins of glycophagy within Metazoa, revealing an overall conserved role of STBD1 within ancient autophagy networks, alongside evidence for structural modifications shaping glycogen binding that can be linked to the distinct metabolic strategies of different metazoan lineages.
The autophagy pathway demonstrates substrate-specific degradation preferences (glycogen or lipid) across species with divergent energy metabolism strategies. We observed that autophagic signal was primarily localized to oyster tissues with glycogen accumulation, consistent with glycogen turnover regulated by nutritional fluctuations. In contrast, the association with lipids was relatively minimal, which was consistent with the observation that glycogen was the primary energy source in oysters17,18,19. This autophagy-mediated metabolic adaptation strategy exhibits a distinct divergence from that of species predominantly relying on lipid-based energy utilization. Phenotype observations in the Atg7 or beclin knockout zebrafish found a remarkable undigested intestinal lipid and triglycerides aggregation, whereas glycogen was gradually depleted45. Similarly, in Drosophila larvae, inhibition of autophagy has been noted to diminish adipocyte differentiation and lipid droplet size, yet without discernible impact on glycogen mobilization16. Comparing the differential contributions of energy substrates (glycogen, lipid or protein) between oysters and other metazoans during the autophagy process indicates that autophagy mediates adaptive adjustments in energy homeostasis. This is tightly linked to the selective degradation functions of the glycophagy- and lipophagy-specific receptors, STBD1 and p6246. In oysters, the expression pattern of STBD1 corresponds to the degradation of autophagosomal components by lysosomal acidic hydrolases following autophagosome-lysosome fusion during autophagy activation47,48. However, p62, a key regulator of lipophagy, remains largely unaltered in response to nutritional fluctuations in oysters. Similar observation was reported in zebrafish, where autophagy promoted p62 turnover instead of STBD149,50, suggesting that autophagy receptors mediate adaptive metabolic regulatory mechanisms across different metazoan lineages.
Functional domains of STBD1 play indispensable roles in facilitating the positioning and transportation of glycogen to the autophagosome23. Oyster STBD1 displays a domain architecture distinct from that of vertebrates, featuring an N-terminal CBM20 domain while lacking the W-X-X-V AIM motif and N-terminal hydrophobic region found in vertebrate STBD1. N-terminal CBM20 in oyster STBD1 enhances its glycogen-binding efficiency relative to vertebrate orthologs and promotes a heightened influx of glycogen into the autophagy pathway for degradation. This functional divergence may be attributed to the Site 2 pocket within N-terminal CBM20, which provides substantial electrostatic interactions, Van der Waals forces, and non-polar solvation contributions for oligosaccharide binding to oyster STBD1. Notably, unlike the conserved Site 1, Site 2 lacks evolutionary conservation. We found that the key residue E85 provides a strong energetic contribution to oligosaccharide binding in oyster STBD1, whereas its effect is weaker in zebrafish and not predicted in mouse. Similar divergence is also observed in comparisons between human STBD1 and its paralogs, EPM2A and GPCPD124, consistent with lineage-specific diversification of the Site 2 binding mode. Additional AIM motifs in oysters were delineated as [Y/F]-X-X-[L/V/I], lacking the classical W-X-X-V motif that interacts with GABARAPL1 in mammals, suggesting an alternative binding preference for distinct Atg8 family members51,52. Indeed, the C. gigas genome encodes only four Atg8 homologs-LC3A, LC3C, GABARAP, and GABARAPL2, with GABARAPL2 functionally compensating as the primary interacting partner of STBD1. The GABARAP ortholog LGG-1 is indispensable for autophagosome formation in C. elegans, whereas loss of the LC3 ortholog LGG-2 exclusively yielded smaller autophagosomes53. Notably, the loss of the lipophagy-associated LC3B further reflects metabolic adaptations in oysters, potentially compensating for the limited lipid utilization pathway by reinforcing a glycogen-dominated energy homeostasis mechanism. Furthermore, vertebrate STBD1 requires the N-terminal hydrophobic region for endoplasmic reticulum localization, but this region is absent in oysters23. Loss of this domain likely underlies the shorter length of molluscan STBD1 compared with its vertebrate counterparts. We speculate that the emergence of the N-terminal hydrophobic region in vertebrates reflects a domain-gain event during evolution, which enabled neofunctionalization towards roles in mediating endoplasmic reticulum (ER) stress responses54 and facilitating physical contacts between the ER and mitochondria55. Notably, although deletion of this region results in a diffused cytoplasmic distribution, it does not impair the binding of glycogen or amylose24,25, further explaining that oyster STBD1 binds directly to glycogen in a free state, leading to a higher glycophagy flux. Collectively, the structural domain composition of STBD1 demonstrates evolutionary diversity and functional specialization across different taxa. Future work should examine 3D domain topology and coupling (e.g., a chimera swapping murine C-terminal CBM20 for the oyster N-terminal CBM20) to determine how interdomain crosstalk affects oligosaccharide binding and degradation.
Protein domain evolution is a key process leading to changes in molecular function within metabolic pathways56. This study reconstructed the evolution of CBM20 domain-containing genes in metazoans. CBM20 was also ancestrally located at the N-terminal of STBD1, when it arose during early bilaterian evolution, potentially from the functionalization of an ancestral CBM20-2 gene. Oyster STBD1 exhibits the ancestral N-terminal CBM20 domain, whereas a domain shuffling event in early chordates led to the derived C-terminal CBM20 of STBD1 found in all vertebrates. The repositioning of CBM20 to the C-terminus of STBD1 thus led to lower binding with glycogen compared to the ancestral condition. It is plausible this reflects adaptive trade-offs in energy mobilization strategies. While vertebrates balance glycogen and lipid utilization via coordinated glycophagy-lipophagy pathways13,22, oysters, as lophotrochozoans, exhibit a glycogen-centric metabolism, with lipophagy evidently absent. Interestingly, among all lophotrochozoans analyzed, some species may have experienced recent events of domain duplication and recruitment, respectively resulting in the STBD1 genes with more than one CBM20 copy and other functional domains. Thus, STBD1 has evolved rapidly in lophotrochozoans, with a high proportion of lineage-specific gene duplications and domain rearrangements, implying ongoing innovation in glycophagy pathways within this lineage. Further work is required to elucidate the functional role of these diverse STBD1 paralogs and determine whether these putative glycophagy receptors also interact with upstream autophagy components. In addition to the bilaterian evolution of STBD1, the origin of CBM20 domain-containing genes in early metazoans remains an open question. While our phylogenetic reconstruction suggests that three ancestral CBM20-containing proteins were likely present in the last common metazoan ancestor, an alternative and equally plausible interpretation is that the similarity between the Hexacorallian LAF-like protein and vertebrate LAF reflects convergent evolution. In this view, the Hexacorallian CBM20-2 protein may have independently acquired a LAF-like domain architecture and a function analogous to that of vertebrate LAF. This scenario would eliminate the need to invoke multiple independent losses of LAF across cnidarians and other metazoan lineages and instead imply repeated, lineage-specific innovations of LAF-associated functions. Further structural and functional analyses, particularly biochemical characterization of the Hexacorallian LAF-like proteins, will be essential to distinguish between these alternative hypotheses.
In conclusion, our demonstration of the conserved function of STBD1 in oysters and vertebrates positions STBD1-mediated glycophagy as an ancestral metabolic pathway in bilaterians (Fig. 7). Lophotrochozoan STBD1 with N-terminal CBM20 retains an ancestral domain composition from the ancestor of bilaterians and exhibits strong binding affinity for glycogen. In vertebrates, a domain shuffling event shaped chordate-specific STBD1 with C-terminal CBM20 domain. This structural characteristic leads to low binding forces between vertebrate STBD1 and glycogen, potentially designating the lipophagy pathway as a compensatory or alternative mechanism to glycophagy for maintaining energy homeostasis. This work further highlights the importance of protein structural evolution in the regulation of energy metabolism, thereby advancing understanding of how distinct metabolic strategies arose in different animal lineages.
STBD1-mediated glycophagy originated before the diversification of bilaterians. In mollusks and other lophotrochozoans, STBD1α represents the ancestral domain architecture of STBD1 proteins. Among them, oyster STBD1α binds glycogen with high affinity and subsequently anchors to GABARAPL2, facilitating glycophagic flux and providing a primary energy source. Vertebrate STBD1β proteins inherited an ancestral chordate rearrangement of the CBM20 domain to the C-terminus, resulting in lower binding affinity with glycogen compared to the N-terminal position, leading to a relatively reduced glycophagic flux through the glycophagy pathway by binding with GABARAPL or GABARAPL1. Consequently, oysters utilize glycophagy to degrade glycogen as a primary energy source, whereas vertebrates rely more strongly on lipophagy. LD lipid droplet, FFA free fat acid. Created in BioRender. Ren, L. (2025) https://BioRender.com/u86c722. Parts of this figure were created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International license. The silhouette images of the animals in this figure were modified from PhyloPic (https://www.phylopic.org/), which is an open database of free silhouette images of animals, plants, and other life forms, available for reuse under Creative Commons licenses.
Materials and Methods
Animals and treatment
Two-year-old healthy C. gigas individuals (n = 200) were collected from an oyster farm in Rongcheng (37.1°N, 122.5°E, Shandong, China) in May, and acclimated in 50 L tanks for 7 days. The oysters were subjected to a 14-day (d) period of consecutive fasting (F), followed by a single refeeding (R) event with approximately 2.52 × 105 cells/mL of concentrated algae fluid, and then observed for 48 hours (h). Mantle, gill, labial palp, digestive gland, adductor muscle and visceral ganglion was dissected from n = 3 individuals after fasting for 14 d and refeeding for 12 h, then fixed for histology. Labial palp was dissected from n = 6 individuals at 1 d, 3 d, 5 d, 7 d, 14 d after fasting, then again, 1 h, 3 h, 6 h, 12 h, 24 h, 48 h after refeeding for molecular work. N = 9 individuals were sampled at 0 d prior to fasting as a control group. Six tissues were dissected from n = 6 individuals for glycogen and lipid quantification. Labial palp was dissected from n = 71 randomly selected individuals for glycogen content quantification, based on the observed distribution, we selected two extreme groups (n = 9 each) with average dry weight glycogen content of 19.15 ± 4.05% (high glycogen) and 10.7 ± 0.45% (low glycogen). Mouse (n = 3, female, 9-12 weeks old) and zebrafish (n = 3, female, 1 year old) were used for recombinant protein preparation. All samples used for molecular work were flash frozen in liquid nitrogen and kept at −80 °C.
Glycogen content measurement and staining
Glycogen content was determined using a Glycogen Content Assay Kit (Solarbio, BC0345) following the manufacturer’s instructions. Prior to measurement, the tissue samples were freeze-dried for 48 h and then pulverized into powder (0.1 g used for measurements). The cell samples were subjected to 20 rounds of sonication for 3 s, at intervals of 10 s. Then, 750 μL of extract solution was added to each sample, before the mixture was boiled for 20 min and centrifuged at 8,000 g for 10 min to obtain the supernatant. Finally, 240 μL of anthranone was added and boiled, before the light absorption value was determined using a Synergy™ H1 (BioTek) plate reader at an optical density (OD) of 620.
Glycogen staining was performed using a Periodic Acid Schiff (PAS) Stain Kit (Solarbio, G1281) according to the manufacturer’s instructions. Briefly, samples were fixed in 4% PFA, dehydrated in gradient alcohol, embedded in paraffin and sectioned at 5 μm thickness using a LEICA RM2016 microtome (Lecia). Sections were hydrated and oxidized with periodic acid for 5 min, stained with Schiff reagent for 15 min, and counterstained in hematoxylin for 1 min. Images were acquired with a Zeiss Axio Scope A1 microscope equipped with an AxioCam MRc5 digital camera. Positive signals were quantified using Image J (v 2.3.0)57.
Lipid content measurement and staining
Lipid content was determined using a Triglyceride Content Assay Kit (Solarbio, BC0625) following the manufacturer’s instructions. A 0.1 g portion of freeze-dried sample was mixed with 1 mL of Solution 1 and homogenized on ice. The homogenate was then centrifuged at 8,000 g for 10 min to obtain the supernatant. Then, Solution 2 was added, followed by repeated shaking and resting to collect the upper layer of the solution. Subsequently, Solution 3 and Solution 4 were added, and the mixture was heated in a 65 °C water bath for 3 min. Then, Solution 5 and Solution 6 were added, and the mixture was further heated in a 65 °C water bath for 15 min. After cooling, 200 µL of the prepared solution was measured using a Synergy™ H1 (BioTek) plate reader at an optical density (OD) of 420 nm.
Lipid staining was done using Oil Red O Saturated Solution (Solarbio, G1260) according to the manufacturer’s instructions. Samples were prepared as described for glycogen staining. Sections were stained with Oil Red O solution for 10 min, differentiated in 60% isopropanol until the interstitial spaces became clear, and then counterstained in hematoxylin for 1 min for nuclear staining. Images acquisition and quantification of positive signals was performed as described for glycogen staining.
Immunohistochemistry assay
Tissue samples were fixed in 4% PFA, dehydrated in gradient alcohol, and then embedded in paraffin and sectioned at 5 μm thickness. Sections were deparaffinized and hydrated followed by antigen retrieval in 0.01 M sodium citrate buffer (pH 6.0) at 95 °C for 10 min. Slides were rinsed by 1 × PBS for 15 min and immersed in 3% H2O2 for 10 min, then blocked with 5% Bovine serum albumin (BSA, Sangon, A602449) in PBS for 30 min. Next, sections were incubated with 1:400 diluted primary antibody rabbit anti-LC3A/B (Cell Signaling Technology, 4108) overnight at 4 °C. After 1 × PBS washing for 30 min, secondary antibody HRP-conjugated goat anti-rabbit (Beyotime, A0208) was added, and incubated for 30 min. Finally, staining was performed with 3,3-diaminobenzidine (DAB, Sigma, D12384) substrate and counterstained with hematoxylin for nuclear differentiation. Negative control without primary antibody treatments were included. Images acquisition and quantification of positive signals was performed as described for glycogen staining.
Quantitative real-time PCR
Total RNA was extracted from samples using FreeZol reagent (Vazyme, R711) according to the manufacturer’s instructions. RNA concentration and purity were assessed using a NanoDrop 2000 spectrophotometer (Thermo Fisher). Total RNA (1 μg) was reverse transcribed with the HiScript III RT SuperMix for qPCR ( + gDNA wiper) (Vazyme, R323). The transcript abundances of genes of interest were measured using ChamQ SYBR qPCR Master Mix (Vazyme, R311) with a LightCycler®480 real-time PCR instrument (Roche). 10.0 μL RT-PCR reactions contained 5.0 μL 2 × SYBR Green PCR Master Mix, 0.7 μL of each forward and reverse primer (10 μM), 1.0 μL diluted cDNA, and 2.6 μL PCR-grade water. The relative expression of target genes was calculated by the 2-ΔΔCt method using EF1α (Elongation factor 1-α) as reference. Target gene specific primers (Supplementary Data 8) were designed using Primer Express software (Applied Biosystems). Differences in gene expression across different points in the time course compared to the baseline at 0 d before fasting.
STBD1 domain analyses
Amino acid sequences of STBD1 were downloaded from NCBI database (https://www.ncbi.nlm.nih.gov) for mouse (GenBank: 52331), zebrafish (GenBank: 792854) and wild-type C. gigas (GenBank: LOC105333128). The domain structures of these proteins were further confirmed by comparison with the InterPro database (https://www.ebi.ac.uk/interpro/). AIM motifs were identified based on the standard sequence [W/Y/F]-X-X-[L/V/I].
Molecular modeling and docking with maltotetraose
To compare STBD1 protein structures carrying N-terminal or C-terminal CBM20 domains, an in silico CgSTBD1 mutant was generated by repositioning the N-terminal CBM20 domain (aa 1-106) to the C-terminus (aa 90-194) (Fig. 4A). Mouse and zebrafish STBD1 were used as comparative references containing a C-terminus CBM20 domain. STBD1 structural models were generated using the online AlphaFold 358 and assessed for predictive confidence based on pLDDT scores (Supplementary Fig. S5A). Further validation of the predicted model was conducted by structural alignment using PyMOL (v 2.5.5)59 with the experimentally resolved crystal structure of human STBD1/CBM20 (PDB ID: 8X8K)24, yielding a root mean square deviation (RMSD) of approximately 0.5 Å, thereby confirming the accuracy of the prediction (Supplementary Fig. S5A).
Molecular docking of branch maltotetraose with STBD1 proteins from mouse, zebrafish, wild-type C. gigas, and the in silico C. gigas STBD1 mutant (Supplementary Data 9–12) was carried out using sugar-optimized AutoDock Vina-Carb (v 1.1.2)60. Protein structures from AlphaFold prediction were refined through the incorporation of hydrogen atoms, integration of non-polar interactions, before computation of kollman charges. The 3D structure of the ligand glycogen was obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/), before being optimized and energy minimized using Chem 3D61, and finally hydrogen atoms were added and the torsion key was set. After pretreatment of the ligand and receptors, parameter files were configured with appropriate grid box dimensions for only the CBM20 domain of STBD1: mouse (Site 1: 21.0 × 18.75 × 17.25; Site 2: 20.0 × 20.0 × 20.0), zebrafish (Site 1: 26.25 × 26.25 × 25.5; Site 2: 19.5 × 15.75 × 18.0), C. gigas (Site 1: 37.5 × 29.25 × 31.5; Site 2: 26.25 × 25.5 × 19.5) and the in silico C. gigas STBD1 mutant (Site 1: 24.75 × 26.25 × 24.0; Site 2: 21.0 × 19.7 × 21.6). Running exhaustiveness was set to 24, and default settings were used for all other parameters. The optimal docking model was defined with reference to the resolved crystal structure of human STBD1/CBM20 (PDB ID: 8X8K) and an RMSD threshold of <2 Å, and was subsequently employed for binding free energy calculations. Protein-ligand interactions were visualized by PyMOL (version 2.5.5).
Molecular dynamics simulation and binding free energy calculation
Molecular dynamics (MD) simulations of STBD1-maltotetraose complexes from mouse, zebrafish, wild-type C. gigas, and an in silico C. gigas STBD1 mutant were systematically analyzed using GROMACS (v2024.5)62. Proteins were parameterized with Amber ff14SB and maltotetraose with GLYCAM_06j-1, using the TIP3P water model throughout. Topologies were prepared in AmberTools (v23.6)63 and converted to GROMACS format via ACPYPE (v2023.10)64. Each protein–ligand complex was solvated in a cubic TIP3P box with ≥10 Å padding in all directions and neutralized with Na⁺/Cl⁻. Energy minimization (steepest-descent, up to 50,000 steps) proceeded until the maximum force was < 1000 kJ·mol⁻¹·nm⁻¹, under periodic boundary conditions with particle-mesh Ewald (PME) electrostatics. Equilibration comprised 1 ns in the NVT ensemble at 300 K (V-rescale thermostat) followed by 1 ns in the NPT ensemble at 1 bar (isotropic barostat) with heavy-atom positional restraints. Production trajectories (100 ns) were run in NPT at 300 K and 1 bar with a 2 fs timestep; bonds to hydrogens were constrained with LINCS. Long-range electrostatics used PME, and Van der Waals and short-range Coulomb interactions employed a 1.2 nm cutoff. Coordinates and energies were written every 10 ps.
Binding free energy was estimated via MM/GBSA using gmx_MMPBSA (v1.6.3)65. The final production trajectories were sampled at every 10th frame. Calculations employed GB model igb=8 with a 1.4 Å solvent probe. Per-residue energy decomposition (idecomp = 3) was performed to identify residues that drive ligand binding.
Phylogenetic analysis of Atg 8 family and metazoan CBM20 domain-containing proteins
We downloaded protein datasets from 74 metazoan genomes representing 14 phyla (Supplementary Data 6), as well as 11 non-metazoan eukaryotic genomes (Supplementary Data 7). Atg8 family proteins (Pfam: PF02991.22) were identified in representative vertebrates (M. musculus, G. gallus, C. mydas, X. laevis, and D. rerio) and molluscan species (C. gigas, C. virginica, M. coruscus, P. maximus, and H. rubra) using hmmsearch (HMMER v.3.3.2)66 with the trusted cutoff parameter “--cut_tc”. Similarly, proteins containing sequence similarity to the CBM20 domain (Pfam: PF00686.24) were retrieved using the same method and cutoff. Domain annotation was achieved using InterProScan (v.5.69) with default parameters67. Transmembrane helices were predicted using TMHMM-2.0 online (https://services.healthtech.dtu.dk/services/TMHMM-2.0/). Whole sequences were aligned using MAFFT with the parameters “--maxiterate 1000 --localpair”. The resulting alignment was trimmed using TrimAl68 with a gap threshold of 0.8 (Supplementary Data 13). Phylogenetic relationships were estimated using maximum likelihood with IQ-Tree (v.2.3.5) and the parameters “-B 5000 -bnni”69. ModelFinder70 from IQ-Tree was used to select the best-fitting model (Q.pfam+F + I + R5). The final phylogenetic trees (Supplementary Data 14) were visualized and labeled using iTOL (v.6.9.1) online (https://itol.embl.de/).
Plasmid construction
Plasmids used for purification of recombinant STBD1 proteins were constructed in the pET32a vector (with 6 × His-tag, Biomed). The complete coding sequences of STBD1 from mouse (GenBank: 52331), zebrafish (GenBank: 792854) and wild-type C. gigas (GenBank: LOC105333128) were PCR amplified using 2 × Phanta Max Master Mix (Vazyme, P525). The in silico C. gigas STBD1 mutant was generated by overlapping PCR based on homologous sequences of fragment-1 (nucleotides 1-318) and fragment-2 (nucleotides 319-585), which resulted in 1-106aa being rearranged to 90-194aa. The purified PCR products were subcloned into pET32a vector pretreated with BamHI and HindIII (New England Biolabs) using ClonExpress® II One Step Cloning Kit (Vazyme, C112) to generate the four pET32a-STBD1 constructs. Plasmids used for overexpression of each STBD1 protein in 293 T cells were constructed in the pcDNA3.1 vector (Invitrogen). A HA-tag (TACCCATACGATGTTCCAGATTACGCT) was added at the C-terminus by PCR amplification. Then, the purified PCR products were subcloned into pcDNA3.1 vector pretreated with NheI and BamHI (New England Biolabs) to generate pcDNA3.1-HA-STBD1 expression plasmids for each of the four proteins. Plasmids used for co-IP of STBD1 from C. gigas with LC3A, LC3C, GABARAP and GABARAPL2 were also constructed in the pcDNA3.1 vector. The expression plasmid used for IP with STBD1 from C. gigas was constructed using the pcDNA3.1-HA plasmid. The LC3A, LC3C, GABARAP and GABARAPL2 expression plasmids were generated by amplifying their coding sequences with a Flag tag (GATTACAAGGACGACGATGACAAG) at N-terminus and cloning into the pcDNA3.1 vector. Primers for the construction of expression plasmids are listed in Supplementary Data 8. All plasmids were extracted using the EndoFree Midi Plasmid Kit (TIANGEN, DP118) and confirmed by Sanger sequencing.
Glycogen co-sedimentation assay
Prior to the co-sedimentation assay, the four STBD1 and pET32a vector proteins were purified using a prokaryotic expression system. Briefly, each pET32a-STBD1 plasmid, additional to a pET32a blank plasmid, was transformed into the E. coil BL21 (DE3) strain (Vazyme, C504) and cultured in LB medium containing ampicillin. The cultured bacteria were proliferated to OD = 0.6 and induced by 1 mM isopropyl β-D-thiogalactoside (IPTG, Sangon, A600168) for 4 h at 37 °C. The bacteria were centrifugated and resuspended in a binding buffer (0.1 M N2HPO4, 0.01 M NaH2PO4, 0.5 M NaCl, 8 M urea and 25 mM imidazole, pH = 7.4) before ultrasonic cracking. Lysed protein supernatant was loaded on Ni-NTA agarose gel (MDBio, HI020) and eluted by 250 mM imidazole. Finally, proteins were refolded and further purified in ultrafiltration tubes containing dialysate buffer (2 mM Reduced Glutathione, 0.2 mM Oxidized Glutathione, 0.1 M Glycine, 5% Glycerine and gradient urea [6 M-4 M-2 M-1 M-0 M]).
The co-sedimentation assay was performed as described previously71. 360 μL of 0.55 mg/mL purified STBD1 proteins were incubated in 1440 μL co-sedimentation buffer (10 mM Tris-HCl, pH 7.5, 1 mM 2-mercaptoethanol, 1 mM PMSF, 1 mM EDTA and 1 mM protease inhibitor cocktail (Beyotime, P1005) with or without 600 μL of 17.6% glycogen from cattle liver (Sigma, G0885) overnight at 4 °C. Then, 4 mL extra buffer was added, and the solution layered over 6.0 mL of 0.25 M sucrose in a 13.2 mL centrifuge tube (Beckman). After centrifugation at 100,000 g for 90 min, the supernatant and pellet were collected for Western blot. The fraction before centrifugation was used as input. pET32a vector protein was used as a negative control and treated in the same way.
Overexpression of STBD1 protein in 293 T cells and EBSS treatment
Human 293 T cells (Beyotime, C6008) were cultured in DMEM (Gibco, 11965092) supplemented with 10% FBS (Hyclone, SH30406) and 1 × penicillin/streptomycin (Sangon, E607011) with 5% CO2 at 37 °C. Before transfection, 1 × 106 cells per well were seeded into 6-well dishes (Corning), allowing for cellular proliferation until reaching 50-70% confluence. Then, 1 μg of each pcDNA3.1-HA-STBD1 expression plasmid was transfected into these cells using Lipofectamine™3000 (Invitrogen, L3000015) according to the manufacturer’s instructions. Cells were cultured for 24 h, followed by treatment with or without the autophagy inducer Earle’s balanced salt solution (EBSS, Beyotime, C0213) for 6 h. After transfection, the protein was extracted by cell lysis buffer (Beyotime, P0013) for Western blot and the glycogen content within the cells determined.
Dual-fluorescence in situ hybridization
The labial palp was used to analyze co-localization of mRNA from STBD1, LC3A, LC3C, GABARAP and GABARAPL2 of C. gigas. Briefly, the samples were fixed in 4% PFA at 4 °C, dehydrated in gradient alcohol, then embedded in paraffin and sectioned at 5 μm thickness. Digoxigenin-labeled STBD1 and biotin-labeled LC3A, LC3C, GABARAP and GABARAPL2 probes were transcribed in vitro using an RNA labeling kit (Roche, 10999644001) from the PCR products containing STBD1(nucleotides 7-418), LC3A (GenBank: LOC105342446, nucleotides 68-358), LC3C (GenBank: LOC105319833, nucleotides 57-320), GABARAP (GenBank: LOC105335545, nucleotides 71-350) and GABARAPL2 (GenBank: LOC105331926, nucleotides 42-302) fragments. Primers for synthesizing RNA probes are listed in Supplementary Data 8. Sections were deparaffinized, hydrated, and then treated with 4 μg/mL proteinase K (Roche, 03115836001) before pre-hybridization in 66% formamide and incubation with a mixture of the STBD1 probe with either LC3A, LC3C, GABARAP or GABARAPL2 probes for 24 h at 65 °C. Slides were rinsed in maleic acid buffer and blocked with 1% blocking reagent (Roche, 11096176001). Subsequently, Alexa488-conjugated anti-digoxigenin (Jackson ImmunoResearch, 200-542-156) and Alexa647-conjugated anti-biotin (Jackson ImmunoResearch, 200-602-211) diluted 1:500 with blocking reagent were added and incubated overnight at 4 °C. Nuclear differentiation was performed using DAPI staining (Beyotime, C1006) solution according to the manufacturer’s instructions. All the images were acquired with a Lecia TCS P8 laser confocal microscope.
Co-immunoprecipitation (co-IP) assay
Protein interactions of C. gigas STBD1 with LC3A, LC3C, GABARAP or GABARAPL2 were analyzed using co-IP assays. Prior to co-IP, 293 T cells were seeded at 5 × 106 cells per 10 cm dish and transfected at 50–70% confluency with the above described pcDNA3.1-HA-STBD1 and either pcDNA3.1-Flag-LC3A, pcDNA3.1-Flag-LC3C, pcDNA3.1-Flag-GABARAP or pcDNA3.1-Flag-GABARAPL2 plasmids using Lipofectamine™3000 according to the manufacturer’s instructions. Transfections were cultured for 48 h at 37 °C, with 5% CO2, before harvest. Cells were lysed using IP lysis buffer (Beyotime, P0013), before co-IP was performed using an immunoprecipitation kit with Flag-tagged protein (Beyotime, P2202). After protein quantification using a BCA protein quantification kit (Vazyme, E112), 5% lysates were used as input, and the remaining samples incubated with Flag-tagged agarose beads overnight at 4 °C. The beads were washed 3 times with lysis buffer and the protein products eluted for Western blot.
Western blot
Protein samples were denatured by boiling in 2 × SDS-PAGE loading buffer (EpiZyme, LT101) at 100 °C for 5 min. Then, proteins were separated by 12% or 15% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred to PVDF membranes (Millipore). Membranes were blocked with 5% skimmed milk dissolved by TBST buffer at 37 °C for 2 h. For the fasting and refeeding samples, anti-STBD1 rabbit pAb (AntibodySystem, PHB57101) or anti-p62 rabbit pAb (AntibodySystem, PHG56101) were added (1:2000 dilutions) and incubated overnight at 4 °C. Other primary antibodies were added with 1:1500 dilution and incubated overnight at 4 °C: anti-His mouse mAb (Beyotime, AF5060) for the glycogen co-sedimentation assay; anti-HA rabbit mAb (Beyotime, AG8057) for STBD1 overexpression experiments; anti-HA rabbit mAb and anti-Flag rabbit mAb (Beyotime, AG8050) for co-IP assays. Next, the membranes were washed 5 times with TBST and incubated with secondary antibody HRP-conjugated goat anti-mouse (Beyotime, A0216) or goat anti-rabbit (Beyotime, A0208) diluted 1:1000 by TBST for 1 h at 37 °C. β-actin mouse mAb (Beyotime, AF0003) was used as a control for the fasting and refeeding samples. GAPDH mouse mAb (Beyotime, AF5009) was used as a control for the STBD1 overexpression experiment and co-IP assays. Finally, the blots were measured with SuperPico ECL chemiluminescence kit (Vazyme, E422) and visualized using the GE ImageQuant LAS4000mini system.
Statistics and reproducibility
The Kolmogorov–Smirnov test was employed to assess the normality of the data, while the Levene test was utilized to evaluate the homogeneity of variance across groups, both of which were essential for determining statistical significance. Data were represented as mean ± standard deviation (SD). A two-tailed unpaired Student’s t-test was applied for comparisons between two groups, one-way or two-way analysis of variance (ANOVA) was employed for comparisons involving three or more groups, and Tukey’s post hoc test was used for pairwise comparisons. Statistical significance was presented below: *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. All experimental data were gathered from at least three independent experiments showing similar results, except for the co-IP assay, which was repeated twice.
Ethics statement
All animal experiment guidelines were approved by the Institutional Animal Care and Use Committee of Ocean University of China (OUC-IACUC), with approval numbers 2020-0032-0517 and 2023-0032-0039.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Caetano, R., Ispolatov, I. & Doebeli, M. Evolution of diversity in metabolic strategies. eLife 10, e67764 (2021).
Braakman, R., Follows, M. & Chisholm, S. Metabolic evolution and the self-organization of ecosystems. Proc. Natl. Acad. Sci. USA 114, E3091–E3100 (2017).
Sacristán, H. et al. Energy reserves mobilization: Strategies of three decapod species. PLoS ONE 12, e0184060 (2017).
Sánchez-Paz, A. et al. Usage of energy reserves in crustaceans during starvation: Status and future directions. Insect Biochem. Mol. Biol. 36, 241–249 (2006).
Yu, L., Chen, Y. & Tooze, S. A. Autophagy pathway: Cellular and molecular mechanisms. Autophagy 14, 207–215 (2017).
Dikic, I. & Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 19, 349–364 (2018).
Kuma, A. & Mizushima, N. Physiological role of autophagy as an intracellular recycling system: With an emphasis on nutrient metabolism. Semin. Cell Dev. Biol. 21, 683–690 (2010).
Lmd, D. et al. Glycophagy – the physiological perspective on a newly characterized glycogen-selective autophagy. Curr. Opin. Physiol. 30, 100598 (2022).
Grabner, G., Xie, H., Schweiger, M. & Zechner, R. Lipolysis: Cellular mechanisms for lipid mobilization from fat stores. Nat. Metab. 3, 1445–1465 (2021).
Moon, T. Glucose intolerance in teleost fish: Fact or fiction? Comp. Biochem. Physiol. B 129, 243–249 (2001).
Arrese, E. & Soulages, J. Insect fat body: Energy, metabolism, and regulation. Annu. Rev. Entomol. 55, 207–225 (2009).
Sheridan, M. Regulation of lipid metabolism in poikilothemic vertebrates. Comp. Biochem. Physiol. B 107, 495–508 (1994).
Wang, J. et al. Lipophagy is essential for lipid metabolism in fish. Sci. Bull. 63, 879–882 (2018).
Vargas, J. N., Hamasaki, M., Kawabata, T., Youle, R. & Yoshimori, T. The mechanisms and roles of selective autophagy in mammals. Nat. Rev. Mol. Cell Biol. 24, 1–19 (2022).
Zhang, S. et al. The regulation, function, and role of lipophagy, a form of selective autophagy, in metabolic disorders. Cell Death Dis. 13, 132 (2022).
Yamada, T., Habara, O., Kubo, H. & Nishimura, T. Fat body glycogen serves as a metabolic safeguard for the maintenance of sugar levels in Drosophila. Development 145, 158865 (2018).
Liu, S., Li, L., Wang, W., Busu, l. & Zhang, G. Characterization, fluctuation and tissue differences in nutrient content in the Pacific oyster (Crassostrea gigas) in Qingdao, northern China. Aquac. Res. 51, 1353–1364 (2020).
Berthelin, C., Kellner, K. & Mathieu, M. Histological characterization and glucose incorporation into glycogen of the Pacific oyster Crassostrea gigas storage cells. Mar. Biotechnol. 2, 136–145 (2000).
Kalachev, A. & Yurchenko, O. Autophagy in nutrient storage cells of the Pacific oyster, Crassostrea gigas. Tissue Cell 61, 30–34 (2019).
Koutsifeli, P. et al. Glycogen-autophagy: Molecular machinery and cellular mechanisms of glycophagy. J. Biol. Chem. 298, 102093 (2022).
Wu, L.-X. et al. Glycophagy mediated glucose-induced changes of hepatic glycogen metabolism via OGT1-AKT1-FOXO1Ser238 pathway. J. Nutr. Biochem. 117, 109337 (2023).
Heden, T., Chow, L., Hughey, C. & Mashek, D. Regulation and role of glycophagy in skeletal muscle energy metabolism. Autophagy 18, 1078–1089 (2021).
Jiang, S. et al. Starch binding domain-containing protein 1/genethonin 1 is a novel participant in glycogen metabolism. J. Biol. Chem. 285, 34960–34971 (2010).
Zhang, Y. et al. Decoding the molecular mechanism of selective autophagy of glycogen mediated by autophagy receptor STBD1. Proc. Natl. Acad. Sci. USA. 121, e2402817121 (2024).
Zhu, Y., Zhang, M., Kelly, A. & Cheng, A. The carbohydrate-binding domain of overexpressed STBD1 is important for its stability and protein–protein interactions. Biosci. Rep. 34, e00117 (2014).
Machovic, M. & Janecek, S. Starch-binding domains in the post-genome era. Cell. Mol. Life Sci. 63, 2710–2724 (2007).
Christiansen, C. et al. The carbohydrate-binding module family 20 - Diversity, structure, and function. FEBS J. 276, 5006–5029 (2009).
Janecek, S., Svensson, B. & MacGregor, E. Structural and evolutionary aspects of two families of non-catalytic domains present in starch and glycogen binding proteins from microbes, plants and animals. Enzym. Microb. Technol. 49, 429–440 (2011).
Kuchtova, A., Gentry, M. & Janecek, S. The unique evolution of the carbohydrate-binding module CBM20 in laforin. FEBS Lett. 592, 586–598 (2018).
Sidar, A., Voshol, G., Vijgenboom, E. & Punt, P. Novel design of an α-Amylase with an N-terminal CBM20 in Aspergillus niger improves binding and processing of a broad range of starches. Molecules 28, 5033 (2023).
Delaporte, M. et al. Impact of food availability on energy storage and defense related hemocyte parameters of the Pacific oyster Crassostera gigas during experimental reproductive cycle. Aquaculture 254, 571–582 (2006).
Yurchenko, O. & Kalachev, A. Morphology of nutrient storage cells in the gonadal area of the Pacific oyster, Crassostrea gigas (Thunberg, 1793). Tissue Cell 56, 7–13 (2019).
Malham, S. et al. Summer mortality of the Pacific oyster, Crassostrea gigas, in the Irish Sea: The influence of temperature and nutrients on health and survival. Aquaculture 287, 128–138 (2009).
Liu, W., Gao, F. & Kong, L. Effect of starvation on biochemical composition and gametogenesis in the Pacific oyster Crassostrea gigas. Fish. Sci. 76, 737–745 (2010).
Berthelin, C., Kellner, K. & Mathieu, M. Storage metabolism in the Pacific oyster (Crassostrea gigas) in relation to summer mortalities and reproductive cycle (West Coast of France). Comp. Biochem. Physiol. B 125, 359–369 (2000).
Lam, T. et al. Reversal of intramyocellular lipid accumulation by lipophagy and a p62-mediated pathway. Cell Death Discov. 2, 16061 (2016).
Noda, N., Ohsumi, Y. & Inagaki, F. Atg8-family interacting motif crucial for selective autophagy. FEBS Lett. 584, 1379–1385 (2010).
Jiang, S., Wells, C. & Roach, P. Starch-binding domain-containing protein 1 (STBD1) and glycogen metabolism: Identification of the Atg8 family interacting motif (AIM) in Stbd1 required for interaction with GABARAPL1. Biochem. Biophys. Res. Commun. 413, 420–425 (2011).
Hess, P., De, C. & Russo, C. An empirical test of the midpoint rooting method. Biol. J. Linn. Soc. 92, 669–674 (2007).
Ceperuelo-Mallafré, V. et al. Adipose tissue glycogen accumulation is associated with obesity-linked inflammation in humans. Mol Metab. 5, 5–18 (2015).
Hofmann, A., Wurm, F. & Frey, H. Rapid access to polyfunctional lipids with complex architecture via oxyanionic ring-opening polymerization. Macromolecules 44, 4648–4657 (2011).
Gáliková, M. & Klepsatel, P. Endocrine control of glycogen and triacylglycerol breakdown in the fly model. Semin. Cell Dev. Biol. 138, 104–116 (2022).
Koyama, M. et al. Gluconeogenesis and glycogen metabolism during development of Pacific abalone, Haliotis discus hannai. Am. J. Physiol. Regul. Integr. Comp. Physiol. 318, R619–R633 (2020).
Gusarov, I. & Nudler, E. Glycogen at the crossroad of stress resistance, energy maintenance, and pathophysiology of aging. Bioessays 40, e1800033 (2018).
Mawed, S., Zhang, J., Ren, F., He, Y. & Mei, J. Atg7 and beclin1 are essential for energy metabolism and survival during the larval-to-juvenile transition stage of zebrafish. Aquac. Fish. 7, 359–372 (2021).
He, C. Balancing nutrient and energy demand and supply via autophagy. Curr. Biol. 32, R684–R696 (2022).
Tang, Q., Liu, M., Zhao, H. & Chen, L. Glycogen-binding protein STBD1: Molecule and role in pathophysiology. J. Cell. Physiol. 238, 2010–2025 (2023).
Bjørkøy, G. et al. Chapter 12 monitoring autophagic degradation of p62/SQSTM1. Methods Enzymol. 452, 181–197 (2009).
Zhao, X. et al. Functional teas from penthorum chinense pursh alleviates ethanol-induced hepatic oxidative stress and autophagy impairment in zebrafish via modulating the AMPK/p62/Nrf2/mTOR signaling axis. Plant Foods Hum. Nutr. 77, 514–520 (2022).
Jia, Z. et al. Hepatotoxicity assessment of Rhizoma Paridis in adult zebrafish through proteomes and metabolome. Biomed. Pharmacother. 121, 109558 (2019).
Johansen, T. & Lamark, T. Selective autophagy: ATG8 family proteins, LIR motifs and cargo receptors. J. Mol. Biol. 432, 80–103 (2019).
Rogov, V. et al. Structural and functional analysis of the GABARAP interaction motif (GIM). EMBO Rep. 19, e47268 (2018).
Varga, V., Keresztes, F., Sigmond, T., Vellai, T. & Kovács, T. The evolutionary and functional divergence of the Atg8 autophagy protein superfamily. Biol. futura 73, 375–384 (2022).
Lytridou, A. et al. STBD1 promotes glycogen clustering during endoplasmic reticulum stress and supports survival of mouse myoblasts. J. Cell Sci. 133, jcs244855 (2020).
Demetriadou, A. et al. Mouse Stbd1 is N -myristoylated and affects ER-mitochondria association and mitochondrial morphology. J. Cell Sci. 130, jcs.195263 (2017).
Suen, S., Lu, H. & Yeang, C.-H. Evolution of domain architectures and catalytic functions of enzymes in metabolic systems. Genome Biol. Evol. 4, 852–869 (2012).
Jensen, E. Quantitative analysis of histological staining and fluorescence using ImageJ. Anat. Rec. 296, 378–381 (2013).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Mooers, B. Shortcuts for faster image creation in PyMOL. Protein Sci. 29, 268–276 (2019).
Nivedha, A., Thieker, D., Hu, H. & Woods, R. Vina-Carb: Improving glycosidic angles during carbohydrate docking. J. Chem. Theory Comput. 12, 892–901 (2016).
McGuire, R. et al. 3D-e-Chem-VM: Structural cheminformatics research infrastructure in a freely available virtual machine. J. Chem. Inf. Model. 57, 115–121 (2017).
Pronk, S. et al. GROMACS 4.5: A high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013).
Case, D. et al. AmberTools. J. Chem. Inf. Model. 63, 6183–6191 (2023).
Silva, A. & Vranken, W. ACPYPE-Antechamber python parser interface. BMC Res. Notes 5, 367 (2012).
Wang, C. et al. Calculating protein–ligand binding affinities with MMPBSA: Method and error analysis. J. Comput. Chem. 37, 2436–2446 (2016).
Eddy, S. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Capella-Gutierrez, S., Silla-Martínez, J. & Gabaldón, T. TrimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Minh, B. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B., Wong, T., von Haeseler, A. & Jermiin, L. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Kerekes, E., Kokai, E., Páldy, F. & Dombrádi, V. Functional analysis of the glycogen binding subunit CG9238/Gbs-70E of protein phosphatase 1 in Drosophila melanogaster. Insect Biochem. Mol. Biol. 49, 70–79 (2014).
Acknowledgements
We acknowledge the support of the High-Performance Biological Supercomputing Center at the Ocean University of China, and the Marine Biodiversity and Evolution Research Institute’s Instrument and Equipment Sharing Platform for providing the high-speed centrifuge (XPN-100) for this research. This work was supported by grants from the National Natural Science Foundation of China (32341060 and 42276112), the Key Research and Development Program of Shandong Province (2021ZLGX03), the National Key Research and Development Program of China (2022YFD2400300), the Fundamental Research Funds for the Central Universities (202461040), and the earmarked fund for the Agriculture Research System of China (CARS-49). DJM received support from an Institute Strategic Programme award (BBS/E/RL/230001 C) from the Biotechnology and Biological Sciences Research Council to The Roslin Institute.
Author information
Authors and Affiliations
Contributions
Liting Ren: Methodology, Investigation, Formal analysis, Data curation, Writing–original draft. Yitian Bai: Investigation, Data curation and Writing–original draft. Chenyu Shi and Zhengwang Hao: Investigation. Qi Li: Supervision and Resources. Daniel J Macqueen: Writing–review & editing, Funding acquisition and Supervision. Shikai Liu: Writing–review & editing, Supervision, Funding acquisition, Formal analysis, Conceptualization.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Bernard Degnan, Fabian Ruperti, Łukasz Sobala and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Joao Valente. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ren, L., Bai, Y., Shi, C. et al. Glycophagy is an ancient bilaterian pathway supporting metabolic adaptation through STBD1 structural evolution. Commun Biol 9, 268 (2026). https://doi.org/10.1038/s42003-026-09546-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09546-6