Abstract
Converging evidence suggests that understanding the human brain requires more than just examining pairwise functional brain interactions. The human brain is a complex, nonlinear system, and focusing solely on linear pairwise functional connectivity often overlooks important nonlinear and higher-order interactions. Infancy is a critical period marked by significant brain development that could contribute to future learning, health, and life success. Exploring higher-order functional relationships in the brain can provide insight into brain function and development. To the best of our knowledge, there is no existing research on multiway, multiscale brain network interactions in infants. In this study, we comprehensively investigate the interactions among brain intrinsic connectivity networks (ICNs), including both pairwise (pair-FNC) and triple relationships (tri-FNC). We focused on a dataset of typically developing infants scanned during the first six months of life-a critical period for brain maturation. In total, 71 infants (aged 4-179 days) contributed 126 scans (76 from 41 males, 50 from 30 females). Our results revealed significant hierarchical, multiway, multiscale brain functional network interactions in the infant brain. These findings suggest that tri-FNC provide additional insights beyond what pairwise interactions reveal during early brain development. The tri-FNC predominantly involve the default mode, sensorimotor, visual, limbic, language, salience, and central executive domains. Notably, these triplet networks align with the classical triple network model of the human brain, which includes the default mode network, the salience network, and the central executive network. This suggests that the brain network system might already be initially established during the first six months of infancy. We also found that pair-FNC were less effective at detecting these networks. The present study suggests that exploring tri-FNC can offer additional insights beyond pair-FNC by capturing higher-order nonlinear interactions, potentially yielding more reliable biomarkers to characterize developmental trajectories.

Similar content being viewed by others
Introduction
The human brain is organized in a hierarchical, multiscale manner that facilitates efficient information processing, which is essential for maintaining functional brain network interactions1,2,3,4. These interactions reflect the core principles that allow brain networks to specialize for specific functions. These principles involve mechanisms that coordinate brain activity across different scales of organization5. During infancy, this developmental period is characterized by the discovery of the underlying rules that govern network interactions. By examining multiscale, multiway interactions during this phase, we can gain valuable insights into how these mechanisms drive the emergence of higher-order interactions and the specialization of brain networks. Notably, this intricate process of network specialization and integration is particularly pronounced during the early developmental stage6. Specifically, the first six months represent a critical period when the brain undergoes substantial growth and reorganization, laying the foundation for the complex network interactions observed in later stages of life7,8,9. A significant aspect of this developmental phase involves uncovering the rules that govern network interactions within functional connectomes10. Gaining insight into how these processes evolve during the perinatal and early postnatal periods is essential for understanding the mechanisms behind both typical and atypical brain developments7,8,11,12.
Previous research using resting-state functional magnetic resonance imaging (rsfMRI) has shown that changes in functional connectivity during early human development are mainly driven by the maturation of primary brain systems, with relatively less impact on higher-level regions such as the default mode, salience, frontoparietal, and executive control networks13,14,15,16. One main reason for this is that by the end of the first year, primary sensorimotor and auditory networks have developed to closely resemble their adult counterparts, whereas higher-level networks, which undergo considerable development and continue to develop17. Studies suggest that whole-brain connectivity efficiency improves significantly by one year and reaches a stable level by age 215,18. After this period, brain development is thought to be characterized mainly by the reorganization and remodeling of established major functional networks, including the refining and optimizing of neural connections and the shaping of functional brain architecture19. Furthermore, by two years of age, previous studies suggest that the default network begins to show some resemblance to that observed in adults, involving regions such as the medial prefrontal cortex, the posterior cingulate cortex/retrosplenial, the inferior parietal lobule, the lateral temporal cortex, and the hippocampus16,18. Furthermore, to explore brain network interactions during early brain development, functional connectivity is widely used to estimate these interactions. Pairwise metrics, such as Pearson correlation and mutual information, are frequently applied to assess both linear and nonlinear pairwise functional connectivity20,21,22. However, brain network interactions extend beyond pairwise connections, involving multiway interactions that these traditional metrics are unable to capture20,23,24,25,26,27.
Consequently, previous research has often faced limitations in both scope and methodology due to a reliance on broad age ranges and pairwise interactions derived from predefined brain atlases to investigate lifespan trajectories. Although data-driven independent component analysis (ICA) has been utilized in lifespan studies, these investigations have typically overlooked the examination of multiscale interactions. As a result, these studies did not provide a comprehensive and precise view of developmental trajectories during the crucial first six months of infant brain development. Additionally, the role of multiway network interactions in connectome growth during this critical period has yet to be studied, as pairwise interactions may overlook significant high-order functional connections in the brain28,29,30,31,32,33.
To address these critical knowledge and technique gaps, this study explored the continuous developmental trajectory of multiway, multiscale network interactions over the first 180 days and examined its relationship with infant age. Firstly, we utilized high-quality longitudinal neuroimaging data, including 126 scans from 71 typically developing infants who underwent resting-state functional MRI at various ages, ranging from birth to six months. Secondly, we employed multi-scale intrinsic connectivity networks (ICNs) derived from these infants, building upon recent foundational work that identified 105 multi-scale ICNs in infants aged 0 to 6 months34. These networks were processed using a group multi-scale ICA approach incorporating distinct model orders31. Higher model orders typically provide finer spatial resolution, whereas lower model orders capture integrated features across larger brain regions. By leveraging data across multiple scales, our methodology enables the modeling of a diverse range of intrinsic connectivity networks. Additionally, to estimate multiway interactions, we used total correlation35, an information-theoretic metric designed to capture beyond pairwise interactions. Previous studies have shown that total correlation provides more information than pairwise metrics and can enhance the diagnosis of brain disorders23,28,29,30.
To comprehensively map the developmental trajectory and capture higher-order functional connections, we assessed both pairwise and triple interactions, along with their respective subspaces, and examined their correlation with infant age to identify key networks. Including both interaction types is crucial, as pairwise interactions alone may not fully capture the complexity of brain connectivity, particularly during early development when networks are rapidly evolving. By investigating both pairwise and triple interactions and their subspace, we are able to reveal a more complete and nuanced understanding of the brain’s functional architecture. This dual approach enables us to identify not only direct interactions between brain networks but also the higher-order relationships that are critical for understanding the complex dynamics of early brain development.
Results
Estimated subject-specific ICNs and their time courses using infant rsfMRI
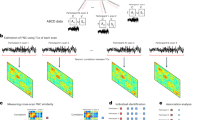
To estimate subject-specific ICNs and their time courses, we applied the spatially constrained ICA method, Multivariate-Objective Optimization (MOO-ICAR)31,36, using the NeuroMark2.1 Template as a reference to obtain subject-specific multi-scale ICN spatial maps and their time courses in infants. Finally, we categorized 105 multi-scale ICNs into the following domains: visual (VI, 12 ICNs, includes visual network, VIS; ventral attention network, VAN), cerebellar (CB, 13 ICNs), temporal (TP/TMN, 13 ICNs, includes paralimbic network, PLN; explicit memory network, EM), subcortical (SC, 23 ICNs, includes limbic network, LIM; explicit memory network, EM; language network, LAN), somatomotor (SM, 13 ICNs), and higher cognitive (HC, with 31 ICNs, includes the default mode network, DMN; language network, LAN; central executive network, CEN; salience network, SN; and dorsal attention network, DAN), as shown in Fig. 1A.
The preprocessed infant resting-state fMRI data were input into Multivariate Objective Optimization ICA with Reference (MOO-ICAR) using the spatially constrained NeuroMark2.1 Template, derived from over 100K subjects31. This process yielded subject-specific estimates of 105 intrinsic connectivity networks (ICNs) and their corresponding time courses in infant rsfMRI. These 105 ICNs were then grouped into six brain domains: visual (VI), cerebellar (CB), temporal (TP/TMN), subcortical (SC), somatomotor (SM), and higher cognitive (HC), as depicted in A. As we know, the network in the human brain is not isolated; interactions occur both pairwise and beyond pairwise for information exchange. In this study, we estimated pairwise (interaction order = 2) and triple interactions (interaction order = 3) among intrinsic connectivity networks (ICNs), shedding light on multiway interactions in the infant brain, as shown in B. We first estimated the pairwise interactions for each subject and then concatenated these pairwise interactions (105 × 105) across subjects (F × Subjs, where F represents the total number of features). For triple interactions, we flattened the 3D interaction tensors (105 × 105 × 105) after estimating them and then concatenated them across subjects. Due to the high dimensionality of triple interactions and to explore the latent space underlying these complex pairwise and triple interactions, we applied group independent component analysis (GICA) to decompose the intricate patterns of both pairwise and triple interaction components. Subsequent analyses then focused on exploring the relationship between age and the decomposed brain network interaction components, as illustrated in C.
Estimated triple and pairwise interactions from infant rsfMRI
Here, we estimated triple and pairwise interactions between specific ICNs using total correlation (for interaction order 3, i.e., TC(ICNx, ICNy, ICNz)), mutual information (for interaction order 2, i.e., MI(ICNx, ICNy)), and Pearson correlation (i.e., PC(ICNx, ICNy)), respectively, as illustrated in Fig. 1B. In total, there are 1053 possible triple interactions when considering all combinations, including \(\left(\begin{array}{l}105\\ 3\end{array}\right)\)=187460 unique sets of triple interactions. Similarly, there are 1052 possible pairwise interactions, which include \(\left(\begin{array}{l}105\\ 2\end{array}\right)\)=5460 unique sets of pairwise interactions. The estimated pairwise interactions for each subject are represented as 2D matrices, while triple interactions are represented as 3D tensors, as shown in Fig. 1C.
Triple and pairwise functional connectivity in the infant brain and their associations with age
We investigate the relationship between brain development and age by examining the association between triple and pairwise functional connectivity and age. Specifically, we analyze the correlation between age and z-scored functional connectivity for both triple (flattened from a 3D tensor and organized based on network community structure) and pairwise interactions is shown on the left side of Fig. 2A,B. Additionally, networks with triple and pairwise interactions showing significant associations with age were identified (p < 0.05, FDR corrected)), as presented on the right side of Fig. 2A,B. Triple functional connectivity analysis identified 80,709 significant networks, while pairwise functional connectivity identified 1506 (MI) and 1465 (PC) networks, indicating a strong association between functional connectivity and participants’ age. In addition, we observed a cluster pattern along the diagonal in triple functional connectivity, with a similar pattern present in pairwise connectivity.
In A, the Pearson correlation was calculated between each z-scored triple functional connectivity (TRI-FNC) matrix, obtained by flattening the 3D triple interaction tensors, and the age of the subjects, with the corresponding functional domains indicated on the left side. Significant networks associated with age were identified based on the correlation coefficients. Significance was determined using p-values corrected for false discovery rate (FDR) at an alpha level of 0.05, which are presented on the right side. A comparable analysis was carried out for MI and PC, employing the same scale (z-scored) to ensure consistency and enable direct comparison, as shown in B.
Triple interactions capture different brain development information than pairwise interactions
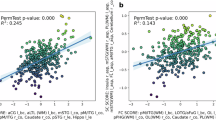
The information about brain development derived from triple interactions differs from that obtained through pairwise interactions alone, as illustrated in Fig. 3A. To explore these differences, we analyzed the highest positive and negative triple correlation networks with age by breaking them down into pairwise interactions. First, we observed that triple interaction networks show substantial positive and negative correlations with age (ρ = 0.650 vs. ρ = −0.549, p < 0.001, Bonferroni corrected), as presented in Fig. 3B. We decomposed the triple interaction into pairwise interactions and then evaluated the average combined pairwise interactions (\(\frac{[IC94+IC102]+[IC90+IC102]+[IC94+IC90]}{3}\) vs. \(\frac{[IC89+IC102]+[IC68+IC102]+[IC89+IC68]}{3}\)) for MI and PC separately. We observed that the average combined pairwise interactions generally exhibit weak negative associations with age (ρ = −0.067 vs. ρ = −0.186 for MI, and ρ = −0.276 vs. ρ = −0.127 for PC). These results, as illustrated on the left side of Fig. 3C, highlight the differing patterns between the two measures. Meanwhile, we also examined pairwise interactions (IC94 vs. IC102; IC94 vs. IC90; IC102 vs. IC90 in the highest positive triple interaction networks, and IC89 vs. IC102; IC89 vs. IC68; IC102 vs. IC68 in the highest negative triple interaction networks) separately using MI and found similar results (ρ = −0.125 vs. ρ = −0.107; ρ = 0.072 vs. ρ = −0.051; ρ = −0.130 vs. ρ = −0.315). In summary, we demonstrated that triple interactions capture different brain development network interactions than pairwise interactions, and that pairwise interactions alone may miss some information that is captured in triple interactions.
The triple interaction (e.g., [A–C]) can be broken down into pairwise interactions ([A, B], [A, C], [B, C]) or their combined average. However, these decomposing pairwise interactions are not equivalent to the original triple interaction, as illustrated in A. To further illustrate this distinction, we examined how these interactions relate to brain development. B shows the strongest positive and negative triple network interactions across different infant ages. In C, we compare these interactions by presenting the associations derived from both the combined average of pairwise interactions and purely pairwise interactions, which are extracted from the z-score pairwise functional connectivity matrix. This comparison highlights the additional brain development information provided by triple interactions that is not captured by pairwise interactions alone. The colored dots represent age information, with colors from left to right corresponding to ages from 0 to 6 months.
Identified latent network connectivity subspaces in triple and pairwise interactions from ICA
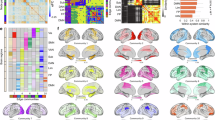
By employing ICA with a model order of 20, selected based on the elbow criterion, we successfully separated the complex pairwise and triple interaction patterns into 20 distinct components, as illustrated in Fig. 4A. This decomposition is essential for gaining a deeper understanding of the independent structures within infant brain networks. Examining both the individual components and the interactions themselves allows us to explore their contributions to brain function. Moreover, this approach enables us to extract latent space information, enriching our analysis of the dynamic connectivity in infant brains.
In panel A, the triple and pairwise subspace latent interactions are identified, with the first and second strongest interactions for each component displayed from left to right. Interactions estimated from TC, MI, and PC are depicted in green, cyan, and blue, respectively. The indices correspond to specific ICNs listed on the left. The reliability of ICA for both pairwise and triple interactions was assessed by analyzing spatial similarities and stability values (IQ) from 20 triple components (TRI-C) obtained through ICA after 20 repetitions. For comparison, similar analyses were performed for pairwise interactions using mutual information and Pearson correlation. The resulting similarities and corresponding IQ values for these 20 components are also presented, as shown in panel B. Furthermore, the strongest triple interactions were identified in components 2, 6, and 8, while the weakest interactions were found in component 18. Components 13 and 15 were selected for pairwise interactions, as shown in panel C. Additionally, pairwise interactions within triple interactions are also presented, with circle and line thickness indicating interaction strength. The association between age and the selected triple and pairwise components is presented, along with the corresponding Akaike Information Criterion (AIC) values to quantify the fitting performance (a lower value means better performance). The strongest shared and distinct weighted brain domains are displayed in panel D. The most shared domains across both interaction types include default mode (DMN), visual (VIS), somatomotor (SM), cerebellar (CB), subcortical (SC), and central executive domain(CEN). Notably, one domain (language, LAN) was detected by both TC and MI, and another domain (salience, SN) was detected exclusively through TC, but neither was observed in pairwise interactions.
To ensure that the components produced by ICA are stable and reliable, we evaluated the reliability of both triple (TRI-C) and pairwise (PA-C: PC and MI) independent components. We report the related spatial similarities for these components (TRI-C vs. PA-C) and their stability values (IQ) (TRI-IQ vs. PA-IQ) in Fig. 4B. The results indicate that all triple and pairwise components exhibit high spatial similarities across the 20 independent components, demonstrating stable reliability of ICA, with triple components generally performing better than pairwise components.
Latent triple and pairwise network connectivity subspaces in the infant brain and their associations with age
The ICA resulted in the decomposition of both the triple and pairwise interaction tensors into 20 distinct latent triple (TRI-C) and pairwise (PA-C) network connectivity subspaces, respectively. These components represent the most independent patterns of triple and pairwise interactions observed in the infant brain. The isolated 20 components for both triple and pairwise interactions were visually represented in the graph shown in Fig. 4A, and we observed that hierarchical multiway network interactions occurred during infant development, involving multiple brain networks. The separate components of triple and pairwise interaction are illustrated in Supplementary Fig. S1 and Fig. S4, respectively.
To thoroughly investigate the decomposed latent TRI-C and PA-C network connectivity subspaces in infant brain development, we analyzed several of the strongest and weakest components, focusing on their distinct patterns and how these patterns relate to developmental trajectories across different infant ages. Specifically, we focused on TRI-C2, TRI-C6, TRI-C8, and TRI-C13, which represent the strongest triple interactions, and TRI-C18, which represents the weakest triple interaction. Additionally, we analyzed PA-C13 and PA-C15 to identify the most significant and weakest pairwise interactions. All TRI-Cs and PA-Cs spatial maps are presented in Supplementary Fig. S2 and S5, respectively.
In the analysis of the strongest triple interactions in TRI-C2, we examined both the components and interactions to gain a comprehensive understanding of early brain development. The components reveal independent connectivity patterns, while the interactions show how these patterns combine to support brain function, capturing the complexity of brain network organization. Our analysis identified that the ICNs with the highest contributions-ICN102, ICN43, and ICN89-correspond to the SM, SM, and TP domains, respectively (see Fig. 4C). This suggests that the strongest interactions within this component are primarily driven by the SM and TP domains, with the other ICNs contributing less to the component. Furthermore, we noted stronger pairwise interactions between ICN102 and ICN43 compared to other pairwise interactions, likely due to both ICs being part of the SM domains, where intra-network interactions are generally more robust than inter-network interactions.
In TRI-C6, we observed that the LAN (IC57) is involved in triple interactions with the SM (IC70) and LIM (IC54). Similar interactions are also observed in TRI-C8 (SM-LAN-EM) and TRI-C18 (VIS-VIS-CEN). Furthermore, in PA-C13, we identified the strongest pairwise interactions between the DMN (IC2 and IC83) and LAN (IC9) and DAN (IC52). Additionally, in PA-C15, the weakest interactions were observed between the LAN (IC82) and SN (IC85), as well as between the LAN and LIM (IC97) and CB (IC91).
To further investigate the relationship between decomposing TRI-Cs and PA-Cs and infant age, we applied a generalized additive model (GAM). The results of this analysis are presented in Fig. 4C. Additionally, to quantify the model fit performance, we reported the Akaike Information Criterion (AIC) value, which indicated that TRI-Cs provide a better fit than PA-Cs. The relationships between all TRI-Cs and PA-Cs with infant age were presented in Supplementary Fig. S3, Figure S6, and Figure S7, respectively.
Based on the hierarchical triple and pairwise interactions identified in Fig. 4A, we observed the engagement of multiple ICNs. To determine which brain domains significantly contribute to infant brain development, we identified several commonly shared brain domains across both triple (by TC) and pairwise interactions (by MI and PC), including the DMN, VIS, SM, CB, LIM, and CEN, as shown in Fig. 4D. This analysis indicates that most of these domains emerge within the first six months of life. Additionally, we noted that the LAN was detected only by TC and MI, but not by PC. One major reason for this is that PC may not capture some nonlinear interactions effectively. Furthermore, MI was also unable to detect the SN compared to TC, as illustrated in Fig. 4D. In summary, triple interactions reveal certain domains in infants that are not captured by pairwise interactions, highlighting the added value of analyzing hierarchical triple interactions for a more comprehensive understanding of early brain development.
Discussion
Investigating multiway multiscale interactions in extensive brain networks during the first 180 days of life provides crucial insights into early brain development and potential early markers for neurodevelopmental disorders. This research, which includes 71 infants within their first 180 days, employs spatially constrained ICA to derive multiscale infant brain networks and then examines multiway interactions within these networks. Pairwise and triple interactions are estimated using PC, MI, and triple interaction models, with their associations to infant age assessed. Additionally, the subspace latent triple and pairwise interactions and their associations with age are explored. The findings from this work are significant, as they lay the groundwork for a deeper understanding of the brain’s early functional organization and its developmental trajectory.
Our studies revealed that multiway network interactions occur during infant brain development. These brain domains encompass high-level cognitive processes as well as visual, attention, and language networks. In our study, we also found several key domains captured by all metrics in the infant brain during the first six months, including the DMN, VIS, SM, CB, LIM, and CEN. Additionally, we discovered that one domain, LAN, was detectable through TC and MI but not through PC. This underscores the importance of considering nonlinearity when estimating network interactions. Furthermore, we found that another domain, SN, was detectable only through TC and not through MI or PC. This highlights the need to estimate high-order network interactions when studying brain function, as pairwise interactions alone do not capture these more complex relationships.
Moreover, after applying ICA to both triple and pairwise functional connectivity, we identified latent network connectivity subspaces within these interactions. We then explored their correlation with age, and our results also demonstrate that the triple interaction networks provide a better explanation for age-related changes than the pairwise interactions. This indicates that triple interactions provide richer information than pairwise interactions, which is valuable for capturing more comprehensive information to deepen our understanding of changes in brain network interactions during early infant brain development.
Taken together, the analysis of multiway interactions in infant brain connectivity, while offering valuable insights into intricate network interactions, is inherently limited by several factors that reflect the broader challenges of studying multi-order interactions in the infant brain.
One major limitation is the computational complexity associated with triple interactions22,24,28. The complexity of analyzing triple interactions is markedly higher than that of pairwise interactions, leading to increased computational demands. As the number of possible interactions grows combinatorially with the number of ICNs, this can strain computational resources and extend analysis time. The interpretation of such interactions is inherently complex; unraveling the contributions of each individual component and understanding their joint effects on infant brain function is often challenging. Unlike pairwise interactions, which provide a relatively straightforward measure of connectivity between ICNs, triple interactions involve understanding how three ICNs influence each other simultaneously. This complexity can obscure the underlying mechanisms of brain function and make it difficult to pinpoint the exact nature of the interactions28. In addition, in our case, the relatively small sample size increases the likelihood of inflated or unstable effect size estimates, particularly for specific triplets. This limitation underscores the need for cautious interpretation. Moreover, the biological relevance of these interactions is not always clear37; Overall, while high-order interaction analyses hold promise for advancing our understanding of brain connectivity, balancing computational demands with the need for clear biological interpretation remains a key challenge. Additionally, triple interaction analyses are highly affected by data quality issues like noise and inaccuracies in preprocessing, which can significantly impact the results37,38,39.
Another critical consideration is the nature of interactions beyond triple interactions26,28,29,37. The brain’s connectivity is not limited to simple triplets but involves a complex interplay of interactions at multiple levels. Higher-order interactions, such as quadruple or quintuple interactions, and the mixing of different types of interactions (e.g., linear and nonlinear) further complicate the analytical landscape. The interplay between various interaction orders and their combined effects can create a tangled web of connectivity that is challenging to untangle and interpret. Furthermore, we will also face computing and memory challenges as we use higher-order interactions, which will require significant computational resources.
In addition, in this study, we utilized TC to estimate triple interactions in infant brain connectivity, which provides valuable insights into the intricate dependencies among three ICNs. TC measures the overall dependence among three variables by quantifying how much the joint distribution deviates from the product of the marginal distributions. However, there are also other methods to estimate high-order interactions, such as hypergraphs40, persistent diagrams41, dual total correlation42, and O/S information43. Each of these metrics offers a different approach to capturing high-order interactions, and exploring these methods in future research could provide additional valuable insights. Specifically, O/S information is noteworthy because it can quantify the balance between redundancy and synergy in the human brain, and it is calculated from TC and dual total correlation43.
In this study, we focused on analyzing triple interactions to understand connectivity patterns within the infant brain. However, it is essential to recognize that these interactions are inherently dynamic rather than static, which has significant implications for interpreting our findings and understanding brain connectivity28,37,44,45,46. By examining how interactions between infant brain regions change over time, dynamic functional connectivity methods can reveal transient states and fluctuations that are not apparent in static models47,48. Therefore, integrating dynamic approaches into the study of high-order interactions is crucial for a more comprehensive understanding of brain connectivity. Additionally, exploring how these dynamic interactions relate to cognitive processes and behaviors will enhance our understanding of the functional network dynamics in the infant brain.
In summary, while analyzing triple interactions provides valuable insights into the development of infant brain connectivity beyond what is revealed by pairwise interactions, it also presents significant challenges that mirror the broader difficulties of studying complex multi-order interactions. Overcoming these challenges through rigorous statistical methods and exploring alternative approaches is essential for deepening our understanding of brain development and its impact on cognitive and behavioral growth.
Methods
Infancy data
Infant scans were conducted at Emory University’s Center for Systems Imaging Core using either a 3T Siemens Tim Trio or a 3T Siemens Prisma scanner, each equipped with a 32-channel head coil49,50. To ensure optimal conditions, all scans were performed while the infants were in a natural sleep state. We collected longitudinal resting-state functional magnetic resonance imaging (rsfMRI) data from 71 typically developing infants at up to 3 pseudorandom time points between birth and 6 months, resulting in a total of 126 scans, as depicted in Fig. 5. The participants comprised 76 scans from 41 male subjects and 50 scans from 30 female infants aged from 4 to 179 days (mean corrected age ± Std. is 86.52 ± 48.05 in days), covering the period from birth to approximately 6 months12, as shown in Table 1. Informed written consent was obtained from the parents of all participants before their participation, and ethical approval for all study procedures was granted by Emory University.
Each dot represents a single participant scan, with longitudinal scans connected by lines (black: first scan; blue: second scan; magenta: third scan).
Data acquisition
The fMRI data were acquired using two different scanner setups, both employing a multiband echo-planar imaging (EPI) sequence to enhance acquisition quality.
The fMRI was collected using a Siemens Tim Trio scanner. The acquisition parameters for this setup were a repetition time (TR) of 720 ms, an echo time (TE) of 33 ms, and a flip angle of 53∘. The multiband factor was set to 6, with a field-of-view (FOV) of 208 × 208 mm and an image matrix of 84 × 84, resulting in a spatial resolution of 2.5 mm isotropic. To cover the entire brain, 48 axial slices were acquired. To improve the signal-to-noise ratio (SNR), 6 dummy scans were performed prior to the actual data acquisition to ensure steady-state magnetization.
The fMRI was acquired using a Siemens Prisma scanner. The parameters for this acquisition included a repetition time (TR) of 800 ms, an echo time (TE) of 37 ms, and a flip angle of 52∘. The multiband factor was increased to 8, with the field-of-view (FOV) maintained at 208 × 208 mm and an image matrix of 104 × 104, providing a spatial resolution of 2 mm isotropic. This setup included 72 axial slices to cover the entire brain. Similar to the previous setup, 6 dummy scans were conducted before the actual data acquisition to ensure steady-state and enhance the SNR.
Data preprocessing
We performed standard NeuroMark preprocessing using the FMRIB Software Library (FSL v6.0, https://fsl.fmrib.ox.ac.uk/fsl/fslwiki/) and the Statistical Parametric Mapping (SPM12, http://www.fil.ion.ucl.ac.uk/spm/) toolboxes within the MATLAB 2020b environment. Initially, we discarded the first ten dummy scans showing significant signal changes. Subsequently, we corrected image distortions using field maps and SBRef data with reverse phase encoding blips, resulting in pairs of images with distortions in opposite directions. The applytopup tool was used to apply the field map coefficients and correct distortion in the fMRI data. After distortion correction, we performed slice timing to address slice-dependent delays using SPM. Following slice timing, we conducted head motion correction and realigned all scans to the reference scan.
Given the unique characteristics of infant fMRI data compared to adults, we implemented a two-step normalization process to align the infant data to a standardized Montreal Neurological Institute (MNI) space. Initially, we acquired the UNC-BCP 4D Infant Brain Template51 (https://www.nitrc.org/projects/uncbcp_4d_atlas/) and registered it to the UNC-BCP template space. Subsequently, we normalized the infant data to the common MNI space using the EPI template, specifically selecting the template corresponding to 3 months of age, which represents the median age of our infant dataset. Finally, the normalized fMRI data underwent spatial smoothing using a Gaussian kernel with a full width at half maximum (FWHM) of 6 mm.
Quality control
To minimize the impact of motion on our dataset, we employed several quality control (QC) strategies throughout the preprocessing and analysis stages. First, head motion was corrected using FSL realignment, which accounted for translations and rotations. We also implemented rigorous NeuroMark spatial quality control to assess and exclude scans with poor spatial normalization, based on group mask similarity. Rather than excluding scans with excessive motion, we retained all scans and included framewise displacement (FD) as a covariate in our statistical models to control for motion-related confounds. This approach preserved statistical power, maintained the original data distribution, and minimized the bias associated with excluding high-motion participants. Our FD metric follows the commonly used L1-norm approach, which sums the absolute head displacement across six motion parameters (translations and rotations). Among the QC-passed scans (FD<0.5 mm), we observed a moderate, though non-significant, correlation (R=-0.4) between FD and age, with FD tending to be higher in younger individuals.
Statistics and reproducibility
Spatially constrained ICA on infancy rsfMRI
A spatially constrained ICA (scICA) method known as Multivariate Objective Optimization ICA with Reference (MOO-ICAR) was implemented using the GIFT software toolbox (http://trendscenter.org/software/gift)46. The MOO-ICAR framework estimates subject-level independent components (ICs) using existing network templates as spatial guides44,45,46,52,53,54. Its main advantage is ensuring consistent correspondence between estimated ICs across subjects. Additionally, the scICA framework allows for customization of the network template used as a spatial reference in the ICA decomposition. This flexibility enables either disease-specific network analyses or more generalized assessments of well-established functional networks suitable for diverse populations31,32,52,55,56,57.
The MOO-ICAR algorithm, which implements scICA, optimizes two objective functions: one to enhance the overall independence of the networks and another to improve the alignment of each subject-specific network with its corresponding template52. Both objective functions, \(J({S}_{l}^{k})F({S}_{l}^{k})\), are listed in the following equation, which summarizes how the lth network can be estimated for the kth subject using the network template Sl as guidance:
Here \({S}_{l}^{k}={({w}_{l}^{k})}^{T}\cdot {X}^{k}\) represents the estimated lth network of the kth, Xk is the whitened fMRI data matrix of the kth subject and \({w}_{l}^{k}\) is the unmixing column vector, to be solved in the optimization functions. The function \(J({S}_{l}^{k})\) serves to optimize the independence of \({S}_{l}^{k}\) via negentropy. The v is a Gaussian variable with mean zero and unit variance G(. ) is a nonquadratic function, and E[. ] denotes the expectation of the variable. The function \(F({S}_{l}^{k})\) serves to optimize the correspondence between the template network Sl and subject network \({S}_{l}^{k}\). The optimization problem is addressed by combining the two objective functions through a linear weighted sum, with each weight set to 0.5. Using scICA with MOO-ICAR on each scan yields subject-specific ICNs for each of the N network templates, along with their associated time courses.
In this study, we used the NeuroMark_fMRI_2.1 template (available for download at https://trendscenter.org/data/) along with the MOO-ICAR framework for scICA on infant rsfMRI data. It enabled us to extract subject-specific ICNs and their associated time courses. This template includes N = 105 high-fidelity ICNs identified and reliably replicated across datasets with over 100K subjects31. These ICNs are organized into six major functional domains: visual domain (VI, 12 sub-networks), cerebellar domain (CB, 13 sub-networks), temporal-parietal domain (TP, 13 sub-networks), sub-cortical domain (SC, 23 sub-networks), sensorimotor domain (SM, 13 sub-networks), and high-level cognitive domain (HC, 31 sub-networks).
Estimating two-way and multiway interactions in infancy rsfMRI
Two-Way Interactions (Pearson Correlation): The infancy rsfMRI signal comprising n ICNs obtained from scICA, denoted as Xi (1≤i≤n and n = 105), each Xi corresponds informally to a time instance ti in a sequence {t1, t2, …, tT} with a constant time interval t. Therefore, the two-way functional connectivity based on Pearson correlation can be estimated as follows:
where \(\widehat{{X}_{i}}\) and \(\widehat{{X}_{j}}\) are the average of the rsfMRI signals at ICNs i and j. When two ICNs share substantial information, they are expected to exhibit a strong correlation, and conversely. In total, there are 1052 pairwise interactions based on Pearson correlation.
Two-Way Interactions (Mutual Information): The mutual information between one ICN Xi and another ICN Xj is given by58.
where the Shannon entropy h(Xi(t)), h(Xj(t)) of a continuous random variable \(X\in {{\mathcal{X}}}\) with probability density function fX, its differential entropy is defined as58,
The mutual information measures the dependency between Xi(t) and Xj(t), and attains its minimum, equal to zero, if two ICNs independent. In total, there are 1052 pairwise interactions based on mutual information.
Although two-way interactions are commonly used to estimate functional connectivity in fMRI studies, we know that information interactions in the human brain usually go beyond two-way interactions. Considering the human brain as a high-dimensional and nonlinear complex system, relying solely on two-way interactions to explain brain function is insufficient and often overlooks higher-order interactions. Therefore, considering the limitations of two-way interactions20,24,26,28,29, we applied total correlation to overcome these limitations and estimate higher-order information interactions in the human brain.
Multiway Interactions (Total Correlation): The Total Correlation (TC) describe the dependence among n variables (X1, ⋯ , Xn) and can be considered as a non-negative generalization of the concept of mutual information from two parties to n parties. Let the definition of total correlation due Watanabe35 be denoted as:
Where \(h({X}^{i})\) is marginal entropy, and \(h({X}^{1},\cdots \,,{X}^{n})\) is joint entropy. The TC will be equal to mutual information if we only have two variables, and TC will be zero if all variables are totally independent.
In real situations, estimating the marginal entropy \({{\bf{h}}}({X}^{i})\) is straightforward, but estimating the joint entropy \({{\bf{h}}}({X}^{1},{X}^{2}\cdots \,,{X}^{n})\) is considerably challenging. To address this challenge, Gaussian information theory is commonly applied to estimate total correlation because the BOLD signals satisfy Gaussian distributions24,30,59,60. For a univariate Gaussian random variable \(X \sim {{\mathcal{N}}}(\mu ,\sigma )\), the entropy (given in nats) will be,
And for a multivariate Gaussian distribution, the joint entropy will be,
where ∣Σ∣ refers to the determinant of the covariance matrix of \(({X}^{1},\cdots \,,{X}^{n})\). For the multivariate Gaussian case, the mutual information between \(({X}^{1},\cdots \,,{X}^{n})\) and \(({Y}^{1},\cdots \,,{Y}^{n})\) is given by,
Finally, the Gaussian estimator for TC can be:
The Gaussian estimator for TC operates without additional hyperparameters, relying solely on the covariance structure to reduce overall model complexity. Now, we can estimate interactions beyond two-way (k = 2) by computing k-way interactions (k > 2) among n = 105 ICNs, iterating over each set of indices used to obtain TC. In total, there are 1053 triple interactions if k = 3 is considered for all possible triple interactions, and there are \(\left(\begin{array}{l}105\\ 3\end{array}\right)\) unique sets of triple interactions for multiscale infancy brain networks.
Decomposing two-way and triple interaction tensors
Two-way interactions using PC and MI, as well as triple interaction tensors based on TC, were computed individually for each subject across 105 ICNs. Regarding triple interactions, we first extracted unique pairwise and triple interactions, flattened them, and then combined them across all subjects. These interactions were then aggregated across subjects and underwent blind ICA to extract the most independent components. To ensure the robustness and reliability of the ICA results, we executed the analysis 100 times using the Infomax optimization algorithm61. Each run included random initialization and bootstrapping techniques to mitigate potential biases and enhance the generalizability of the findings. This rigorous approach aimed to validate the stability and consistency of the identified components across multiple iterations, thereby reinforcing the overall validity of the ICA results62,63.
The decomposed components from the two-way and triple interactions were then ranked in descending order based on their self-weights. We focused on the top 10 weights and determined their corresponding indices to highlight the most distinctive triple-brain network interactions. We proceeded by evaluating the strength of these interactions across each ICN. Further analysis led us to identify the two strongest networks based on their aggregated weights. Subsequently, we pinpointed the networks that contributed most significantly to infancy development by examining interactions between the 105 ICNs and these top two networks.
Assessing the reliability of ICA for two-way and triple interactions in infants
To measure reliability, we computed the average spatial similarity between corresponding ICNs from the two independent half-splits using Pearson correlation, as illustrated in Fig. 6. Assessing the reliability of ICA for concatenated two-way (PC and MI) and triple (TRI) interactions across subjects is crucial for evaluating the stability and robustness of ICA55. In this study, we assessed the reliability of ICA for two-way and triple interactions by initially dividing the dataset randomly into two distinct subsets. Each subset underwent independent ICA processing, repeated 20 times. Each iteration produced components and their respective stability values (IQ), and the average spatial similarities were reported to evaluate the performance of ICA.
The entire infant dataset was randomly split in half. ICA was applied to each half-split, concatenating two-way and triple interactions across subjects to generate 20 components and their corresponding IQ. This process was independently repeated 20 times. Finally, the average spatial similarity was calculated.
Analyzing linear and nonlinear association with infant age using the general linear models and generalized additive models
To investigate age-related changes in functional connectivity, we employed the general linear models (GLMs)64 to assess the age effect on multiscale brain network interactions in infancy. We included gender, site, head motion parameters (mean FD) and TR as covariates. The general linear model is defined as follows:
where YFC represents the connectome measures of each infant, βage denotes the age effect. Bonferroni correction was applied to adjust for multiple comparisons across the measures, and significance was determined using a Bonferroni corrected p < 0.05.
To test the nonlinearity between brain networks and infant age, we applied generalized additive models (GAMs)65. GAMs are a flexible extension of generalized linear models (GLMs) that allow for nonlinear relationships between brain network interactions and infant age. In this context, GAMs are applied to model how functional connectivity changes with age, while also accounting for additional covariates such as gender, site, motion and TR. Mathematically, a GAM for modeling brain functional connectivity YFC can be expressed as:
where YFC represents the functional connectivity measure for the ith ICN. β0 is the intercept term. f1(agei) is a smooth function of age, capturing the nonlinear relationship between age and functional connectivity. f2(covariatei) is a smooth function of other covariates (e.g., gender, site, motion and TR). ϵi represents the residual error term, assumed to be normally distributed. The functions f1 and f2 are typically estimated nonparametrically using spline smoothing techniques, allowing for the flexible modeling of nonlinear effects. This approach enables the detection of complex patterns in functional connectivity that might be overlooked in traditional linear models, providing a more accurate understanding of how brain networks evolve over the lifespan. Here, we used Restricted Maximum Likelihood (REML) to select the optimal level of smoothing, which estimates smoothing parameters while accounting for the fixed effects. The method was implemented using the “mgcv” library in the R environment (https://cran.r-project.org/web/packages/mgcv/index.html). To quantify fitting performance, we calculated the Akaike Information Criterion (AIC)66. AIC assesses each model’s quality relative to others by balancing goodness of fit with model complexity, penalizing models with excessive complexity to prevent overfitting. Lower AIC values indicate a better balance between fit and complexity66.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Ethics Statement
The infant datasets used in this work in under the study protocol approved by the Emory University Institutional Review Board (IRB). Legal guardians provided written consent for their infant’s participation at time of study enrollment. All ethical regulations relevant to human research participants were followed.
Data availability
Data collected from National Institute of Mental Health (NIMH) 2P50MH100029, R01MH118285, and R01MH119251 are available from the NIMH Data Archive (NDA). NDA is a collaborative informatics system created by the National Institutes of Health to provide a national resource to support and accelerate research in mental health. Dataset identifiers: https://doi.org/10.15154/vjcy-f58967. NeuroMark 2.1 templates are accessible on our lab website (https://trendscenter.org/data/) and GitHub (https://github.com/trendscenter/gift/tree/master/GroupICAT/icatb/icatb_templates). The UNC-BCP 4D Infant Brain Template is available at https://www.nitrc.org/projects/uncbcp_4d_atlas/.
Code availability
The codes of the GICA, and MOO-ICAR have been integrated into the group ICA Toolbox (GIFT 4.0c, https://trendscenter.org/software/gift/). The GAM model from the “mgcv" library in the R environment (https://cran.r-project.org/web/packages/mgcv/index.html).
References
Power, J. D. et al. Functional network organization of the human brain. Neuron 72, 665–678 (2011).
Park, H.-J. & Friston, K. Structural and functional brain networks: from connections to cognition. Science 342, 1238411 (2013).
Iraji, A. et al. Multi-spatial scale dynamic interactions between functional sources reveal sex-specific changes in schizophrenia. Netw. Neurosci. 6, 1–48 (2021).
Iraji, A. et al. Spatial dynamics within and between brain functional domains: A hierarchical approach to study time-varying brain function. Hum. Brain Mapp. 40, 1969–1986 (2018).
Breukelaar, I. A. et al. Cognitive ability is associated with changes in the functional organization of the cognitive control brain network. Hum. Brain Mapp. 39, 5028–5038 (2018).
Haartsen, R., Jones, E. J. H. & Johnson, M. H. Human brain development over the early years. Curr. Opin. Behav. Sci. 10, 149–154 (2016).
Tsang, T. et al. Salience network connectivity is altered in 6-week-old infants at heightened likelihood for developing autism. Commun. Biol. 7, 485 (2024).
Liu, J. et al. Altered lateralization of dorsal language tracts in 6-week-old infants at risk for autism. Dev. Sci. 22 3, e12768 (2018).
Tran, X. A. et al. Functional connectivity during language processing in 3-month-old infants at familial risk for autism spectrum disorder. Eur. J. Neurosci. 53, 1621 – 1637 (2020).
Bruchhage, M. M. K., Ngo, G.-C., Schneider, N., D’Sa, V. & Deoni, S. C. L. Functional connectivity correlates of infant and early childhood cognitive development. Brain Struct. Funct. 225, 669 – 681 (2020).
Blass, E. M. & Camp, C. A. Biological bases of face preference in 6-week-old infants. Dev. Sci. 6, 524–536 (2003).
Seraji, M. et al. Spatial development of brain networks during the first six postnatal months. Commun. Biol. 8, 1514 (2025).
Fransson, P. et al. Resting-state networks in the infant brain. Proc. Natl. Acad. Sci. USA 104, 15531–15536 (2007).
Doria, V. et al. Emergence of resting state networks in the preterm human brain. Proc. Natl. Acad. Sci. USA 107, 20015–20020 (2010).
Fransson, P., Åden, U., Blennow, M. & Lagercrantz, H. The functional architecture of the infant brain as revealed by resting-state fmri. Cereb. Cortex 21, 145–154 (2011).
Gao, W. et al. Evidence on the emergence of the brain’s default network from 2-week-old to 2-year-old healthy pediatric subjects. Proc. Natl. Acad. Sci. USA 106, 6790–5 (2009).
Gao, W. et al. Functional network development during the first year: relative sequence and socioeconomic correlations. Cereb. Cortex 25, 2919–2928 (2015).
Gao, W. et al. Temporal and spatial evolution of brain network topology during the first two years of life. PloS One 6, e25278 (2011).
Gilmore, J. H., Knickmeyer, R. C. & Gao, W. Imaging structural and functional brain development in early childhood. Nat. Rev. Neurosci. 19, 123–137 (2018).
Li, Q. Functional connectivity inference from fmri data using multivariate information measures. Neural Netw. 146, 85–97 (2022).
Ashrafi, M. & Soltanian-Zadeh, H. Multivariate Gaussian copula mutual information to estimate functional connectivity with less random architecture. Entropy 24, 631 (2022).
Li, Q., Yu, S., Madsen, K. H., Calhoun, V. D. & Iraji, A. Higher-order organization in the human brain from matrix-based rényi’s entropy. In 2023 IEEE International Conference on Acoustics, Speech, and Signal Processing Workshops (ICASSPW), 1–5 (2023).
Herzog, R. et al. Genuine high-order interactions in brain networks and neurodegeneration. Neurobiol. Dis. 175, 105918 (2022).
Gatica, M. et al. High-order interdependencies in the aging brain. Brain Connect. 11, 734–744 (2021).
Xie, Q. et al. Constructing high-order functional connectivity network based on central moment features for diagnosis of autism spectrum disorder. PeerJ 9, e11692 (2021).
Li, Q., Steeg, G. V., Yu, S. & Malo, J. Functional connectome of the human brain with total correlation. Entropy 24, 1725 (2022).
Iraji, A. et al. Functional multi-connectivity: A novel approach to assess multi-way entanglement between networks and voxels. In 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI), 1698–1701 (2020).
Battiston, F. et al. Networks beyond pairwise interactions: structure and dynamics. Phys. Rep. 874, 1–92 (2020).
Li, Q., Ver Steeg, G. & Malo, J. Functional connectivity via total correlation: analytical results in visual areas. Neurocomputing 571, 127143 (2023).
Li, Q. et al. Aberrant high-order dependencies in schizophrenia resting-state functional MRI networks. In NeurIPS 2023 workshop: Information-Theoretic Principles in Cognitive Systems (2023).
Iraji, A. et al. Identifying canonical and replicable multi-scale intrinsic connectivity networks in 100k+ resting-state fmri datasets. Hum. Brain Mapp. 44, 5729–5748 (2023).
Calhoun, V. D., Adalı, T., Pearlson, G. D. & Pekar, J. J. A method for making group inferences from functional mri data using independent component analysis. Hum. Brain Mapp.14 (2001).
Faghiri, A., Iraji, A., Adalı, T. & Calhoun, V. D. Analysis of high-order brain networks resolved in time and frequency using cp decomposition. ICASSP 2024 - 2024 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP) (2024).
Bajracharya, P. et al. Born connected: Do infants already have adult-like multi-scale connectivity networks? bioRxiv (2024).
Watanabe, S. Information theoretical analysis of multivariate correlation. IBM J. Res. Dev. 4, 66–82 (1960).
Du, Y. et al. Artifact removal in the context of group ica: a comparison of single-subject and group approaches. Hum. Brain Mapp. 37, 1005–1025 (2015).
Torres, L., Blevins, A. S., Bassett, D. & Eliassi-Rad, T. The why, how, and when of representations for complex systems. SIAM Rev. 63, 435–485 (2021).
Benson, A. R., Gleich, D. F. & Higham, D. J. Higher-order network analysis takes off, fueled by classical ideas and new data. https://arxiv.org/abs/2103.05031 (2021).
Luppi, A. et al. Systematic evaluation of fmri data-processing pipelines for consistent functional connectomics. Nat. Commun. 15, 4745 (2024).
Zhang, Y., Lucas, M. & Battiston, F. Higher-order interactions shape collective dynamics differently in hypergraphs and simplicial complexes. Nat. Commun. 14, 1605 (2023).
Rieck, B. et al. Uncovering the topology of time-varying fmri data using cubical persistence. In NeurIPS (2020).
Han, T. S. Nonnegative entropy measures of multivariate symmetric correlations. Inf. Control 36, 133–156 (1978).
Rosas, F. E., Mediano, P. A., Gastpar, M. & Jensen, H. J. Quantifying high-order interdependencies via multivariate extensions of the mutual information. Phys. Rev. E 100, 032305 (2019).
Iraji, A., Miller, R., Adali, T. & Calhoun, V. Space: A missing piece of the dynamic puzzle. Trends Cogn. Sci.24 (2020).
Allen, E. et al. Tracking whole-brain connectivity dynamics in the resting state. Cerebral Cortex (New York, N.Y.: 1991) (2012).
Iraji, A. et al. Tools of the trade: Estimating time-varying connectivity patterns from fmri data. Soc. Cogn. Affect. Neurosci. 16, 849–874 (2020).
Vidaurre, D., Smith, S. M. & Woolrich, M. W. Brain network dynamics are hierarchically organized in time. Proc. Natl. Acad. Sci. USA 114, 12827 – 12832 (2017).
Iraji, A. et al. The spatial chronnectome reveals a dynamic interplay between functional segregation and integration. Hum. Brain Mapp. 40, 3058–3077 (2019).
Shultz, S., Klin, A. & Jones, W. Neonatal transitions in social behavior and their implications for autism. Trends Cogn. Sci. 22, 452–469 (2018).
Ford, A. et al. Caregiver greeting to infants under 6 months already reflects emerging differences in those later diagnosed with autism. Proc. B 291, 20232494 (2024).
Chen, L. et al. A 4d infant brain volumetric atlas based on the unc/umn baby connectome project (bcp) cohort. NeuroImage 253, 119097 (2022).
Du, Y. et al. Neuromark: An automated and adaptive ica based pipeline to identify reproducible fmri markers of brain disorders. NeuroImage: Clin. 28, 102375 (2020).
Meng, X. et al. Multi-model order spatially constrained ica reveals highly replicable group differences and consistent predictive results from resting data: a large n fmri schizophrenia study. NeuroImage: Clin. 38, 103434 (2023).
Calhoun, V. et al. Exploring the psychosis functional connectome: Aberrant intrinsic networks in schizophrenia and bipolar disorder. Front. Psychiatry Front. Res. Found. 2, 75 (2011).
Erhardt, E. et al. Comparison of multi-subject ica methods for analysis of fmri data. Hum. Brain Mapp. 32, 2075–95 (2011).
Lin, Q.-H., Liu, J., Zheng, Y.-R., Liang, H. & Calhoun, V. Semiblind spatial ICA of fmri using spatial constraints. Hum. Brain Mapp. 31, 1076–88 (2009).
Lu, W. & Rajapakse, J. Approach and applications of constrained ica. IEEE Trans. Neural Netw. 16, 203–212 (2005).
T. M., C. & J. A., T. Elements of Information Theory 2 edn (Wiley-Interscience, Hoboken, NJ, USA, 2006).
Hlinka, J., Paluš, M., Vejmelka, M., Mantini, D. & Corbetta, M. Functional connectivity in resting-state fmri: is linear correlation sufficient?. Neuroimage 54, 2218–2225 (2011).
Liégeois, R., Yeo, B. T. & Van De Ville, D. Interpreting null models of resting-state functional mri dynamics: not throwing the model out with the hypothesis. NeuroImage 243, 118518 (2021).
Bell, A. J. & Sejnowski, T. J. An information-maximization approach to blind separation and blind deconvolution. Neural Comput. 7, 1129–1159 (1995).
Abou Elseoud, A. et al. The effect of model order selection in group pica. Hum. Brain Mapp. 31, 1207–16 (2009).
Ray, K. et al. ICA model order selection of task co-activation networks. Front. Neurosci. 7, 237 (2013).
Nelder, J. A. & Wedderburn, R. W. M. Generalized linear models. J. R. Stat. Soc. Ser. A (Gen.) 135, 370–384 (1972).
Hastie, T. & Tibshirani, R. Generalized Additive Models (Wiley Online Library, 1990).
Cavanaugh, J. E. & Neath, A. A. The akaike information criterion: Background, derivation, properties, application, interpretation, and refinements. WIREs Comput. Stat. 11, e1460 (2019).
Seraji, M. et al. Spatial development of brain networks during the first six postnatal months. [Data set]. NIMH Data Archive https://doi.org/10.15154/vjcy-f589 (2026).
Acknowledgements
We are deeply grateful to the families and their infants who generously volunteered to participate in this research. We also extend our sincere thanks to the research coordinators, assistants, and fellows at the Marcus Autism Center-Brittney Sholar, Carly Reineri, Joanna Beugnon, Lindsey Evans, Jordan Pincus, Jennifer Gutierrez, Tristan Ponzo, and Adriana Mendez-for their invaluable efforts in data collection. We further thank the MRI technologists at the Emory Center for Systems Imaging Core-Michael White, Sarah Basadre, and Samira Yeboah-for their skilled technical support. Finally, we acknowledge Dr.Lei Zhou and Michael Valente for their contributions to equipment development and data acquisition protocols.
This work was supported by the National Institutes of Health (NIH) grant number R01MH119251 to Sarah Shultz and Armin Iraji, R01EB027147 to Sarah Shultz and Vince Calhoun, P50MH100029 to Sarah Shultz, and National Science Foundation (NSF) grant number 2112455 to Vince Calhoun.
Author information
Authors and Affiliations
Contributions
Q.L. served as the first author, with primary responsibility for the conception, design, and overall execution of the study, as well as for drafting the manuscript. S.S. contributed critically to data collection and manuscript preparation, ensuring the accuracy and integrity of the collected data. Z.F., H.W., M.S., and P.B. were responsible for data preprocessing and made significant contributions to manuscript drafting. A.I., S.S., and V.C. provided expert supervision throughout the research process, offering essential guidance, critical feedback, and substantive contributions to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Michel Thiebaut de Schotten and Jasmine Pan.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Q., Fu, Z., Walum, H. et al. Deciphering multiway multiscale brain network connectivity from birth to 6 months. Commun Biol 9, 271 (2026). https://doi.org/10.1038/s42003-026-09549-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s42003-026-09549-3