Abstract
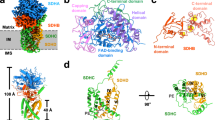
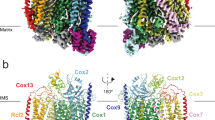
Respiratory complex II (CII), or succinate dehydrogenase, couples succinate oxidation in the Krebs cycle with electron transfer to the respiratory chain. Owing to this pivotal role, CII inhibitors are widely used fungicides globally; however, their development has largely proceeded without structural insights from fungal targets. Here, we report cryo-electron microscopy structures of the 128 kDa mitochondrial CII from Saccharomyces cerevisiae in two states: active, with endogenous ubiquinone-6 bound (3.15 Å), and inhibited with the fungicide bixafen (3.00 Å). Although closely related to the mammalian type C enzyme, our structures show that the yeast CII has lost the canonical heme cofactor. They also reveal how clade-specific sequence extensions of the membrane subunits Sdh3 and Sdh4 - conserved in pathogenic fungi - uniquely contribute to complex stability and fungicide binding. Our findings provide a foundation for rational design of next-generation CII inhibitors and combatting resistance, in both agriculture and human health.
Similar content being viewed by others
Data availability
E.M. maps and model coordinates for CII have been deposited in the EM Data Bank and Protein Data Bank, respectively, under accession numbers EM-53029 and PDB-9QDL for CII as purified (CII-nat), and accession numbers EM-53030 and PDB-9QLM for CII with bixafen bound (CII-bix). Raw data presented in Supplementary Fig. 1 for the characterization of the final purified CII sample used for structure determination are available in the Supplementary Data file, with uncropped gels provided in Supplementary Fig. 15.
References
Rich, P. R. & Maréchal, A. Electron transfer chains: structures, mechanisms and energy coupling In Comprehensive Biophysics Vol. 8 (ed Ferguson S. J.) 72-93 (Elsevier Inc., 2012).
Luttik, M. A. et al. The Saccharomyces cerevisiae NDE1 and NDE2 genes encode separate mitochondrial NADH dehydrogenases catalyzing the oxidation of cytosolic NADH. J. Biol. Chem. 273, 24529–24534 (1998).
Feng, Y. et al. Structural insight into the type-II mitochondrial NADH dehydrogenases. Nature 491, 478–482 (2012).
Hartley, A. M. et al. Structure of yeast cytochrome c oxidase in a supercomplex with cytochrome bc1. Nat. Struct. Mol. Biol. 26, 78–83 (2019).
Lange, C. & Hunte, C. Crystal structure of the yeast cytochrome bc1 complex with its bound substrate cytochrome c. Proc. Natl. Acad. Sci. USA 99, 2800–2805 (2002).
Guo, H., Bueler, S. A. & Rubinstein, J. L. Atomic model for the dimeric FO region of mitochondrial ATP synthase. Science 358, 936–940 (2017).
Rathore, S. et al. Cryo-EM structure of the yeast respiratory supercomplex. Nat. Struct. Mol. Biol. 26, 50–57 (2019).
Hartley, A. M., Meunier, B., Pinotsis, N. & Maréchal, A. Rcf2 revealed in cryo-EM structures of hypoxic isoforms of mature mitochondrial III-IV supercomplexes. Proc. Natl. Acad. Sci. USA 117, 9329–9337 (2020).
Iverson, T. M., Singh, P. K. & Cecchini, G. An evolving view of complex II-noncanonical complexes, megacomplexes, respiration, signaling, and beyond. J. Biol. Chem. 299, 104761 (2023).
Karavaeva, V. & Sousa, F. L. Modular structure of complex II: An evolutionary perspective. Biochim Biophys. Acta Bioenerg. 1864, 148916 (2023).
Spinelli, J. B. et al. Fumarate is a terminal electron acceptor in the mammalian electron transport chain. Science 374, 1227–1237 (2021).
Murphy, M. P. & Chouchani, E. T. Why succinate? Physiological regulation by a mitochondrial coenzyme Q sentinel. Nat. Chem. Biol. 18, 461–469 (2022).
Hägerhäll, C. Succinate: quinone oxidoreductases. Variations on a conserved theme. Biochim Biophys. Acta 1320, 107–141 (1997).
Hägerhäll, C. & Hederstedt, L. A structural model for the membrane-integral domain of succinate: quinone oxidoreductases. FEBS Lett. 389, 25–31 (1996).
Gong, H. et al. Cryo-EM structure of trimeric Mycobacterium smegmatis succinate dehydrogenase with a membrane-anchor SdhF. Nat. Commun. 11, 4245 (2020).
Lancaster, C. R., Kroger, A., Auer, M. & Michel, H. Structure of fumarate reductase from Wolinella succinogenes at 2.2 A resolution. Nature 402, 377–385 (1999).
Yankovskaya, V. et al. Architecture of succinate dehydrogenase and reactive oxygen species generation. Science 299, 700–704 (2003).
Iverson, T. M., Luna-Chavez, C., Cecchini, G. & Rees, D. C. Structure of the Escherichia coli fumarate reductase respiratory complex. Science 284, 1961–1966 (1999).
Zhou, X. et al. Architecture of the mycobacterial succinate dehydrogenase with a membrane-embedded Rieske FeS cluster. Proc. Natl. Acad. Sci. USA 118, https://doi.org/10.1073/pnas.2022308118 (2021).
Sun, F. et al. Crystal structure of mitochondrial respiratory membrane protein Complex II. Cell 121, 1043–1057 (2005).
Huang, L. S., Shen, J. T., Wang, A. C. & Berry, E. A. Crystallographic studies of the binding of ligands to the dicarboxylate site of Complex II, and the identity of the ligand in the “oxaloacetate-inhibited” state. Biochim Biophys. Acta 1757, 1073–1083 (2006).
Shimizu, H. et al. Crystal structure of mitochondrial quinol-fumarate reductase from the parasitic nematode Ascaris suum. J. Biochem 151, 589–592 (2012).
Du, Z. et al. Structure of the human respiratory complex II. Proc. Natl. Acad. Sci. USA 120, e2216713120 (2023).
Mühleip, A. et al. Structural basis of mitochondrial membrane bending by the I-II-III2-IV2 supercomplex. Nature 615, 934–938 (2023).
Han, F. et al. Structures of Tetrahymena thermophila respiratory megacomplexes on the tubular mitochondrial cristae. Nat. Commun. 14, 2542 (2023).
Wú, F. M. et al. Structure of the II2-III2-IV2 mitochondrial supercomplex from the parasite Perkinsus marinus. bioRXiv https://doi.org/10.1101/2024.1105.1125.595893 (2024).
Hoekstra, A. S. & Bayley, J. P. The role of complex II in disease. Biochim Biophys. Acta 1827, 543–551 (2013).
Benit, P. et al. Succinate dehydrogenase, succinate, and superoxides: a genetic, epigenetic, metabolic, environmental explosive crossroad. Biomedicines 10, https://doi.org/10.3390/biomedicines10081788 (2022).
Chouchani, E. T. et al. Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature 515, 431–435 (2014).
Valls-Lacalle, L. et al. Succinate dehydrogenase inhibition with malonate during reperfusion reduces infarct size by preventing mitochondrial permeability transition. Cardiovasc. Res. 109, 374–384 (2016).
Adolph, C. et al. Identification of chemical scaffolds that inhibit the Mycobacterium tuberculosis respiratory complex succinate dehydrogenase. ACS Infect. Dis. 10, 3496–3515 (2024).
Sierotzki, H. & Scalliet, G. A review of current knowledge of resistance aspects for the next-generation succinate dehydrogenase inhibitor fungicides. Phytopathology 103, 880–887 (2013).
Desbordes, P. et al. Isoflucypram, the first representative of a new succinate dehydrogenase inhibitor fungicide subclass: Its chemical discovery and unusual binding mode. Pest Manag Sci. 76, 3340–3347 (2020).
Duarte Hospital, C. et al. SDHi fungicides: An example of mitotoxic pesticides targeting the succinate dehydrogenase complex. Environ. Int 180, 108219 (2023).
Esterio, M. et al. Control of Botrytis cinerea from Chilean grapevines by pydiflumetofen: baseline and carboxamide-mutant sensitivity. Plant Dis. 109, 445–453 (2024).
Li, S., Li, X., Zhang, H., Wang, Z. & Xu, H. The research progress in and perspective of potential fungicides: Succinate dehydrogenase inhibitors. Bioorg. Med. Chem. 50, 116476 (2021).
Huang, L.-S., Luemmen, P. & Berry, E. Crystallographic investigation of the ubiquinone binding site of respiratory Complex II and its inhibitors. Biochim Biophys. Acta Proteins Proteom. 1869, 140679 (2021).
Inaoka, D. K. et al. Structural insights into the molecular design of flutolanil derivatives targeted for fumarate respiration of parasite mitochondria. Int J. Mol. Sci. 16, 15287–15308 (2015).
Robinson, K. M. & Lemire, B. D. Isolation and nucleotide sequence of the Saccharomyces cerevisiae gene for the succinate dehydrogenase flavoprotein subunit. J. Biol. Chem. 267, 10101–10107 (1992).
Lombardo, A., Carine, K. & Scheffler, I. E. Cloning and characterization of the iron-sulfur subunit gene of succinate dehydrogenase from Saccharomyces cerevisiae. J. Biol. Chem. 265, 10419–10423 (1990).
Daignan-Fornier, B., Valens, M., Lemire, B. D. & Bolotin-Fukuhara, M. Structure and regulation of SDH3, the yeast gene encoding the cytochrome b560 subunit of respiratory complex II. J. Biol. Chem. 269, 15469–15472 (1994).
Bullis, B. L. & Lemire, B. D. Isolation and characterization of the Saccharomyces cerevisiae SDH4 gene encoding a membrane anchor subunit of succinate dehydrogenase. J. Biol. Chem. 269, 6543–6549 (1994).
Maklashina, E., Rajagukguk, S., McIntire, W. S. & Cecchini, G. Mutation of the heme axial ligand of Escherichia coli succinate–quinone reductase: Implications for heme ligation in mitochondrial complex II from yeast. Biochim Biophys. Acta Bioenerg. 1797, 747–754 (2010).
Crichton, P. G., Harding, M., Ruprecht, J. J., Lee, Y. & Kunji, E. R. S. Lipid, detergent, and Coomassie blue G-250 affect the migration of small membrane proteins in blue native gels: mitochondrial carriers migrate as monomers not dimers. J. Biol. Chem. 288, 22163–22173 (2013).
Yamashita, M. & Fraaije, B. Non-target site SDHI resistance is present as standing genetic variation in field populations of Zymoseptoria tritici. Pest Manag Sci. 74, 672–681 (2018).
Zhang, A. et al. Discovery of N-(4-fluoro-2-(phenylamino)phenyl)-pyrazole-4-carboxamides as potential succinate dehydrogenase inhibitors. Pestic. Biochem. Physiol. 158, 175–184 (2019).
Harding, M. M. Small revisions to predicted distances around metal sites in proteins. Acta Crystallogr D. Biol. Crystallogr 62, 678–682 (2006).
Zhang, Y. et al. Structure of the mitochondrial TIM22 complex from yeast. Cell Res 31, 366–368 (2021).
Maklashina, E., Rothery, R. A., Weiner, J. H. & Cecchini, G. Retention of heme in axial ligand mutants of succinate-ubiquinone xxidoreductase (complex II) from Escherichia coli. J. Biol. Chem. 276, 18968–18976 (2001).
Oyedotun, K. S., Yau, P. F. & Lemire, B. D. Identification of the heme axial ligands in the cytochrome b562 of the Saccharomyces cerevisiae succinate dehydrogenase. J. Biol. Chem. 279, 9432–9439 (2004).
Broomfield, P. L. & Hargreaves, J. A. A single amino-acid change in the iron-sulphur protein subunit of succinate dehydrogenase confers resistance to carboxin in Ustilago maydis. Curr. Genet. 22, 117–121 (1992).
Skinner, W. et al. A single amino-acid substitution in the iron-sulphur protein subunit of succinate dehydrogenase determines resistance to carboxin in Mycosphaerella graminicola. Curr. Genet. 34, 393–398 (1998).
Matsson, M., Ackrell, B. A., Cochran, B. & Hederstedt, L. Carboxin resistance in Paracoccus denitrificans conferred by a mutation in the membrane-anchor domain of succinate:quinone reductase. Arch. Microbiol. 170, 27–37 (1998).
Matsson, M. & Hederstedt, L. The carboxin-binding site on Paracoccus denitrificans succinate:quinone reductase identified by mutations. J. Bioenerg. Biomembr. 33, 99–105 (2001).
Shima, Y., Ito, Y., Hatabayashi, H., Koma, A. & Yabe, K. Five carboxin-resistant mutants exhibited various responses to carboxin and related fungicides. Biosci. Biotechnol. Biochem 75, 181–184 (2011).
Yin, Y. N., Kim, Y. K. & Xiao, C. L. Molecular characterization of boscalid resistance in field isolates of Botrytis cinerea from apple. Phytopathology 101, 986–995 (2011).
Lucas, J. A., Hawkins, N. J. & Fraaije, B. A. The evolution of fungicide resistance. Adv. Appl Microbiol 90, 29–92 (2015).
Vielba-Fernandez, A. et al. Resistance to the SDHI fungicides boscalid and fluopyram in Podosphaera xanthii populations from commercial cucurbit fields in Spain. J. Fungi (Basel) 7, https://doi.org/10.3390/jof7090733 (2021).
Miyadera, H. et al. Atpenins, potent and specific inhibitors of mitochondrial complex II (succinate-ubiquinone oxidoreductase). Proc. Natl. Acad. Sci. USA 100, 473–477 (2003).
Maklashina, E. & Cecchini, G. Comparison of catalytic activity and inhibitors of quinone reactions of succinate dehydrogenase (Succinate-ubiquinone oxidoreductase) and fumarate reductase (Menaquinol-fumarate oxidoreductase) from Escherichia coli. Arch. Biochem Biophys. 369, 223–232 (1999).
Schilling, R. J., Baldwin, T. & Palmer, G. The characterization of highly purified Complex II from baker’s yeast. Federation Proc. 41, 896 (1982).
Tran, Q. M., Rothery, R. A., Maklashina, E., Cecchini, G. & Weiner, J. H. Escherichia coli succinate dehydrogenase variant lacking the heme b. Proc. Natl. Acad. Sci. USA 104, 18007–18012 (2007).
Hassan, Y., Chew, S. Y. & Than, L. T. L. Candida glabrata: pathogenicity and resistance mechanisms for adaptation and survival. J. Fungi (Basel) 7, https://doi.org/10.3390/jof7080667 (2021).
Rodriguez-Cerdeira, C. et al. Virulence and resistance factors of Nakaseomyces glabratus (formerly known as Candida glabrata) in Europe: A systematic review. J. Eur. Acad. Dermatol. Venereol. https://doi.org/10.1111/jdv.20273 (2024).
Barns, S. M., Lane, D. J., Sogin, M. L., Bibeau, C. & Weisburg, W. G. Evolutionary relationships among pathogenic Candida species and relatives. J. Bacteriol. 173, 2250–2255 (1991).
Kellis, M., Birren, B. W. & Lander, E. S. Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428, 617–624 (2004).
Gebert, N. et al. Dual function of Sdh3 in the respiratory chain and TIM22 protein translocase of the mitochondrial inner membrane. Mol. Cell 44, 811–818 (2011).
Szeto, S. S. W., Reinke, S. N., Oyedotun, K. S., Sykes, B. D. & Lemire, B. D. Expression of Saccharomyces cerevisiae Sdh3p and Sdh4p paralogs results in catalytically active succinate dehydrogenase isoenzymes. J. Biol. Chem. 287, 22509–22520 (2012).
Chang, Y. L. et al. Instability of succinate dehydrogenase in SDHD polymorphism connects reactive oxygen species production to nuclear and mitochondrial genomic mutations in yeast. Antioxid. Redox Signal. 22, 587–602 (2015).
Kim, H. J., Khalimonchuk, O., Smith, P. M. & Winge, D. R. Structure, function, and assembly of heme centers in mitochondrial respiratory complexes. Biochim Biophys. Acta 1823, 1604–1616 (2012).
Meunier, B., Maréchal, A. & Rich, P. R. Construction of histidine-tagged yeast mitochondrial cytochrome c oxidase for facile purification of mutant forms. Biochem. J. 444, 199–204 (2012).
Aliverti, A., Curti, B. & Vanoni, M. A. Identifying and quantitating FAD and FMN in simple and in iron-sulfur-containing flavoproteins. Methods Mol. Biol. 131, 9–23 (1999).
Wittig, I., Karas, M. & Schägger, H. High resolution clear native electrophoresis for in-gel functional assays and fluorescence studies of membrane protein complexes. Mol. Cell. Proteom. 6, 1215–1225 (2007).
Armstrong, J. M. The molar extinction coefficient of 2,6-dichlorophenol indophenol. Biochim Biophys. Acta 86, 194–197 (1964).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. Sect. D: Struct. Biol. 75, 861–877 (2019).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Computat. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D: Biol. Crystallogr. 66, 486–501 (2010).
Zheng, H. et al. Validation of metal-binding sites in macromolecular structures with the CheckMyMetal web server. Nat. Protoc. 9, 156–170 (2014).
Gouet, P., Robert, X. & Courcelle, E. ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res 31, 3320–3323 (2003).
Acknowledgements
This work was supported by the Medical Research Council UK (Transition Support MR/T032154/1 to A.M.). Cryo-EM data were collected at the ISMB EM facility (Birkbeck College, University of London) with financial support from the Wellcome Trust (202679/Z/16/Z and 206166/Z/17/Z). We thank Dr D. Houldershaw for his support with computing.
Author information
Authors and Affiliations
Contributions
A.M. funded, designed, and supervised the research. B.M. produced the yeast mutant strain. C.B.-L. grew cells, prepared mitochondria, and purified complex II, with support from S.J. C.B.-L. performed activity measurements with input from E.A.B., S.C., and N.L., prepared and optimized cryo-EM grids, and collected cryo-EM data. N.P. and A.M. processed the cryo-EM images. N.P. and E.A.B. built the models. A.M., N.P., and E.A.B. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Masatoshi Murai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Janesh Kumar and Laura Rodríguez. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pinotsis, N., Burn-Leefe, C., Jones, S. et al. Cryo-EM structure of bixafen-bound S. cerevisiae complex II unravels SDHI specificity against pathogenic fungi. Commun Biol (2026). https://doi.org/10.1038/s42003-026-09617-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-026-09617-8